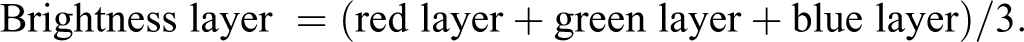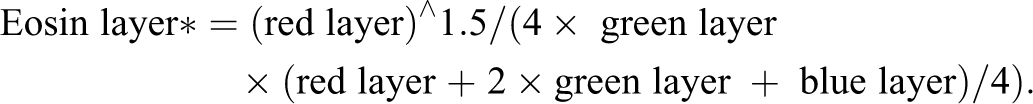Abstract
Bone marrow toxicity is a common finding when assessing safety of drug candidate molecules. Standard hematoxylin and eosin (H&E) marrow tissue sections are typically manually evaluated to provide a semiquantitative assessment of overall cellularity. Here, we developed an automated image analysis method that allows quantitative assessment of changes in bone marrow cell population in sternal bone. In order to test whether the method was repeatable and sensitive, we compared the automated method with manual subjective histopathology scoring of total cellularity in rat sternal bone marrow samples across 17 independently run studies. The automated method was consistent with manual scoring methodology for detecting altered bone marrow cellularity and, in multiple cases, identified changes at lower doses. The image analysis method allows rapid and more quantitative assessment of bone marrow toxicity compared to manual examination of H&E slides, making it an excellent tool to aid detection of bone marrow cell depletion in preclinical toxicologic studies.
Introduction
The bone marrow is a major site of toxicity in drug safety assessment (Travlos 2006). Histological examination of bone marrow provides crucial information regarding the state of the hematopoietic system. Drugs that target the immune system or rapidly dividing cells are likely to cause depletion of hematopoietic cell lineages.
For most preclinical safety assessment studies in rats, sternal bone marrow is assessed by manual, subjective microscopic examination of standard hematoxylin and eosin (H&E) histology slides. Manual examination typically involves a descriptive or semiquantitative estimation of total bone marrow cell depletion (Haley et al. 2005). It would, however, be useful to have a procedure that could quantify bone marrow cellularity on the same slides, without pathologist-dependent subjectivity.
To achieve this goal, we took advantage of the recent advances in whole-slide imaging and digital imaging technology (Webster and Dunstan 2014). We developed an automated image analysis algorithm, which quantifies the entire bone marrow cell population density within the slide tissue sample. The algorithm was successfully applied to images of bone marrow slides from rats in 17 studies performed over 9 years at multiple sites, demonstrating its reliability. In addition, the automated bone marrow cellularity analysis was applicable to sternal or femoral/tibial bone marrow samples and bone marrow of multiple species. We demonstrate that the automated method is reliable and sensitive, making it well suited for rapid screening, and use as a complimentary assay to standard manual subjective evaluation of bone marrow toxicity in preclinical toxicology models.
Method
Study Selection
For rat studies, 17 anonymized studies from preclinical animal safety experiments performed over 9 years were used for data analysis. Studies were selected with the following properties: A control group (no treatment except vehicle) was available. Sternal bone marrow cellularity had been scored on H&E slides by a pathologist on a six-point scale (0–5) of increasing cellular depletion. Only adult male and female Sprague-Dawley rats were used. Histologic preparations were of high enough quality (tissues had been fixed properly and did not have severe artifacts such as large tears) for automated analysis.
We also selected one historical study each of mouse, dog, and cynomolgus monkey, where sternal bone cellularity was manually scored to compare with automated scoring. Sections were 5 μm in thickness.
For experimental and sample preparation details for the study used to compare sternal and femoral/tibial bone, as well as for reproducibility analysis, see our companion article (Kozlowski et al. In Press).
Microscopic Evaluation
A veterinary pathologist recorded cellular changes in femoral/tibial (a preparation of left long bone stifle joints that included areas of femur and tibial cancellous and medullary bone marrow tissues) and sternal bone marrow H&E-stained sections. Cellular depletion in H&E sections was subjectively assessed on a six-point scale (0–5) in increasing severity: normal, minimal, mild, moderate, marked, and severe.
Whole-slide Imaging and Automated Image Analysis
All slides were scanned at 20× (0.46 μm/pixel) using the Nanozoomer HT 2.0 Slide scanner with an 8-bit camera. Definiens Developer v2.6 (Definiens AG, Munich, Germany) software (Baatz, Zimmermann, and Blackmore 2009) was used for image analysis.
This algorithm is available upon request in Definiens native .dcp format. We provide here a high-level overview and key features of the algorithm. The algorithm could be transferred to other software, provided the powerful “multi-resolution segmentation algorithm” in Definiens is also implemented (Benz et al. 2004). The images were analyzed in the RGB color space. For each color, a Definiens “layer” was assigned where the pixel intensity for that color was stored in 8 bit, from 0 (darkest) to 255 (brightest). Various layers were created by combining the pixel information from the different color layers. The specific components of the layers were visually tuned. The primary layers created in this way were the following:
*Note that the “eosin layer” and “hematoxylin layer” are so-called because they approximately correspond to these components of the H&E image but do not describe a truly color-deconvolved image of the two stains.
Scanned H&E slide images were analyzed at low magnification (0.2×) on a downsampled (reduced resolution) image (Figure 1A–D). The downsampled image was convolved with the Definiens “Gaussian blur” algorithm with kernel of 11 × 11 pixels on the red layer, producing a convolved red layer (conv-red). The tissue area was distinguished from the background by a threshold on conv-red, using the “auto-threshold” algorithm. Afterward, the Definiens “multi-resolution segmentation” algorithm was run with the scale parameter set to 30, to split similar looking tissues into cohesive pixel units. Evaluation classes were created for bone and the marrow. In Definiens Developer, evaluation classes are created by drawing a function shape describing the contributions of specific object features used to determine object classification. The standard deviation of the pixels in the normalized red layer (norm-red), the norm-red pixel values, and the pixel values of the green layer affected the classification of bone tissue. The marrow classification depended on the balance between the “hematoxylin layer” and “eosin layer” as well as the standard deviation in the blue layer. Once marrow and bone were separated, the average values of the “hematoxylin layer” and “eosin layer”, as well as the standard deviation of the red and blue layers were recorded, and used for evaluation classes in the higher magnification analysis.
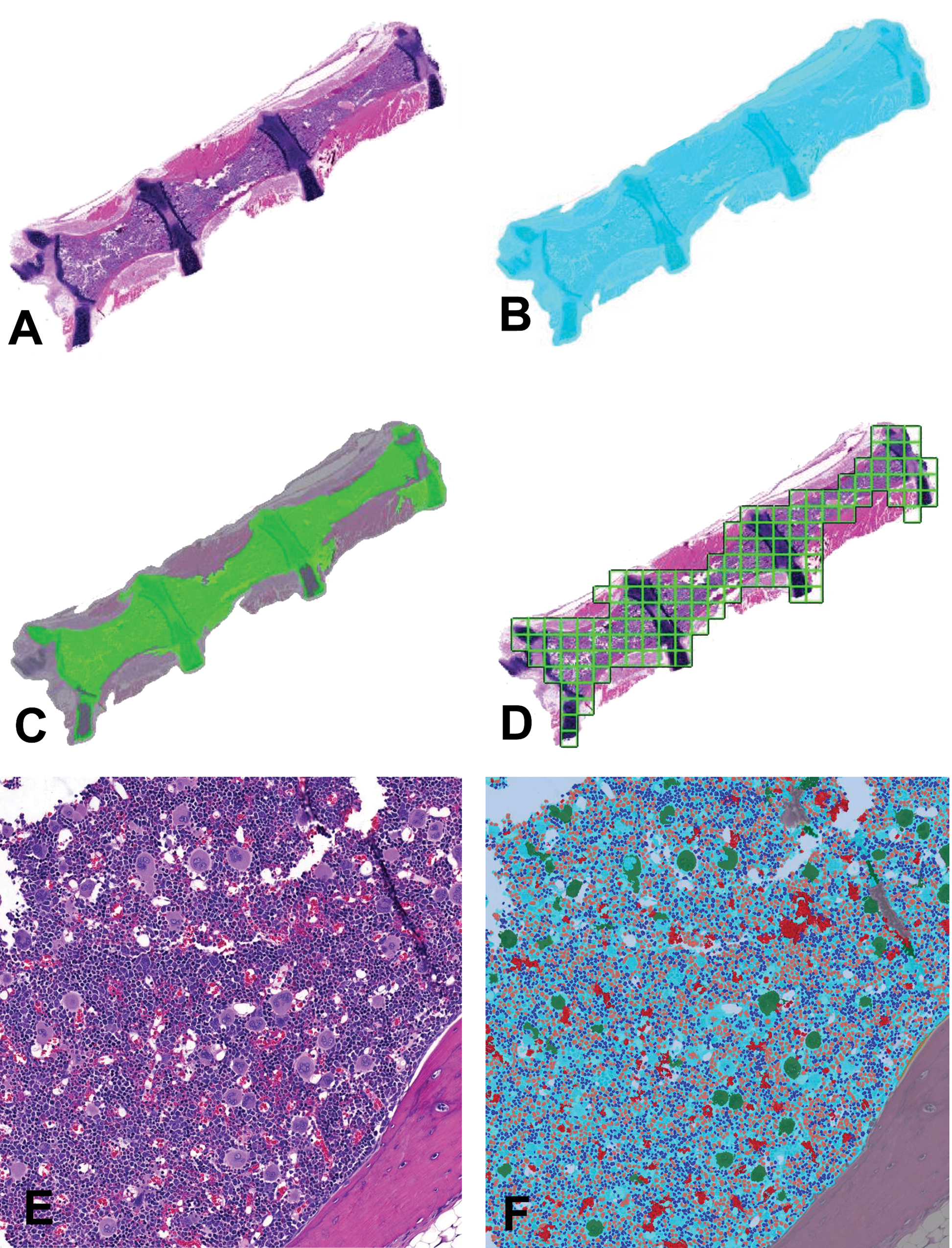
Image analysis method of low- and high-magnification images. (A) Low-magnification original hematoxylin and eosin (H&E) bone marrow sternum image from a control group rat. (B) Segmentation of tissue versus background. Detected tissue: cyan color transparency overlaid on image in A. (C) Within the detected tissue in B, marrow is identified. Marrow: green overlay, bone: gray overlay. (D) Marrow area in C is analyzed tile by tile in a grid pattern (grid outline shown in green). (E) High magnification tile section of original H&E bone marrow, equivalent to one square illustrated in D. (F) Image analysis results shown as color overlays of the type of objects that were segmented and classified, on the original image E. Bone and cartilage tissue are excluded from analysis of marrow cell density at this stage. Bone: gray overlay, megakaryocytes: green overlay, mature red blood cells: red overlay, other marrow: cyan overlay, adipose tissue: white overlay, myeloid cell nuclei: orange overlay, and erythroid/lymphoid cell nucleus: blue overlay.
Once the marrow and bone were classified in the downsampled image, the images were analyzed at a high magnification (20×) tile by tile (Figure 1E and F), where classification of objects was further refined. At this magnification, the entire tissue was segmented using “multi-resolution segmentation” with scale parameter 85, which captured cohesive objects in the image such as individual nuclei. Bone was detected using evaluation classes based on the values from the low-magnification analysis. White spaces that were irregular in shape and large were classified as artifacts, while round white spaces were classified as adipose, and small spaces (vascular sinuses) were classified as part of the overall tissue area. Since H&E does not stain cell membranes, nuclei are easier to identify and count than entire cells, except for megakaryocytes where their large cytoplasm make them easier to identify as whole cells. Both nuclei and megakaryocyte cells were segmented using the Definiens filter “pixel min/max filter.” The total number of detected nuclei and megakaryocytes divided by the marrow area analyzed (which excluded bone and artifact but included adipose tissue and vascular sinuses) was reported as a measure of total cell density. See companion article (Kozlowski et al. In Press) for detailed description of how we classified myeloid nuclei and erythroid/lymphoid nuclei.
To apply the algorithm to dog, mouse, and monkey bone marrow images, minor modifications were made in the low-resolution analysis because under the H&E stain, the color of bone is different in these species from rat (Figure 7A, C, and E). We made no modifications to the algorithm for high-magnification analysis of bone marrow in these species. The species-specific modifications are included in the Definiens algorithm that will be shared upon request.
Automated analysis took approximately 40 min per slide on servers running Windows 7 Definiens Developer Server v2.6, but only ∼30 sec of manual operation by a technician loading the slides, confirming that the image was scanned properly, and initiating the computational analysis.
Statistical Analysis
MATLAB (MathWorks version R2016a, Natick, MA) software was used to perform all statistical analyses and graphing. In all studies, the control group of the study was compared to all treatment groups. The Wilcoxon rank-sum test was used to compare groups, since it is a general statistical test that may be applied to parametric (automated cell number computation) and nonparametric (the pathologist generated manual scores) data. When indicated in figures or text, we report the linear Pearson’s cross-correlation coefficient.
Results
Automated Measures of Bone Marrow Cellular Depletion Are More Sensitive Than Manual Scores
In order to test if the cell density measurements generated from the algorithm could improve upon manual scoring, we performed image analysis on a total of 17 studies, utilizing 565 Sprague-Dawley rats. In all these studies, H&E slides of sternal bone had been manually scored for bone marrow depletion. All studies except one were dose–response studies, and the exception was a time-course study where animals were terminated after different durations of dosing. The goal of all these studies was the identification of treatment groups that statistically differed from the control group, and estimation of the effect size, or the magnitude of the difference. We wished to evaluate if the algorithm could be used with at least equivalent or better performance compared to manual pathologist scores.
The algorithm computes the density of cells identified on the slide per area analyzed and reports an absolute quantitation. When pathologists manually score the extent of bone marrow depletion, they typically examine control samples, and then estimate the relative extent of bone marrow depletion in animals in the treatment groups, on a scoring scale of increasing severity of the finding. To make the automatic cell density measurements comparable to manual scores, we normalized the cell density measurements to the mean cell density of the animals in the control group, and computed the relative change in cell density for all animals (henceforth, these normalized cell density measures will be referred to as automated measures). We illustrate an example in one study (Kozlowski, et al. In Press), where rats were treated with varying doses of Monomethyl auristatin E (MMAE; Jununtula et al. 2008), and one group was treated with a single dose of a Bcl-xL inhibitor (Kile 2014). The raw cell density measurements inversely correlate with manual scores (Figure 2A), but when normalized, they trend in the same direction (Figure 2B), making them easier to compare.
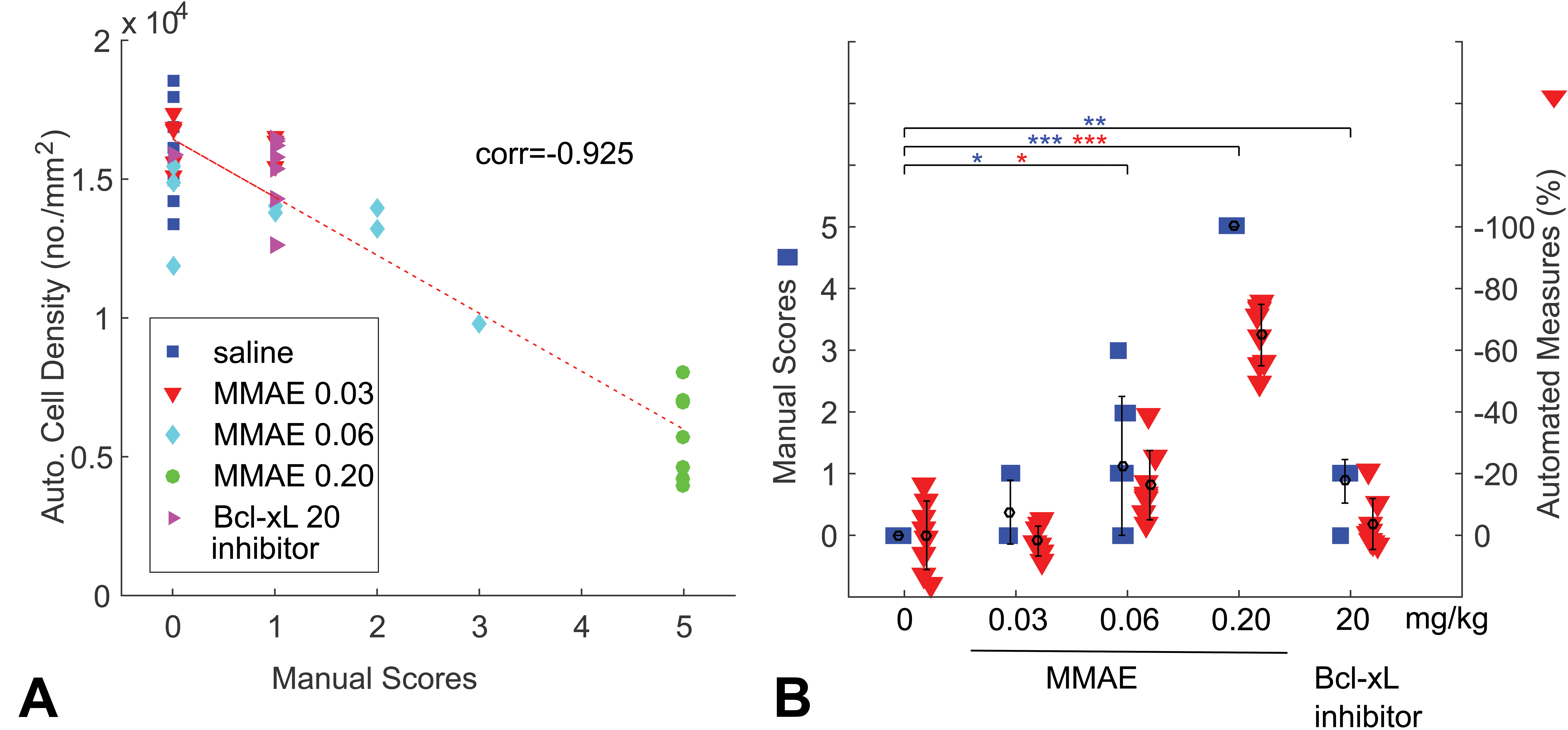
Comparison of manual scores and automated methods of sternal and femoral/tibial marrow images. (A) Absolute measure of overall cellular density is plotted against manual scores. The value “corr” indicates the Pearson’s correlation coefficient between the two scores. (B) The manual scores are plotted using the Y-axis on the left. The normalized image analysis data points or “automated measures” are plotted on the right, as the percentage of cell depletion relative to the mean of control animals. All treatment groups were compared to the control group for both methods, and asterisks (*) are placed when a statistical significant difference was detected using manual scores (blue) or automated analysis (red). No asterisks (*) between groups in figures indicates the differences are statistically insignificant (p ≥ .05). Otherwise, *p < .05, **p < .01, and ***p < .001. Automated methods and manual scores appear to have comparable ability to resolve differences from control in the test article (Monomethyl auristatin E) group. For Bcl-xL inhibitor, we cannot conclude whether manual scores or automated scores are more accurate since only one dose is present.
In order to select the statistical test appropriate for comparing the control group with treatment groups within each study, it was important to understand the distribution of the data. To this end, we plotted the distribution of the manual scores and automated measures from all animals from 17 studies (Figure 3A and B). While the discrete manual scores are in a left-skewed distribution, the continuous automated measures behaved more similarly to a Gaussian distribution. We determined from the analyses that while parametric statistical tests may be applied to compare differences between automated measures, nonparametric tests should be used to analyze the manual scores (Qualls, Pallin, and Schuur 2010). Although nonparametric tests are less powerful than parametric tests, the nonparametric test (the Wilcoxon rank-sum test) was applied for both manual scores and automated measures for further analysis to enable a fair comparison.
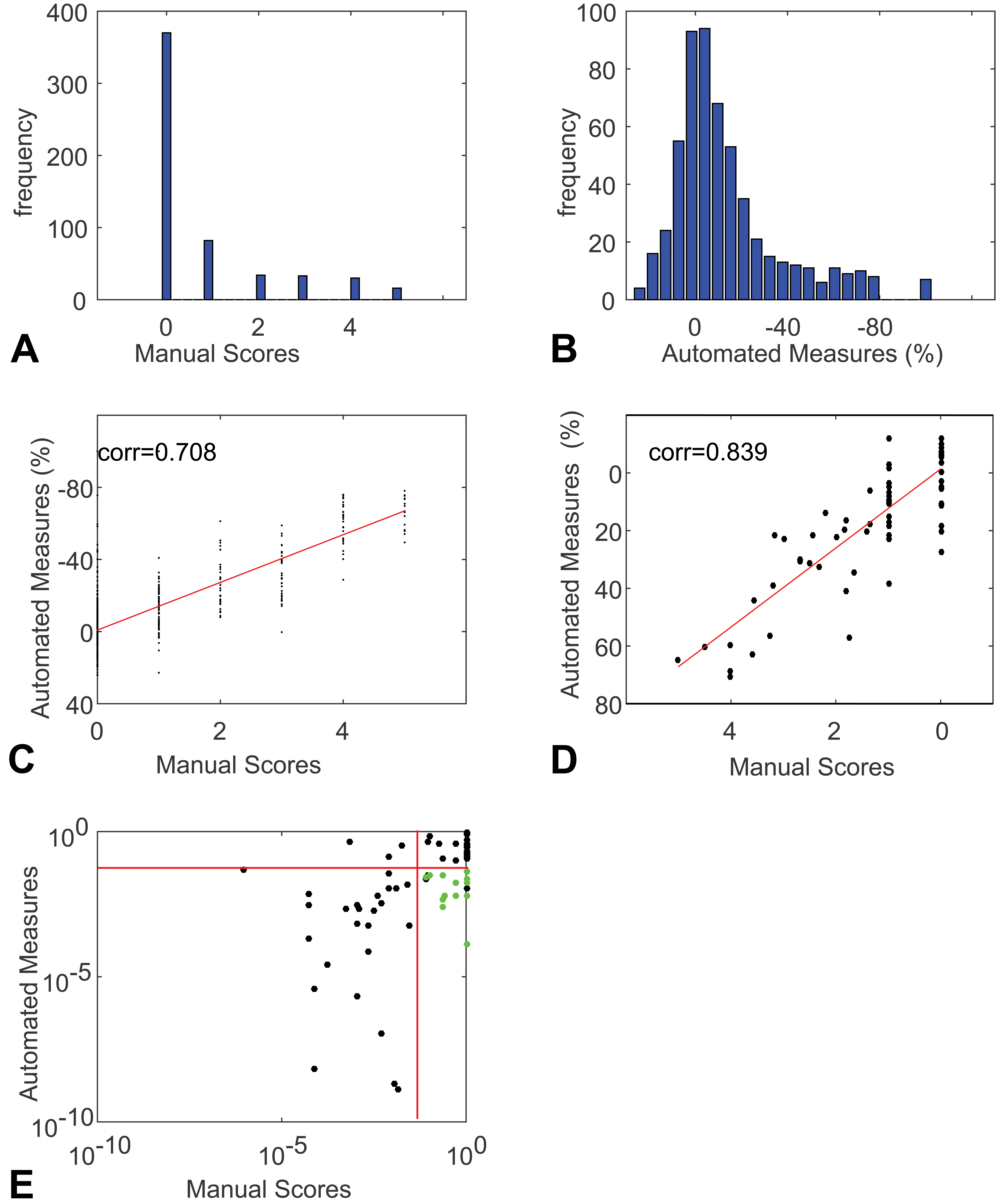
Meta-analysis of 17 rat studies indicates automated measures are more sensitive than manual scores. Histogram of scores of all animals in all studies by (A) manual scoring and (B) automated measures scaled relative to the control group in each study. The histogram in A reveals a markedly left-skewed, discrete distribution, while B resembles a continuous Gaussian distribution. It is visually apparent that the automated measures are more likely to meet the assumptions for a parametric statistical test, compared to manual scores. (C) Scatter plot of manual scores and automated measures in all animals. Red line indicates line of best linear fit. (D) Scatter plot of effect size: the difference in means between the control group and treatment groups for all studies, using manual scores or automated measures. Red line indicates line of best fit. The X- and Y-axes are plotted to show the groups with the greatest cell depletion toward the left bottom corner, and the groups most similar to controls on the right top corner of the graph, for clarity. (E) The scatter plot of p values of the results of the Wilcoxon rank-sum test between control group and treatment groups. The dots in green are the treatment groups where we have confirmed that the automated method identified a statistically significant difference, while the manual scores did not. Red lines indicate threshold of statistical significance (p < .05). All green dots are in the right bottom quadrant, an indication that all confirmed cases when scores disagreed corresponded to situations where the automated measures identified a group difference where the manual scores did not.
Despite the differences in the overall distribution of data, there was a linear correlation of .708 between the manual scores and automated measures for all animals across all studies (Figure 3C). To determine if any of the disconcordance on an individual animal level affected the interpretation of the study results, we compared the “effect size” of all treatment groups, which is the magnitude of difference between the means of each treatment group with its corresponding control. We found that the linear correlation between the computed effect sizes was .839, indicating a high degree of concordance between the two methods in assessing magnitude of treatment effect (Figure 3D).
We then compared the p values resulting from the Wilcoxon rank-sum test between the control group and each treatment group using both methods. Overall, automatic measures identified greater statistically significant differences (i.e., smaller p values) between each treatment group and corresponding control group in a study, compared to manual scores (Figure 3E). Setting p ≤ .05 as the threshold for significance, there were 16 cases where the automated measures produced a statistically significant difference but manual scores did not, and 3 cases with the opposite situation. The rest were concordant: in 21 cases, the treatment groups were statistically significantly different from their corresponding control, both by automated measures and manual scores, and in 26 cases, neither method identified a difference between the treatment group and control.
In order to understand which method was more sensitive, we reasoned as follows: if the manual scores and the automatic measures did not concur, we looked to see if in this same study, there was a treatment group with the same compound, but at a higher dose. If both methods identified the higher dose treatment to be significantly different from control, it was more likely that the mid- or low-dose effect was “real”, even if only one method identified it. For the 16 cases where a significant change was identified by the automated measures but not by manual scores, we found 10 cases in 8 studies (these 8 studies are shown in Figure 4) that we could confirm a higher dose with an effect identified by both methods (the p values for these cases are colored green in Figure 4E). There was no confirmed instance of the opposite situation. From these data, we reasoned that automated measures are more accurate at predicting dose-dependent changes compared to manual scores.
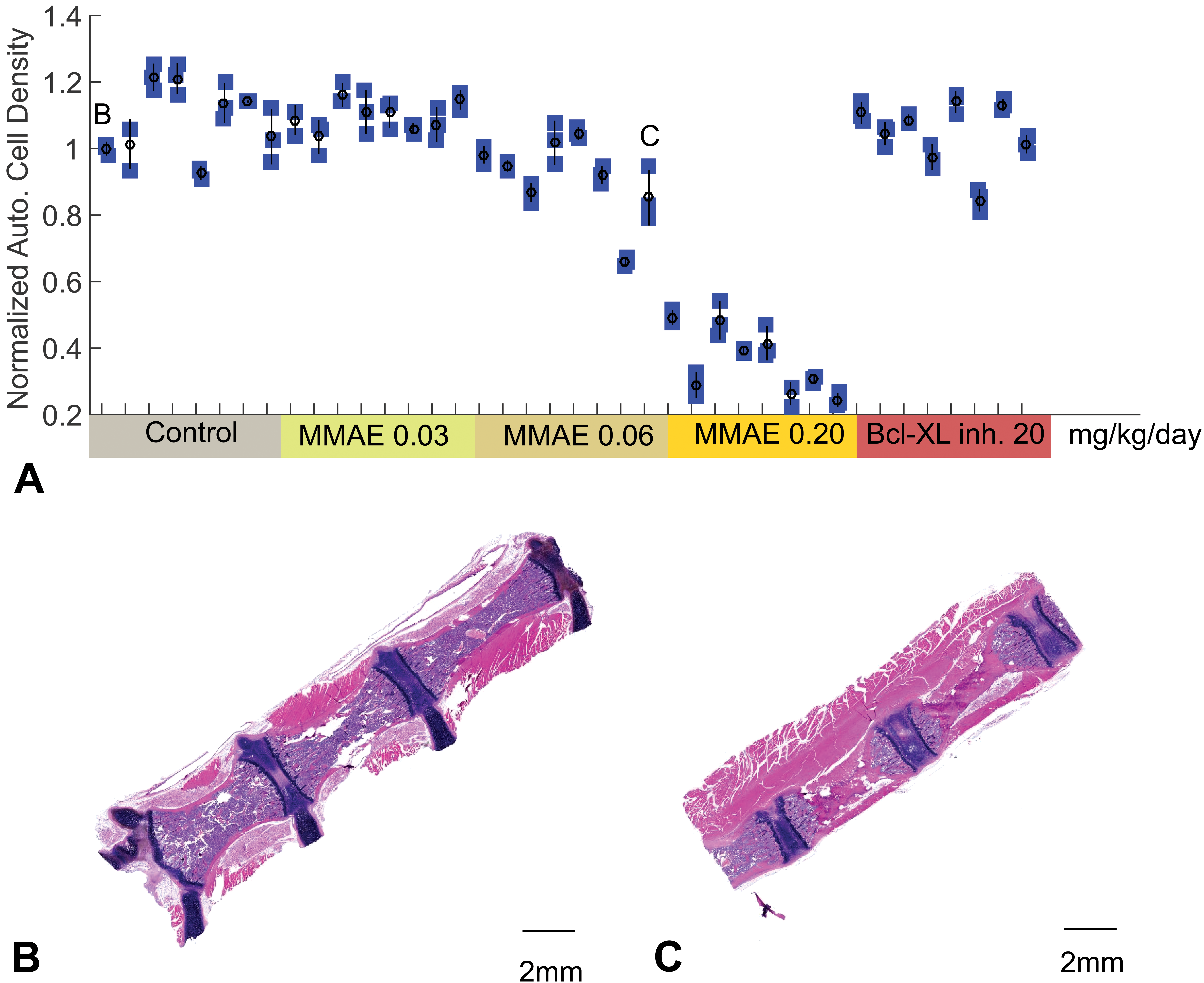
Image analysis reproducibility analysis. (A) For all 40 rats in the study, three serial sections were analyzed for each animal by the image analysis algorithm; and the results were plotted normalized to the first animal. The low-magnification hematoxylin and eosin image of an animal where there was low intersection variation (B) and where there was high variation (C) are also shown. Note that there is far less analyzable marrow area in C compared to B, which is the most likely cause of variability in the analysis of serial sections of C.
Image Analysis Is Reproducible in Serial Sections
In order to measure the reproducibility of the image analysis method in the same animal, we analyzed three serial sternal sections stained on different H&E runs, acquired from every animal in the study presented in Figure 2A and B. We normalized the cell density values computed for each slide relative to the mean value of the first control animal in the study. For all 35 animals that were used in this study, the average coefficient of variation of automated cell density was 4.4 ± 3.3%. Visual examination of sections where variation between serial sections was low (Figure 5B) and high (Figure 5C) revealed that in general, high variation in cell density estimation between serial sections was the result of smaller amounts of marrow tissue available for analysis.
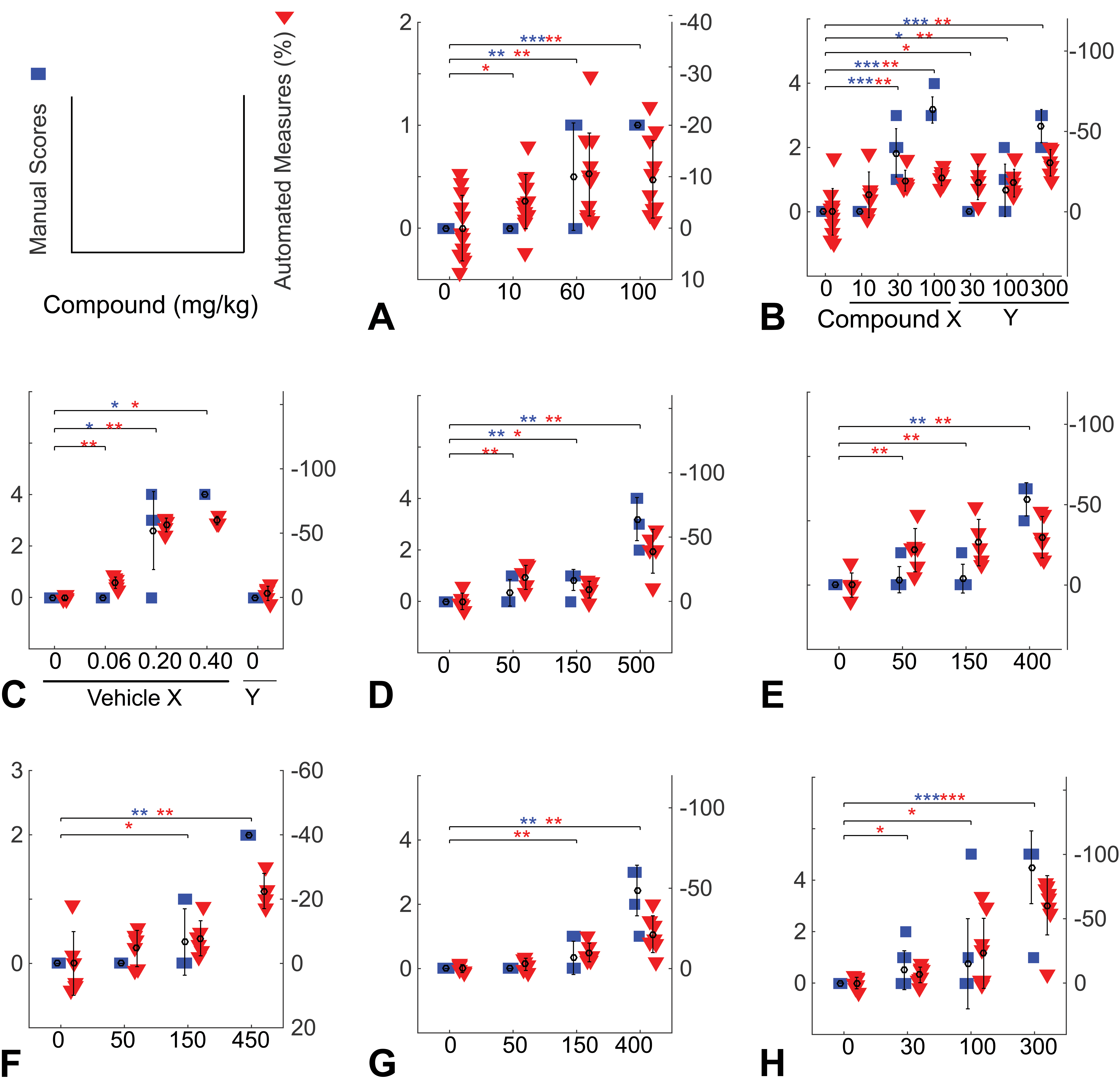
Studies where image analysis detected a change at higher sensitivity than manual scores. Manual scores (left axis) and automated measures (right axis) are plotted for eight rat studies. The comparison between a treatment group and control has a blue asterisk(s) (*) for manual scores, red asterisk(s) for automated measures. No asterisk (*) between groups in figures indicates the differences are statistically insignificant (p ≥ .05). Otherwise, *p < .05, **p < .01, and ***p < .001. In each these studies, there is at least one dose group where the automated measures, but not manual scores, showed a statistically significant difference to control (red asterisk(s) present but no blue stars). Furthermore, a higher dose existed for the same compound, where both methods identified a difference from control (both blue asterisk(s) and red asterisk(s) present). Compound names are anonymized, but most studies were simple dose-finding studies, where the same compound was given at multiple doses. However, study B investigated 2 different studies, while study C contained an extra group that was used to test a different vehicle from the other groups.
Image Analysis Is Comparable in Sternum and Femur/Tibia
We examined if similar results could be obtained using femoral/tibial (Figure 6A) and sternal (Figure 1A) bone marrow. The correlation between manual scores and automated measures obtained from sternum (.925) was comparable to that between manual scores and automated measures obtained from femur/tibia (.919). The total cell densities predicted by the algorithm on femur/tibia and sternum had a correlation of .938 (Figure 6B); however, the analysis in sternum, but not in the femur/tibia, detected a statistically significant difference between control and .06 mg/kg doses of MMAE (Figure 6C).
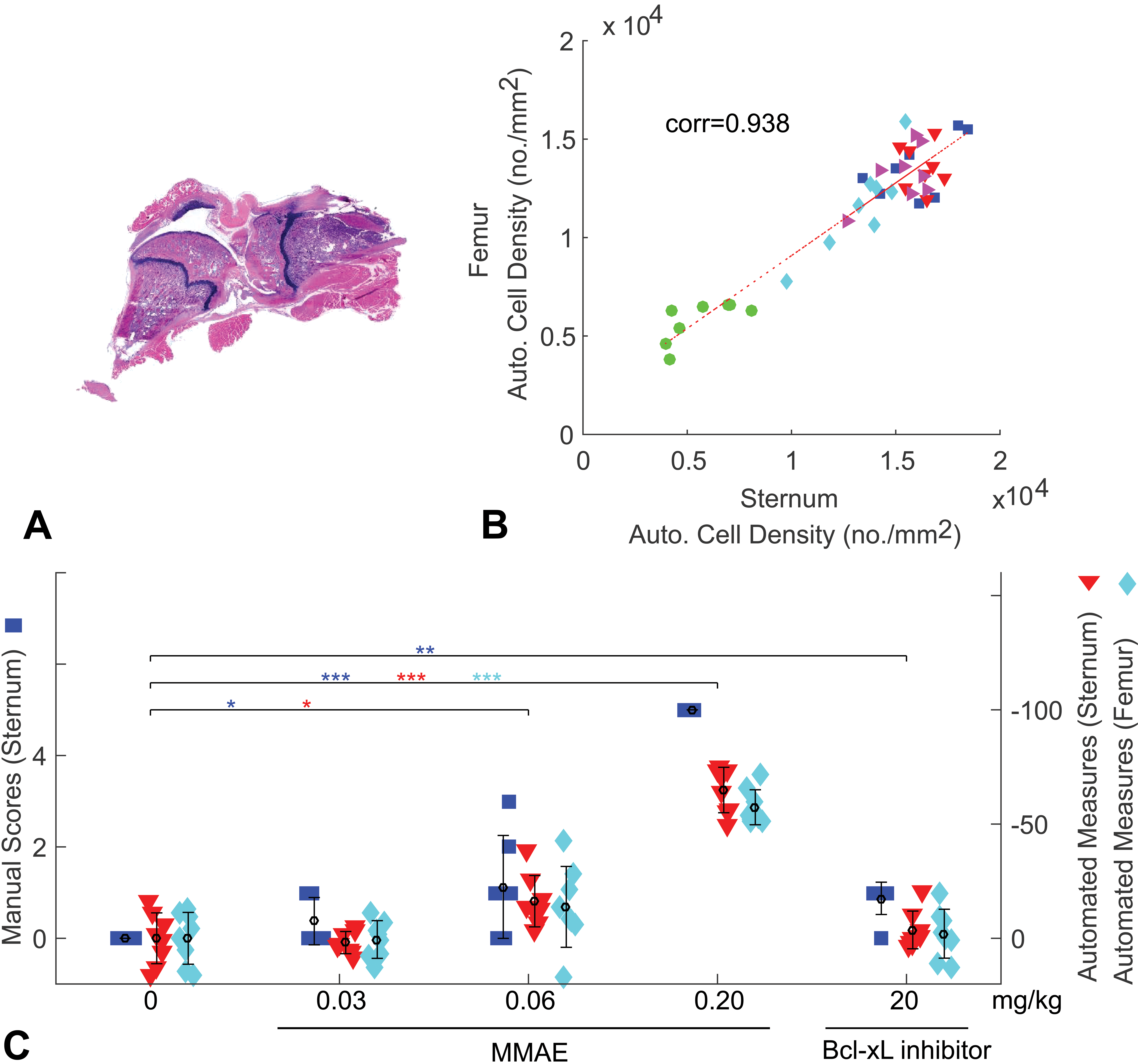
Comparison between sternal and femoral/tibial bone marrow. (A) Hematoxylin and eosin image of a femoral/tibial bone marrow section from rat shown here to contrast with sternal bone preparation shown in Figure 1A. (B) Scatter plot between absolute cell density measurements from femoral/tibial and sternal bone marrow from the same animals. (C) Comparison of manual scores (sternum) and automated measures for femoral/tibial and sternal bone marrow. The comparison between a treatment group and control has a blue asterisk(s) (*) for manual scores (sternum), red asterisk(s) for automated measures on sternum, cyan asterisk (s) for automated measures on femoral/tibial bone. No asterisk (*) between groups in a particular color indicates the differences are statistically insignificant (p ≥ .05) using that method. Otherwise, *p < .05, **p < .01, and ***p < .001. Note that for the Monomethyl auristatin E (.06 mg/kg) group, there is no significant difference with control detected using the automated scores based on femur/tibia, but the difference is significant using manual scores and automated scores based on sternum. This suggests that the femoral/tibial bone preparation may have not had enough tissue for analysis compared to the sternal bone preparation.
Image Analysis May be Applied to Multiple Species
The image analysis algorithm was then applied to scanned images of mouse, dog, and cynomolgus monkey sternal bone marrow slides. Although the sizes of these bone marrow images were quite different from each other, the algorithm required only minor adjustment for it to be adequately applied to different species. For dog (Figure 7A and B) and mouse (Figure 7C and D), both the manual scores and automated measures identified the highest dose group of a compound to be a dose significantly different from control. For monkey (Figure 7E and F), while no groups were statistically different from control by either method, all three dose groups had evidence of a recovery response (identified by reduced cell depletion), between the terminal and recovery animals, by both automated measures and manual scores.
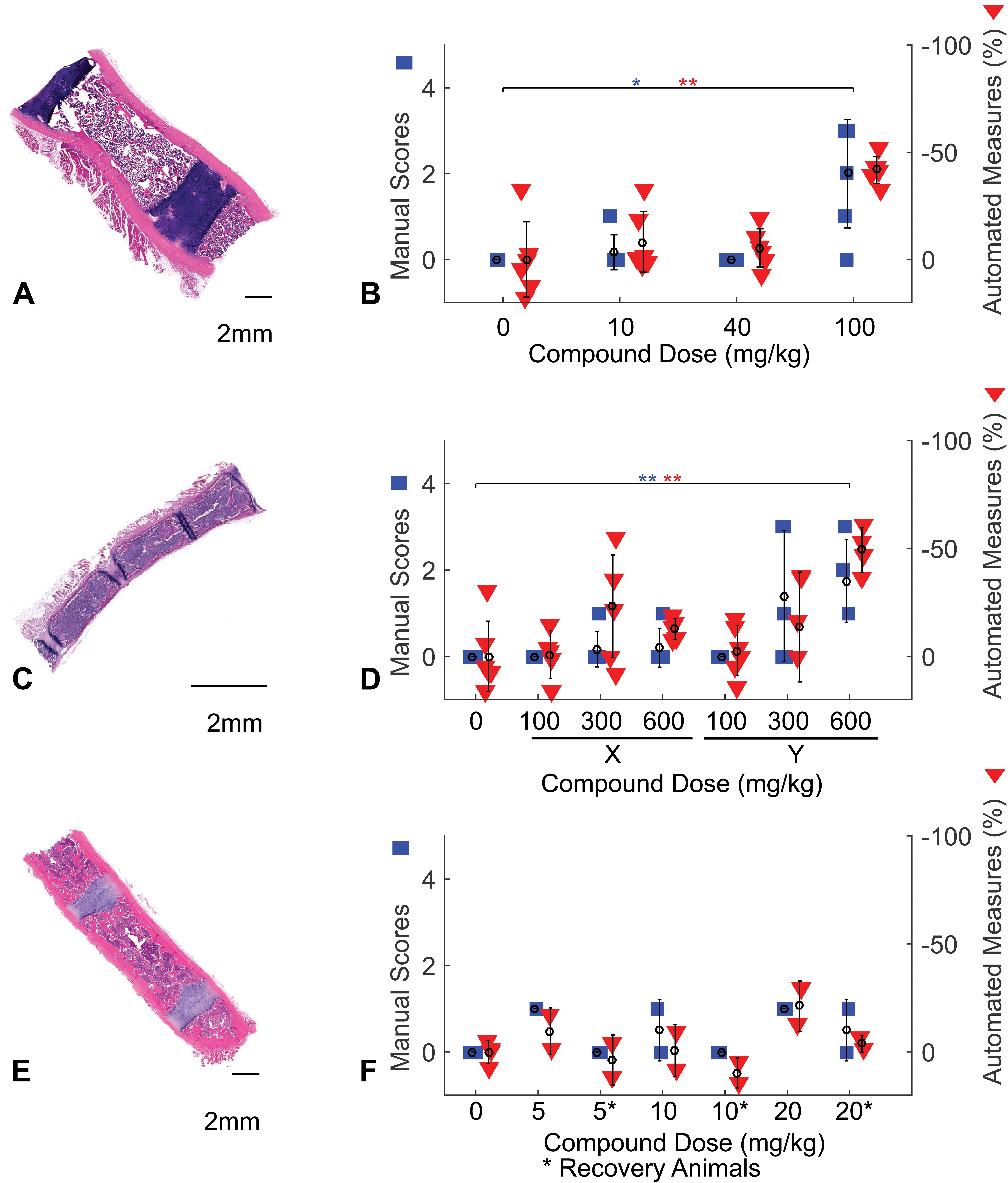
Image analysis algorithm applied to mouse, dog, and monkey. On the left panel, low-magnification hematoxylin and eosin figures from a control animal are presented for bone marrow toxicity studies that were performed either with dog (A), mouse (C), or cynomolgus monkey (E). Scale bars are placed to emphasize large differences in image size. Note also the different coloring of the encapsulating bone. Despite the differences, the image analysis algorithm originally designed for rat appears to have performed correctly. On the right panel, manual scores from the corresponding study are plotted together with automated measures, in dog (B), mouse (D), and monkey (F). In study B, multiple doses of the same compound were tested, whereas for D, two compounds with multiple doses each were tested. In F, animal groups were given varying doses of a compound and split into terminal and recovery groups. The significant difference detected between the control group and treatment group is illustrated with a blue asterisk (*) when the manual scores were used, and a red asterisk when automated measures were used. No asterisk (*) between groups in figures indicates the differences are statistically insignificant (p ≥ .05). Otherwise, *p < .05, **p < .01, and ***p < .001. There is an alignment between manual scores and automated measures in all studies.
Discussions
We have developed a novel image analysis method that allows rapid screening for bone marrow toxicity, using scanned images of bone marrow slides from preclinical safety models. We demonstrate that standard 20× images of H&E sections, which are readily obtained and quickly scanned, may be utilized to extract more quantitative information compared to standard microscopic examination utilizing automated digital image analysis. The image analysis methodology performed consistently in analysis of serial sections, can be applied to femoral/tibial as well as sternal bone marrow samples, and is applicable in multiple species including rat, mouse, dog, and monkey.
Manual microscopic scoring is labor-intensive and susceptible to inter- and intraoperator variability (Aeffner et al. 2017). Unlike manual assessment, our image analysis method is automated and reproducible. By performing a meta-analysis of multiple rat bone marrow dose-finding studies, we demonstrated that the image analysis method was able to resolve group differences at lower doses compared to manual scoring. Here, we used a nonparametric test to compare both methods fairly, but since the automated measures are continuous and close to a normal distribution, for future use it is appropriate to use a powerful parametric test such as the Student’s t-test to compare groups analyzed automatically.
Although we use manual assessment as a gold standard for evaluating bone marrow cell depletion, the image analysis method differs in one important aspect: it does not attempt to establish a range of “normal” cellularity. While a pathologist designates a score of “0” for all animals that are within this range, the algorithm simply enumerates the cell density. This contributes to the statistical power of the automated measures, since parametric tests such as the Student’s t-test cannot be applied when all values for animals in the control group are 0 (no variance), as was the case for manual scores in all our studies. We normalize and compare all treatment groups relative to the control group, with the assumption that a large departure from control is a likely flag for bone marrow toxicity. The biological relevance of this depletion must be interpreted by the pathologist on the study.
Our method may be further refined in the future by using slides scanned and analyzed at 40×. Currently, the slide scanning and computational time required for analysis at this magnification are prohibitive for high-throughput analysis, but our existing framework would allow us to take advantage of technological improvement in these areas.
To our knowledge, this is the first instance of an algorithm designed to enumerate bone marrow cells in situ at relatively low magnification (20×). Several published image analysis algorithms have been described that segment images of human blood cells in smears or peripheral blood, using shape and color information (MoradiAmin et al. 2016; Reta et al. 2015; Arslan, Ozyurek, and Gunduz-Demir 2014) at high magnification (100×). But in our case, because the marrow cells are still encapsulated in bone, a major challenge was to accurately exclude non-marrow tissue by using shape features of objects identified on a lower magnification image (0.4×). This process can be sensitive to sample preparation artifacts. Some artifacts arise during the decalcification of bone (Reagan et al. 2011), leading to tissue tearing and shrinkage, which create empty spaces within the bone marrow. Empty bone marrow spaces can be confounding as well, when bone marrow is pharmaceutically depleted. With drug-induced marrow depletion, marrow sinuses start to empty due to hypocellularity (Haschek, Rousseaux, and Wallig 2009). When tissue shrinkage causes artificially large empty spaces in the marrow area, or when poorly fixed tissue falls out of the sample, it may cause an underestimation of cell density in these areas. The algorithm was designed to correct for these effects to an extent. Circular objects were classified as adipose tissue, whereas large, angular tear-like structures were considered fixation artifacts, and excluded from analysis. Yet, if there is complete loss of tissue integrity in the imaged slide, the analysis is unlikely to be accurate. Another source of error is small evaluable sample area. Analysis in femoral/tibial samples appeared slightly less accurate compared to sternum, most likely due to the presence of bone spicules and a larger growth plate area in this particular preparation, which restricted the amount of tissue available for image analysis. Analysis of bone marrow serial sections with unusually low marrow tissue area led to higher variability. This is due to a low but consistent level of errors in cell classification, which become relatively more significant with smaller sample sizes. For example, if only the tip of a megakaryocyte cell has been captured in an image section, it may be excluded from analysis (leading to underestimation of cell number) or misclassified as multiple myeloid cells (leading to an overestimation). For these reasons, before performing image analysis, it is important to visually assess the scanned tissue section quality and tissue availability visually, as is recommended for any whole-slide imaging effort (Aeffner et al. 2016).
Despite these known sources of sample preparation–related errors, most studies that we examined from our archival samples were acceptable for accurate image analysis. Since these samples were prepared in different labs over 9 years, it provides evidence that the algorithm is fairly robust to sample preparation variation. In summary, our method proved to be repeatable across different H&E stain runs and allows more accurate determination of bone marrow toxicity in dose-range finding studies, compared to standard, manual scoring methodology.
Footnotes
Acknowledgments
We would like to thank Jacqueline Tarrant for helpful comments and suggestions on the article.
Author Contributions
Authors contributed to conception or design (CK, GC); data acquisition, analysis, or interpretation (CK, JB, GC); drafting the manuscript (CK); and critically revising the manuscript (CK, JB, GC). All authors gave final approval, and agreed to be accountable for all aspects of work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Declaration of Conflicting Interests
All authors are employers and stockholders of Genentech Inc.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The work was fully funded and supported by Genentech Inc.