Abstract
Brain trimming through defined neuroanatomical landmarks is recommended to obtain consistent sections in rat toxicity studies. In this article, we describe a matrix-guided trimming protocol that uses channels to reproduce coronal levels of anatomical landmarks. Both setup phase and validation study were performed on Han Wistar male rats (Crl:WI(Han)), 10-week-old, with bodyweight of 298 ± 29 (SD) g, using a matrix (ASI-Instruments®, Houston, TX) fitted for brains of rats with 200 to 400 g bodyweight. In the setup phase, we identified eight channels, that is, 6, 8, 10, 12, 14, 16, 19, and 21, matching the recommended landmarks midway to the optic chiasm, frontal pole, optic chiasm, infundibulum, mamillary bodies, midbrain, middle cerebellum, and posterior cerebellum, respectively. In the validation study, we trimmed the immersion-fixed brains of 60 rats using the selected channels to determine how consistently the channels reproduced anatomical landmarks. Percentage of success (i.e., presence of expected targets for each level) ranged from 89 to 100%. Where 100% success was not achieved, it was noted that the shift in brain trimming was toward the caudal pole. In conclusion, we developed and validated a trimming protocol for the rat brain that allow comparable extensiveness, homology, and relevance of coronal sections as the landmark-guided trimming with the advantage of being quickly learned by technicians.
Introduction
The challenges of microscopic evaluation of the brain arise from the complex anatomy and physiology of this organ and on the wide variability of coronal levels when its evaluation is conducted in toxicity studies. In fact, one of the brain peculiarities is its complex organization in different neuroanatomic regions associated with high pleomorphism of neurons and glial cells (Garman 2011). This complexity is further enhanced considering that each neuroanatomic region has a specific neurologic function, a unique physiology, and energy requirements contributing to the marked brain heterogeneity in the sensitivity to pathologic stimuli (Moser et al. 2008; Sokoloff 2008; Moser 2011). For these reasons, even in the presence of clear signs of toxicity, such as with known neurotoxicants, the anatomical localization of damage, the identification of cerebral regions receiving afferent and efferent connections that may be secondarily affected, and the correlation of the damage with a specific neurologic deficit can be a challenge (Fix et al. 2000; Lester et al. 2000; Garman 2011; Switzer, Lowry-Frassen, and Benkovic 2011).
Given the anatomical complexity and heterogeneity of the brain, a proper neuropathology assessment can be further complicated if microscopic sections lack consistency between different animals. In fact, the trimming protocol of the brain is essential to guarantee adequate homology of sections and the relevance of the sampled anatomical structures. In Good Laboratories Practices (GLP)-type rodent general toxicity studies, guidelines are not prescriptive regarding the trimming protocol applied to the brain. In the past, in these types of screening tests, the brain was routinely sampled at 3 coronal levels. These levels broadly included rostral forebrain (cerebral cortex and basal nuclei), caudal forebrain (cerebral cortex, hippocampus with either diencephalon or midbrain), and hindbrain (cerebellum and medulla oblongata; Morawietz et al. 2004). However, a free hand-trimming method can result in wide variability of coronal levels and inconsistent sampling of neuroanatomic structures when performed by less experienced technicians.
A nonstandard approach is the tailored trimming for the microscopic evaluation of specific brain regions, if the area of potential neurotoxicity has been previously identified or suspected. In fact, the updated version of European guidelines for repeated dose toxicity testing recommends that for central nervous system (CNS)-active substances the histopathologic evaluation of the brain should be more extensive and target-oriented European Medicine Evaluation Agency/Committee on Human Medicinal Products/Safety Working Party (EMEA/CHMP/SWP/488313/2007). In these cases, the microscopic evaluation should be extended to brain sections including the cells or the CNS regions affected directly or indirectly by treatment because of the substance-receptor binding profile or substance-related pharmacodynamic effects. Thus, to assure consistency and relevance of sections to be examined, the brain trimming should be based on consolidated knowledge of macroscopic and microscopic anatomy.
A brain trimming method that assures adequate homology and relevance of sections between control and treated rats is the one based on anatomical landmarks. Moreover, a landmark-guided trimming approach has been recently suggested for general toxicity studies by the Working Group on Nervous System Sampling established by the Society of Toxicologic Pathology (STP; Bolon et al. 2013). In the past decade, several anatomical landmark-guided trimming methods for the rat brain have been published, both for developmental neurotoxicity testing (Bolon et al. 2006) and for the adult rat brain in general toxicity studies (Jordan et al. 2011; Rao et al. 2011; Rao, Little, and Sills 2014). Even if differences exist between the levels of trimming between these methods, all protocols recommend the sampling of the brain through a variable number of coronal cuts identified by target anatomical landmarks recognizable mainly on the ventral aspect of the brain (Bolon et al. 2006). However, the application of a landmark-guided trimming can be time-consuming, and the identification of anatomical landmarks requires a high level of training for the histology technical staff.
In this context, we have developed a new trimming protocol for rat brain, taking advantage of the accuracy and anatomical awareness of the landmark-guided trimming, but improving its feasibility. In this new trimming protocol, we have used a brain matrix (a device developed for the correct positioning of the rat brain to allow slicing of discrete coronal sections through grooves, specifically called channels, set 1 mm apart) and identified a set of matrix channels that reproduces the coronal levels of the anatomical landmark-guided trimming. This “matrix-guided” trimming approach allowed comparable extent, homology, and relevance of coronal sections to the landmark-guided trimming, with the advantages of reducing the asymmetry often encountered in coronal sections during manual trimming and the fact that is easily learned by less experienced technicians. The aim of this article is to present the experimental steps we covered to develop this trimming protocol (setup phase) and the results from its application on a large number of rats to demonstrate its effectiveness (validation study).
Materials and Methods
Animals
Both the setup phase and the validation study were performed using male Han Wistar (Crl:WI(Han)) rats, 10 weeks old on the day of necropsy with an average body weight of 298 ± 29 (SD) g. Ten rats were used for the setup phase and 60 rats were used for the validation study. Rats were obtained from 2 breeding sites of Charles River Laboratories: Lentilly, Rhône-Alpes, France (setup phase and validation study) and Margate, Kent, UK (only validation study).
After 1-week acclimatization, rats were housed in pairs in solid bottom plastic cages, each with sawdust litter and an external watering system. Environmental enrichment, including 1 isoblox/animal and 1 fun-tunnel/cage, was also provided. Animal room temperature was maintained at 21 ± 1°C, and relative humidity ranges were generally maintained at 55 ± 10%. Rats were given free access to filtered tap water and ad libitum diet (Gamma Irradiated Rat and Mouse No 1 Expanded Diet, Special Diet Service Ltd., Whitam, Essex, UK), except for the day prior to necropsy, when food was removed overnight. All the work on animals was carried out in accordance with Italian legislation governing use of experimental animals and the European Directive 2013/63/EU.
Terminal Necropsy and Histological Procedures
Terminal necropsy and histological procedures were the same in both the setup phase and validation study. To collect the brain, rats were euthanized by exsanguination via abdominal aorta, under deep isoflurane anesthesia (Isoba, MSD Animal Health, Milton Keynes, Buckinghamshire, UK). At necropsy, full macroscopic examination was performed; the head was removed by sectioning at the atlantooccipital joint; the dorsal cranial bones of the skull were removed using roungeurs; and the brain was gently taken out from the cranial cavity, weighed (for the validation study only), and immersed in 100 ml of 10% neutral buffered formalin for at least 48 to 72 hr. Trimming of the brain was performed using a brain matrix (RBM-4000C, ASI-Instruments®, Houston, TX) designed for brains of rats with a bodyweight of 200 to 400 g. Brain sections obtained were paraffin embedded lying on their rostral surface, cut with the rotatory microtome (5 μm in thickness), stained with hematoxylin and eosin, and microscopically examined by the same trained pathologist.
Setup Phase
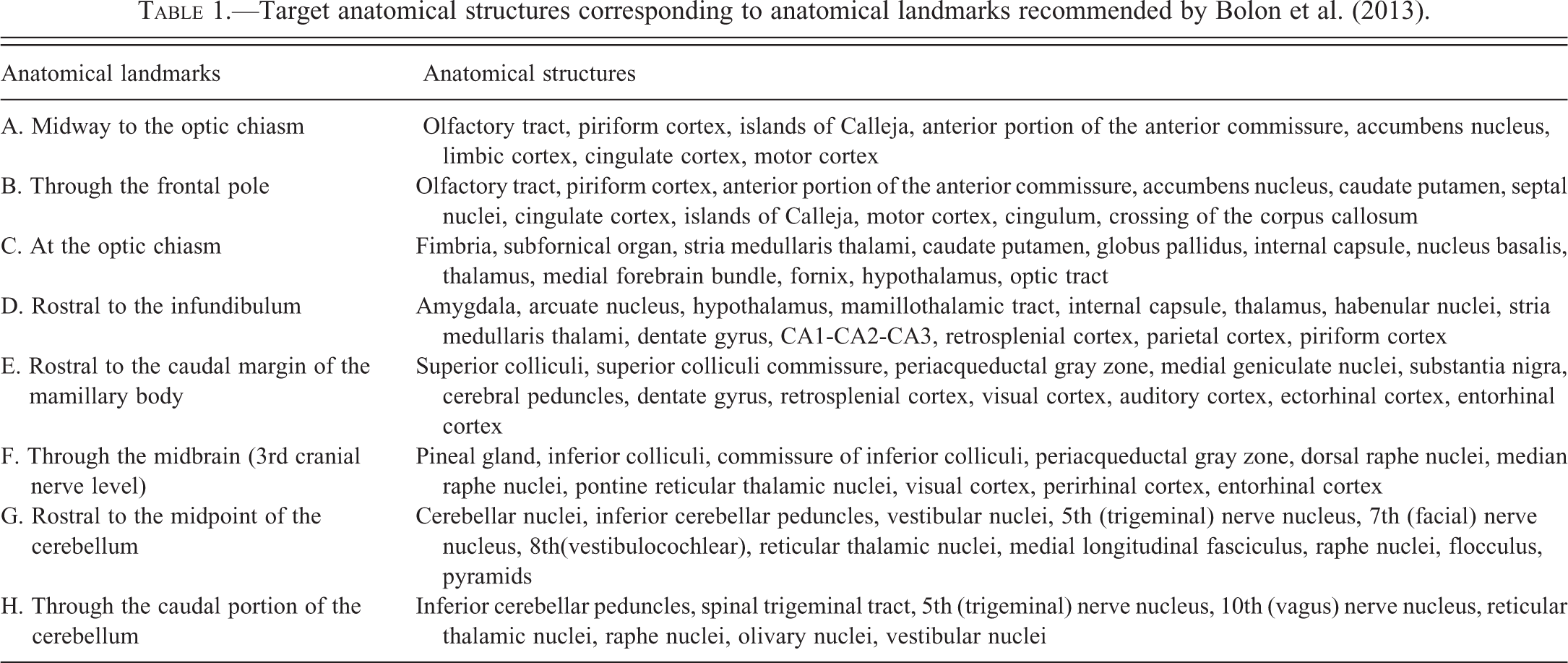
The setup phase aimed to identify a set of matrix channels that allows reproduction of the same coronal levels obtained using the classical approach, based on the visual identification of anatomical landmarks. To reach a systematic histological evaluation of the brain, neuroanatomical targets of interest were selected based on published recommendations (Bolon et al. 2013) and other targets specifically investigated in our institution. Anatomical landmarks and associated neuroanatomical structures are listed in Table 1.
Target anatomical structures corresponding to anatomical landmarks recommended by Bolon et al. (2013).

In the process of landmark to channel translation, we first determined, for all the nuclei and anatomical regions of interest, their extension along the brain longitudinal axis. For this purpose, we referred to The Rat Brain in Stereotaxic Coordinates (Paxinos and Watson 2007) and, in particular, we considered the distance from Bregma. In the stereotaxic reference system of Paxinos and Watson, the Bregma is a vertical plane of reference passing through the anatomical point on the rat skull at which the coronal suture and sagittal suture intersect perpendicularly. The “Bregma coordinate” represents the rostro-caudal distance of a specific point in the brain from this reference plane.
This Bregma coordinate is expressed in millimeters (mm) and characterized by positive or negative values according to their position rostral or caudally, respectively, to the zero Bregma plane. In particular, we used the Bregma coordinates of the rostral and caudal boundaries of the anatomic structures and the intercurrent interval to extrapolate their longitudinal extension and their reciprocal positions (Figure 1). From the complete mapping of the structures expected for each coronal level of the landmark-guided trimming, we determined the approximate Bregma coordinate of the landmarks (Table 2) corresponding to the coronal plane intersecting all the structures expected for that level (Figure 1). We then identified the brain matrix channel matching the Bregma coordinate of the most caudal anatomical landmark (i.e., caudal portion of the cerebellum localized at Bregma −12 mm). The choice of starting caudally, rather than rostrally, in the process of landmark to channel translation was arbitrarily decided by the authors. For this purpose, we trimmed the fixed brains from 10 rats cutting through the 6 caudal channels of the matrix.
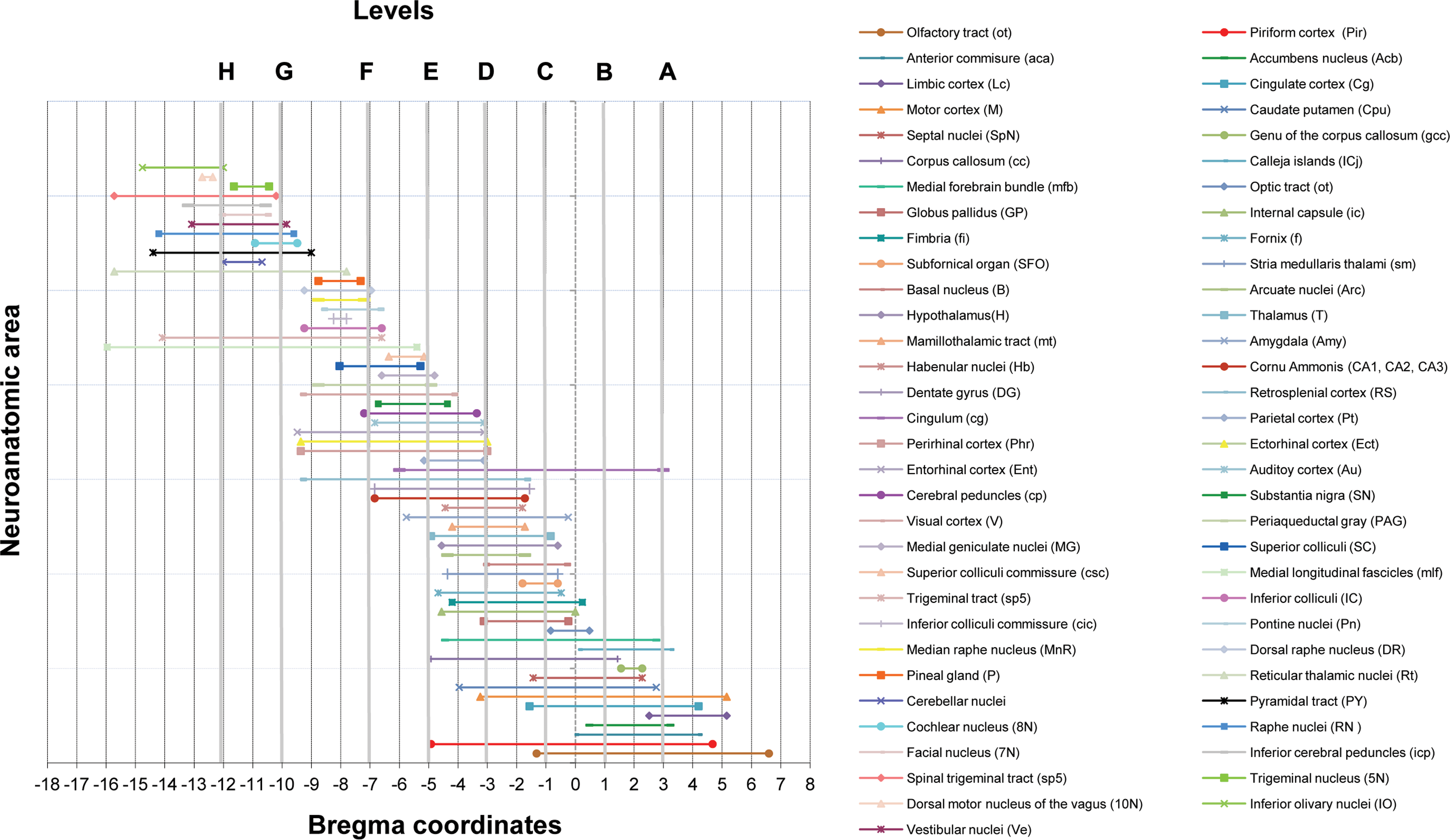
Complete mapping of the structures expected for each coronal levels and approximate Bregma coordinate of the landmarks.
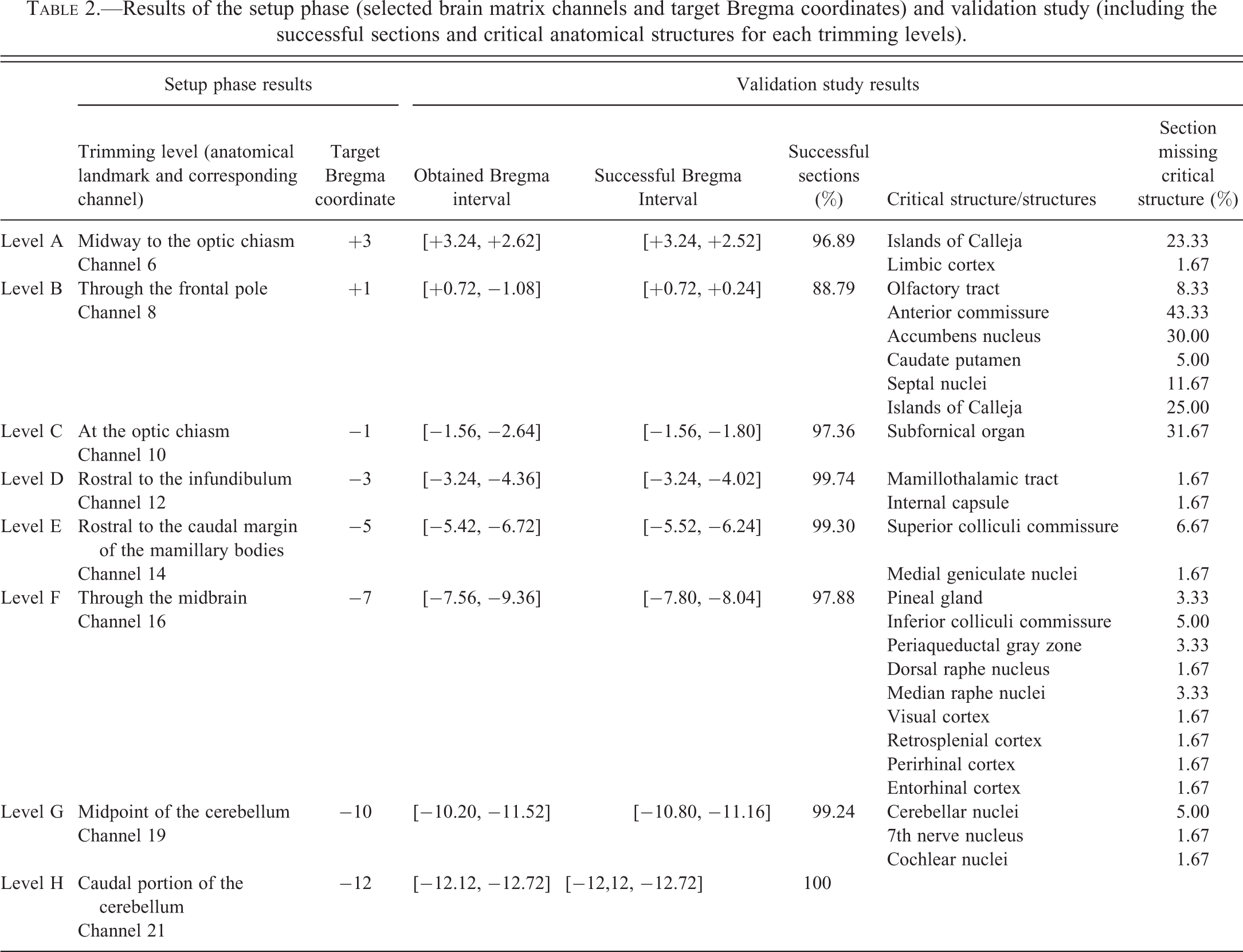
Results of the setup phase (selected brain matrix channels and target Bregma coordinates) and validation study (including the successful sections and critical anatomical structures for each trimming levels).
Validation Study
A validation study was then performed to determine whether and how consistently the brain matrix channels selected in the setup phase reproduced the coronal levels of the corresponding anatomical landmark, thus resulting in microscopic sections characterized by high homology between rats and including all the relevant anatomical structures. With this purpose, the brain matrix–guided trimming was applied to the brain of 60 male Han Wistar (Crl:WI(Han)) rats. Slices (2- or 3-mm thick) obtained from channel trimming levels were identified following rostro-caudal sense by alphabetic letters (levels from A to H). Slices were embedded in pairs (A/B; C/D; E/F; G/H) in the cassette lying down on their rostral surface.
Brain weights were expressed as mean ± SD. Brain weight from 2 different sources (France and UK) was compared using a Student’s t-test. A p value ≤ .05 was considered significant.
The 8 coronal sections per rat obtained from the application of matrix-guided trimming were evaluated microscopically. The evaluation consisted in the determination of the Bregma coordinate for each section using the Paxino’s atlas as a reference (Paxinos and Watson 2007). Based on the Bregma coordinate determined, the presence or absence of expected anatomical structures was recorded using a binary system (1 = presence; 0 = absence). Once microscopic evaluation of the sections was completed, for each trimming level, Bregma coordinates determined were expressed as an interval (Obtained Bregma interval). The overall absence of each anatomical structure was calculated as the number of negative results obtained for that specific structure over the total number of sections expected to contain it (% sections missing critical structure). This allows us to determine a “Successful Bregma interval,” that is, the interval of Bregma coordinates including brain sections with all the target anatomical structures present.
For each trimming level, the percentage of successful sections was calculated as the weighted average of the presence of anatomical structures included in that level. This value was indicative of the frequency with which the trimming at that selected matrix channel reproduced the coronal levels of the corresponding anatomic landmark.
Results
Setup Phase
Microscopic evaluation of the sections obtained by this preliminary trimming indicated that the third caudal channel of the matrix (channel 21) was the one most effectively reproducing the coronal levels around Bregma −12 mm and, consistently, the most caudal landmark (i.e., caudal portion of the cerebellum). Once we had determined that Bregma −12 mm could be found trimming the brain through third caudal channel of the matrix, we determined the Bregma coordinates of the other matrix channels, taking into account that they are 1 mm apart. Then we were able to select a set of matrix channels corresponding to the Bregma coordinates of the anatomical landmarks of interest identified initially (Figures 1 and 2; Table 2).
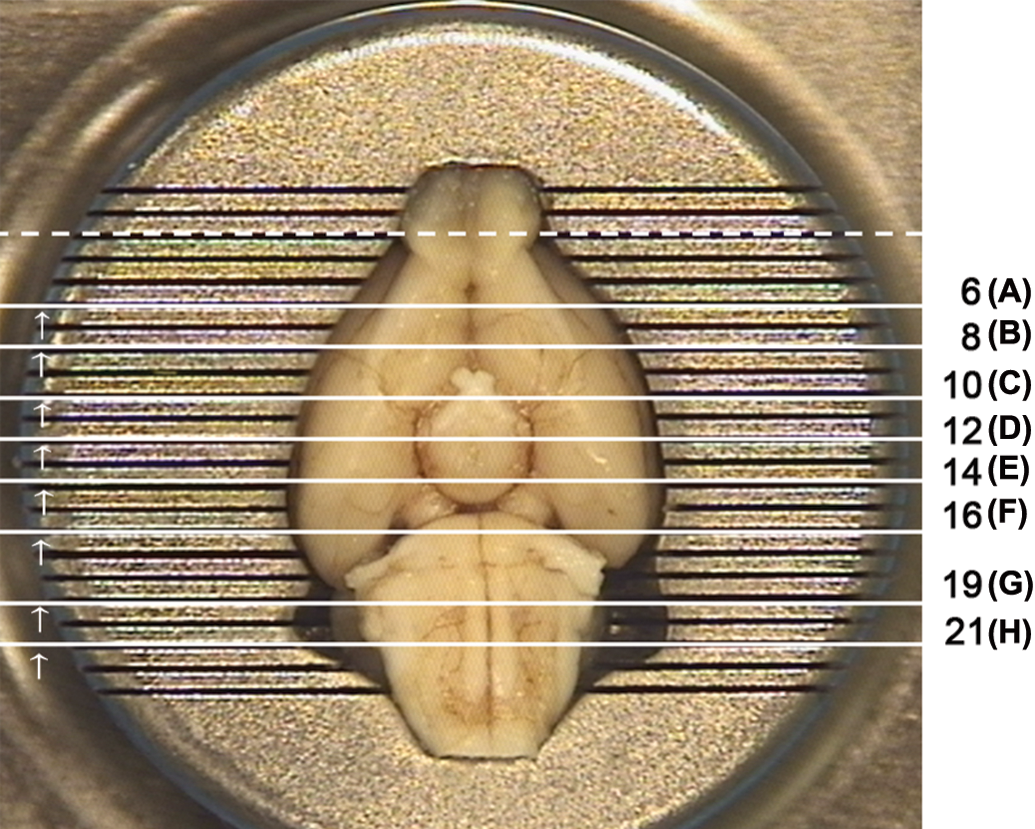

The brain placed dorsoventrally in the brain matrix. Solid white lines indicate the position of the selected matrix channels; channel numbers with corresponding trimming level in parentheses are shown on the side. For each trimming level, an arrow indicates the face of the block to be placed down in the cassette. Dashed white line indicates the trimming level that can be used for additional evaluation of the olfactory bulbs.
Validation Study
Macroscopic Examination
No macroscopic abnormalities were detected in the brain or in any other organs or tissues.
Brain Weight
The mean value of absolute brain weight was 1.92 ± 0.08 (mean ± SD) g for rats obtained from France (n = 30) and 1.93 ± 0.10 (mean ± SD) g for rats from the United Kingdom (n = 30). There were no significant differences between the weights of the brain of rats from the 2 sources, and for this reason both sets of data were presented together.
Microscopic Evaluation Results
The results of the microscopic evaluation of brain sections, including their success in terms of presence or absence of target anatomical structures, are summarized in Table 2. Microscopic pictures of all brain levels (A–H), including target anatomical structures are represented in Figure 3.
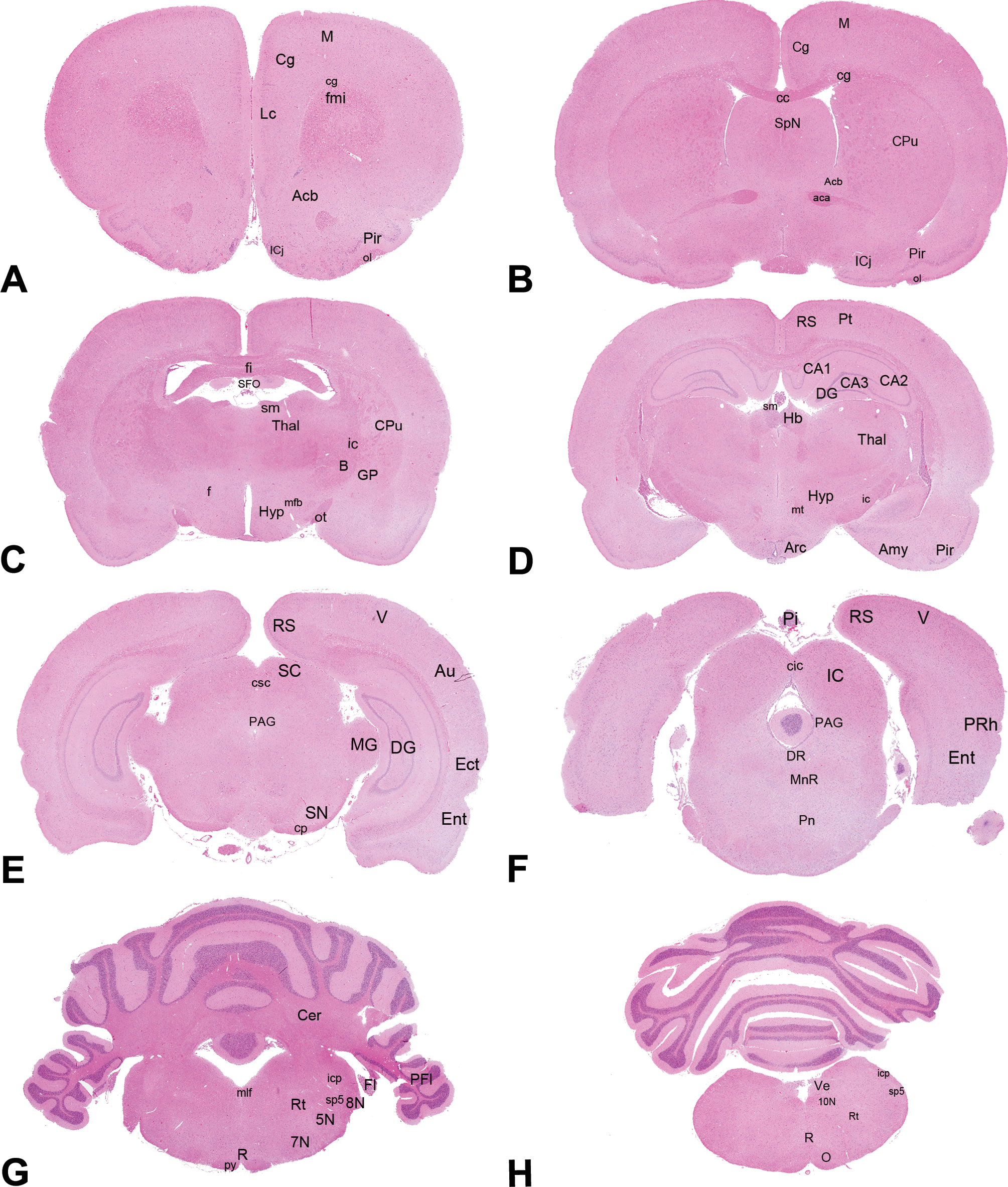
Hematoxylin and eosin coronal sections of the rat brain obtained at a level midway to the optic chiasm (level A), through the frontal pole (level B), at the optic chiasm (level C), level rostral to the infundibulum (level D), level rostral to the caudal margin of the mamillary body (level E), through the midbrain (3rd cranial nerve level; level F), rostral to the midpoint of the cerebellum (level G), and through the caudal portion of the cerebellum (level H). For each level, anatomical structures of interest are shown: aca = anterior commissure; Acb = accumbens nucleus; Amy = amygdala; Arc = arcuate nuclei; Au = auditory cortex; B = basal nucleus; CA1, CA2, CA3 = cornu ammonis 1, 2 and 3; cc = corpus callosum; Cer = cerebellum; Cg = cingulate cortex; cg = cingulum; cic = inferior colliculi commissure; cp = cerebral peduncles; Cpu = caudate putamen; csc = superior colliculi commissure; DG = dentate gyrus; DR = dorsal raphe nucleus; Ect = ectorhinal cortex; Ent = entorhinal cortex; Fl = flocculus; fmi = forceps minor of the corpus callosum; f = fornix; fi = fimbria; GP = globus pallidus; Hb = habenular nuclei; Hyp = hypothalamus; ic = internal capsule; IC = inferior colliculi; ICj = Calleja islands; icp = inferior cerebral peduncles; Lc = limbic cortex; mfb = medial forebrain bundle; M = motor cortex; MG = medial geniculate nuclei; mlf = medial longitudinal fascicles; MnR = median raphe nuclei; O = olivary nuclei; ol = olfactory tract; PAG = periaqueductal gray; PFl = paraflocculus; PRh = perirhinal cortex; Pi = pineal gland; Pir = piriform cortex; Pn = pontine nuclei; Pt = parietal cortex; py = pyramidal tract; RN = raphe nuclei; RS = retrosplenial cortex; Rt = reticular thalamic nuclei; SC = superior colliculi; SFO = subfornical organ; sm = stria medullaris thalami; SN = substantia nigra; SpN = septal nuclei; sp5 = spinal trigeminal tract; Thal = thalamus; V = visual cortex; Ve = vestibular nuclei; 5N = trigeminal nerve nucleus; 7N = facial nucleus; 8N = cochlear nerve nucleus; 10N = vagus nerve nucleus.
For all levels, obtained Bregma intervals included the target Bregma coordinate forecasted in the setup phase. As expected, the obtained Bregma interval was wider than the successful Bregma interval for almost all levels, resulting in a percentage of success lower than 100%. Percentage of success ranged from 89 to 100%, with level H being the most successful (100%). In those levels where 100% success was not achieved, there was a shift (0.1–1.32 mm) in the caudal sense.
Lack of success was determined by the absence of a variable number of neuroanatomical structures for each level (i.e., critical structures). The frequency with which each of these structures was missing was determined as a percentage. This percentage was variable, reaching 43% for the anterior commissure in level B sections. No histopathological abnormalities of the brain parenchyma were noted.
Discussion
In this study, we have proposed a brain-matrix trimming method resulting in coronal sections that reproduced successfully the coronal levels of landmark-guided trimming (high reproducibility). All the neuroanatomical structures requested for an extended brain examination were present in a high percentage of sections (high homology). The proposed matrix-guided trimming includes neuroanatomical targets recommended by recently published literature (Bolon et al. 2013) and other targets specifically investigated in our institution. For example, the nucleus accumbens and the limbic cortex were included in our trimming method because they are key components of the limbic system and their examination is essential in studies of drug dependence, but are not routinely sampled in general toxicology studies.
A caudal shift was observed at specific coronal levels resulting in a lower percentage of success due to the lack of one or more anatomical structures in the microscopic sections. Examples include Calleja Islands (Figure 3B), subfornical organ (Figure 3D), and cerebellar nuclei (Figure 3H). These critical structures shared peculiar features, namely the limited extension along the longitudinal brain axis and/or their localization in close proximity to the slice-cutting plane, which can be easily skipped during the trimming or microtome-sectioning procedures.
In one case (level F), lower reproducibility of coronal levels was likely due to a procedural error, such as embedding of the brain slice lying with the caudal surface down. In this case, the error was recognizable by its sporadic occurrence in few rats and the associated abnormally wide Bregma interval. Other potential procedural errors (e.g., inaccurate sampling at necropsy with excessive tissue damaging/loss, trimming through a wrong channel, or embedding of the slices with the caudal surface down) may have a severe impact on the reproducibility of the method and must be carefully avoided through strict adherence to procedure and training of staff.
The routine use of a brain matrix has been questioned since adequate reproducibility of sections could be influenced by variations in brain size (Bolon et al. 2013). Nevertheless, in our study, good reproducibility has been shown for most target neuroanatomical structures in male rats with a body weight of 299 ± 29 g using a brain matrix fitted for rats weighing 200 to 400 g, suggesting that the trimming method described is applicable to general toxicity studies using rats of this age and body weight range. Unpublished data from our laboratory confirm the accuracy of this method when used in large scale in male and female rats of approximately 3 months of age (corresponding to a body weight range of 200–400 g). In fact, rats of this age are in a phase of steep increase in body weight, but brain growth rate is reported to reach a plateau around 20 days, with only further minimal increases up to 275 days of age (Kishimoto, Davies, and Radin 1965). In addition, most of the neuroanatomical structures evaluated are characterized by consistent rostro-caudal extension, so that minimal variation in brain size/weight is not critical for the reproducibility of the method.
Most recent recommendations (Bolon et al. 2013) for neural sampling in preclinical studies recommend the importance of olfactory bulbs evaluation as a potential target for inhaled and ingested toxicants, as well as a source of neural stem cells. Although olfactory bulbs are not included in our trimming protocol, they can be easily identified macroscopically and trimmed accordingly using the most rostral brain matrix channels.
Microscopic evaluation can be initially limited to 3 levels, such as B, C, and G, which overall represent the same levels, used as a common screening technique in general toxicity studies (Morawietz et al. 2004) with the advantage that our matrix-guided trimming allows a higher degree of homology between sections. A more extensive microscopic evaluation could be chosen on the basis of the pharmacodynamic profile of the test compound or on the basis of unexpected events occurring during the course of the study (e.g., neurological signs developing during the course of the in-life phase, gross lesions in the neural organs detected by noninvasive imaging, or at macroscopic examination; Bolon et al. 2011). In fact, the application of this method considers the preservation in formalin of sliced brain tissues properly identified to promptly perform additional investigations at other levels, if a more extensive neuropathological examination is needed.
One point to consider when choosing a subset of coronal levels is that some anatomical structures, given their extension along the longitudinal axis, can be present in 2 adjacent coronal levels with different probability to be sampled at trimming (e.g., nucleus accumbens). When evaluating a selected panel of brain structures, careful consideration on the highest probability to detect the structure of interest (especially for those present at different levels) should drive the selection of the appropriate trimming channel.
In conclusion, a standardized trimming protocol for the rat brain was developed and validated to assist the toxicologic pathologist in the extensive and neuroanatomic-aware brain evaluation. This protocol offers the advantage of being applicable routinely on large numbers of animals, being standard and not necessitated of extensive training in neuroanatomy, as required for anatomic landmark recognition. Moreover, the use of a specifically designed device to slice discrete coronal levels, such as the brain matrix, should significantly reduce the manipulation of an organ so prone to artifacts. The number of slices to be examined microscopically should be tuned on the basis of specific criteria, established case-by-case.
Footnotes
Acknowledgments
The authors wish to thank Alessandro Poffe, Renzo Carletti, and Erica Zambello for technical support and Manolo Mugnaini for critical review of the manuscript.
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
The author(s) received no financial support for the research, authorship, and/or publication of this article.
