Abstract
A 2-stage algorithmic framework was developed to automatically classify digitized photomicrographs of tissues obtained from bovine liver, lung, spleen, and kidney into different histologic categories. The categories included normal tissue, acute necrosis, and inflammation (acute suppurative; chronic). In the current study, a total of 60 images per category (normal; acute necrosis; acute suppurative inflammation) were obtained from liver samples, 60 images per category (normal; acute suppurative inflammation) were obtained from spleen and lung samples, and 60 images per category (normal; chronic inflammation) were obtained from kidney samples. An automated support vector machine (SVM) classifier was trained to assign each test image to a specific category. Using 10 training images/category/organ, 40 test images/category/organ were examined. Employing confusion matrices to represent category-specific classification accuracy, the classifier-attained accuracies were found to be in the 74–90% range. The same set of test images was evaluated using a SVM classifier trained on 20 images/category/organ. The average classification accuracies were noted to be in the 84–95% range. The accuracy in correctly identifying normal tissue and specific tissue lesions was markedly improved by a small increase in the number of training images. The preliminary results from the study indicate the importance and potential use of automated image classification systems in the histologic identification of normal tissues and specific tissue lesions.
Numerous advances in image processing and machine learning techniques along with major improvements in computing and digital data storage abilities have occurred in recent decades. With these improved techniques, automated digital image classification has become an important component of digital image processing in applications such as biometric security using face recognition, 12 fingerprint identification, 1 and medical images acquired by tomography techniques. 9 The use of digital mammography has directed attention toward computer-aided diagnosis (CAD) techniques in the field of radiology.4,5 The potential of increased efficiency in the evaluation of digitized histology slides using image analysis techniques has generated a great amount of interest among pathologists. The combination of histology and improvements in algorithmic image analysis and computational systems can effectively complement the pathologist in histologic slide evaluation. Computer-aided diagnosis systems have been employed in the detection and grading of cervical, prostate, breast, and lung carcinomas using histology and immunohistochemical image analysis and in the identification of oral tumors.6,8,10 In the current study, a CAD system for histologic image analysis was developed and evaluated using normal bovine tissue and tissue containing specific histologic lesions.
Images can be automatically classified either in a supervised or in an unsupervised learning paradigm. In supervised learning, a limited set of representative training images from each of the classes is available along with the associated class labels. Given a new test image, the goal is to automatically assign the correct class label from among the available categories. In unsupervised learning, the challenge lies in learning class structure from unlabeled data (i.e., training data whose class affiliations are not known beforehand). In the current study, a supervised classification framework was proposed.
Supervised image classification is primarily a 2-stage framework. The first stage consists of the extraction of important, representative characteristics from training images.7,11 The capture of discriminative information between various classes is a highly desirable property of the feature design. For example, the normal lung image (Fig. 1a) is characterized by large clear openings of the alveoli. In the lung image displaying acute suppurative inflammation (Fig. 1b), the alveoli contain large numbers of inflammatory cells (neutrophils and occasional macrophages). This discriminative information should be captured in the features. Additionally, in image classification, it is often necessary to have an economical way of encapsulating image information for reduced computational complexity. For example, a raw histological image of size 1,000 × 1,000 pixels is represented by a million coefficients. Extracted features, on the other hand, are of the order of a few hundred coefficients.
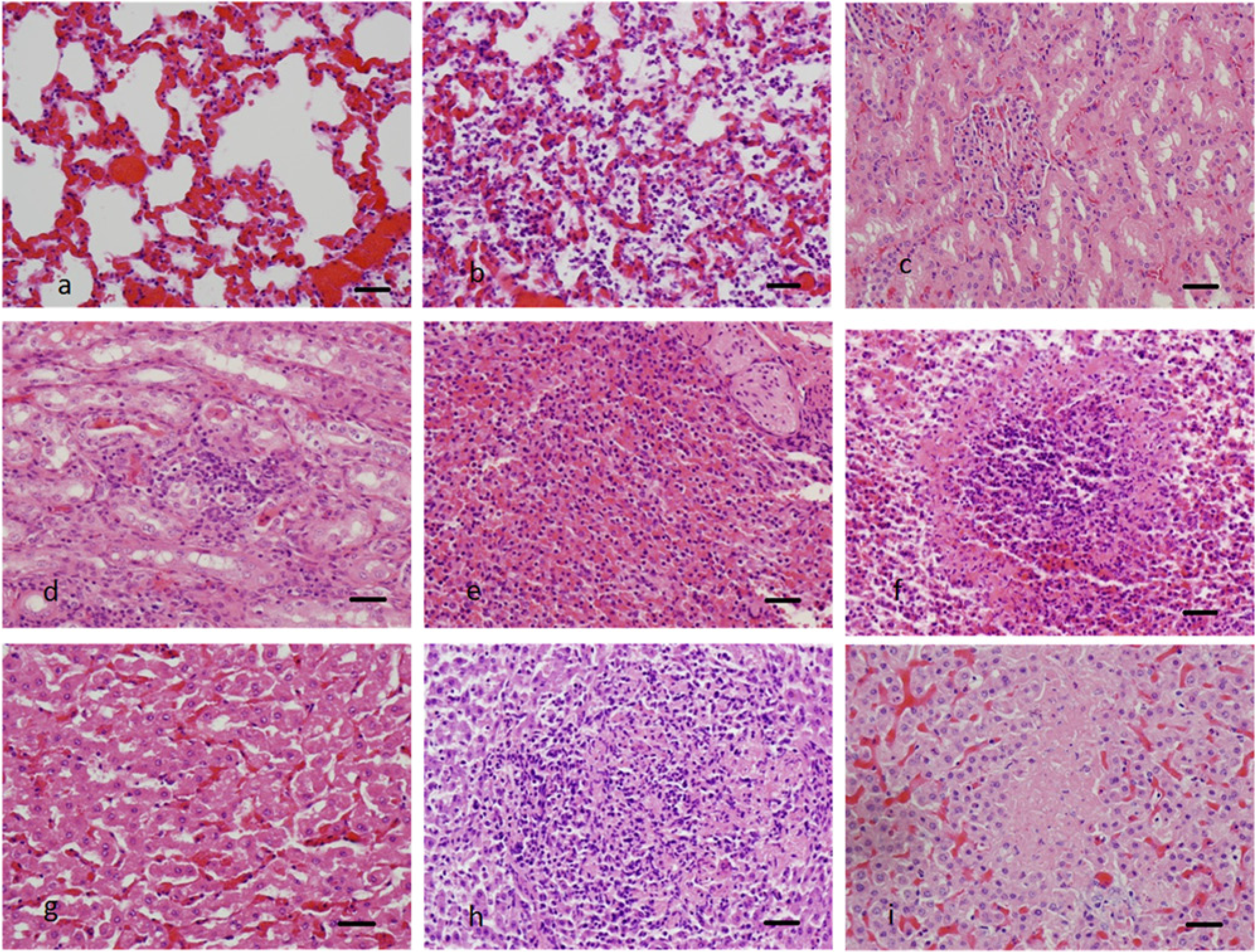
Examples of digitized photomicrographs from different bovine organs. Normal lung (
The redundancy in image pixel information can be exploited to design features of much smaller dimension. In the framework described herein, well-known image features such as color image pixel statistics (mean, median, maximum, minimum, and standard deviation of pixel intensities) histograms, gray level co-occurrence matrix, and wavelet coefficients, 7 as well as novel features specifically designed for the 4 organs under consideration, were extracted from each image.
In the second stage, the features acquired from the training images are used to train a classifier or a decision engine that learns a classification rule based on which any new test image is assigned to a particular category (Fig. 2). A standard linear support vector machine (SVM) was used as the decision engine for the framework used in the current study. 2 The SVM is a widely used machine learning technique and has demonstrated significant success in image classification. 3
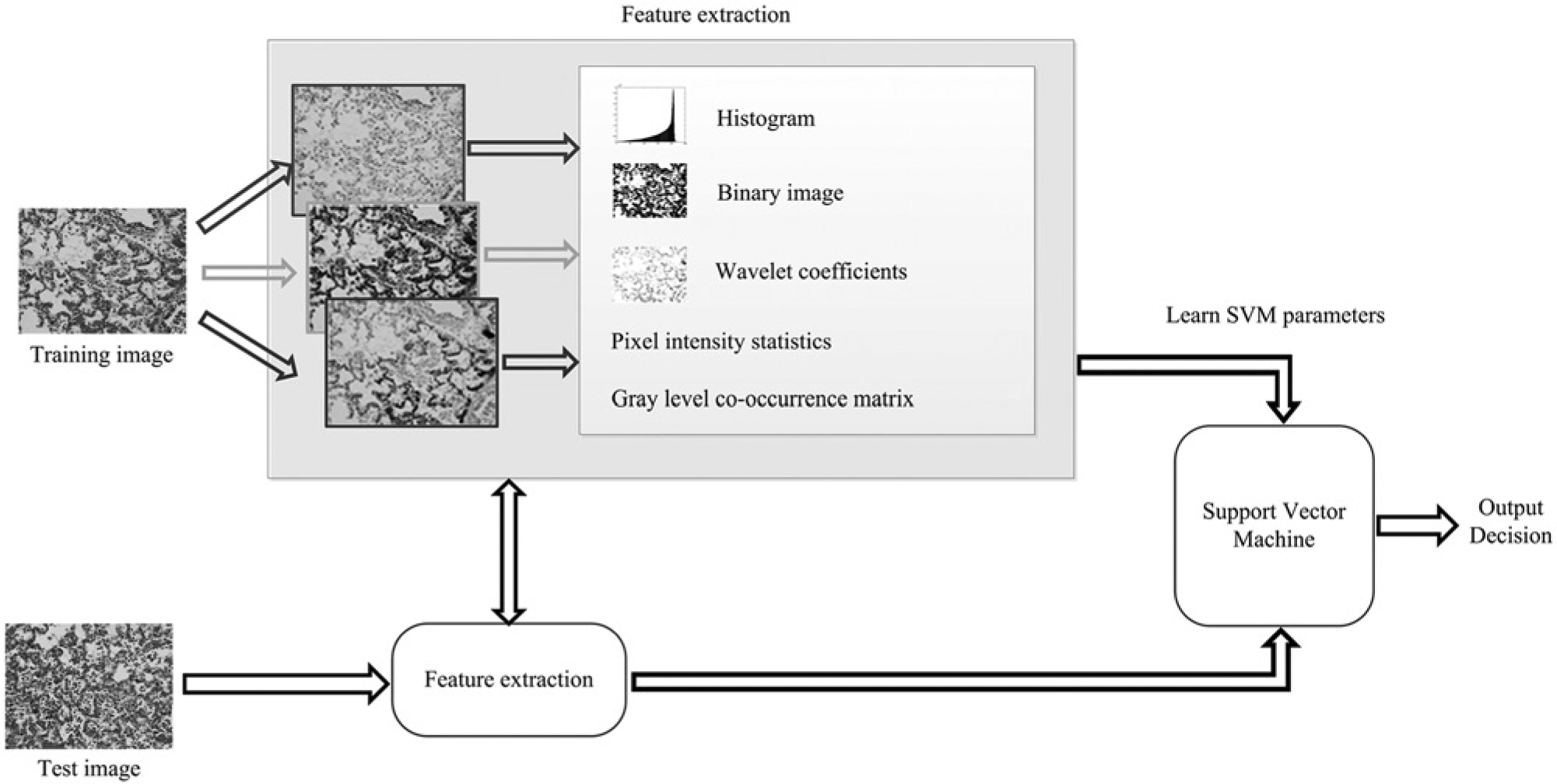
Proposed histologic image classification system using a support vector machine for the identification and classification of digitized photomicrographs.
A total of 60 images per category of normal tissue and tissue with known lesions (acute necrosis; acute suppurative inflammation) were obtained from bovine liver samples, 60 images per category of normal tissue and tissue with known lesions (acute suppurative inflammation) were obtained from bovine spleen and lung samples, and 60 images per category of normal tissue and tissue with known lesions (chronic inflammation) were obtained from bovine kidney samples (Fig. 1a–i). The samples of liver used as examples of acute necrosis exhibited areas of multifocal acute hepatocellular coagulative necrosis with no inflammatory cells present within the necrotic foci. The liver and spleen tissues used as examples of acute suppurative inflammation contained multifocal accumulations of neutrophils scattered throughout the tissues. The lung tissues used as examples of acute suppurative inflammation were characterized by coalescing intra-alveolar and intra-bronchiolar accumulations of neutrophils and small numbers of macrophages. Kidney tissues used as examples of chronic inflammation contained multifocal interstitial accumulations of lymphocytes and plasma cells within the cortical and medullary tissues. In the majority of cases, the tissue lesions present within the liver and spleen were secondary to septicemia produced by a variety of bacterial infections. Lesions present within the lung were secondary to a variety of pulmonary pathogenic bacteria, including Pasteurella sp. The chronic inflammation lesions present within the kidney were of unknown etiology. Each tissue was fixed in 10% buffered formalin, routinely processed, paraffin embedded, sectioned at 5 µm, and routinely stained with hematoxylin and eosin. The corresponding digitized color photomicrographs were obtained at 40× magnification in the standard TIF image format.
Two different experiments were performed in the current study, corresponding to the different number of images used to train the classifier. Initially, 10 images/category/organ were randomly identified from the original 60 slides and were used as training images for the SVM classifier. In the second case, 20 images/category/organ were chosen from the original 60 images and used as training images. In each case, however, a fixed set of 40 images/category/organ, different from the set of training images, was identified for testing.
In order to minimize selection bias sensitivity of classification performance to a specific choice of training images, the procedure was repeated 10 times, each time with a different random selection of images for training. Confusion matrices were employed to represent the accuracy of classification. Each row in the matrix identifies the correct category of the test image, and each column identifies the category as predicted by the SVM classifier. In the first experiment, the classifier-attained accuracies were found to be in the 74–90% range using 10 images/category/organ training images (Table 1). In the next experiment, 20 images/category/organ were identified as training images for the classification system. The average classification accuracies were noted to be in the 84–95% range (Table 1). In addition to the average classification accuracy, the standard deviation values were calculated by repeating the experiment with multiple random selections of training images. The relatively small standard deviation (approximately 4%) demonstrates that the proposed system is robust to the particular choice of 10 or 20 training images and that the classification accuracy values are statistically reliable.
Confusion matrix values for the correct histologic identification and classification of normal tissue and specific lesions within the liver, lung, kidney, and spleen.*
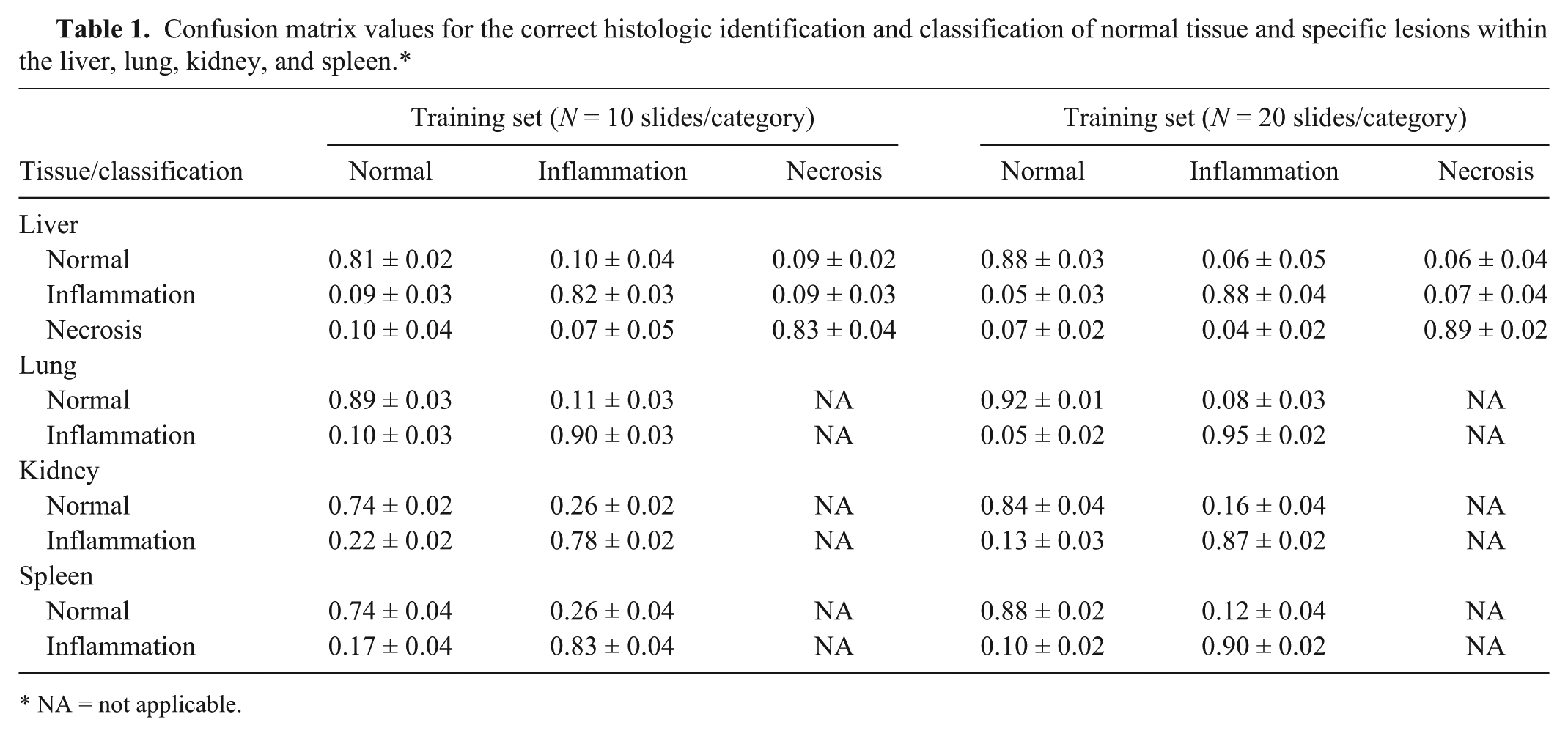
NA = not applicable.
Critical examination of histologic slides can be a time-consuming endeavor, with much of the pathologist’s time spent looking at normal tissue. In diagnostic and toxicology settings, the development of an accurate automated system to identify normal tissue and identify and classify specific histologic lesions would be of great benefit to the pathologist. Identification of normal tissue images by this system would eliminate such slides from further evaluation and enable the pathologist to spend more time in the critical evaluation of histologic slides containing tissue lesions. As opposed to medical image processing and evaluation techniques that are currently in widespread use in such fields as radiology, automated image processing for histology imaging is not routinely employed to any large extent. The use of similar systems used in radiology for the evaluation of histology images is somewhat problematic because of the different imaging techniques and image characteristics required for histology images as compared with radiology images, including the widely distributed tissue types, different staining colors, and larger image size (approximately 109 pixels) inherent in histology images. Although medical imaging techniques have been employed in the identification, grading, and classification of a variety of neoplasms, including urinary bladder, mammary, prostatic and pulmonary carcinomas, follicular lymphomas, and melanomas, the authors are not aware of previously published studies using CAD to specifically identify acute necrosis and inflammation in selected tissues similar to those used in this series of experiments.
The findings in the current study indicate that the classification performance of the SVM system for the correct identification of normal lung, spleen, liver, and kidney tissues and specific tissue lesions, including acute necrosis and inflammation (acute suppurative, chronic), was improved from an accuracy range of 74–90% to an accuracy range of 84–95% by using a larger set of training images. A small increase in the number of training images, from 10 to 20, produced a significant increase in the correct identification of normal tissue and specific tissue lesions. The greatest increase in accuracy was evident in the correct identification of chronic kidney inflammation (78–87%), normal kidney (74–84%), and normal spleen (74–88%). Identification accuracy rates of 92% and 95% were achieved in correctly identifying normal lung and acute suppurative lung inflammation, respectively, with limited training of the SVM system. The classification accuracy for the normal kidney images seems relatively low (74%) when 10 training images per category are used. There are 2 main reasons to explain this finding. First, as noted earlier, a common set of features that capture the essence of histologic images in order to distinguish between different medical conditions was used. In future investigations, design features will be customized for images from each organ, and it is believed that this can improve the classification accuracy. Second, improvements in accuracy are expected as the number of training images increases. An immediate validation of this conjecture is seen in Table 1 for the case when 20 training images per category are used, resulting in significant performance gain. In fact, in other domains such as radiology where automated image classification systems have been successfully deployed, a much larger number of training images is routinely required. The improved accuracy in the correct identification of both normal tissues and tissues with selected lesions with the use of increased numbers of training images and employing the CAD system developed in this preliminary study indicates a strong potential for this methodology to enable the pathologist to efficiently and accurately identify not only normal tissues but also specific tissue lesions in large numbers of tissue slides. As the system is further evaluated and perfected by using larger training and testing image sets, improved accuracy and a greater range of specific tissue lesions in additional types of organs may be accurately identified with confidence.
Footnotes
Declaration of conflicting interests
The author(s) declare no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by a research grant from the Pennsylvania Department of Agriculture (Hattel, ME 44113120).
