Abstract
We report the identification, pathogenesis, and transmission of a novel polyomavirus in severe combined immunodeficient F344 rats with null Prkdc and interleukin 2 receptor gamma genes. Infected rats experienced weight loss, decreased fecundity, and mortality. Large basophilic intranuclear inclusions were observed in epithelium of the respiratory tract, salivary and lacrimal glands, uterus, and prostate gland. Unbiased viral metagenomic sequencing of lesioned tissues identified a novel polyomavirus, provisionally named Rattus norvegicus polyomavirus 2 (RatPyV2), which clustered with Washington University (WU) polyomavirus in the Wuki clade of the Betapolyomavirus genus. In situ hybridization analyses and quantitative polymerase chain reaction (PCR) results demonstrated viral nucleic acids in epithelium of respiratory, glandular, and reproductive tissues. Polyomaviral disease was reproduced in Foxn1rnu nude rats cohoused with infected rats or experimentally inoculated with virus. After development of RatPyV2-specific diagnostic assays, a survey of immune-competent rats from North American research institutions revealed detection of RatPyV2 in 7 of 1,000 fecal samples by PCR and anti-RatPyV2 antibodies in 480 of 1,500 serum samples. These findings suggest widespread infection in laboratory rat populations, which may have profound implications for established models of respiratory injury. Additionally, RatPyV2 infection studies may provide an important system to investigate the pathogenesis of WU polyomavirus diseases of man.
Keywords
The Polyomaviridae family was revised in 2016 to accommodate a rapidly growing number of newly discovered viruses in diverse animal species (Polyomaviridae Study Group of the International Committee on Taxonomy of Viruses et al. 2016; DeCaprio and Garcea 2013). A new criterion for the creation of novel polyomavirus species is based on >15% difference in sequence identities of the large T antigen (LTAg) coding sequence compared to the most closely related species. Four genera, Alphapolyomavirus, Betapolyomavirus, Gammapolyomavirus, and Deltapolyomavirus, were designated to include all currently recognized polyomavirus species. The genera Alphapolyomavirus, Betapolyomavirus, and Deltapolyomavirus infect mammals, whereas Gammapolyomavirus infects birds. Seven distinct polyomaviruses (PyVs) have been described in rodents including bank vole PyV, common vole PyV, hamster PyV, Mastomys PyV, mouse PyV, mouse pneumotropic PyV, and Rattus norvegicus PyV (RnorPyV1). Phylogenetic analyses based on the LTAg sequences reveal hamster PyV, mouse PyV, and RnorPyV1 are related taxons of which mouse and hamster PyVs are assigned members of the Alphapolyomavirus genus. Mastomys and mouse pneumotropic PyVs are members of the Betapolyomavirus genera (Polyomaviridae Study Group of the International Committee on Taxonomy et al. 2016). Recently discovered RnorPyV1 (Ehlers et al. 2015) along with bank vole PyV and common vole PyV (Nainys et al. 2015) is currently unclassified. Based on LTAg and VP1 sequence analyses, RnorPyV1 is closely related to mouse PyV within the Alphapolyomavirus genus, and the vole PyVs are close relatives of human polyomavirus WU (WUPyV) within the Betapolyomavirus genus (Buck et al. 2016).
RnorPyV1, the first polyomavirus from rats to be genetically characterized, was detected in the spleens of asymptomatic feral Norway rats. However, the first report of a polyomavirus infection involving rats described a wasting disease in immunodeficient nude (Foxn1rnu) research rats (Ward et al. 1984). Intranuclear inclusions suggestive of viral infection were observed primarily in ductal and acinar cells of the parotid salivary gland and less often in epithelium of Harderian gland, laryngeal submucosal glands, and bronchi. The virus was determined to be a polyomavirus based on virus morphology from evaluation of electron micrographs of infected tissue and demonstration of reactivity of infected cells with antibodies to Simian Virus 40, a polyomavirus of macaques belonging to the Betapolyomaviridae genus. This rat polyomavirus was antigenically distinct from mouse PyV and mouse pneumotropic PyV as determined by serologic screening of serum antibodies from nude and related heterozygous nude rats. Since genetic sequences of this virus were not available, no comparison to RnorPyV1 was possible.
Herein, we report a naturally occurring polyomaviral disease in a breeding colony of immune-deficient rats lacking Prkdc and IL2rg gene function (F344-scid gamma or FSG; Mashimo et al. 2012). Using nucleic acid random library generation from infected tissues and unbiased viral metagenomic sequencing, we identified a novel polyomavirus, provisionally named Rattus norvegicus polyomavirus 2 (RatPyV2), which is distinct from RnorPyV1. RatPyV2 is phylogenetically related to WUPyV within the Betapolyomavirus genus. Histopathological lesions and in situ hybridization (ISH) studies verified that RatPyV2 targets epithelial cells of multiple tissues, including the respiratory tract, lacrimal glands, and salivary glands. These target organs are the same as those described in the first report of polyomavirus infection in nude rats. Similarly, WUPyV has been identified in respiratory epithelium from immunocompromised patients with pneumonia (Siebrasse et al. 2016). RatPyV2 infection also occurred in the uterus of female rats and in secondary sex glands of the male, which suggests potential for both vertical and sexual transmission. We determine that RatPyV2 infection occurs by both lateral (contact) transmission and experimental inoculation and causes similar patterns of disease in infected nude rats. Lastly, we identify anti-RatPyV2 antibodies and virus shedding in samples from immunocompetent research rats, which suggest widespread prevalence of viral infection.
Material and Method
Animals and Study Samples
Animal studies were carried out in accordance with the guidelines outlined in the National Research Council’s Guide for the Care and Use of Laboratory Animals and were approved by the Institutional Animal Care and Use Committee of the University of Missouri.
Male and female adult immune-deficient rats (Rattus norvegicus), lacking the Prkdc and IL2rg genes (F344-scid gamma or FSG; Mashimo et al. 2012), were from a breeding colony with a history of decreased fecundity and mortality. These rats as well as tissue and serum samples from cohort FSG rats (n = 12) and immune-competent F344 rats (F344 female rats bred to FSG rats and their heterozygous offspring [n = 16]) were submitted to IDEXX BioResearch and evaluated by a comprehensive panel of serological and PCR assays designed to detect known infectious agents of rats, including Sendai virus, pneumonia virus of mice, rat coronavirus, parvoviruses (Kilham rat virus, Toolan’s H-1 virus, rat parvovirus, and rat minute virus), reovirus, rat theilovirus, lymphocytic choriomeningitis virus, hantavirus, mouse adenovirus, Rattus novegicus polyomavirus 1 (RnorPyV1), Bordetella bronchiseptica, Filobacterium rodentium (cilia-associated respiratory bacillus), Clostridium piliforme, Corynebacterium kutscheri, Mycoplasma pulmonis, Pasteurella multocida, Pasteurella pneumotropica, Pneumocystis carinii, Pseudomonas aeruginosa, Salmonella spp., Streptococcus pneumoniae, Encephalitozoon cuniculi, ectoparasites, endoparasites, and enteric protozoa.
Four nude rats (Foxn1rnu, Rattus norvegicus, Envigo, Indianapolis, IN) were used for experimental infection studies. Cohort nude rats were determined to be polyomavirus free by testing salivary glands, lung, and feces by PCR using RnorPyV1 and RatPyV2-specific PCR tests. Three nude female rats were cohoused with 2 FSG female rats for 10 to 15 weeks. One nude rat was inoculated with clarified supernatant derived from homogenized FSG respiratory and glandular tissues containing viral inclusion bodies. The inoculated rat was euthanized for sample collection at week 12 postinoculation. From 1 cohoused and the inoculated nude rat, a comprehensive battery of tissues was collected for histopathologic examination and molecular testing using RatPyV2 PCR. Tissues from the other cohoused nude rats were collected for histopathologic examination or frozen at −80°C.
Rats were housed in autoclaved individually ventilated cages (Thoren Caging Systems Inc., Hazelton, PA). Irradiated commercial diet (Teklad diet, Envigo, Indianapolis, IN) and autoclaved acidified water were provided ad libitum. The room was maintained with constant environmental conditions (14:10-hr light:dark cycle, an average daily relative humidity of 30–70% and a temperature range of 20.6–21.7°C). All cage changes took place in a laminar flow hood.
Histopathology, ISH, and Immunohistochemistry
Tissues either collected at the time of necropsy or submitted from the FSG colony at the originating institution to IDEXX BioResearch were evaluated. Tissues were placed in 10% buffered formalin for 24 to 48 hr and processed routinely for paraffin embedding, sectioning, and staining with hematoxylin and eosin. Bones were decalcified in a 50% formic acid/20% sodium citrate solution prior to trimming and processing for hematoxylin and eosin slide preparation.
Two ISH probe sets were designed to detect RatPyV2. V-RatPyV2ID-LT contains 30 probe pairs complementary to nucleotides 1,835 to 31 of the LTAg coding region, covering approximately 1.8 kb of total sequence. A second probe set, V-RatPyV2ID-VP1, includes 19 paired probe sets covering the complementary sequence of nucleotides 7 to 1,032 in the VP1 coding region. Probes as designed hybridized to both messenger RNA and viral genomic DNA. Colorimetric ISH was performed manually on Superfrost Plus slides (Thermo Fisher Scientific, Pittsburgh, PA) using the RNAscope kit (Advanced Cell Diagnostics, Hayward, CA) according to the manufacturer’s instructions. Prior to probe hybridization, each 5 µm section of formalin-fixed, paraffin-embedded tissue was pretreated by incubation at 37°C for 10 min in 0.25% pepsin in 1× Tris-buffered saline (pH 2.0); pepsin activity was stopped by a 5-min incubation at 105°C. Scrambled probes of similar length and complexity as well as a probe designed to detect bacterial dapB were used as controls on all slides. A horseradish peroxidase–based signal amplification system was hybridized to the target probes followed by color development with 3,3-diaminobenzidine (DAB; Advanced Cell Diagnostics). Slides were counterstained with Meyer’s hematoxylin, and coverslips were mounted with xylene-based SHUR/Mount (Triangle Biomedical Sciences, Durham, NC). Positive staining was identified as brown, punctate dots. Slides were digitized using an Olympus VS120 scanner and a 40× objective with bright-field illumination.
For immunohistochemical assays, tissue sections of 4 to 5 µm thickness were cut, transferred to positively charged glass slides, dried overnight at 37°C, dewaxed, and rehydrated by routine methods. Sections were pretreated with proteinase K for 15 min for the CD68 assay, while those slated for Ki67 immunoreactivity were subjected to heat-induced antigen retrieval in citrate buffer (0.1M, pH 6.0) for 30 min. Tissue sections were treated with 5% bovine serum albumin (BSA) to block nonspecific protein binding prior to incubation with these primary antibodies: anti-CD68 (1:200; clone ED1, MCA341R; AbD Serotec, Raleigh, NC) for alveolar macrophages and anti-Ki67 (1:1,000; ab15580; Abcam, Cambridge, MA) for cell proliferation. Bound anti-CD68 antibody was detected with a biotinylated secondary antibody (Jackson ImmunoResearch Labs, West Grove, PA) and a streptavidin: horseradish peroxidase amplification system using DAB as the chromogen. Bound anti-Ki67 was detected with a biotinylated secondary antibody (4Plus Detection System, Biocare Medical, Concord, CA) and streptavidin: horseradish peroxidase amplification system (Biocare Medical) with a NovaRED chromagen (Vector Laboratories, Burlingame, CA). Slides were counterstained with Meyer’s hematoxylin and permanently coverslipped. CD68-positive alveolar macrophages were labeled with brown cytoplasmic deposits. Actively replicating cells expressing Ki67 had red cell nuclei.
Sample Processing for Next-generation Sequencing
A pool of tissue samples (∼25 mg) including lung, salivary glands, lacrimal glands, and prostate glands from each of the 4 FSG rats were immersed in 1 ml of Hank’s balanced saline solution and disrupted with a tissue homogenizer for 30 sec. The resulting homogenates were placed on dry ice for 5 min and thawed at room temperature. Freezing and thawing steps were then repeated 2 times. The tissue homogenates were then vortexed and centrifuged at 12,000 × g for 5 min (Li et al. 2015). The supernatant was then collected and passed through a 0.45-μm pores filter to exclude large cell and microorganism particles. The filtrate was treated with a mixture of nuclease enzymes including Turbo DNase (Ambion, Thermo Fisher Scientific), Baseline-ZERO (Epicentre, Madison, WI), Benzonase (Novagen, EMD Millipore, Bellerica, MA), and RNase One (Promega, Madison, WI) in DNase buffer (Ambion) at 37°C for 1.5 hr to further reduce the background from host and bacterial genetic material (Victoria et al. 2009), while the viral nucleic acids remain protected within their capsids (Allander et al. 2001). Nucleic acid extraction was performed by Maxwell 16 Viral Total Nucleic Acid Purification Kit (Promega) following manufacturer’s instructions.
Viral complementary DNA (cDNA) synthesis was performed by incubation of 10 µl extracted viral nucleic acids with 100 pmol of a primer containing a fixed 18 bp sequence plus a random nonamer at the 3′ end (GCCGACTAATGCGTAGTCNNNNNNNNN) at 72°C for 2 min. Then, 200U SuperScript III reverse transcriptase (Invitrogen, Carlsbad, CA), 0.5 mM of each deoxynucloside triphosphate (dNTP), 10 mM dithiothreitol, and 1× first-strand extension buffer were added to the mixture and incubated at 25°C for 10 min followed by 50°C incubation for 1 hr and 70°C for 15 min. The second-strand DNA synthesis was performed by incubation of reverse transcribed (RT) product with 5U Klenow Fragment (New England Biolabs, Ipswich, MA) at 37°C for 1 hr followed by 75°C for 20 min. The resulted cDNA was PCR amplified using 5 μl of the RT-Klenow DNA products and 2.5 μM primer consisting the fixed 18 bp portion of the random primer (GCCGACTAATGCGTAGTC) with 1U AmpliTaq Gold DNA polymerase (Life Technologies, Thermo Fisher Scientific, Waltham, MA), 2.5 mM MgCl2, 0.2 mM dNTPs, and 1× PCR Gold buffer in a reaction volume of 50 µl. The cycling temperature was used as follows: for initial denaturation, 1 cycle of 95°C for 5 min, continued by 30 cycles of 95°C for 30 sec, 55°C for 30 sec, 72°C for 1.5 min with final extension at 72°C for 10 min.
Five nanograms of randomly amplified DNA product was used as input for the construction of DNA libraries using Nextera XT DNA sample preparation kit (Illumina, San Diego, CA). The samples were then pooled, and the final library concentration was estimated using 35 PCR cycles and KAPA Library Quant Kit (Kapa Biosystems, Wilmington, MA) following manufacturer’s instructions. The resulting library was then sequenced on HiSeq 3000 platform (2 × 150 cycle).
Bioinformatics Pipeline
Paired-end next-generation sequencing reads of 250 bp generated by HiSeq were debarcoded using vendor software from Illumina and then merged using Fast Length Adjustment of SHort reads (FLASH; Magoc and Salzberg 2011). Host reads and bacterial reads were subtracted using bowtie2 (Langmead and Salzberg 2012). Low sequencing quality tails were trimmed using Phred quality score 10 as the threshold. Adaptor and primer sequences were trimmed using the default parameters of VecScreen. The cleaned reads were de novo assembled using EnsembleAssembler (Deng et al. 2015). The assembled contiguous sequences along with singlets were aligned to Blood Systems Research Institute in-house viral proteome database using Basic Local Alignment Search Tool (BLASTx) using E value cutoff .001. A web-based graphical user interface was used to access the results along with taxonomy information and processing meta-information.
Phylogenetic Analysis
The complete polyomavirus reference viral nucleic acid and amino acid sequences for LTAg and VP1 were obtained from GenBank. Sequence alignments were performed using MUltiple Sequence Comparison by Log-Expectation (MUSCLE) with the default settings, and phylogenies were generated by neighbor-joining method (p-distance model) integrated in MEGA package, version 6.0 (Tamura et al. 2013). Statistical significance of tree topologies was evaluated by 1,000 bootstrap resampling iterations, and a bootstrap value of 50% was used as the cutoff point for cluster analysis.
Serological Assay
The RatPyV2 VP1 gene was amplified by PCR using LongAmp polymerase (New England Biolabs) according to the manufacturer’s protocol. Each 25 μl reaction mixture contained 2.5U of polymerase, 100 ng of template, and primers RatPyV2 VP1 forward (5′-GTCCAGGGTGACATGC-3′) and RatPyV2 VP1 reverse (5′-TCAGCCCTGCACATT-3′). Thermocycling parameters consisted of an initial denaturation step (94°C, 30 sec) followed by 35 cycles of denaturation (94°C, 30 sec), annealing (59°C, 45 sec), and extension (65°C, 1 min).
The purified RatPyV2 VP1 PCR product was digested and directionally cloned into the His-tagged fusion vector pOET1N_6xHis (Oxford Expression Technologies, Oxford, UK). Plasmids were sequenced to insure proper insertion and then used to construct recombinant baculovirus expression vectors. High-titer stocks of recombinant baculovirus were generated in Spodoptera frugiperda (Sf-9) insect cells (Invitrogen). Standard insect cell and baculovirus propagation protocols were followed. Pilot experiments were performed to determine the kinetics for maximal production of each recombinant protein. To generate large quantities of RatPyV2 VP1 protein for use as antigens, 1-L shaker flasks were seeded with 500 ml of High Five insect cells (Invitrogen) at a density of 1 × 106 cells per ml in insect cell medium (Hyclone Laboratories, Logan, UT). At 65 hr postinoculation, the cells were pelleted by centrifugation at 10,000 × g for 15 min at 4°C, suspended in 50 ml of phosphate-buffered saline (PBS), and disrupted by six 30-sec cycles of sonication (Thermo Fisher Scientific) on ice. Insoluble material was pelleted by centrifugation at 10,000 × g for 15 min at 4°C. The clarified supernatant was harvested and purified over a cesium chloride gradients and centrifuged at 26,500 rpm for 16 hr at 4°C. Following centrifugation, the gradient fraction containing the recombinant protein was visualized and harvested. The recovered protein was buffer exchange against PBS. The RatPyV2 VP1 recombinant protein was verified for correct size, purity, and antigenicity by polyacrylamide gel electrophoresis and Western blot. The concentrations of the recovered proteins were determined by Pierce 660 nm Protein Assay procedure (Pierce Endogen, Rockford, IL), and the proteins were stored at −80°C until use.
A RatPyV2 multiplex fluorescent immunoassay (MFI) was created by covalently coupling purified recombinant RatPyV2 VP1 to 5 × 106 carboxylated polystyrene microspheres (Luminex Corp., Austin, TX) according to manufacturer’s recommended protocols. Microspheres were stored at 4°C in the dark until use. Testing of rat serum samples for anti-RatPyV2 VP1 antibody was performed using the Luminex LX200 (Luminex Corp.). Microspheres coupled to RatPyV2 VP1 protein were suspended by vortexing and sonication. Approximately 1,500 microspheres in 100 μl PBS-BSA (PBS, 1% BSA, 0.05% sodium azide; Sigma-Aldrich, St. Louis, MO) along with the test serum at a final dilution of 1:200 were added to each well of an AcroPrep 96-well filter-bottom plates (Pall Corporation, Port Washington, NY). Plates were covered and incubated for 60 min on an orbital shaker at 400 rpm at room temperature in the dark. Each well was washed 4 times by adding 100 μl of PBS-BSA, shaking at 900 rpm for 1 min, and then removing the fluid with a vacuum manifold. The microspheres were suspended in 100 μl of PBS-BSA containing 1:200 diluted F(ab′)2 fragment goat anti-rat immunoglobulin G (IgG; heavy plus light chains [H+L]) phycoerythrin-conjugated secondary antibody (Jackson ImmunoResearch Laboratories, Inc., West Grove, PA). The plate was covered and incubated for 60 min on an orbital shaker at 400 rpm at room temperature in the dark. Plates were washed 4 times as described above, and microspheres were suspended in 100 μl of fresh PBS-BSA and analyzed on the Luminex workstation. For each serum sample, the median fluorescent intensity of 100 RatPyV2 VP1-coated microspheres was recorded. Twenty known uninfected and 16 RatPyV2-infected rat serum samples, as determined by Western blot as the reference standard, were used to plot a receiver operating characteristic curve for the RatPyV2 VP1 MFI. Fluorescence values corresponding to >99% sensitivity and >99% specificity were used as the threshold values to discriminate negative or positive test results in all subsequent serum samples tested by MFI.
RatPyV2 PCR
Total nucleic acids were extracted from tissues and feces with standard protocols using a commercially available platform (One-For-All Vet Kit, Qiagen, Valencia, CA). The RatPyV2 PCR tests were based on the IDEXX BioResearch proprietary service platform (IDEXX Laboratories, Inc., Westbrook, ME). Briefly, the RatPyV2 real-time PCR assay targets a region of the LTAg gene with sequence conservation among strains of other PyVs but was designed to be specific for RatPyV2 with 5 mismatches in each primer and 7 mismatches in the 6-carboxyfluorescein/tetramethylrhodamine (FAM/TAMRA)-labeled hydrolysis probe as compared with the closest related polyomavirus (KR612373). Hydrolysis probe-based real-time PCR assays targeting a mammalian gene (18S rRNA) or bacterial gene (16S rRNA) were used to ensure DNA recovery and the absence of PCR inhibitors in nucleic acid extracted from tissues and feces, respectively. Real-time PCR was performed with standard primer and probe concentrations using a commercially available mastermix (LC480 ProbesMaster, Roche Applied Science, Indianapolis, IN) on a commercially available real-time PCR platform (Roche LightCycler 480). To confirm the specificity of the RatPyV2 LTAg PCR assay, another PCR assay targeting the RatPyV2 VP1 gene was designed and used to retest positive RatPyV2 rat samples from 2 geographically different locations. These RatPyV2 VP1 amplicons were sequenced and compared to that of the RatPyV2 type strain. For quantitative analysis, standard curves were generated using log fold dilutions of known amounts of synthetic RatPyV2 LTAg and 18S rRNA genes. The mean crossing point value of 3 technical replicate PCR assays was used to calculate the copy number of RatPyV2 LTAg. Virus genome copy number was then normalized to 1 × 106 copies of the 18S rRNA gene.
Results
Pathology
Immune-deficient FSG rats that are functionally devoid of B cells, T cells, and natural killer (NK) cells had a history of adult-onset wasting, low fecundity, and associated mortality of dams and pups. Clinical signs included emaciation, hunched posture, and/or dyspnea. Necropsy examination of FSG rats (n = 12) revealed severe atrophy of the submandibular, sublingual and parotid salivary glands, the exorbital lacrimal glands, and Harderian glands. Histopathological examination of hematoxylin and eosin-stained tissue sections identified multifocal ductal and acinar degeneration with acinar epithelial dysplasia and hyperplasia and interstitial mixed inflammatory cell infiltrates. Large basophilic homogeneous intranuclear inclusions were within acinar and ductal epithelium of the salivary (Figure 1A), exorbital lacrimal, Harderian, thyroid, and prostate glands as well as endometrium and endometrial glands. Intranuclear inclusions were common in bronchiolar epithelium (Figure 1B) and pneumocytes and less frequently observed in renal pelvic urothelium, nasolacrimal ductal epithelium, and nasoturbinate respiratory and subepithelial glandular epithelium. No epithelial nuclear inclusions were observed in other abdominal organs, brain, pituitary gland, cardiac or skeletal muscle, or gonads. Cellular distribution and inclusion characteristics were consistent with an epitheliotropic DNA virus.
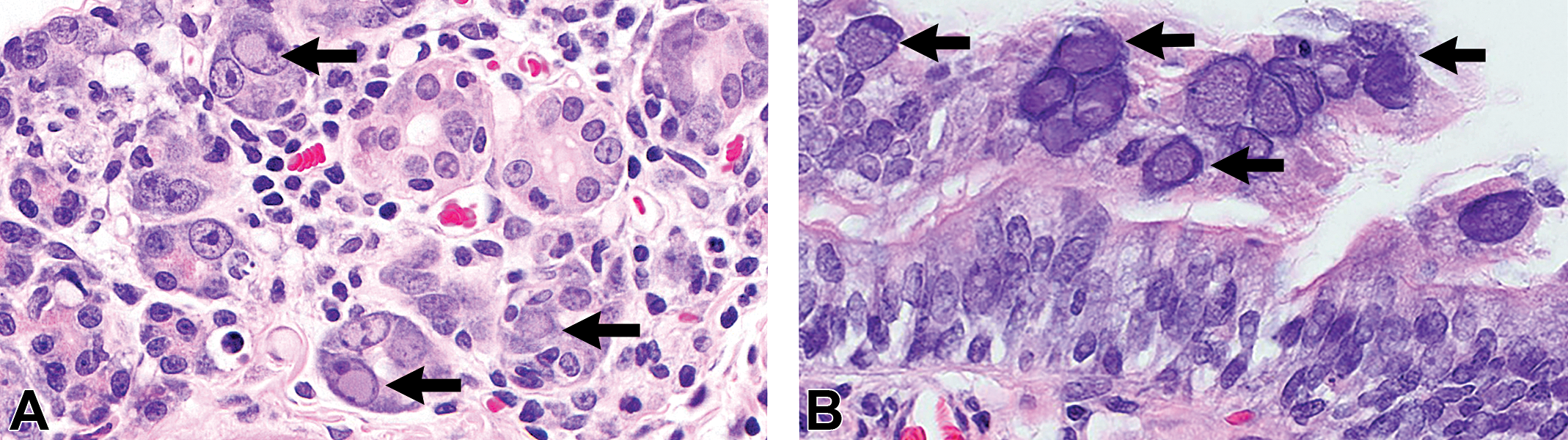
Intranuclear inclusions (arrows) are within (A) acinar and ductal cells of parotid salivary gland and (B) bronchiolar epithelium in the lung. Hematoxylin and eosin-stained sections, original objective at 40×.
Antibodies to viral and bacterial pathogens of rats were not detected in serum samples from cohort immune-competent F344 rats (female rats that were bred to FSG male rats and their heterozygous offspring, n = 16). Similarly, testing of tissues from the immune-deficient FSG rat (n = 1) with PCR assays designed to detect known viral and bacterial pathogens of rats yielded negative results. Samples also tested negative with RnorPyV1 MFI and PCR assays. Pneumocystis carinii was detected by PCR in lung samples from FSG rats and by MFI in serum samples from the F344 breeder rats. These findings were considered incidental to the clinical disease spectrum. The histopathologic lesions in the FSG rat lungs, associated with the fungal infection, were minimal to mild, with few alveolar macrophage aggregates associated with foamy eosinophilic material variably containing argyrophilic cysts and mild thickening of alveolar septa. Cumulatively, the histopathologic findings and the diagnostic test results suggested a novel viral infection of the FSG rats.
Viral Metagenomics
From pooled tissues from each of the 4 affected FSG rats, nucleic acids were extracted, randomly amplified, and subjected to deep-sequencing analyses. Using protein sequence similarity searches against all known eukaryotic viral proteins (VPs), polyomavirus sequences were detected in all 4 pooled tissue samples and were used to assemble the entire 5,108 bp genome. No other viral sequences were detected. Open reading frame (ORF) analysis revealed a genome organization consistent with the Polyomaviridae family, with an early region encoding the putative small T antigen and LTAg and a late region coding for the structural viral proteins VP1, VP2, and VP3. A putative ORF at map position 4462 to 4112 was predicted to encode an Alternate frame of the Large T Open reading frame (ALTO) gene (Figure 2A). The proposed name Rattus norvegicus polyomavirus 2 (RatPyV2, GenBank ID NC_032005.1) is applied for reference. Phylogenetic analysis of the LTAg (Figure 2B) and VP1 (Figure 2C) amino acid sequences demonstrates that RatPyV2 is divergent from RnorPyV1 of the Alphapolyomavirus genus and clusters with bank and common vole PyVs and human polyomavirus 4 (WUPyV) of the Betapolyomavirus genus (Gaynor et al. 2007; Nainys et al. 2015). Importantly, RatPyV2 is identical to another rat polyomavirus that was independently and recently described as causing disease in genetically altered rats with X-linked severe combined immunodeficiency (Rigatti et al. 2016). RatPyV2 shares 61.87% nucleotide identities of the LTAg coding sequence with the most closely related virus species qualifying it as a novel species based on criteria from the International Committee on Taxonomy of Viruses (Polyomaviridae Study Group of the International Committee on Taxonomy of et al. 2016; Table 1). The genome coverage depth is shown in Figure 2D.
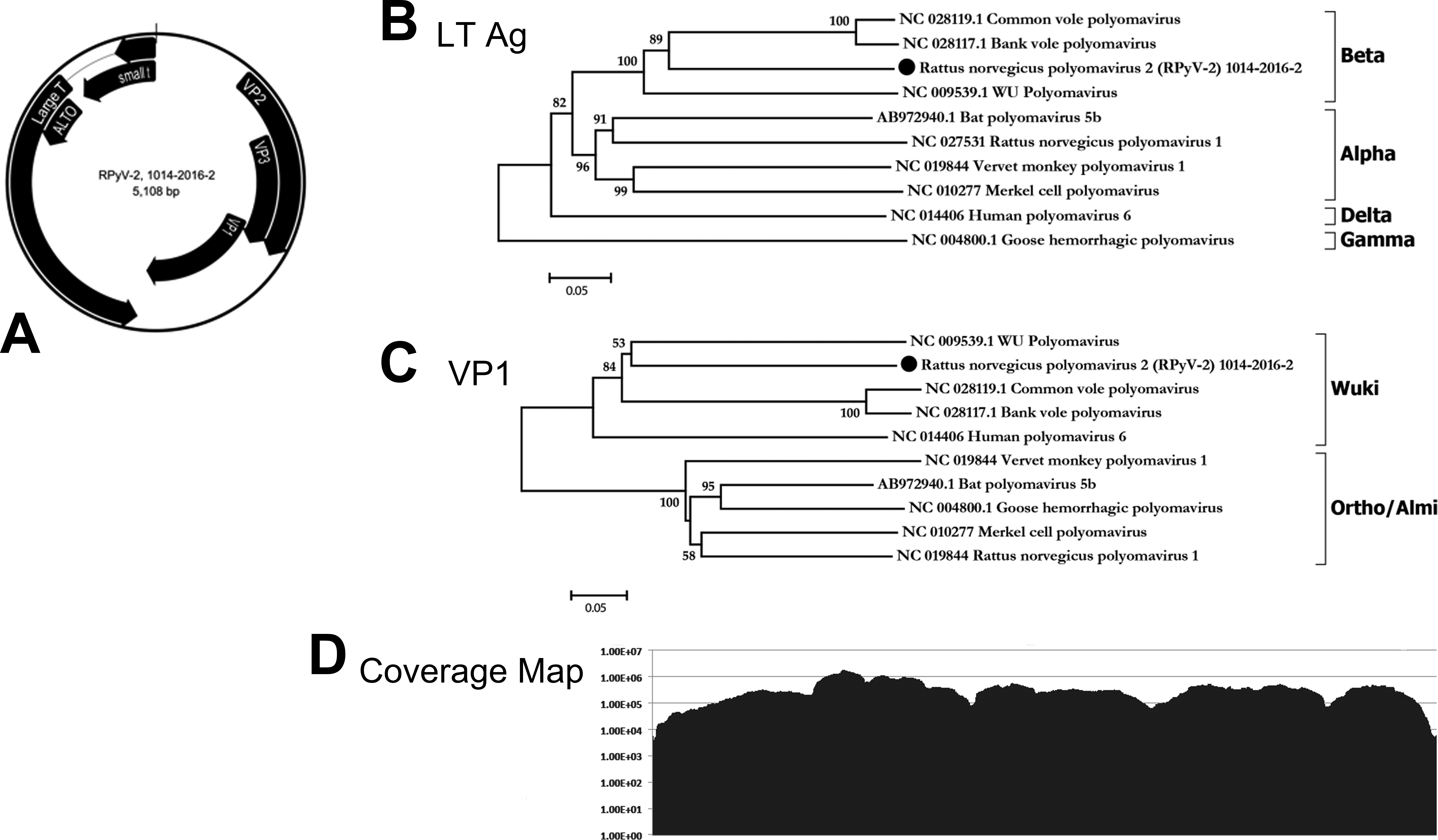
Genome organization of Rattus norvegicus polyomavirus 2 (RatPyV2) with putative coding regions for the structural (VP1 and VP2 proteins) and the nonstructural (small T antigen and large T antigens [LTAg]) are separated by a noncoding regulatory region (A). Phylogenetic analysis of the (B) LTAg and (C) VP1 proteins demonstrates that RatPyV2 is a novel Betapolyomavirus whose VP1 clusters in the Wuki clade. (D) The number of sequence reads supporting each base of the RatPyV-2 genome.
Percentage of Nucleotide Identities of the LTAg Coding Sequence.
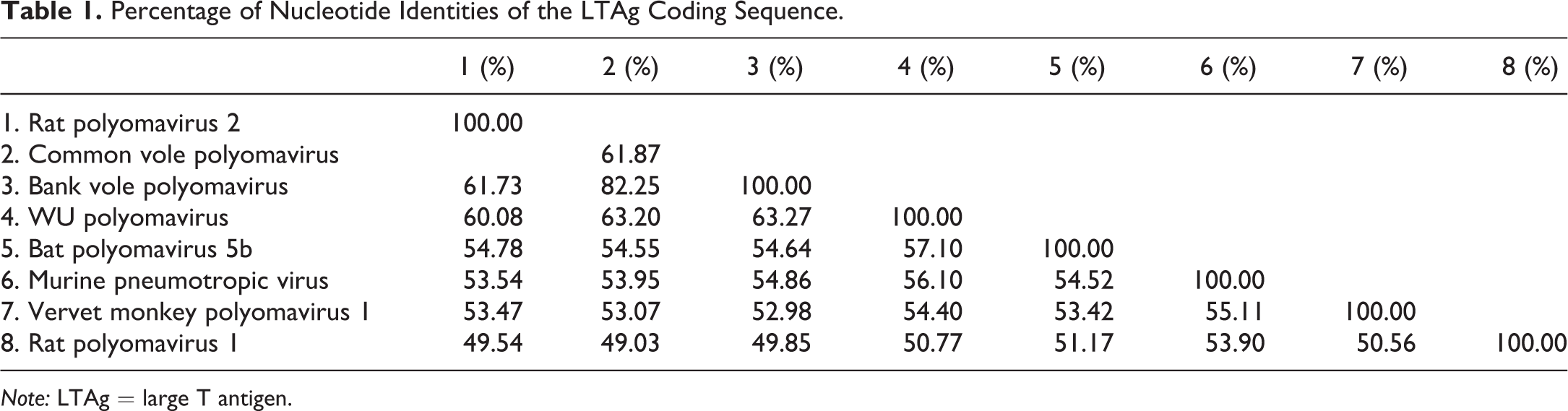
Note: LTAg = large T antigen.
Retrospective Study of RatPyV2 Tissue Distribution
Representative tissues from 3 naturally infected FSG rats from the original colony and 1 nude rat that was cohoused with FSG rats were selected for RatPyV2 ISH testing. Tissues tested in all animals included submandibular and parotid salivary glands, exorbital and Harderian lacrimal glands, lung, and kidney. Examined in a subset of animals was uterus (n = 3), prostate/seminal vesicle/coagulating glands (n = 1), anterior and posterior nasal cavity (n = 1), brain (n = 3), liver (n = 2), trachea (n = 2), esophagus (n = 2), stomach (n = 2), intestine (n = 3), and skin (n = 1).
Both V-RatPyV2ID-LT and VP probes gave similar results. Epithelial cells in multiple organ systems were targeted. In order of decreasing signal intensity, viral nucleic acid was detected in glands > respiratory > reproductive > urinary > gastrointestinal tissues. Within the upper respiratory tract, probe hybridization was present in nuclei of scattered respiratory and transitional epithelia, vomeronasal epithelium, and subepithelial glands. Infected cells were abundant in the trachea, bronchi, and bronchioles, where RatPyV2-infected cells, often with intranuclear inclusions, bulged and sloughed into airways (Figure 3A and B). In patchy regions of alveoli, viral laden cells were present lining alveolar walls and free within the airspaces. In the normal rat lung, turnover of bronchiolar epithelium (as assessed by Ki67 expression) is quite low (between 1% and 2% of cells). In the RatPyV2-infected animals of this study, replicating cells were 15% to 30%, and RatPyV2-infected cells were part of the dividing cell population (Figure 3C). Alveolar macrophages, labeled with antibody to CD68, were negative for viral inclusions, which suggest that virus-infected cells in alveolar spaces and walls are pneumocytes (Figure 3D). In FSG rats, all glands were heavily targeted (Figure 4A and B), and the pattern of hybridization in the parotid, exorbital, and Harderian was similar, albeit attenuated, in the nude rat. Male reproductive tissues analyzed were limited to 1 FSG rat, with virus heavily present within the epithelium lining the prostate (Figure 4C and D), and less abundant, but present as well, within the seminal vesicles, coagulating gland, and urethral lining. Many sloughed cells present within the urethral plug contained detectable virus. Within the uterus of a FSG female rat, scattered surface and glandular epithelial cells contained viral nucleic acid (Figure 4E and F). Probe hybridization was not observed in brain; and despite the general epitheliotropism, it was not observed in intestine, liver, or skin.
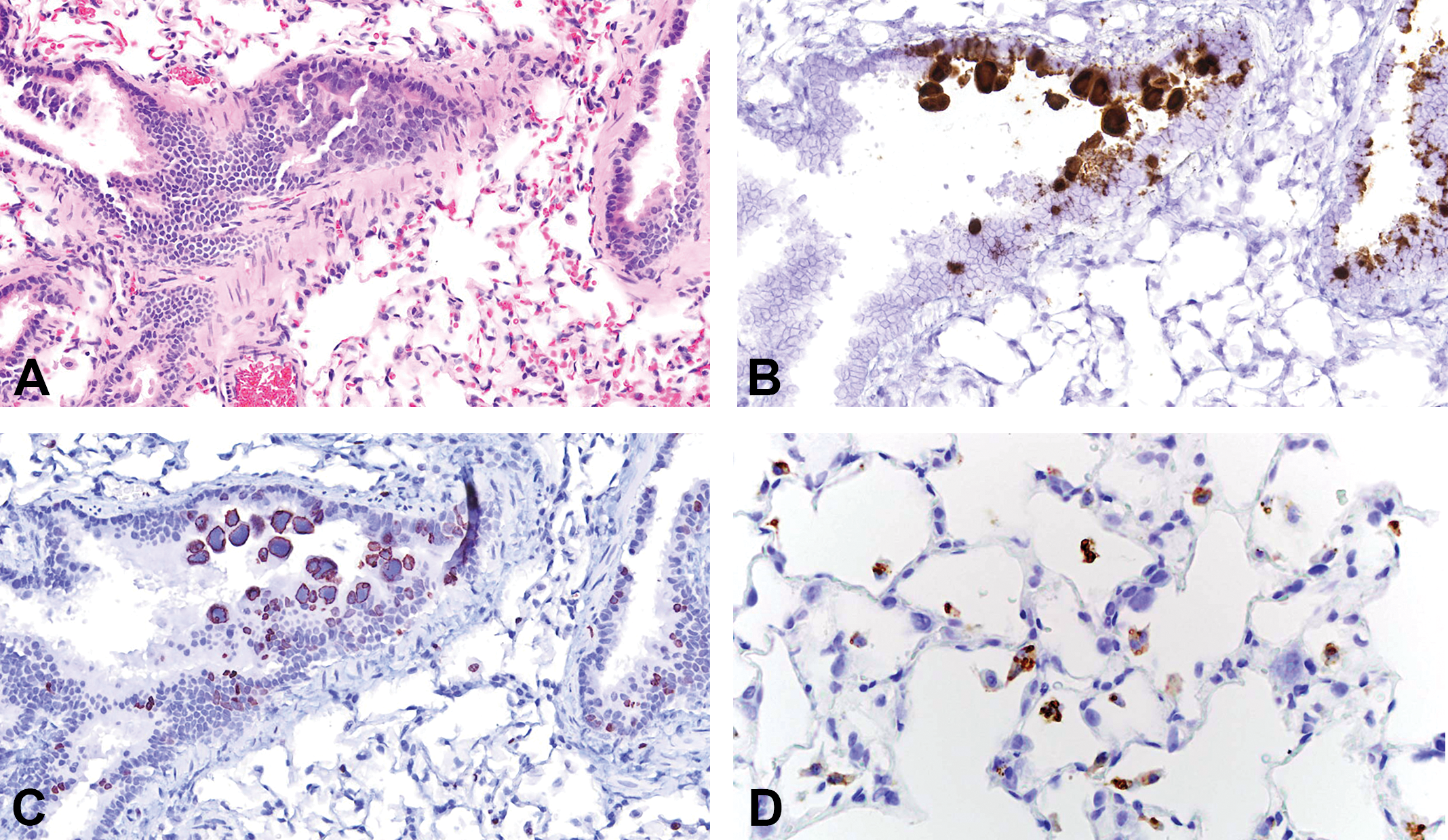
(A) On hematoxylin and eosin-stained lung sections, homogeneous basophilic intranuclear viral inclusions were present in bronchiolar epithelial cells. (B) Lung tissue incubated with nucleic acid probes to Rattus norvegicus polyomavirus 2 (RatPyV2) large T antigen via in situ hybridization verified RatPyV2 localization in nuclei of bronchiolar epithelium. (C) Bronchiolar epithelial turnover is increased in rats with RatPyV2 pulmonary infection as assessed by expression of Ki67, a nuclear protein used as a marker for cell proliferation. In infected nuclei, chromatin marginated by viral inclusions is labeled by anti-Ki67 antibodies (anti-Ki67 immunohistochemistry). (D) In pulmonary tissue, hypertrophic pneumocytes bearing intranuclear inclusions are not labeled by antibody to the macrophage antigen CD68, which verifies that alveolar macrophages do not support viral replication. Original objective at 40×.
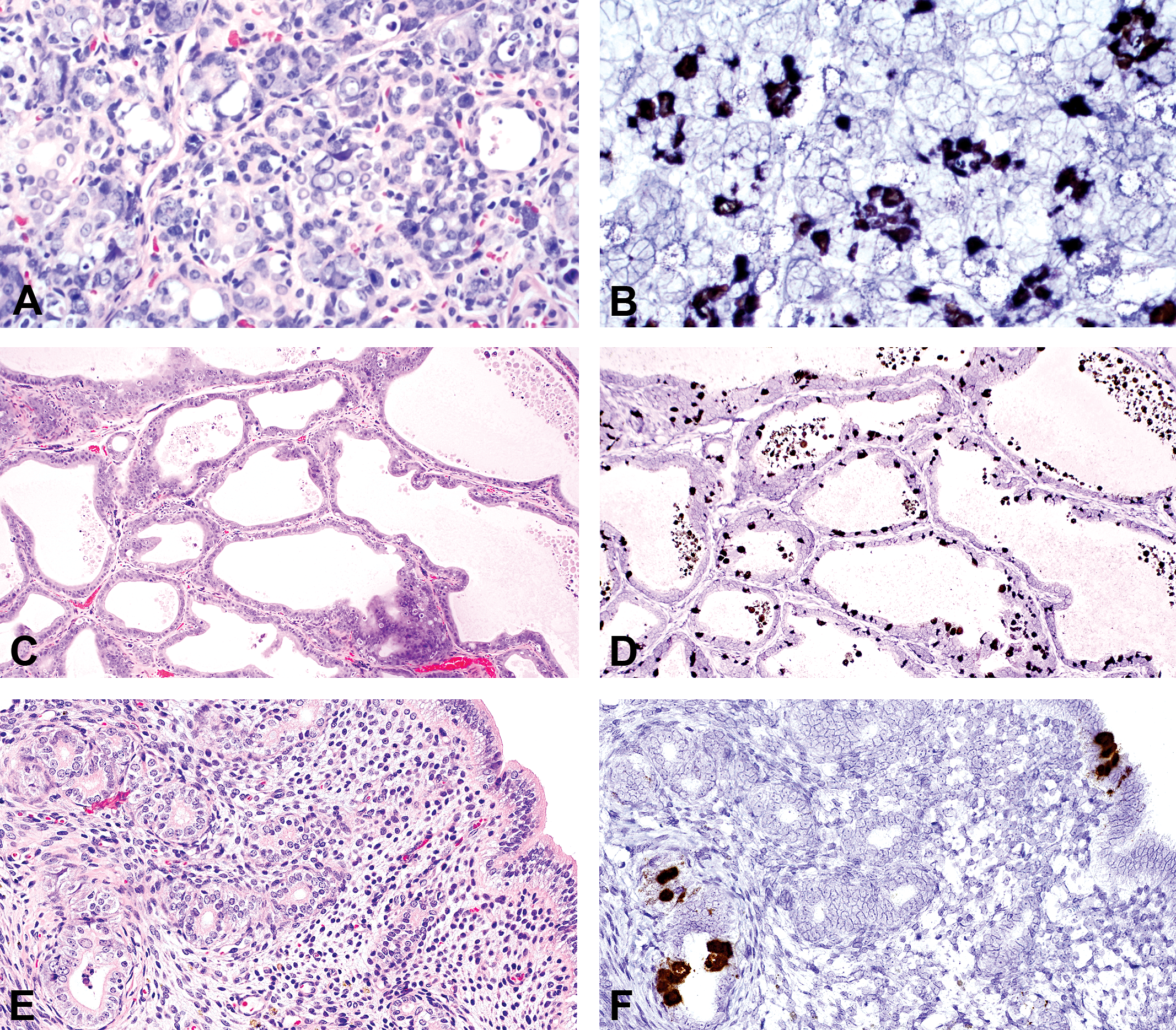
Rattus norvegicus polyomavirus 2 (RatPyV2) replicated in acinar and ductal epithelium of the parotid salivary gland (A, B), epithelium of the ventral prostate gland (C, D), and endometrial and glandular epithelium of the uterus (E, F). A, C, and E are hematoxylin and eosin-stained tissues sections, and B, D, and F are sister sections probed with nucleic acid probes to RatPyV2 large T antigen. Original objective at 40×.
RatPyV2 Transmission, Tissue Distribution, and Prevalence
RatPyV2 transmission was evaluated in nude rats by cohousing 3 nude rats with infected FSG rats for 10 to 15 weeks or by intraperitoneal plus intranasal administration in 1 nude rat of clarified tissue homogenates from RatPyV2 naturally infected FSG rats. At week 15 postexposure, 1 cohoused nude rat and the experimentally inoculated rat were euthanatized, and paired tissue samples were collected for histopathologic exam and evaluation of viral load by PCR. Tissues from the other 2 cohoused nude rats (cohoused for 10 to 14 weeks) were examined by histopathologic exam only. Viral-induced lesion distribution as determined by histopathologic evaluation is listed in Table 2. Tissue changes in the experimentally inoculated nude rat included exorbital lacrimal gland degeneration without presence of viral inclusions and inclusion body formation in the bronchiolar epithelium and pneumocytes. Further, intranuclear inclusions were observed in the endometrial glandular epithelium of the uterus. Histopathologic evaluation of tissues from the nude rat cohoused with FSG rats for 15 weeks demonstrated multifocal acinar and ductal epithelial degeneration with intranuclear inclusion body formation in the parotid salivary and exorbital lacrimal glands. Similar viral-induced tissue changes were observed in the salivary and lacrimal glands from the other 2 cohoused nude rats. Viral inclusions were observed in bronchiolar epithelium in only 1 of the 3 cohoused rats and none in the lung or uterus, which are findings in contrast to those observed in the experimentally inoculated rat. Both inoculated and cohoused rats had intranuclear inclusions in the respiratory epithelium of the nasal cavity.
RatPyV2 Viral Load and Lesion Prevalence in Tissues from Experimentally Infected Nude Rats.
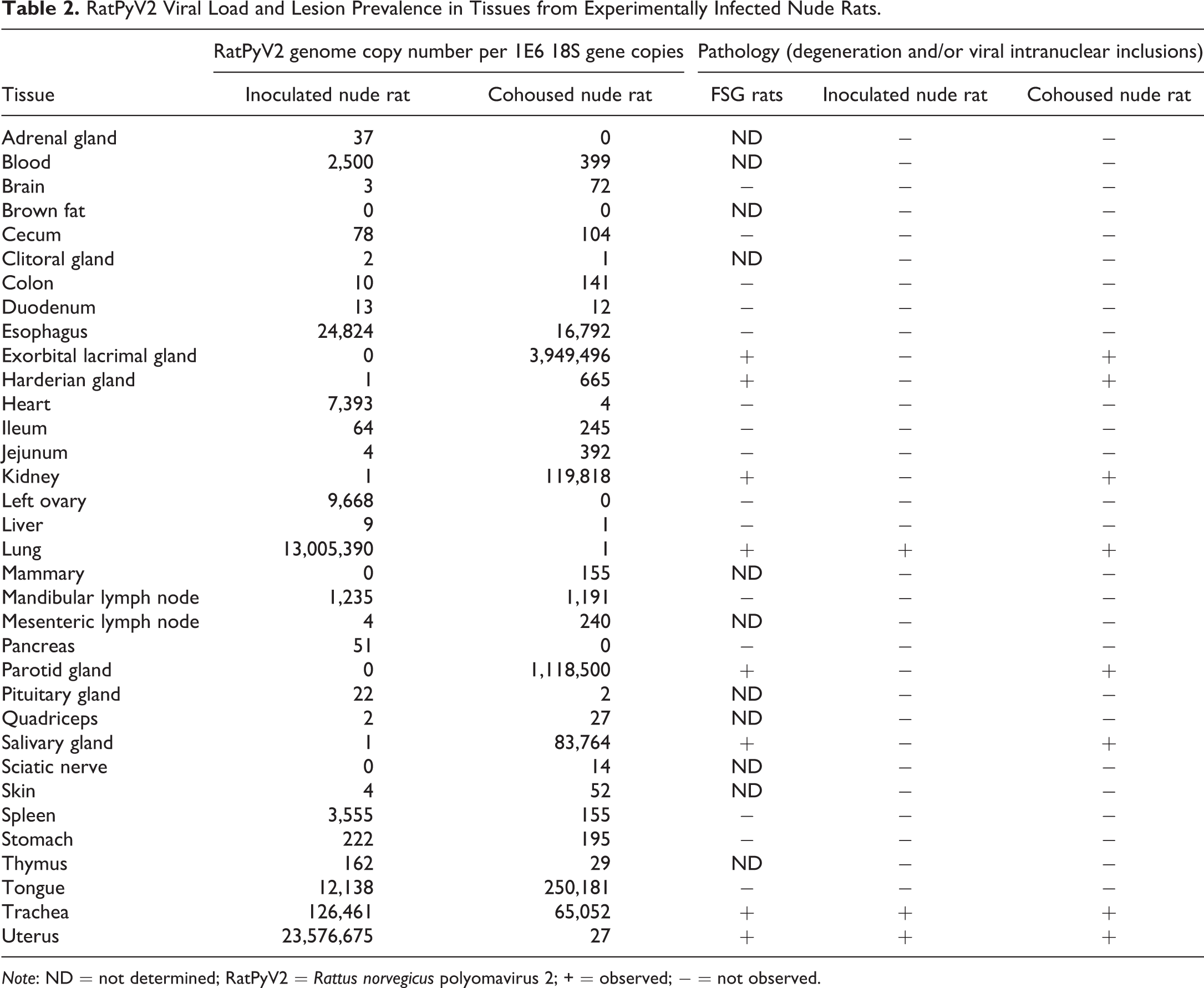
Note: ND = not determined; RatPyV2 = Rattus norvegicus polyomavirus 2; + = observed; − = not observed.
To determine viral load in the tissues, paired tissues samples from 2 of the experimentally infected nude rats were evaluated by quantitative real-time PCR. RatPyV2 genome copy number per tissue sample was normalized to 1 × 106 copies of the 18S rRNA gene, and results are listed in Table 2. The experimentally inoculated nude rat had the highest viral load in the uterus followed by the lung, trachea, esophagus, and tongue. The cohoused nude rat had the highest viral load in the exorbital lacrimal and parotid salivary glands followed by the tongue, kidney, and submandibular/sublingual salivary glands. Cumulatively, the tissue viral load and distribution data correlated with the histological findings of inclusion body formation and degenerative changes in tissues from the naturally infected FSG rats (Table 2).
To determine the prevalence of RatPyV2 in biomedical research rats, sera (n = 1,500) and fecal samples (n = 1,000) submitted to IDEXX BioResearch for routine health monitoring were selected for RatPyV2 surveillance testing. Clinical samples were randomly selected and represented male and female rats of various ages, stocks/strains, and diverse geographical regions. RatPyV2 serum antibody was present in 32% of samples evaluated by MFI using the RatPyV2 VP1 antigen. RatPyV2 genome was detected in 0.70% of fecal samples evaluated by RatPyV2 LTAg PCR. A 446 bp region of the VP1 gene from 2 RatPyV2 PCR positive fecal samples from different geographical locations was sequenced. The nucleotide sequences were 100% identical to each other and 99% identical (441 of 446 identical bases) to the type strain RatPyV2, indicating that there are minor nucleotide differences in the VP1 gene among distinct RatPyV2 isolates (data not shown).
Discussion
We describe a naturally occurring disease in severely immune-compromised FSG rats caused by RatPyV2. FSG rats had experienced reduced fecundity with shortened life spans and displayed clinical signs of emaciation, hunched posture, and/or dyspnea. Intranuclear inclusion bodies were observed in multiple tissues, most often within epithelial cells of the respiratory tract and glandular tissues and also in the epithelium of the uterus and male secondary sex glands. Although a similar disease with intranuclear inclusion bodies attributed to polyomavirus was first reported in immunodeficient rats over 30 years ago, the lack of successful viral characterization impeded the ability to study polyomaviral pathogenesis in rats (Ward et al. 1984). In this study, metagenomic analysis yielded the complete 5,108 bp genome of a novel viral species in the Betapolyomavirus genus. No other viral sequences were detected. RatPyV2 is divergent from RnorPyV1 from the Alphapolyomavirus genus. RatPyV2 shares 61.87% nucleotide identities of the putative LTAg coding sequence with its nearest neighbor in the Betapolyomavirus genus, indicating that RatPyV2 is a novel virus species. Phylogenetic analysis of the VP1 protein shows that it clusters in the Wuki polyomavirus clade along with WUPyV.
RatPyV2 from FSG rats is genetically identical to an independently identified polyomavirus from X-linked severe combined immunodeficient (X-linked SCID) rats with a null IL2rg gene mutation (Rigatti et al. 2016). Infected X-linked SCID rats expressed a similar clinical disease history of adult-onset wasting, decreased fecundity and an identical spectrum of virus-infected tissues. Polyomaviral inclusions in tissue sections were initially identified by immunoreactivity with a cocktail of monoclonal antibodies known to recognize all human PyVs. Polyomavirus identity was verified using rolling circle DNA amplification with PCR primers to conserved polyomavirus VP1 and LTAg sequences, genome walking, and sequence analyses. Viral sequences were identical to those determined in our investigation. The X-linked SCID and FSG rats were obtained from the same rodent repository, which suggests the severely immunocompromised rats may have acquired this polyomavirus from a common source.
RatPyV2 was present in the respiratory tract of all 12 index case FSG and the experimentally exposed (directly inoculated and cohoused) nude rats. Intranuclear inclusion body formation was evident in the nasal epithelium of multiple FSG rats and in the experimentally inoculated and cohoused nude rats. In lungs, from all FSG rats and the experimentally inoculated nude rat were intranuclear inclusions of the bronchiolar epithelium. Similarly, all FSG rats and the cohoused nude rat had intranuclear inclusions and variable degrees of degeneration and repair in the submandibular salivary gland, parotid salivary gland, Harderian gland, and/or exorbital lacrimal glands. Consistent with the histologic and ISH data, quantitative PCR demonstrated that RatPyV2 load was the highest in the exorbital lacrimal gland followed by the parotid salivary gland, tongue, and trachea in the cohoused nude rat. The viral load in the lung of the experimentally inoculated rat was high with little to no virus detected in the glandular tissues. The differences in viral tissue load between the cohoused and experimentally inoculated nude rats may be due to the route of RatPyV2 exposure. Given these data and an understanding of the social and allogrooming behaviors of rats, we hypothesize that the natural mode of RatPyV2 transmission may be through oral, nasal, and respiratory secretions with primary replication in the epithelial cells of glandular and respiratory tissues prior to systemic spread.
In man, detection of polyomavirus in respiratory secretions is common, and viral infection has been associated with and without clinical respiratory disease (Gaynor et al. 2007). Recently, polyomavirus identified as WUPyV was detected within respiratory cells from 2 immunocompromised patients diagnosed with pneumonia. Infected respiratory cells had intranuclear inclusions, and virus was characterized by application of deep-sequencing methodology or virus-specific PCR and with anti-WUPyV VP1 immunohistochemistry on respiratory samples (Siebrasse et al. 2015, 2016). Findings in these case reports correlate well with those of RatPyV2 distribution in the respiratory tract in immunocompromised rats in this study.
Vertical transmission of PyVs has been demonstrated in experimental rodent models and, although still controversial, vertical transmission of JC virus and BK virus has been reported in man (Patel et al. 2009; Zhang, McNees, and Butel 2005; Boldorini et al. 2011). In the present study, intranuclear inclusions were observed in prostatic epithelial cells of all male FSG rats (n = 7/7) and uterine epithelial cells in 2 female FSG rats (n = 2/5). Furthermore, intranuclear inclusions were observed in endometrial, vaginal, and cervical epithelial cells of the experimentally inoculated nude rat. The viral load in the uterus of this rat was extremely high, providing further support for uterine replication. The presence of virus in the prostate and uterus of naturally infected FSG rats suggests that vertical and congenital transmission is plausible and should be further investigated.
The mechanism for human polyomavirus transmission has not been fully established. PyVs WU, BK, and JC have been found in raw sewage and stool samples from children and adults, suggesting that fecal–oral transmission may be an important mode of infection (Bofill-Mas et al. 2001, 2010; Vanchiere et al. 2009; Ren et al. 2009). RatPyV2 was detected by PCR in fecal samples of infected FSG and experimentally exposed nude rats. These rats ranged in age from 10 to 40 weeks, suggesting that fecal viral shedding in immune-deficient rats is prolonged if not persistent. In contrast, virus was detected in 0.70% of feces from immune-competent rats. Given the seroprevalence rate of 32%, these data suggest that viral shedding is transient in immune-competent rats. Curiously, little RatPyV2 genome was detected in the intestinal tract of rats by qPCR and no virus was detected by ISH. Virus detected in the feces may have originated from virus-infected secretions and cell debris from the tongue, respiratory, glandular and urogenital tissues, and not from virus-infected intestinal mucosa and/or secretions.
In summary, RatPyV2 is a novel virus species belonging to the Wuki polyomavirus clade. Infection in immune-deficient rats is associated with clinical disease and virus targets epithelial cells of multiple tissues, including the respiratory tract, lacrimal and salivary glands, and the female and male reproductive tracts, which suggests the potential for horizontal as well as vertical and sexual transmission. RatPyV2 infection was experimentally replicated by exposure of nude rats via direct contact with infected rats and parenteral viral inoculation. The 32% prevalence of anti-RatPyV2 antibodies in samples from immunocompetent rats suggests widespread prevalence of viral infection in laboratory rats that may have significant and possibly confounding implications for established rat models. Finally, RatPyV2 infection in rats may provide a unique system for studies on the pathogenesis polyomaviral diseases in man.
Footnotes
Author Contribution
Authors contributed to conception or design (CB, PP, SH, BB, BK, TP, ED, RL, SC, RW, SL, DB, MM); data acquisition, analysis, or interpretation (CB, PP, SH, BB, BK, TP, ED, RL, SC, RW, SL, DB, MM); drafting the manuscript (CB, PP, BK, ED, RL, SC, RW, SL, DB, MM); and critically revising the manuscript (CB, PP, SH, BB, BK, TP, ED, RL, MM). All authors gave final approval and agreed to be accountable for all aspects of work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported the Blood Systems Research Institute.
