Abstract
In the past few decades, it has become increasingly evident that sensitivity to ionising radiation is variable. This is true for tissue reactions (deterministic effects) after high doses of radiation, for stochastic effects following moderate and possibly low doses, and conceivably also for non-cancer effects such as cardiovascular disease, the causal pathway(s) of which are not yet fully understood. A high sensitivity to deterministic effects is not necessarily correlated with a high sensitivity to stochastic effects. The concept of individual sensitivity to high and low doses of radiation has long been supported by data from patients with certain rare hereditary conditions. However, these syndromes only affect a small proportion of the general population. More relevant to the majority of the population is the notion that some part of the genetic contribution defining radiation sensitivity may follow a polygenic model, which predicts elevated risk resulting from the inheritance of many low-penetrance risk-modulating alleles. Can the different forms of individual radiation sensitivities be inferred from the reaction of cells exposed ex vivo to ionising radiation? Can they be inferred from analyses of individual genotypes? This paper reviews current evidence from studies of late adverse tissue reactions after radiotherapy in potentially sensitive groups, including data from functional assays, candidate gene approaches, and genome-wide association studies. It focuses on studies published in 2013 or later because a comprehensive review of earlier studies was published previously in a report by the UK Advisory Group on Ionising Radiation.
1. CHARACTERISATION AND MECHANISMS OF RADIATION-INDUCED HEALTH EFFECTS
Exposure to ionising radiation in humans occurs in various settings, including fractionated high-dose exposures, such as those from cancer radiotherapy; a range of acute exposures, such as those experienced by the Japanese atomic bomb survivors; chronic and low-level exposures, such as those received by radiation workers; and fractionated low-dose exposures from diagnostic medical examinations. Radiation-induced health effects can be divided into stochastic effects, such as cancer, which are understood to be induced even after low-dose exposure; and deterministic tissue reactions, such as necrosis, which are only observed after relatively high-dose exposure, and cardiovascular disease and lens opacities, for which much less is known regarding the dose–response relationships. The mechanisms of these effects are diverse. As such, a radiosensitive phenotype towards one type of effect does not necessarily indicate radiosensitivity towards another type (Foray et al., 2016).
Late adverse tissue reactions to high doses of radiation have a complex aetiology. For a long time, they were believed to be solely related to the death of parenchymal cells in tissues (Thames and Hendry, 1987). Today, it is understood that they result both from cell death and interaction among surviving cells from multiple lineages, as well as from the action of various biologically active extracellular molecules (Denham and Hauer-Jensen, 2002). The reaction of normal tissues to high doses of radiation, especially when given in a fractionated manner, involves two complex and partially interacting components: (1) the healing of traumatic wounds which is perturbed by the fractionated radiation exposure; and (2) a combination of specific injuries, not only to the parenchymal cells, but also to connective tissue and, above all, the vascular system. The second component involves chronic inflammation and is believed to be responsible for the progression of injury over a period of many years, resulting in a steadily increasing risk of manifestation as adverse tissue reaction (Jung et al., 2001).
Longer-term effects of radiation exposure include elevated risk of cancer and, at higher levels of exposure, cardiovascular disease. The development of cancer is characterised by the acquisition of distinct biological properties over a long period of time. These hallmarks of cancer are driven by genomic instability (which generates genetic diversity leading to the acquisition of hallmark features) and inflammation (Hanahan and Weinberg, 2011). The carcinogenic effects of ionising radiation are generally understood to result from various types of damage to DNA, including damage to nucleotide bases, single-strand breaks, double-strand breaks (DSBs), and DNA crosslinks (BEIR VII, 2006), with DSBs playing a prominent role (Bernstein et al., 2013). Under normal circumstances, cells undergo cell cycle arrest in response to DNA damage, and the majority of the damage is repaired during this time. When the damage cannot be repaired, the cell dies within a few cell divisions by means of one of several pathways, including necrosis or apoptosis. Incorrectly repaired DNA damage, on the other hand, can cause genomic instability, whereby replication of damaged sites can lead to mutations contributing to cancer development following a lag period of many years.
The range of radiation-associated cardiovascular effects that have been described likely have complex aetiologies which remain poorly understood (Hamada et al., 2014). An important element is endothelial dysfunction, which is associated with increased expression of chemokines, increased endothelial permeability, and adhesion molecules which can result in a pro-fibrotic and pro-inflammatory environment (Stewart et al., 2010; Lowe and Raj, 2014; Kabacik and Raj, 2017). Further factors which are involved are: (1) transforming growth factor beta (Dobaczewski et al., 2011); (2) the renin–angiotensin system (Fyhrquist and Saijonmaa, 2008); (3) cytokine release from mast cells (Engels et al., 1995); and (4) the cardiac sensory nervous system (Sridharan et al., 2014). These various mechanisms interact, leading to different circulatory system pathologies such as myocardial infarction, stroke, and congestive heart failure (Stewart et al., 2010).
2. DETERMINANTS OF INDIVIDUAL RESPONSE TO IONISING RADIATION
Humans respond to ionising radiation in an individually variable manner. This is true for stochastic effects (ICRP, 2007) and tissue reactions (Joiner and van der Kogel, 2009). Individual responses at the tissue level and the stochastic level have been shown (with varying levels of evidence) to be influenced by various factors including age, smoking, diabetes, collagen vascular disease, and genotype (AGIR, 2013). Indeed, it is well established that individuals with certain rare hereditary cancer-prone disorders (e.g. ataxia telangiectasia and Nijmegen breakage syndrome) are particularly radiosensitive (Taylor et al., 1975; Weemaes et al., 1981). However, these cancer susceptibility syndromes only affect a small proportion of the general population (less than one in 10,000). More relevant for the majority of the population is the theory that some part of the genetic contribution defining radiation sensitivity is likely to follow a polygenic model, whereby inheritance of several low-penetrance risk alleles can lead to a larger increase in cancer risk following exposure to a certain radiation dose compared with subjects lacking those risk alleles. This theory (the ‘common disease–common variant’ model) is also plausible for late adverse tissue reactions following radiotherapy, supported by findings that multiple genetic pathways have been implicated, including DNA damage repair, radiation fibrogenesis, oxidative stress, and endothelial cell damage (Barnett et al., 2009).
3. PROSPECTS FOR PREDICTING INDIVIDUAL RESPONSE TO IONISING RADIATION
The fact that individual radiation sensitivity to deterministic and stochastic effects is partly determined genetically raises the question whether it can be inferred from the reaction of cells exposed ex vivo to ionising radiation. Similarly, can individual response be inferred from analyses of the genotype? Ideally, a proposed biomarker would have natural and meaningful variation in the human population and, in the sense of a binary classification test, would identify the most radiosensitive patients (high sensitivity) and few non-radiosensitive patients (high specificity), and be highly reproducible among different laboratories (Bentzen, 1997). For use in human populations, the biomarker would preferably be easily obtained with minimal discomfort or risk to the patient (Hall et al., 2017). Given that levels of a biomarker can change over time and can differ between cells or tissues, the source and appropriate timing of sample collection is also important. Finally, for large-scale population testing, a rapid readout of results is preferable.
This paper reviews the current evidence on assays proposed to predict individual radiation sensitivity, with particular focus on adverse tissue reactions after radiotherapy. It focuses on studies published in 2013 or later because a comprehensive review of earlier studies was published previously in a report by the UK Advisory Group on Ionising Radiation (AGIR, 2013).
3.1. Prediction of individual radiation sensitivity to radiotherapy-related toxicity
Studies using the gamma-H2AX/53BP1/ATM focus assay to investigate whether cellular ex-vivo radiosensitivity predicts clinical radiosensitivity.
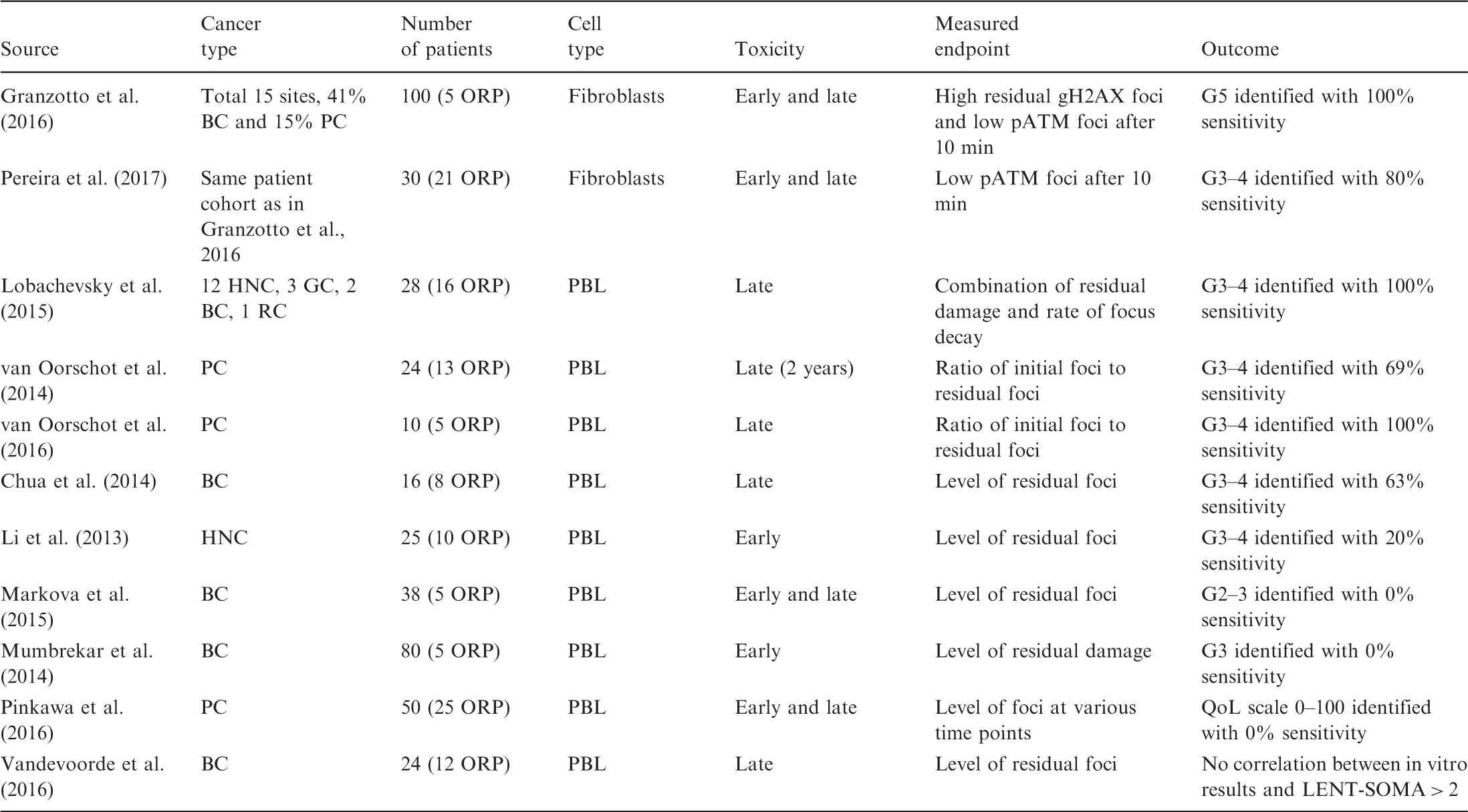
BC, breast cancer; G, Radiation Therapy Oncology Group grade; GC, gynaecological cancer; HNC, head and neck cancer; LENT-SOMA, scale for side effects in all anatomic sites; ORP, over-responding patients; pATM, phosphorylated ATM protein; PBL, peripheral blood lymphocytes; PC, prostate cancer; QoL, quality-of-life grade; RC, rectal cancer. Sensitivity indicates the percentage of patients identified by an assay relative to patients showing a given grade of adverse effect (true positives).
Studies investigating whether chromosomal aberrations, micronuclei, apoptosis, and oxidative stress markers predict clinical radiosensitivity.
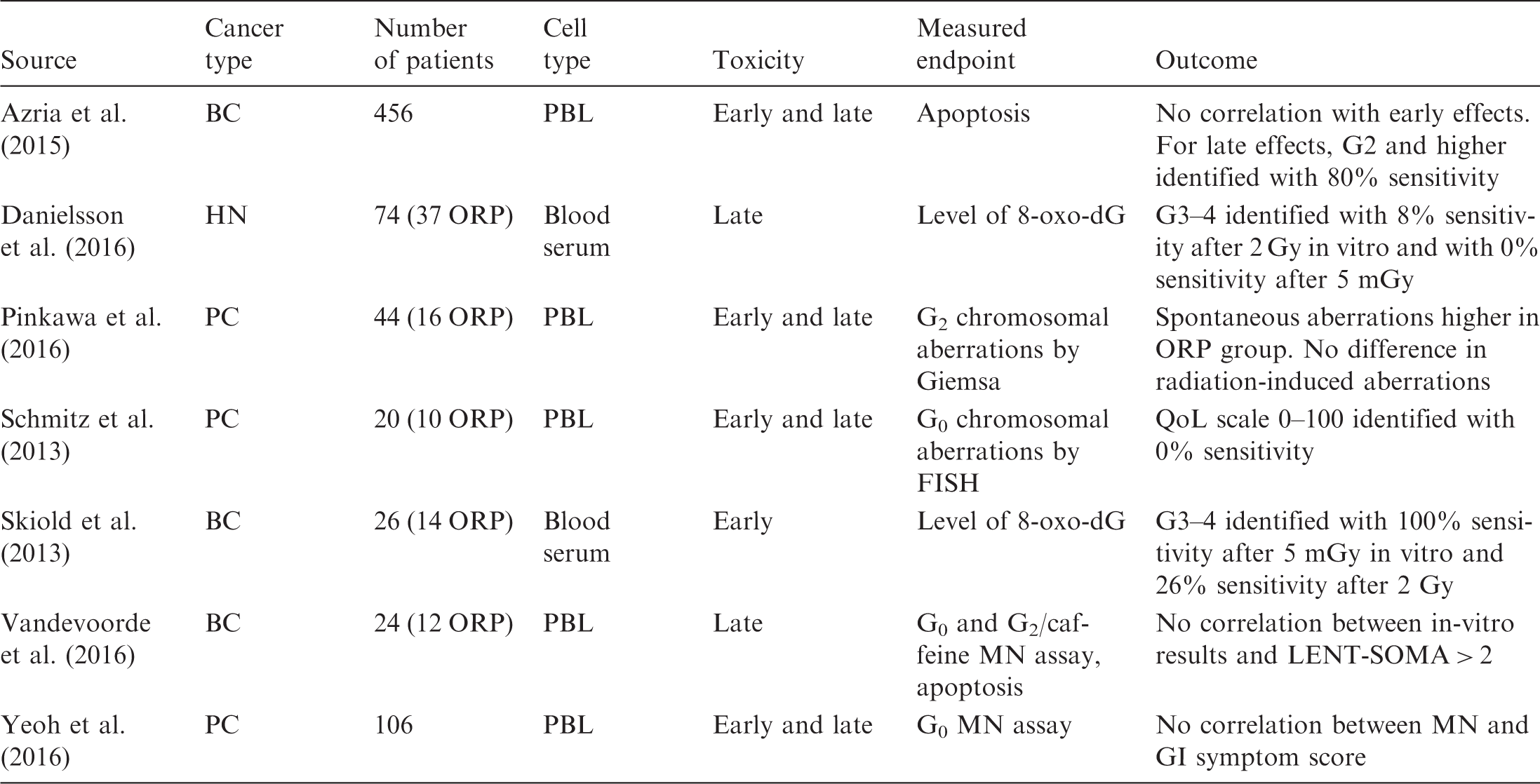
FISH, fluorescence in-situ hybridisation; G, Radiation Therapy Oncology Group grade; GI, gastrointestinal; MN, micronuclei; ORP, over-responding patients; PBL, peripheral blood lymphocytes; QoL, quality-of-life grade; LENT-SOMA, scale for side effects in all anatomic sites; 8-oxo-dG, 8-oxo-2'-deoxyguanosine. Sensitivity indicates the percentage of patients identified by an assay relative to patients showing a given grade of adverse effect (true positives).
Recent developments in molecular biology have made it possible to analyse single nucleotide polymorphisms (SNPs) in peripheral blood lymphocytes or saliva samples of patients in an attempt to assess the association of genotype with the probability of developing adverse effects to radiotherapy. Two experimental approaches have been applied: (1) quantitative polymerase chain reaction to identify a limited number of known SNPs for which there is some prior evidence; and (2) DNA sequencing to evaluate a larger number of genotyping markers in an agnostic manner. While earlier genetic studies focused on a handful of candidate genes, the development of technologies that can rapidly analyse thousands of genetic markers at relatively low cost, along with the mapping of linkage disequilibrium between common SNPs across the genome (Altshuler et al., 2010) and the definition of functional elements critical for regulation and genomic stability (Dunham and Kundaje, 2012), have allowed the comprehensive examination of the approximately 25,000 coding genes and associated functional elements in humans. The latter technique, known as the ‘genome-wide association study’ approach, uses markers across the genome and is a hypothesis-free approach.
Studies investigating whether single nucleotide polymorphisms (SNPs) in selected genes predict clinical radiosensitivity.

BC, breast cancer; DDR, DNA damage response; HNC, head and neck cancer; NSCLC, non-small cell lung cancer; ORP, over-responding patients (defined as having grade 3 or higher side effects); PC, prostate cancer; STAT, standardised total average toxicity.
In contrast to candidate SNP studies, measured SNPs in genome-wide association studies characterise the entire genome with more or less dense coverage. In addition, further unmeasured SNPs located between measured SNPs can be imputed using sequencing data from the 1000 genomes reference panel. However, the number of comparisons is generally much larger than the number of patients, and the signal-to-noise ratio is much lower than for candidate SNP studies. Also, the risk of false-positive findings increases substantially.
Genome-wide association studies to identify single nucleotide polymorphisms (SNPs) predicting clinical radiosensitivity.
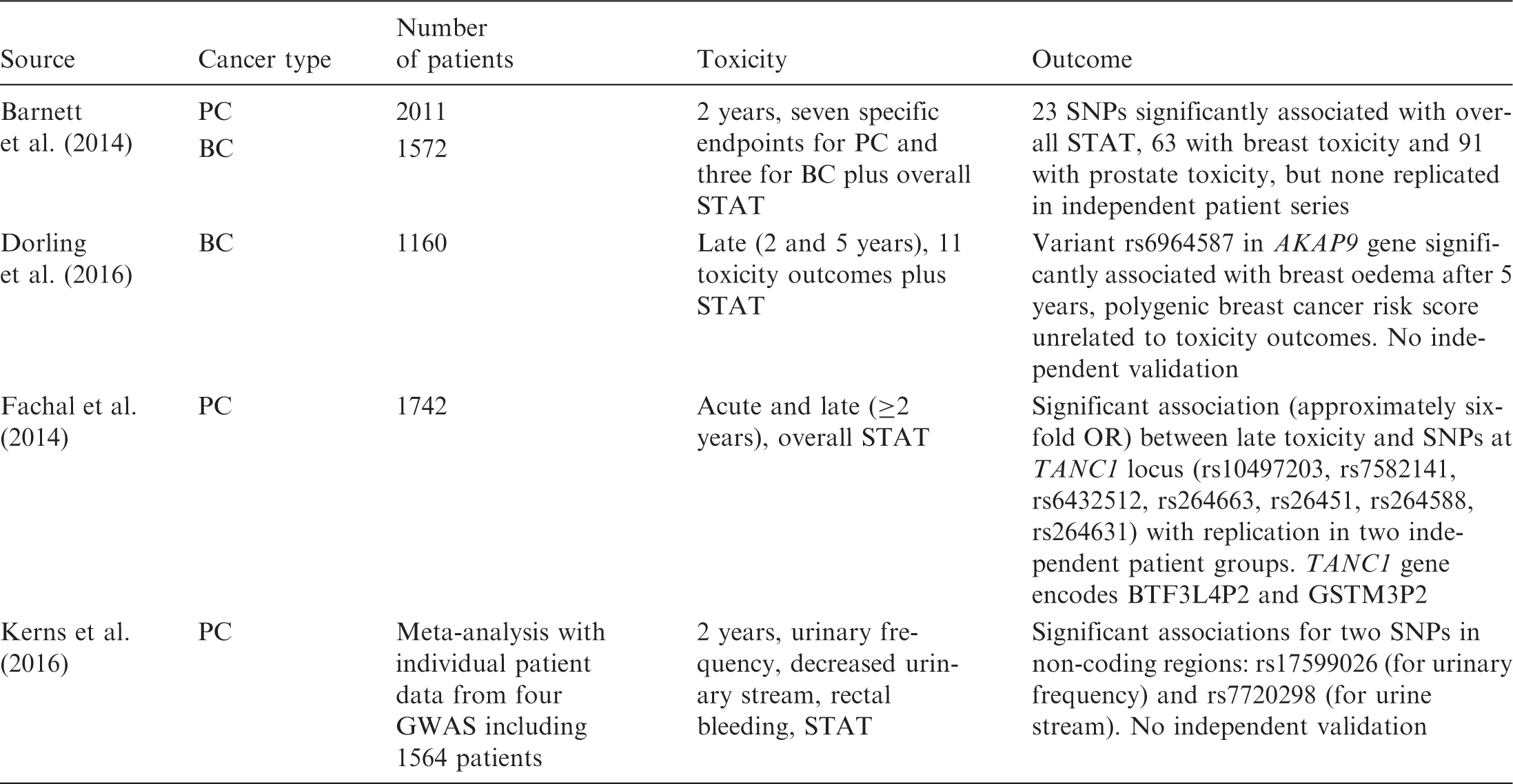
BC, breast cancer; PC, prostate cancer; STAT, standardised total average toxicity; GWAS, genome-wide association study; OR, odds ratio. Number of patients represents the sum of patients from discovery and validation steps.
Despite the large sample sizes, all studies suffer from a substantial lack of power. For example, Dorling et al. (2016) evaluated the association between over 15 million SNPs and 13 late toxicity outcomes (telangiectasia, oedema, shrinkage, induration) and overall toxicity (standardised total average toxicity) at 2 and 5 years plus pigmentation, pain, and sensitivity at 2 years] using data from 1160 patients. This corresponds to almost 200 million comparisons, or 170,000 comparisons per patient. In contrast, the latest genome-wide association study investigating breast cancer risk SNPs included approximately 150,000 breast cancer cases and 120,000 controls, and investigated only one outcome (Michailidou et al., 2017). In addition, all four studies described in Table 4 include at least some patients from the RAPPER (Radiogenomics: Assessment of Polymorphisms for Predicting the Effects of Radiotherapy) study (Burnet et al., 2013). Overlap between studies might lead to false-positive conclusions.
3.2. Prediction of individual sensitivity towards radiation-induced cancer
With respect to radiosensitivity towards cancer induction, a number of population-based epidemiological studies conducted prior to 2013 have examined radiation-related risk of cancer with respect to genes in the DNA repair and other relevant biological pathways. However, results from functional SNPs have not been convincingly replicated to date (AGIR, 2013). The genome-wide association study approach described previously has successfully identified hundreds of risk loci in germline DNA for various cancer forms (Chung and Chanock, 2011). However, the assessment of gene–environment interaction for most environmental carcinogens, including radiation, has remained elusive. A genome-wide association study of 100 Hodgkin lymphoma survivors treated with radiotherapy identified two loci in the
4. CONCLUSIONS
The aim of this paper was to review the current state of knowledge about individual radiosensitivity and the possibility of its prediction. It focused on reports describing the outcome of functional assays, candidate gene approaches, and genome-wide association studies which have been published since 2013, because a comprehensive review of earlier studies is given in a report by the UK Advisory Group on Ionising Radiation (AGIR, 2013).
The conclusions are similar to those reached by the authors of the earlier report. Although theoretical and empirical considerations suggest that individuals differ in their response to radiation exposure, no strong and consistently validated biomarkers of either tissue or stochastic effects have been identified to date. Studies of functional assays and candidate SNPs have been largely inconclusive. As pointed out by the authors of the AGIR report (AGIR, 2013), there may be several reasons for this:
functional assays are not standardised and there has been little attempt to ensure transferability across laboratories. The studies involve different radiation doses, dose rates, parameters, and assay conditions; replication and validation studies are rarely carried out; patient cohorts are heterogeneous, and different scales are often used to quantify adverse tissue effects; and study designs vary considerably, and few involve power calculation and multi-variate analysis.
Developments in high-throughput molecular biology techniques make it possible to apply whole-genome sequencing to rapidly analyse thousands of genetic markers at relatively low cost, along with the mapping of linkage disequilibrium between common SNPs across the genome. The genome-wide association study approach has identified some promising leads, such as the
Although these studies are faced with several challenges, such as the need for large sample sizes, high-quality exposure assessment, and meaningful replication and validation, initial results indicate that the integrated assessment of radiation exposure and genetic and epigenetic alterations may lead to a more nuanced characterisation of radiation-related detriment. In tandem with studying the role of biomarkers of sensitivity, better characterisation of the role of non-radiation risk factors, such as smoking and body mass index, may offer an opportunity to mitigate risk following radiation exposure.
