Abstract
Recent experimental evidence indicates significant pulmonary toxicity of multiwalled carbon nanotubes (MWCNTs), such as inflammation, interstitial fibrosis, granuloma formation, and carcinogenicity. Although numerous studies explored the adverse potential of various CNTs, their comparability is often limited. This is due to differences in administered dose, physicochemical characteristics, exposure methods, and end points monitored. Here, we addressed the problem through sparse classification method, a supervised machine learning approach that can reduce the noise contained in redundant variables for discriminating among MWCNT-exposed and MWCNT-unexposed groups. A panel of proteins measured from bronchoalveolar lavage fluid (BAL) samples was used to predict exposure to various MWCNT and determine markers that are attributable to MWCNT exposure and toxicity in mice. Using sparse support vector machine–based classification technique, we identified a small subset of proteins clearly distinguishing each exposure. Macrophage-derived chemokine (MDC/CCL22), in particular, was associated with various MWCNT exposures and was independent of exposure method employed, that is, oropharyngeal aspiration versus inhalation exposure. Sustained expression of some of the selected protein markers identified also suggests their potential role in MWCNT-induced toxicity and proposes hypotheses for future mechanistic studies. Such approaches can be used more broadly for nanomaterial risk profiling studies to evaluate decisions related to dose/time–response relationships that could delineate experimental variables from exposure markers.
Keywords
With more than 1,000 registered products incorporated with engineered nanoparticles (NPs) to date, nanotechnology-enabled applications have revolutionized many aspects of our lives. Incorporation of engineered NPs into various industrial (e.g., electronics, manufacturing, construction), consumer (e.g., cosmetics, food packaging), and biomedical products may pose an increased exposure risk to humans. It has been estimated that ∼4 million workers worldwide, with ∼2 million in the United States, could be potentially exposed to nanomaterials (Roco 2011). Human exposure to NPs can happen through different ways, including in products that are for intended human use (e.g. diagnosis and treatment of diseases) and those that may be incidental during manufacturing and in other occupational or environmental settings. The human health effects associated with exposure to NPs are, to a large extent unknown. To date, there are no reports of any definitive human disease that can be phenotypically attributed to NP exposure. Thus, there is an urgent need for identification of candidate biological markers useful for assessing NP exposures and early biological effects or responses associated with adverse health outcomes in humans.
Short-term and subchronic exposure studies in rodents have indicated adverse pulmonary outcomes such as inflammation, formation of granulomas, and fibrosis associated with carbon nanotube (CNT) exposures. Despite the many existing studies exploring pulmonary exposures to CNT, development of biological markers that can detect NP exposure and biological effects, particularly focusing on mammalian exposures and toxicity, is almost nonexistent. This is due to the lack of methodological consistency and relatively large number of variables between experiments adopted by different groups as well as the type of CNTs employed by each group. The differences in administered dose, physicochemical characteristics (e.g., agglomeration/aggregation state, metal impurities, stiffness, length, aspect ratio) of the CNTs studied, dispersion methods including vehicle, and exposure methods employed in each study, further complicate the process of finding reliable biological markers across different studies (Wick et al. 2007, 2011; Johnston et al. 2010). Therefore, the current efforts toward monitoring NP exposure in humans suggest use of a panel of biomarkers of inflammation, oxidative stress, systemic effects, and/or disease (e.g., fibrosis, cancer), rather than those related to a particular pathway or mechanism. This is because the biological and pathological responses to different NPs depend greatly on the material characteristics as well as its biological persistence in the target organ and tissue. Considering that it is almost impractical to test the potential biological effects in vivo for all the anticipated large number of nanoscale products, as an initial step, it might be useful to select biomarkers that share a common pathway or mechanism of toxicity or biological response across different NPs (e.g., fibrosis, inflammation).
One approach for obtaining such panel of select biomarkers is the application of a supervised machine learning approaches, which analyzes the labeled training data and produces an inferred function with selected features that can be used for predicting new exposure data. Supervised learning is composed of 2 categories of algorithms: (a) classification and (b) regression. Importantly, classification, an example of pattern recognition, deals with categorical response values where data are separated into defined classes (e.g., “disease” vs. “healthy,” “toxic” vs. “nontoxic,” “exposed” vs. “unexposed,” “spam” and vs. “nonspam”). A variety of supervised learning classification algorithms, including artificial neural networks, Bayesian networks, support vector machines (SVMs), and decision trees, have been widely used in biological applications, such as disease modeling and cancer research (Vapnik 1982; Chen et al. 2011; Cortes and Vapnik 1995; Sundaramurthy and Eghbalnia 2015; Vidyasagar 2014; Guo and Wan 2014). Most of these studies involving “-omics” or high-content analysis used some variant of the SVM to identify the biomarkers. SVM-based methods are more robust and have the ability to identify a small subset of highly predictive markers (Maulik and Chakraborty 2014; Huang et al. 2013; Bevilacqua et al. 2012; Duan et al. 2005; Guyon et al. 2002; Misganaw et al. 2015; Ahsen et al. 2017). In this article, we selected and identified protein markers of exposed and/or early responses of a class of carbonaceous NPs, that is, multiwalled carbon nanotubes (MWCNTs). A panel of proteins measured in bronchoalveolar lavage fluid (BAL) collected at various postexposure time points (0, 24 hr, 3 days, 28 days, 56 days, and 84 days) and exposure concentrations from mice exposed to 2 different pristine as-produced (AP) MWCNT, their polymer-coated (PC) counterparts, or a well-studied reference material, MWCNT-7, were analyzed. Analysis of NPs with different physicochemical characteristics studied under various experimental conditions is critical for enabling discovery of convergent/common and divergent markers of exposure to NPs. A sparse classification algorithm that implements a combined l 1- and l 2-norm SVM t-test with recursive feature elimination (RFE), in short “lone-star” (Ahsen et al. 2012, 2017; Vidyasagar 2014), was applied to select optimal protein markers that can predict exposure or biological effects of MWCNT (Figure 1). The sparse classification algorithm lone-star implements various optimization methods to overcome some of the issues inherent to nanotoxicity modeling studies, such as dealing with large number of variables, noisy/inseparable data sets, unequal distribution of sample sizes between classes, and unknown relationships between different experimental conditions. Thus, this approach was chosen to take advantage of the power of combined l 1- and l 2-norm SVM’s ability to select a small set of relevant features and ignore redundant features that distinguish between MWCNT-exposed and MWCNT-unexposed animals across studies employing different types of MWCNT and exposure conditions (Figure 1). Another important advantage of the sparse feature selection-based classification approach compared to traditional SVM is its ability to both achieve the classification goal and selection of a small set of correlated features (i.e., # of predictive features ≤ # of samples) simultaneously (Bradley and Mangasarian 1998). Once identified/validated, a selective predictive marker panel could serve as a valuable initial screen to guide detection of exposure to MWCNT. The methodology described in this study can be extended to future screening strategies for hazard identification of other emerging engineered nanomaterials.
Schematic representation of the overall objective and study design.
Method
Study Design, Approach, and Data Collection
The data used were collected from studies conducted at the National Institute for Occupational Safety and Health (NIOSH). All studies referenced in the article were conducted in the Association for Assessment and Accreditation of Laboratory Animal Care International–accredited NIOSH animal facility in accordance with the NIOSH Institutional Animal Care and Use Committee. The various training data sets employed in this study consisted of 2 types of pristine AP MWCNT and their PC counterparts from 2 separate companies. A well-characterized reference material, Mitsui-7 (MWCNT-7), was also evaluated. A panel of ∼51 to 54 proteins were measured in the BAL collected from mice exposed by oropharyngeal aspiration to 4 and 40 µg of each particle. The 4 and 40 µg were based on previous human dose extrapolations and known in vivo outcomes. The 40 µg dose was chosen because it is a dose known to cause pathology and offers historical reference for toxicology outcomes. The 4 µg dose offers an order of magnitude lower dose and is closer to the potential cumulative exposure of individuals working in CNT facilities at an average workplace exposure of the 10.6 µg/m3 of inhalable elemental carbon (Erdely et al. 2013). An additional set of mice were exposed to MWCNT-7 by inhalation at 0.5 and 5 mg/m3 for 5 hr/days for 19 days. The details of the exposures and various postexposure time points analyzed can be found in Table 1. General physicochemical characteristics of the materials can be found in Table S1. The lungs from a separate set of mice were referenced for pathological outcomes at 84-day postexposure to each type of MWCNT investigated as part of this study. The detailed quantification of various pulmonary pathological outcomes using morphometry techniques and/or severity-based scoring/grading is and/or will be provided as part of the main pulmonary toxicity assessment studies (Bishop et al. 2017; Kodali et al. 2016; Roberts et al. 2016), and only a qualitative analysis describing pulmonary pathology is provided as part of this study. For all the referenced histopathology, at sacrifice, lungs were inflated with buffered formalin, processed into paraffin blocks, sectioned at 5 microns, and stained with hematoxylin and eosin (H&E) for routing histopathology or picrosirius red (PSR) for fibrosis. H&E-stained slides were examined by a veterinary pathologist using bright field microscopy for evaluating morphologic changes and PSR-stained slides by polarizing light microscopy for evaluating collagen. Photomicrographs were captured using an Olympus BX53 microscope equipped with a DP73 camera.
Details of the Training and Test Data Sets for Building the Classification Models of Exposure to MWCNTs.
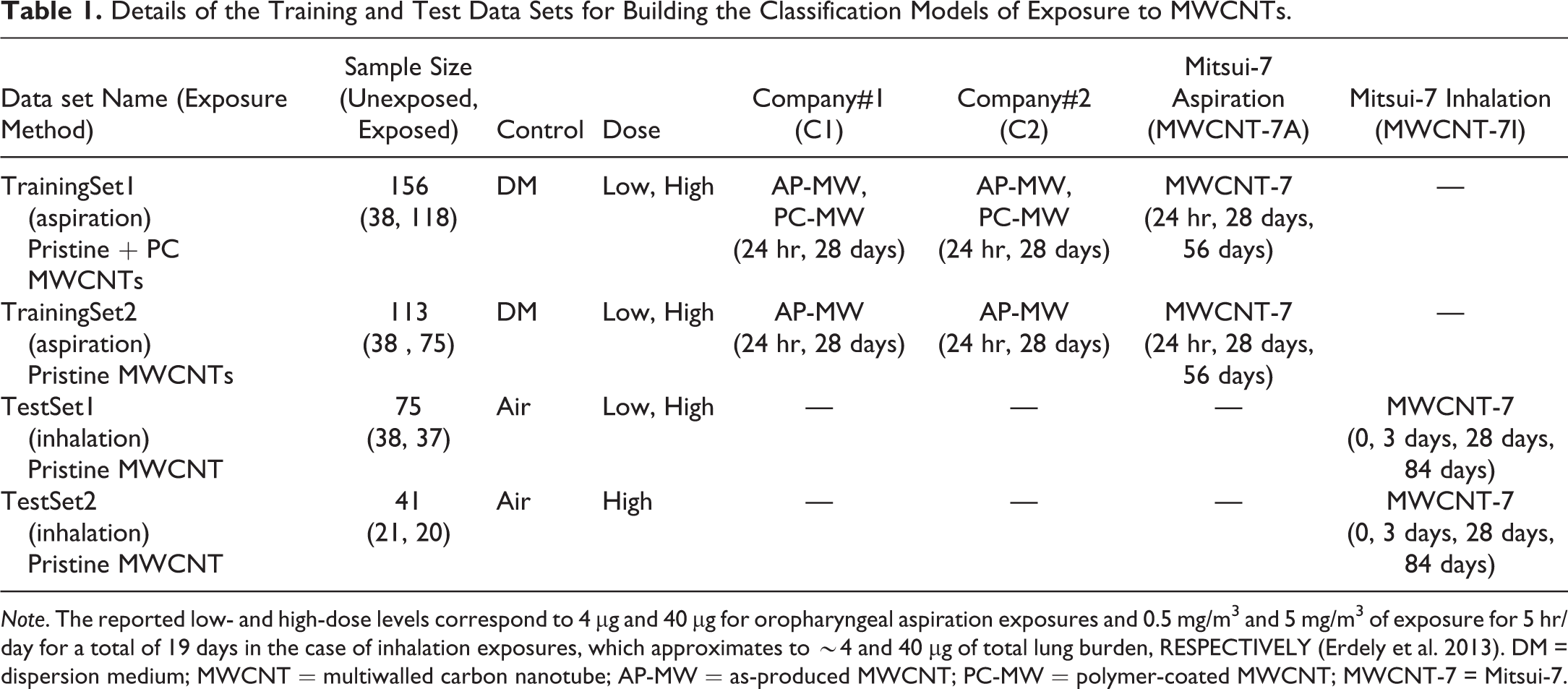
Note. The reported low- and high-dose levels correspond to 4 µg and 40 µg for oropharyngeal aspiration exposures and 0.5 mg/m3 and 5 mg/m3 of exposure for 5 hr/day for a total of 19 days in the case of inhalation exposures, which approximates to ∼4 and 40 µg of total lung burden, RESPECTIVELY (Erdely et al. 2013). DM = dispersion medium; MWCNT = multiwalled carbon nanotube; AP-MW = as-produced MWCNT; PC-MW = polymer-coated MWCNT; MWCNT-7 = Mitsui-7.
Data Prefiltering/Processing
Due to differences in the detection limits and the selection of proteins among different RodentMAP® versions across the chosen studies, all data were prefiltered and normalized to select proteins that were commonly represented. As a first step, the proteins with values below the limit of detection (LOD) or “quantity not sufficient” to analyze in all corresponding samples for each study were removed. Further, samples with no corresponding measurements across all studies considered were removed from each data set. Following this, proteins with values less than the LOD for only certain samples, but not all samples in the study, were imputed by replacing with ½ the lower limit of quantification values or lower detectable dose, depending on the version of RodentMAP. This resulted in a total of 46 proteins that were commonly analyzed across all chosen studies. The detailed comparison of the set of proteins analyzed in each study and the selected list of proteins along with those that were removed (highlighted in red font) is provided in File S1. Similarly, analysis of only the MWCNT-7 particles between oropharyngeal aspiration and inhalation studies resulted in the selection of 51 common protein features across the 2 studies (File S2). Such prefiltered data sets for each study was further normalized to convert raw values to fold-change values compared to their corresponding controls at each time point and/or concentration investigated. Thus, normalized data were combined to form input to the algorithm. This included a set of explanatory variables, a measure of the abundance (in this case, fold-change) of proteins in the lung lavage fluid, and the response variables as class labels, a value of either “1” or “2” corresponding to the exposure treatment. In this study, the MWCNT-exposed samples were labeled 2 or negative class examples, and samples in the control group were assigned class 1 or “positive” class.
Sparse Supervised Machine Learning-based Classification of MWCNT-exposed and Control Mice
To detect or identify potential feature sets (in our case, proteins) in lung lavage fluid that can discriminate between MWCNT-exposed and control groups, we applied a sparse classification algorithm, referred to as “lone-star” developed recently (Ahsen et al. 2012, 2017). The details of the “lone-star” or “l 1-, l 2-norm SVM t-test with RFE” algorithm have been described previously and were successfully applied to several cancer-related problems (Ahsen et al. 2017; Misganaw et al. 2015; Vidyasagar 2014). The selection of lone-star algorithm for this study was motivated by several advantages that are inherent to SVM-based classification models and the other characteristics of the lone-star algorithm implementation such as its ability to deal with noisy/sparse data and robustness to variations in experimental conditions as well as those with unequal distribution of samples between classes. There are several considerations when dealing with modeling toxicity data from NP studies: (a) nonlinearly separable and/or noisy data depending on the dose/time points (e.g., responses at earlier time points can be different from later times), (b) unknown relationships between different doses/time points, (c) different ratios of samples in each class making it difficult to build a general set of parameters, and (d) large number of features compared to samples in some cases. The lone-star algorithm implements several machine-learning ideas to overcome these problems and to identify a small number of highly predictive features. These include (a) combined l 1-, l 2-norm SVM (Bradley and Mangasarian 1998)—which guarantees a sparse solution and allows robust selection of correlated features, (b) RFE (Guyon et al. 2002)—to reduce the size of feature sets, (c) stability selection (Meinshausen and Bühlmann 2010)—to ensure the selected feature set is relatively insensitive to noise, and (d) implements slack variables to penalize false positives and false negatives differently (Veropoulos, Campbell, and Cristianini 1999).
For each of the situations described in Tables 2 to 4, the lone-star algorithm was used to develop a binary classifier to identify a set of highly predictive protein markers that could be used to distinguish between the different conditions explored in each case (e.g., exposed vs. unexposed, AP-MW vs. PC-MW, C1 vs. C2). A flow chart briefly describing the various steps of the algorithm is shown in Figure 2. A detailed description along with mathematical derivation has been reported previously (Ahsen et al. 2012, 2017). The following section is intended to only provide a brief general description of various steps of the algorithm. The initial step of the algorithm involves a preprocessing step to only retain features (in our case, proteins) that show a statistically significant difference in mean fold-change values between the 2 classes. This is accomplished using the “student t-test.” Following this, a set of k 1 samples from class 1 and k 2 from class 2 were randomly chosen as a training set to compute an optimal w using the l 1-norm SVM. In our case, the size of each training set chosen at this step corresponded to half the total samples within each class. This was repeated N (default = 80) times with different randomized samples to select features that did not vary much across random splits. In the next step, all the optimal weight vectors from across the N randomized runs were averaged to choose “k” largest entries and the corresponding feature set (in this case, proteins). This further leads to the reduction in the original n significant features to m features (Figure 2). The process was repeated several times with several randomly split training sets, but with a reduced feature set at each stage, RFE, until no reduction in the number of features was obtained. This resulted in a final set of protein markers to be used for classification. Once selected, a 2-fold cross-validation was carried out by running l 1-norm SVM, using N randomly split samples from the training data set and assessing the performance of each of the resulting classifiers on the remainder of the samples at each split. The top X (e.g., 20) best performing classifier models were further averaged to compute the final overall classifier. The generated final classifier was run on the entire training data set to estimate the accuracy, sensitivity, and specificity values. For the evaluation measures employed, sensitivity corresponds to the proportion of true positives that were correctly observed by the classifier, sensitivity was the proportion of true negatives that were correctly identified, and accuracy, often used for assessing the overall performance of a classifier, estimated the percentage of correctly classified samples.
Predictive Common Markers Selected by the Trained Models for Optimal Classification of the MWCNT-exposed Group from Unexposed Groups.
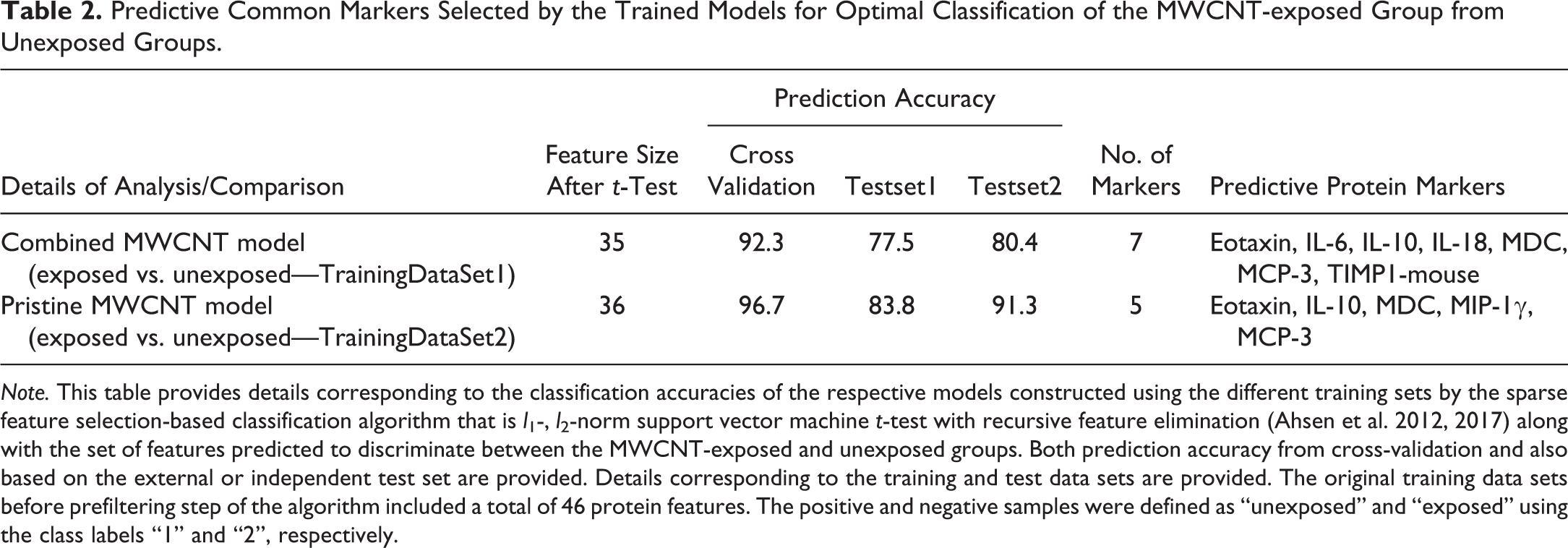
Note. This table provides details corresponding to the classification accuracies of the respective models constructed using the different training sets by the sparse feature selection-based classification algorithm that is l 1-, l 2-norm support vector machine t-test with recursive feature elimination (Ahsen et al. 2012, 2017) along with the set of features predicted to discriminate between the MWCNT-exposed and unexposed groups. Both prediction accuracy from cross-validation and also based on the external or independent test set are provided. Details corresponding to the training and test data sets are provided. The original training data sets before prefiltering step of the algorithm included a total of 46 protein features. The positive and negative samples were defined as “unexposed” and “exposed” using the class labels “1” and “2”, respectively.
Independent Validation of the Selected Predictive Markers.
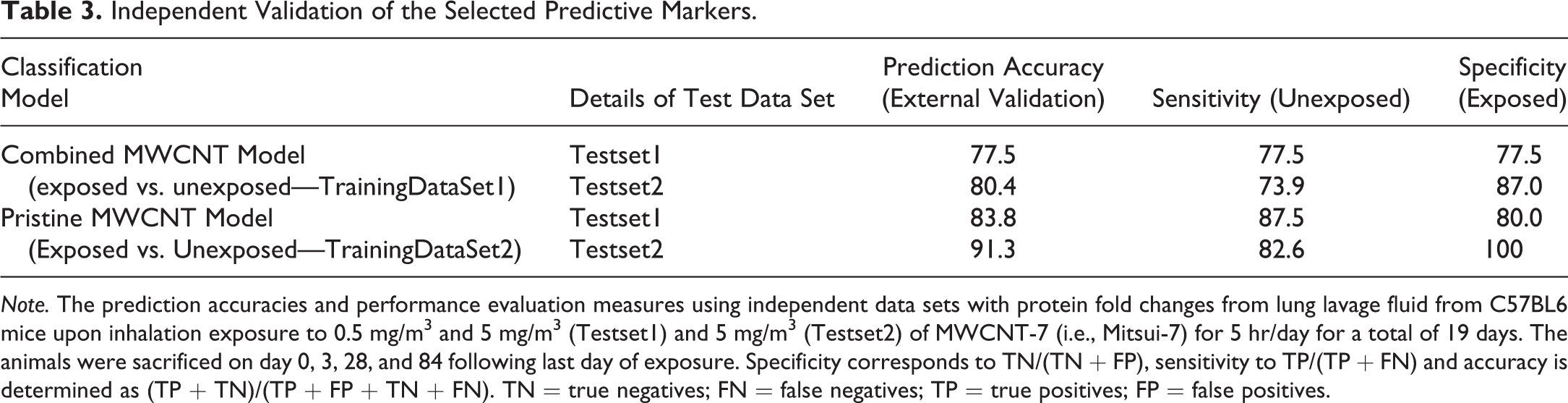
Note. The prediction accuracies and performance evaluation measures using independent data sets with protein fold changes from lung lavage fluid from C57BL6 mice upon inhalation exposure to 0.5 mg/m3 and 5 mg/m3 (Testset1) and 5 mg/m3 (Testset2) of MWCNT-7 (i.e., Mitsui-7) for 5 hr/day for a total of 19 days. The animals were sacrificed on day 0, 3, 28, and 84 following last day of exposure. Specificity corresponds to TN/(TN + FP), sensitivity to TP/(TP + FN) and accuracy is determined as (TP + TN)/(TP + FP + TN + FN). TN = true negatives; FN = false negatives; TP = true positives; FP = false positives.
The Putative Protein Markers of the Lung Lavage Fluid That Could Predict MWCNT-exposed Mice Independent of Exposure Method Employed.
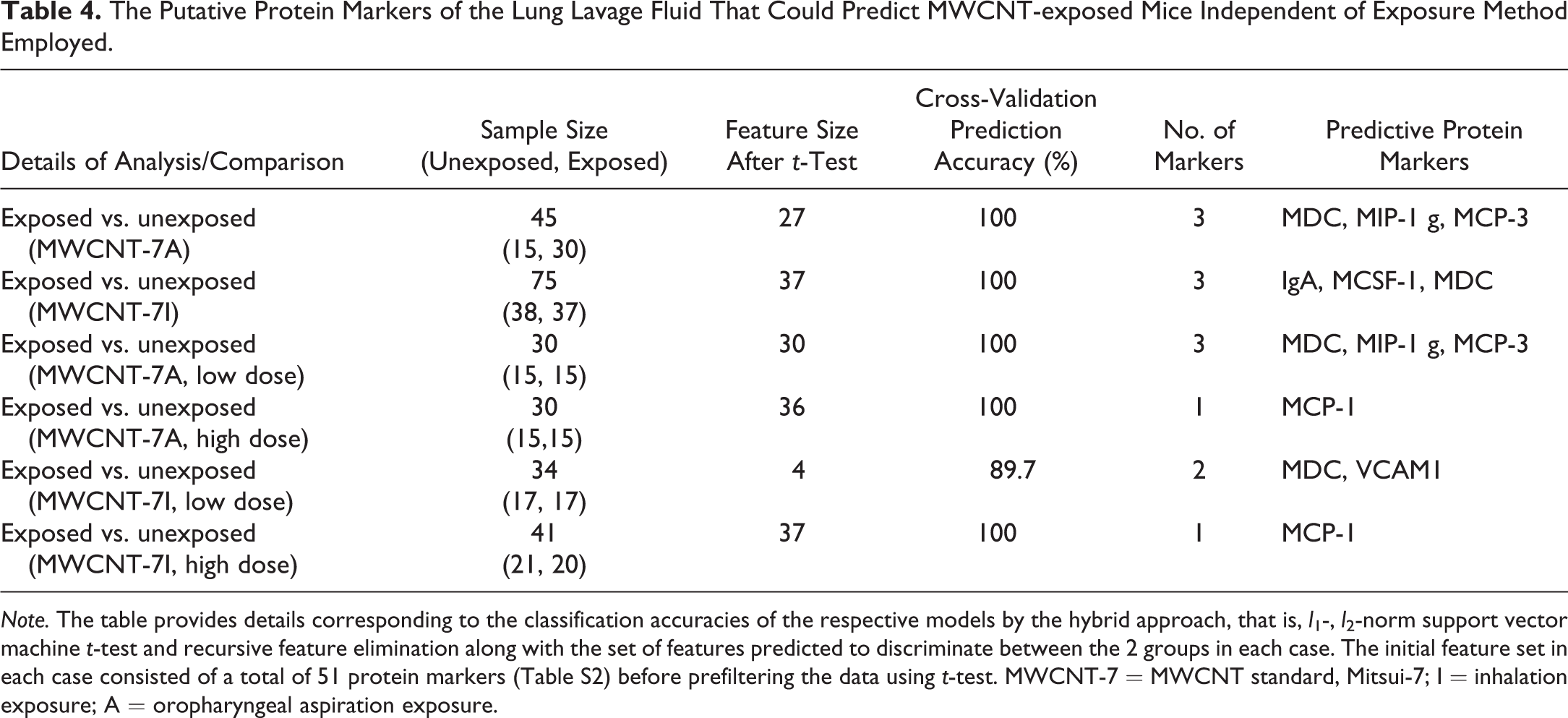
Note. The table provides details corresponding to the classification accuracies of the respective models by the hybrid approach, that is, l 1-, l 2-norm support vector machine t-test and recursive feature elimination along with the set of features predicted to discriminate between the 2 groups in each case. The initial feature set in each case consisted of a total of 51 protein markers (Table S2) before prefiltering the data using t-test. MWCNT-7 = MWCNT standard, Mitsui-7; I = inhalation exposure; A = oropharyngeal aspiration exposure.
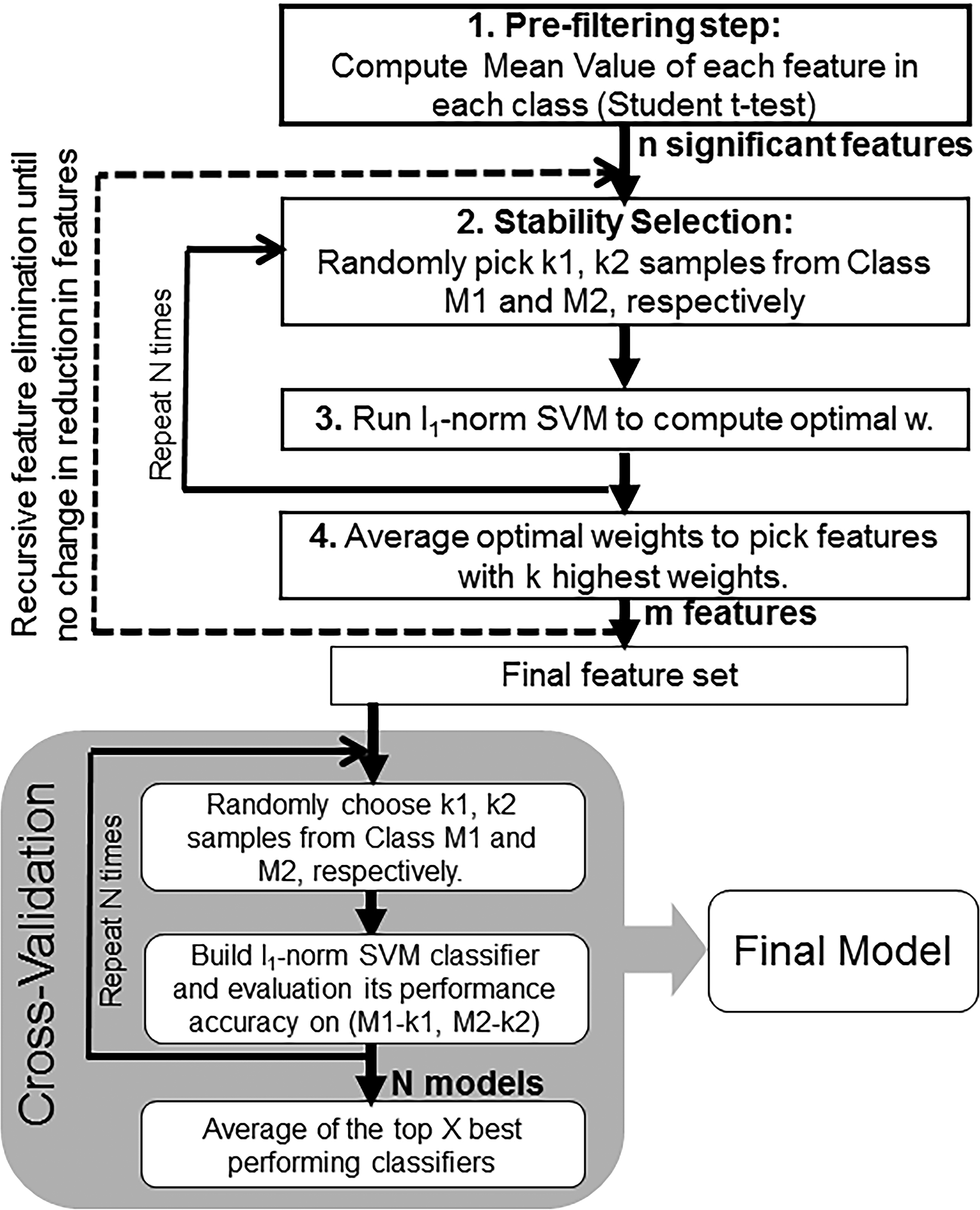
The flowchart with the various steps involved in building the support vector machines–based classifier models.
Validation of Classification Model Using an Independent Test Data Set
The important goal of generated classification models is to correctly fit or predict unseen instances/samples rather than those that are included during training. Thus, it is important to ensure that the models have the ability to generalize well to the new data. To rigorously test the MWCNT classification models developed (Tables 2 and 4), an independent data set of protein changes in BAL fluid was evaluated from mice exposed to MWCNT-7 by inhalation. The use of never seen test data allowed the assessment of overall performance and accuracy of the algorithm in predicting new data, potentially avoiding overfitting. The external validation test set considered for performance evaluation included BAL fluid samples from C57BL/6 mice exposed by inhalation to 0.5 or 5 mg/m3 of MWCNT-7 (5 hr/day for 19 days; MWCNT-7I) for a cumulative lung burden of ∼4 µg (0.5 mg/m3) or ∼40 µg (5 mg/m3) and sacrificed at 0, 3, 28, or 84 d postexposure (Erdely et al. 2013) for determining protein concentrations in lung lavage fluid. An earlier and a later time point, and a different type of exposure method, that is, inhalation instead of aspiration, was considered as part of the validation or new test data set (Table 1) to ensure that the predictive power of the developed models was not limited to the exact conditions represented by the training sets and can be easily generalized. This study comprised a total of 40 BAL fluid samples from MWCNT-7I-exposed and MWCNT-40-unexposed mice (controls; air). Two data sets containing samples from both concentrations, testset1, and only those representing high dose (5 mg/m3), testset2, were created to evaluate and assess the performance of the generated models in classifying the new test data (Tables 1 and 2).
Results
Supervised Machine-learning Approaches Successfully Identified Predictive Markers Upon Exposure to Various MWCNTs
To test the hypothesis that analysis of proteins in lung lavage fluid of mice exposed to MWCNT with different characteristics could allow selection of common and divergent markers that can predict exposure and/or effect of MWCNTs, a supervised machine learning-based approach was implemented to identify features that distinguish samples from exposed and unexposed groups. The details corresponding to the various training and test data sets employed in this study are reported in Table 1. In this study, we specifically chose data sets from materials relevant to nano-enabled manufacturing, that is, toxicological evaluation of pristine MWCNT and their PC counterparts (Bishop et al. 2017). Several strategies have been used to improve dispersion quality of NPs, including chemical and PCs, for various nano-enabled applications. The comparison of pristine/AP and PC nanomaterial toxicological data sets to identify potential markers, as employed in this study, not only provided an additional layer of comparison but can be useful in linking exposures at various points in the occupational life cycle. The training data set for building the classification model included male C57BL/6 mice treated with 0, 4, or 40 µg of a total of 5 different MWCNT materials, with 3 of them (C1 AP-MW, C2 AP-MW, and MWCNT-7A) corresponding to pristine or AP MWCNT and other 2 representing PC forms of 2 of these MWCNT (C1 PC-MW and C2 PC-MW). Mice were exposed via oropharyngeal aspiration and sacrificed at 1, 28, and/or 56 days post exposure for protein determinations. Trainingset1 for building the combined MWCNT model included data from 118 MWCNT-exposed and 38 unexposed mice (vehicle controls; dispersion medium [DM]) and the pristine MWCNT model developed using trainingset2 contained data from 75 pristine MWNCT-exposed mice (C1 AP-MW, C2 AP-MW, and MWCNT-7A) and 38 unexposed mice (vehicle controls; DM). The data collected from an independent study, C57BL/6 mice exposed by inhalation to 0.5 or 5 mg/m3 of MWCNT-7 (5 hr/days for 19 days over 4-week period; MWCNT-7I), were used to evaluate the accuracy of the algorithm in predicting new data. Further, to ensure that the generated models are capable of generalizing to unseen data, the test data also comprised samples from different time points (0, 3, and 84 days) with exposure conditions that were not reflected in the training data sets. The 2 data sets, testset1 and testset2, contained samples from 40 exposed and 40 unexposed mice (controls; air) and 23 exposed and 23 unexposed mice, respectively. In addition to the pristine and combined MWCNT models (Table 2), classification models comparing exposure to the different MWCNT considered was performed. This was done by further splitting the various MWCNT-exposed samples in the training data sets to allow the identification of putative markers that are common and those that differentiate between the different pristine and PC MWCNT investigated.
Common markers of various MWCNT exposures
In order to identify putative protein markers that can predict exposure to any of the 5 different MWCNT materials, the fold-change values of 46 proteins from both vehicle (unexposed) and MWCNT-treated (exposed) mice were analyzed using a sparse classification algorithm (Ahsen et al. 2012, 2017) to select markers with high prediction accuracy. The prediction accuracy results from cross-validation are shown in Table 2. A 92.3% accuracy of prediction was obtained by using a total of 7 proteins selected as optimal for prediction by SVM out of 35 proteins from the prefiltering step. The predictive markers selected were eotaxin/CCL11, interleukin-6 (IL-6), IL-10, IL-18, MDC/CCL22, MCP-3/CCL7, and TIMP1-mouse (Table 2). However, a better prediction accuracy rate (96.7%) was achieved when the classifier was trained using only the pristine AP MWCNT (i.e., C1 AP-MW, C2-AP-MW, and MWCNT-7A), suggesting a differential response of one or both of the PC-MW groups. This resulted in the selection of 5 predictive protein markers, which included eotaxin/CCL11, IL-10, MDC/CCL22, MIP-1γ, and MCP-3/CCL7 (Table 2). Four of these protein markers, MDC/CCL22, eotaxin/CCL11, IL-10, and MCP-3/CCL7, were in common with the combined MWCNT model (Tables 1 and 2). Further, the predictive power of the 2 classification models and the selected feature sets were evaluated using an independent data set, which was never used in training the models. The prediction accuracy results along with performance evaluation measures (e.g., sensitivity and specificity) are reported in Table 3. The classifier model constructed using only pristine MWCNT (MWCNT-7, C1 AP-MW, and C2 AP-MW) samples with 5 predictive markers resulted in overall higher accuracy of prediction than the combined MWCNT classifier model with 7 protein markers. Moreover, the accuracy of the testset2 with data from mice exposed to a high dose (5 mg/m3) of MWCNT-7 via inhalation exposure was consistently higher compared to the testset1 that considered both low- and high-dose exposures (Tables 1 and 3). Despite these differences, the markers selected based on aspiration exposure studies were able to reliably predict exposure to MWCNT in an inhalation study. This suggests that the identified and selected markers could serve as predictive markers of exposure irrespective of postexposure time point or exposure method.
Exposure route/method independent markers of pulmonary exposure or effect of MWCNT
Various experimental routes of pulmonary exposure, including oropharyngeal aspiration, inhalation, and intratracheal instillation, have been used to evaluate and assess biological responses of MWCNTs. However, the route of exposure has been a debate when it comes to determining the common toxicological profiles of MWCNT across different studies. To test if it is possible to select putative signatures common to different routes of pulmonary exposure that can distinguish MWCNT-exposed mice from unexposed controls, we further constructed 2 classification models using training data sets from (a) oropharyngeal aspiration of MWCNT-7 and (b) inhalation exposure to MWCNT-7. The selected feature sets or markers were cross compared in each case to determine whether protein markers were (in)consistent between the 2 types of exposures. The list of selected protein markers that can predict pulmonary exposure or responses to MWCNT-7 along with the predicted accuracies is shown in Table 4. The levels of MDC, MIP-1γ, and MCP-3, and IgA, M-CSF1, and MDC in the lung lavage fluid were selected as potential markers that can predict oropharyngeal and inhalation exposure to MWCNT-7, respectively. External validation of the selected markers using the inhalation data set as a test set for the oropharyngeal aspiration and vice versa resulted in high prediction accuracies, 97.4% and 93.3%, respectively (Table 4). This clearly suggests that irrespective of the exposure method employed, it is possible to precisely predict exposure to MWCNTs, in particular MWCNT-7. Of the 3 protein markers selected by the classification models developed, MDC was common to both exposure methods compared. While the feature sets selected using either the entire data set or low-dose MWCNT-7 aspiration exposure were identical in predicting MDC/CCL22, only 1 protein, that is, MCP-1, was selected as a putative marker that can distinguish (with 100% accuracy) high-dose oropharyngeal aspiration exposures only to MWCNT-7 (Table 4). Similarly, a classifier model that was developed based on the data set from inhalation exposure to 5 mg/m3 (i.e., high dose) of MWCNT-7 also resulted in the selection of MCP-1 as a potential protein marker in the lung lavage fluid identical to aspiration exposures.
Selective markers of each MWCNT exposure
Previous attempts to identify signatures of toxicity responses of NPs, especially MWCNTs, have primarily focused on pristine AP CNTs. However, their application in nano-enabled manufacturing and commercialization requires evaluation and selection of toxicological profiles that are beyond the pristine or AP nanomaterials. Thus, to enable the identification and selection of protein markers of exposure to pristine (AP) and PC counterparts from various facilities, we constructed 4 different classification models to select features that can distinguish (a) exposure to AP- and PC-MW materials of C1 from C2, (b) exposure between AP-MW and PC-MW-exposed mice within C1 and (c) C2, and (d) pristine or AP-MW particle-exposed mice of C1 from C2. A summary of cross-validation accuracies along with the protein markers selected to be optimal for prediction in each case is listed in Table 5. As can be seen from the predicted accuracies of the cross-validation, it was possible to reliably predict, often with high accuracy (97.2% when comparing AP-/PC-MW or 94.3% for AP-MWs), exposure to materials from 1 company versus another (Table 5). While eotaxin/CCL11, FGF-Basic, IgA, MMP-9, MCP-3/CCL7, and SCF were selected as optimal for predicting exposure to AP-/PC-MWs of C1 from C2, the protein levels of IP-10, IL-6, MMP-9, and MCP-3/CCL7 in the lung lavage fluid were selected as putative markers that can distinguish exposure to AP-MW of C1 from C2. However, it was difficult to discriminate, with relatively high accuracy, between the exposures and/or effects of AP-MW and PC-MW particles within each company (Table 5). The prediction accuracy of classifying AP-MW from PC-MW of C1 was lower compared to C2. Moreover, only 2 proteins, haptoglobin and IL-4, after initial prefiltering step, were found to be significantly (p < .05) different between the pristine or AP-MW and PC-MW particles of C1 (Table 5, row 2). This suggests that the biological responses, that is, changes in protein levels in the lavage fluid, could be similar upon exposure to AP-MW and PC-MW particles of C1 in mice. In contrast, a total of 17 proteins exhibited significant differences in their fold-change values between AP-MW and PC-MW particles of C2. Of these markers, LIF, MDC, MIP-1β, MIP-2, and PAI-1, were selected as discriminating features (Table 5, row 3), implying that biological responses of exposure to AP-MW and PC-MW materials from C2 could be different. Importantly, the selection of MDC/CCL22 as a discriminating feature between exposure to AP-MW and PC-MW materials of only C2 is quite interesting, as it was selected as a common marker across several classification models that can distinguish exposure to MWCNT from unexposed controls. This further suggests that PC of pristine MWCNT employed by C2 may be leading to attenuated pulmonary toxicity responses in mice.
Summary of Accuracies and Selected Protein Markers That Can Distinguish Exposure to Different AP-/PC-MWs Particles.
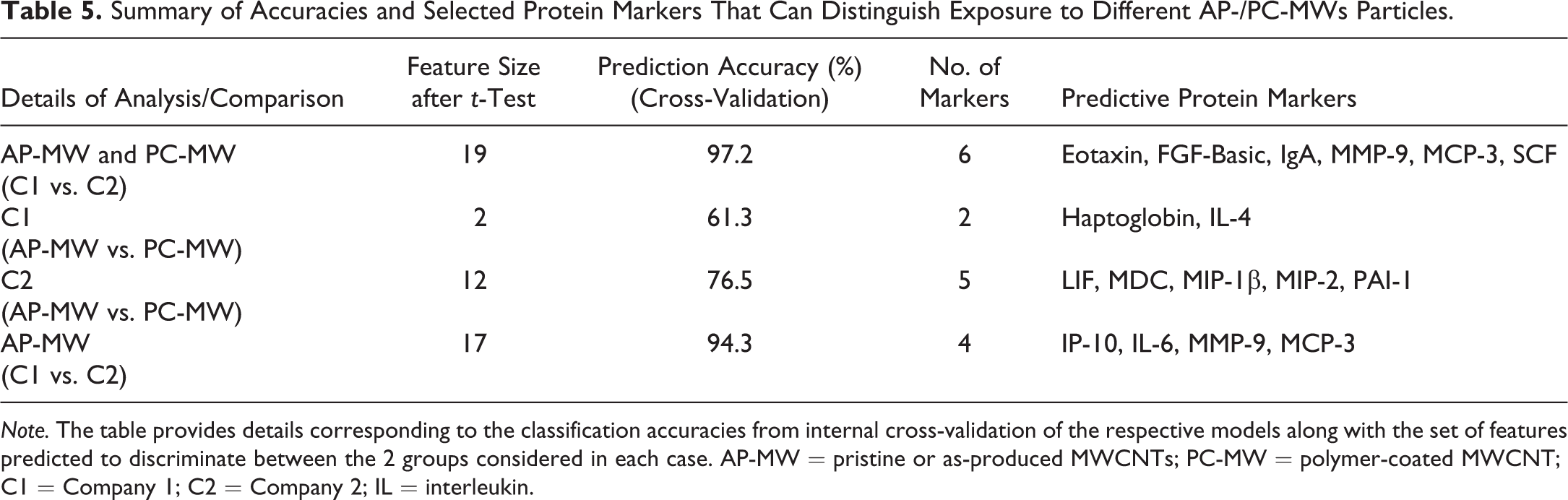
Note. The table provides details corresponding to the classification accuracies from internal cross-validation of the respective models along with the set of features predicted to discriminate between the 2 groups considered in each case. AP-MW = pristine or as-produced MWCNTs; PC-MW = polymer-coated MWCNT; C1 = Company 1; C2 = Company 2; IL = interleukin.
Pathological Outcomes of Exposure to Various MWCNTs
To explore the possibility that the selected markers of MWCNT exposure identified by our supervised machine-learning approach might be predictive of a specific histopathological outcome, murine lung tissues stained with H&E or PSR were evaluated by a veterinary pathologist for evidence of inflammation (H&E) and fibrosis (PSR). Oropharyngeal aspiration exposure to MWCNT-7, C1-AP MW, C1-PC MW, and C2-AP MW was associated with an adverse outcome in the lungs at 84 days postexposure (Figure 3). This outcome was characterized by the presence of small granulomas in association with particle deposits and the deposition of extracellular collagen/fibrosis at 84 days postexposure. In contrast, aspiration of C2 PC-MW was associated with minimal histopathology, notably the attenuation of particle-associated granulomas or fibrosis. Microscopic lung sections from the control animals revealed normal histology.
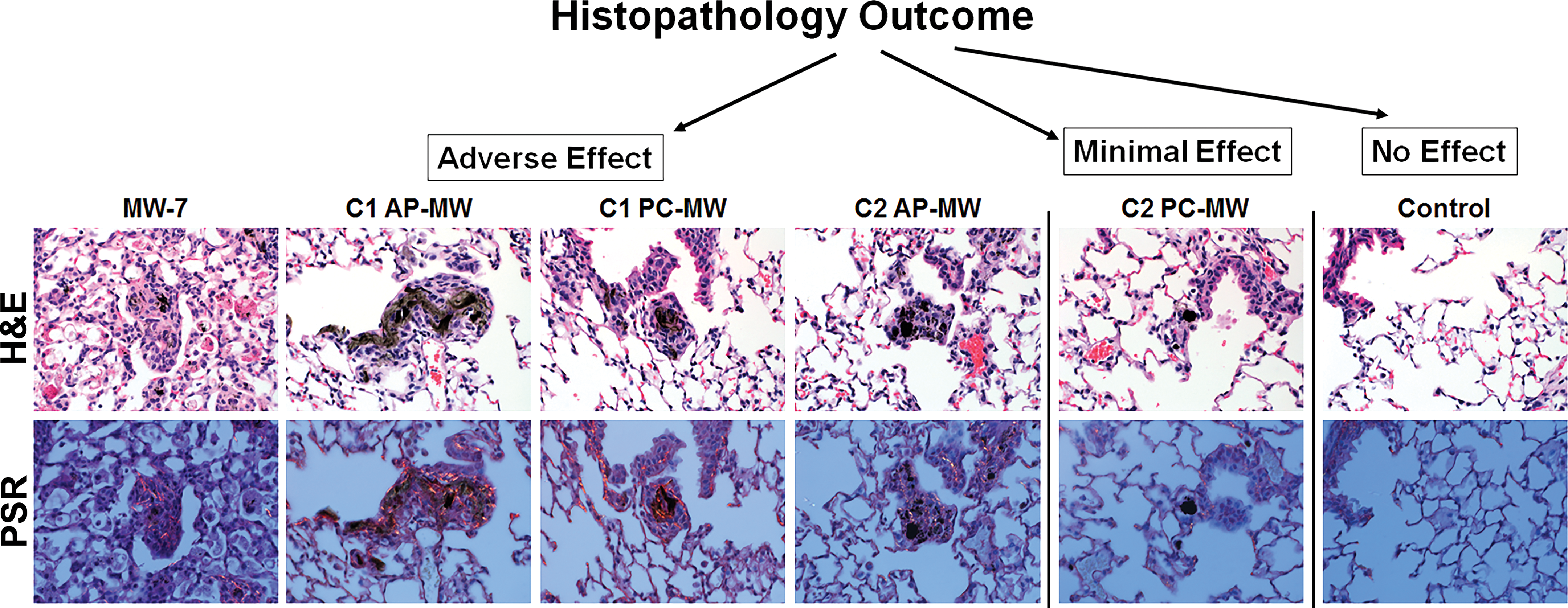
Hematoxylin and eosin and picrosirius red (PSR)-stained lung sections show particle depositions at 84d postexposure. All as-produced–multiwalled carbon nanotubes (MWCNTs) particles (AP-MW) investigated including Mitsui-7 (MW-7) and polymer-coated-MWCNT (PC-MW) particles from company 1 (C1) showed the presence of small granulomas in association with particle deposits and collagen deposition/fibrosis. Interestingly, the PC-MW particles from company 2 (C2) agglomerated in vivo and particle-associated granulomas or fibrosis was attenuated in comparison to other groups. The morphological changes in the lungs were examined using bright-field microscopy, and collagen deposition was visualized in PSR-stained slides by polarizing light microscopy.
Selected Protein Markers Suggest Mechanism(s) Associated with Adverse Outcomes
Most often, the predictive biomarkers are those that can provide mechanistic insights into the etiology of disease or adverse outcomes of toxicological responses. Both models generated using pristine and pristine MWCNT together with their PC counterparts had considerable overlap in their predictive features (Table 2). While the pristine MWCNT model with high predictive accuracy (Table 2) consisted of 5 proteins, the combined MWCNT model with pristine and PC MWCNT led to the selection of 7 proteins. Of the selected proteins, MDC/CCL22, eotaxin/CCL11, MCP-3/CCL7, and IL-10 were found to be common between the 2 models (Figure 4a). While IL-6, IL-18, and TIMP1mouse were selected only by the combined MWCNT model, MIP-1γ/CCL9 was selected solely by the pristine model. Further, we used these common as well as the specific proteins selected in each case to infer mechanistic insights that may be associated with MWCNT-induced adverse outcomes, such as inflammation, granulomatous lesions, and fibrosis. The proteins that were selected exclusively by each models, in particular IL-6, IL-18, and TIMP1, have been implicated in the inflammasome activation upon exposure to high-aspect ratio materials including asbestos and CNTs (Cassel et al. 2008; Dolinay et al. 2012; Dostert et al. 2008; Palomäki et al. 2011; Oberdorster et al. 2015). Of the protein markers common to both models, eotaxin/CCL11, IL-10, and MCP-3/CCL7 are known to contribute to Th2-related immune and/or inflammatory responses during pulmonary fibrosis. These findings together with the reported increase in the expression of MDC levels in various rodent models of human fibrotic lung diseases (Belperio et al. 2002; Ritter et al. 2005) further highlight the biological meaning of the selected markers in this study.
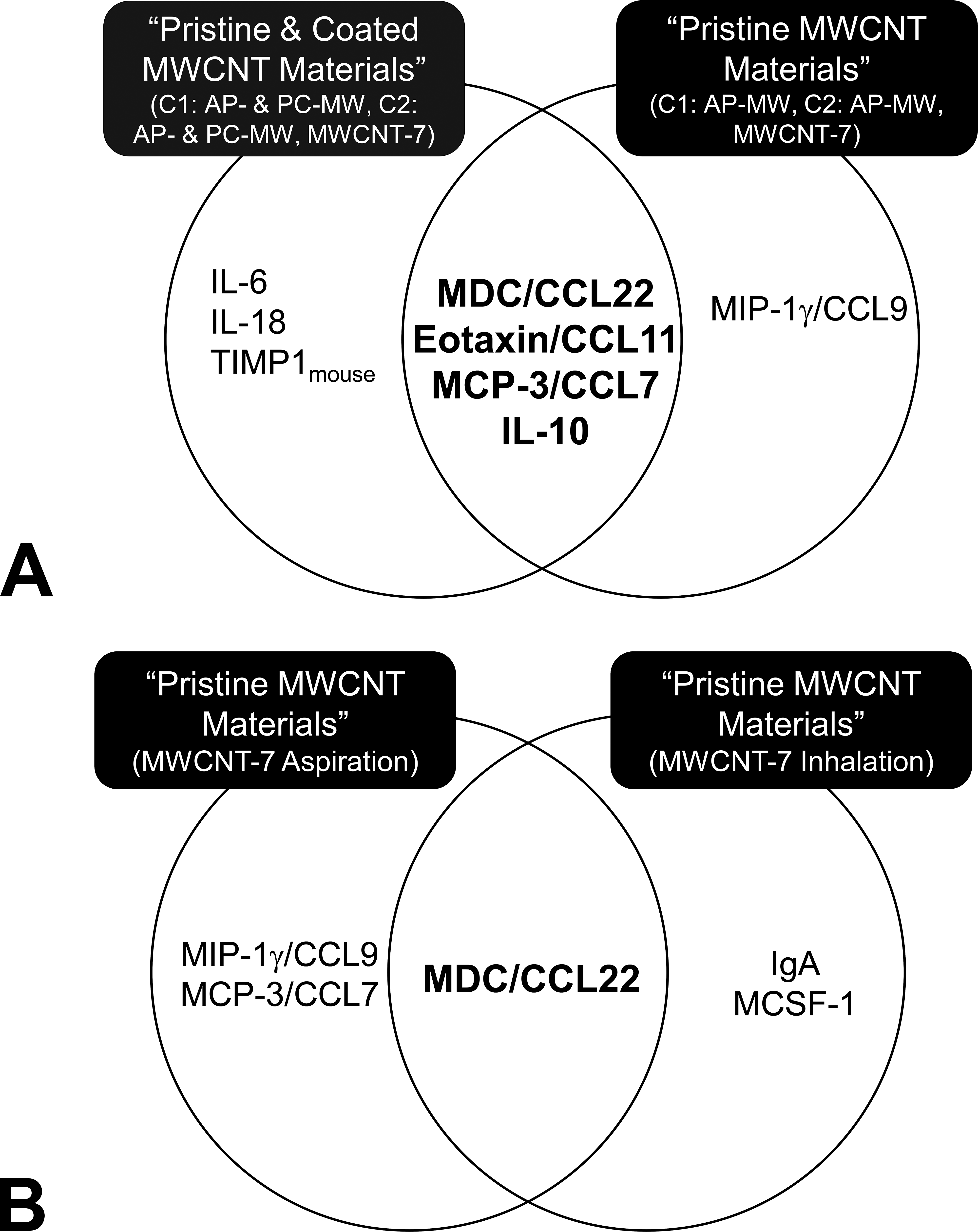
Some protein markers selected were highly predictive of pathological outcomes associated with multiwalled carbon nanotubes (MWCNT) exposures. (a) Maximum predictive power was achieved in the classifier model using “pristine” or as-produced (AP) MWCNT (C1/C2 AP-MW, MWCNT-7) with 5 proteins compared to the combined MWCNT model which used both pristine and polymer-coated (PC) MWCNT (C1/C2 AP- & PC-MW, MWCNT-7) resulting in 7 proteins, with 4 of them in common. These 4 proteins are known to play a key role in explaining the biological processes underlying fibrosis. (b) Comparison of the features selected by oropharyngeal aspiration and inhalation MWCNT-7 classification models. A single protein marker, MDC/CCL22, selected independent of exposure method was also commonly selected by MWCNT models as a material group.
Discussion
In this study, protein levels in the lung lavage fluid were used to discriminate between NP exposed and unexposed controls using male C57BL6 mice exposed by oropharyngeal aspiration to subtoxic and toxic doses of various MWCNT relevant to nano-enabled manufacturing processes. This study was designed and conducted based on the results from several previous and ongoing studies (Bishop et al. 2017; Kodali et al. 2016; Roberts et al. 2016), albeit all using MWCNT particles, but having different physicochemical characteristics and/or PCs (Table S1), by independent investigators at NIOSH. We constructed sparse SVM-based classification models to determine or select proteins in the lung lavage fluid that allow prediction of exposure and/or pulmonary responses to MWCNT as a material group (Figure 1) as well as those that can discriminate between the different forms of MWCNT investigated (Table 5). Importantly, a panel of 3 to 7 proteins identified, within eotaxin/CCL11, IL-6, IL-10, IL-18, MDC/CCL22, MCP-3/CCL7, TIMP1mouse, MIP-1γ, MCP-1, IgA, and VCAM1, using this approach were found to discriminate between MWCNT-exposed versus unexposed groups (Tables 2 and 4). These selected markers were further tested for prediction accuracy on blinded independent test data sets during the training step. Although the test sets were comprised of samples from different time points (0, 3, and 84 days), subchronic inhalation exposures as opposed to oropharyngeal aspiration and different dose rates (∼0.2 and 2 µg per day at 0.5 mg/m3 and 5 mg/m3, respectively, at 5 hr/day for 19 day) that were not reflected by the training data sets, the identified protein features were able to distinguish MWCNT-7 inhalation exposure with high accuracies (∼78–91%, Table 3). This suggests that identified protein patterns in lavage fluid have the ability to successfully classify samples into the 2 groups (unexposed—1, exposed—2) irrespective of exposure method or postexposure time points employed. Additionally, these findings highlight the potential relevance of the identified markers in the lung lavage fluid to serve as predictors of lung pathology.
Applying coatings such as chemicals or polymers to the pristine AP MWCNT has increased recently in order to decrease human exposure (ease in handling and potentially reduce dustiness), increase dispersion to use less material, and enhance the quality of the end product. Previous in vivo toxicity studies indicate that such modifications of NPs, specifically CNTs, for downstream nano-enabled applications can affect pulmonary toxicity (Bishop et. al. 2017; Tabet et al. 2011). Interestingly, the prediction accuracies associated with the combined classification model that incorporated PC-MW exposures together with AP-MWs was consistently less, compared to the model that used only pristine or AP-MW materials (i.e., C1 AP-MW, C2 AP-MW, and MWCNT-7). To explore whether PC-MW of C1 and/or C2 led to decreased prediction accuracies (Table 2 and S1), a detailed cross comparison of the protein features selected that can classify AP-MW from PC-MW exposures of C1 with C2 was performed. Only 2 proteins (haptoglobin and IL4) exhibited significant differences between AP- and PC-MW exposures of C1 after prefiltering using the Student’s t-test, suggesting that the biological responses upon exposure to AP- and PC-MW nanomaterials from C1 are similar and cannot be easily distinguished (Table 5). This is in line with the findings that exposure to both AP- and PC-MW particles from C1 induced similar histopathological outcomes (Figure 3) such as granulomas with particle deposits and fibrosis (Bishop et al. 2017). In contrast, 5 proteins (LIF, MDC, MIP-1β, MIP-2, and PAI-1) were selected, of a total of 12 initial protein features that were significantly different (p < .05) between the 2 groups, to distinguish the AP-MW exposure or effects from PC-MW materials of C2. In fact, granuloma formation or fibrosis, as observed with AP-MW, was attenuated following exposure PC-MW from C2 (Bishop et al. 2017), indicating that PCs employed by C2 altered the toxicity responses of MWCNT leading to overall decreased prediction accuracies of models that incorporated both AP- and PC-MWs data during training (Table 2).
Further analysis investigated the correlation of the common and specific protein markers selected by different classifier models of various MWCNT, in conjunction with pathological responses in the lungs, highlighted the biological significance of the discovered markers. One mediator in particular, MDC/CCL22, was associated with most exposures, irrespective of particle characteristics and exposure methods employed (Tables 2 and 4). In this study, selection of MDC/CCL22 as a putative marker that can discriminate exposures or pulmonary effects was concomitant with an adverse outcome, deposition of extracellular collagen and fibrosis, in the lungs at 84 days postexposure following aspiration of MWCNT-7, C1 AP-MW, C1 PC-MW, and C2 AP-MW. This result is consistent with several previous studies that reported increased expression of MDC in rodent models of human fibrotic lungs disease including murine bleomycin-induced pulmonary fibrosis (Belperio et al. 2002), rat radiation pneumonitis/pulmonary fibrosis (Inoue et al. 2004), and cigarette smoke-treated rats (Ritter et al. 2005). Previous evidence also suggests that MDC/CCL22 plays a role in the pathogenesis of several human inflammatory and fibrotic lung diseases including idiopathic pulmonary fibrosis (Shinoda et al. 2009), World Trade Center (WTC) dust exposure (Caplan-Shaw et al. 2011; Wu et al. 2010; Weiden et al. 2012; Nolan et al. 2012), and smoking-related lung diseases including chronic obstructive pulmonary disease (Kim et al. 2015; Frankenberger et al. 2011). Importantly, MDC protein levels in the serum and lavage samples have been shown to be very sensitive in predicting declining lung function in humans following particulate exposure including smoking (Kim et al. 2015; Shinoda et al. 2009) and WTC dust (Nolan et al. 2012; Weiden et al. 2012). An increase in MDC protein levels in the serum in first responders exposed to WTC dust is further supported by the presence of CNTs-like structures of various sizes and lengths in patient lung samples as well as those in WTC dust samples (Wu et al. 2010). Similar to the significant exposures associated with the WTC dust, if the deposited dose of MWCNT is high enough (e.g., 40 µg), serum levels of MDC increase (Erdely et al. 2009). Moreover, the selection of MDC to be discriminating between AP-MW and PC-MW samples of C2 further supports its role in the development lung pathology, as aspiration exposure to C2 PC-MW is associated with minimal histopathology and no significant fibrosis at the 40 µg exposure dose.
Fibrosis is often the end result of chronic inflammatory responses. Several stimuli including persistent infections, allergic responses, and chemical irritants can trigger fibrosis in lungs. Previous studies, including the current study, indicate that exposure to different variants of MWCNT induces pulmonary fibrosis (Ma-Hock et al. 2009; Mercer et al. 2011; Porter et al. 2010; Ryman-Rasmussen et al. 2009; Pauluhn 2010). Both inflammation and Th2 immune responses are shown to play a key role in the progression of pulmonary fibrosis (Wynn 2004; Dong and Ma 2016; Dong et al. 2016; Tajima et al. 2007). This is further corroborated by the set of protein markers identified and selected in the current study upon exposure to various MWCNTs. Of the protein markers selected, while MDC/CCL22 is crucial for the recruitment of Th2 cells during allergic inflammation, eotaxin/CCL11 and MCP-3/CCL7 are both chemokines that contribute to Th2-related allergic responses. Additionally, a study comparing the pulmonary responses of short-term inhalation exposure of MWCNT-7 with repeated oropharyngeal aspiration exposure reported the expression of chemokines CCL7 and CCL11 and their involvement in cellular chemotaxis (Kinaret et al. 2017). A recent study based on meta-analysis of transcriptomics expression profiles also reported that while early responses (24 hr, 3 days) after exposure to MWCNT in rodents were similar to bacterial- or bleomycin-induced injury responses, the responses 28 days postexposure were suggestive of Th2-mediated responses (Nikota et al. 2016). Interestingly, exposure to MWCNT was associated with the presence of small granulomatous lesions, in addition to fibrosis. While MDC plays a role during both innate and adaptive granuloma formation, increased levels, together with the characteristic profile of the other markers selected, suggest a Th2-mediated adaptive granuloma formation. Exacerbation of Th2 granulomatous inflammation was associated with significant increases in the expression of MDC/CCL22, MCP-3/CCL7, and eotaxin/CCL11 (Chiu et al. 2003; Chensue 2013). Taken together, these findings suggest MDC/CCL22, in conjunction with other markers selected in lung lavage fluid, could serve as an early response marker of exposure and associated adverse outcomes.
In occupational and manufacturing settings, inhalation is widely accepted as the primary means of entry for NPs and ambient particulate materials into the human body (Arora, Rajwade, and Paknikar 2012). In experimental studies, the inhalation exposures are often preferred over oropharyngeal aspiration, which delivers particles as a bolus dose, to assess NP toxicity as it closely mimics real-life exposure scenarios. However, it has been argued that the aspiration technique could be an alternative to inhalation if experiment conditions and doses are properly assessed (Kinaret et al. 2017; Shvedova et al. 2008; Rao et al. 2003). Similar to inhalation, aspiration exposure was shown to lead to even deposition of particles in different regions of the lungs (Rao et al. 2003). Moreover, recent studies comparing inhalation and aspiration exposure to respirable CNTs assessing pulmonary toxicity responses further support the use of aspiration as a surrogate to inhalation exposure (Kinaret et al. 2017; Shvedova et al. 2008). This is further corroborated by our study where the protein markers selected that can classify MWCNT-7 exposed from controls exhibited similarities between aspiration and inhalation exposure techniques. In particular, MDC/CCL22 was commonly selected as a predictive marker for the 2 exposure methods, in addition to MIP-1γ and MCP-3/CCL7 for aspiration and IgA and MCSF-1 for inhalation. Even at low-dose exposures, MDC/CCL22 consistently discriminated between MWCNT-7 exposed and unexposed groups (Table 4). Interestingly, while the classification algorithm trained only using high-dose exposure to MWCNT-7 did not result in the selection of MDC, a single marker, MCP-1/CCL2, was selected that can discriminate both aspiration and inhalation MWCNT-exposed mice from unexposed controls. The overall predicted accuracies of MWCNT-7A classifier in predicting MWCNT-7I exposures and vice versa further reflect the similarity in MWCNT-7-induced pulmonary responses irrespective of the exposure method employed. This is in complete agreement with a recent study which reported that, when the doses are adequately adjusted, aspiration and inhalation exposures to MWCNT-7 exacerbate very similar histological, immunological, and molecular responses (Kinaret et al. 2017). Further, the selection of MCP-1 only in the case of exposure to high doses of MWCNT-7 highlights that it could be a dose-related marker. Given that exposures in the workplace for MWCNT could be relatively low in comparison to in vivo studies (Erdely et al. 2013; Dahm et al. 2015), it is critical to develop predictive discriminators over a dose range representative of occupational exposures.
Several groups have previously attempted to identify toxicogenomic signatures predictive of biological responses of engineered nanomaterials (Guo et al. 2012; Pacurari et al. 2011; Dymacek et al. 2015; Snyder-Talkington et al. 2013, 2016a, 2016b; Shvedova et al. 2016). These studies only focused on one type of a carbon nanomaterial and/or involved preselection of genes related to either cancer or a particular biological response (e.g., inflammation, fibrosis) or pathological outcome in the lungs (e.g., cancer, fibrosis). The current study differs from these studies in several ways and focused on the identification of protein markers that can identify exposure to various MWCNT with different PCs and physical characteristics (Table S1) as opposed to a specific single particle. The advantage of such approach is its ability to detect markers that are reminiscent of biological responses common to MWCNT in general. Moreover, the binary classification approach presented in this study combining various concentrations and postexposure time points spooled together into a single class allowed us to select markers that were consistently dysregulated across all doses and time points as opposed to markers that correspond to dominant biological responses at a given time point. The incorporation of other earlier and later responses, not included during training, in the validation test set allowed us to further test the ability of selected markers in predicting responses that are not limited to the exact conditions of the training set. In occupational settings, such markers may be useful in evaluating exposure and/or early biological effects in workers, thus supporting a more adequate assessment and management of risk in respect of nanomaterial exposure and/or effect (Schulte and Hauser 2012). Despite a limited set of proteins analyzed, as opposed to genome-wide studies, the overall approach and the application of sparse classification methods described in this study was able to select relevant features associated with biological responses in the lungs. However, a study was able to distinguish air (unexposed control) versus mice exposed to MWCNT-7 by inhalation by analyzing miRNA patterns in whole blood samples 17 months postexposure (Snyder-Talkington et al. 2016b). They have suggested that blood miRNA measurements might be useful in screening workers for MWCNT-induced lung pathology. A follow-up study by NIOSH investigators, albeit using a small cohort of workers from MWCNT manufacturing facility, investigating aberrant changes in mRNA and ncRNA expression profiles in the whole blood of humans potentially exposed to MWCNT (Shvedova et al. 2016), also supported the potential role of miRNA, together with mRNA, as relevant blood markers for monitoring MWCNT exposure in humans. As a next logical step, the methodology and approaches presented in this study will be applied and evaluated further to select features involving genome wide studies. Lastly, it has to be noted that there are several limitations in this study including limited dose range (4–40 µg), a predefined protein panel, consideration of unmodified or pristine MWCNT with varied diameters and/or their PC counterparts, lack of particle controls such as carbon black, and that all data were generated by different laboratories within NIOSH. Large-scale meta-analysis studies focusing on MWCNT exhibiting diverse physicochemical characteristics such as varied coatings, altered dissolution, and different oxidative potentials (e.g., metals, chemicals, and oxidized groups) are needed to corroborate our findings and to support the broader application of proposed markers to MWCNT exposures.
Footnotes
Authors’ Note
The findings and conclusions in this report are those of the authors and do not necessarily represent the views of the National Institute for Occupational Safety and Health.
Author Contribution
Authors contributed to conception or design (NY, PZ, JR, AE); data acquisition, analysis, or interpretation (NY, MO, VK, LB, PZ, JR, VC, AE); drafting the manuscript (NY, MO, VK, AE); and critically revising the manuscript (NY, MO, LB, PZ, VC, JR, AE). All authors gave final approval and agreed to be accountable for all aspects of work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by National Institute for Occupational Safety and Health (NIOSH) 939051E.
Supplemental Material
Supplementary material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
