Abstract
Current standards for radiological protection of the public have been uniformly established. However, individual differences in radiosensitivity are suggested to exist in human populations, which could be caused by nucleotide variants of DNA repair genes. In order to verify if such genetic variants are responsible for individual differences in radiosensitivity, they could be introduced into cultured human cells for evaluation. This strategy would make it possible to analyse the effect of candidate nucleotide variants on individual radiosensitivity, independent of the diverse genetic background. However, efficient gene targeting in cultured human cells is difficult due to the low frequency of homologous recombination (HR) repair. The development of artificial nucleases has enabled efficient HR-mediated genome editing to be performed in cultured human cells. A novel genome editing strategy, ‘transcription activator-like effector nuclease (TALEN)-mediated two-step single base pair editing’, has been developed, and this was used to introduce a nucleotide variant associated with a chromosomal instability syndrome bi-allelically into cultured human cells to demonstrate that it is the causative mutation. It is proposed that this editing technique will be useful to investigate individual radiosensitivity.
1. INTRODUCTION
Following the Fukushima Daiichi nuclear power plant accident on 11 March 2011, caused by the Great East Japan Earthquake, social anxiety about the human health effects of ionising radiation has increased. Current radiological protection systems are based on the uniform standards established for protection of the public. However, it has been postulated that individual differences in radiosensitivity exist within the human population (Scott et al., 1998). Therefore, the next generation of radiological protection systems should consider the heterogeneity of individual radiosensitivity.
The cytokinesis-block micronucleus (CBMN) assay has been used previously to measure cellular radiosensitivity (Fenech, 2007). DNA double-strand breaks (DSBs) induced by ionising radiation are recognised and rejoined by the DNA repair system. Therefore, an attenuated DNA repair system results in chromosomal gaps or breaks. When the cell enters the mitotic phase in the absence of DNA repair, chromosomal fragments form one or more secondary nuclei [i.e. micronuclei (MN)]. The CBMN assay has shown that individual differences in radiosensitivity exist, and has identified mild radiosensitive cases in a small population of patients with breast cancer (Scott et al., 1998). This individual radiosensitivity may be attributed to single nucleotide polymorphisms (SNPs) or mutations in DNA repair genes (Iarmarcovai et al., 2008). To verify this, it would be useful to measure the radiosensitivity of peripheral blood lymphocytes from individuals carrying candidate SNPs or mutations. However, using this strategy, CBMN assays may be affected by confounding factors such as smoking (Huang et al., 2009), or influenced by the diverse genetic backgrounds of the human population. It is therefore necessary to establish an evaluation system of radiosensitivity in a uniform genetic background with model human cells. This paper presents a novel genome editing strategy, ‘transcription activator-like effector nuclease (TALEN)-mediated two-step single base pair editing’, and it is proposed that this editing technique will be useful to investigate individual radiosensitivity.
2. CBMN ASSAY USING B-CELL LINES
In order to verify that variants in DNA repair genes are involved in individual differences in radiosensitivity, radiation-induced MN formation was studied in B-cell lines from family members with Nijmegen breakage syndrome (NBS). NBS is an autosomal-recessive disorder characterised by microcephaly, short stature, immunodeficiency, and a high incidence of cancer. NBS is caused by germline mutations in the DSB repair gene, NBS1 (Matsuura et al., 1998). Cultured cells from patients with NBS show chromosomal instability, increased sensitivity to radiation-induced cell killing, and abnormal cell-cycle regulation after irradiation (Chrzanowska et al., 2012).
Epstein-Barr (EB) virus-transformed lymphoblastoid cell lines were collected from homozygous patients with NBS 7998 and 8026 (NBS1−/−), heterozygous carriers 8000, 8001, 8002, 8004 and 8005 (NBS1+/−), and a control individual (NBS1+/+). Radiosensitivity was compared by CBMN assay. Cells (2.5 × 105 cells ml−1) were irradiated with 1 Gy of γ rays, and treated with 3 µg ml−1 cytochalacin-B for 48 h to form binucleated (BN) cells. The cells were grown on slides, fixed with cold methanol, and nuclei were counterstained with Hoechst33258. The number of MN in at least 1000 BN cells was counted manually under fluorescence microscopy. Control cells (NBS1+/+) showed no appreciable increase in MN formation, whereas cells from patients (NBS1−/−) exhibited an increase in MN formation after irradiation (Fig. 1). All cells showed an intermediate response between the control individual and the patients with NBS. These results suggest that the heterozygous mutation may underlie the individual differences in radiosensitivity. This study also found that the proportion of MN formation in cells varied considerably within genotypes, particularly for the NBS1+/− heterozygotes, and possibly for the patients with NBS to a lesser extent. These results suggest that the CBMN assay may be affected by the diverse genetic backgrounds of the human population.
Radiosensitivity of the Epstein-Barr virus-transformed lymphoblastoid cells established from family members with Nijmegen breakage syndrome (NBS). Male patient cell line 7998 was established from Family Kl in Germany, and heterozygous cell lines 8001 and 8000 were established from his father and mother, respectively. Male patient cell line 8026 was established from Family W in Germany. Heterozygous cell lines 8005, 8002 and 8004 were established from the patient’s siblings from Family 1 in the Netherlands (Matsuura et al., 1998). A founder mutation, 657-661del ACAAA, was detected in all three families (Matsuura et al., 1998). A control cell line was established from a healthy male individual. Cells from NBS heterozygous carriers showed intermediate frequency of micronucleus formation between that of control cells and the cells of patients with NBS. BN, binucleated cells.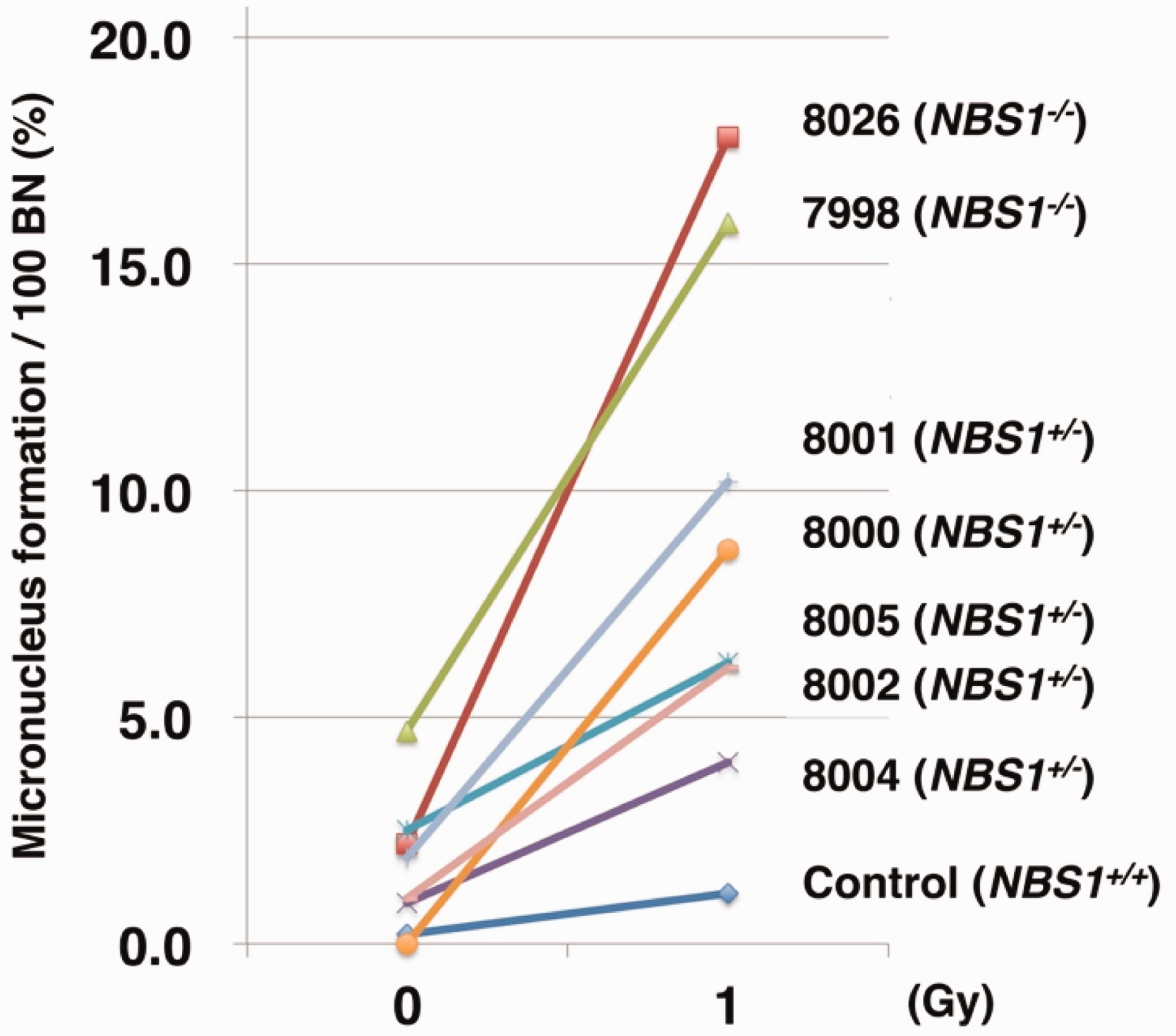
3. TALEN-MEDIATED TWO-STEP SINGLE BASE PAIR EDITING
Reverse genetics in cultured human cells with a uniform genetic background could be a direct approach to quantitative measurement of genotype-dependent radiosensitivity. Targeted gene modification in cultured human cells has been used previously to investigate gene function. However, this technique proved difficult due to the extremely low frequency of homologous recombination (HR) in human cells. Artificial nucleases have been developed recently, including zinc finger nucleases, TALENs, and clustered regulatory interspaced short palindromic repeat/Cas9-based RNA-guided DNA endonucleases (CRISPR/Cas9). The co-introduction of artificial nucleases with targeting vectors and subsequent antibiotic selection enabled efficient HR-mediated genome editing to be performed in mammalian cells (Soldner et al., 2011; Yusa et al., 2011).
A single base pair editing technique in cultured human cells was developed recently, and used to identify a causal mutation of a cancer-prone genetic disorder, premature chromatid separation with mosaic variegated aneuploidy [PCS (MVA)] syndrome (Ochiai et al., 2014). This is a rare human autosomal-recessive disorder characterised by constitutional aneuploidy and a high risk of childhood cancer (Matsuura et al., 2000). Both bi-allelic and mono-allelic mutations of the BUB1B gene encoding BubR1 have been identified in affected individuals. Mono-allelic mutations in BUB1B were identified in seven Japanese families with the syndrome (Matsuura et al., 2006). No second mutation was found in the opposite allele of any of the families studied, although a conserved BUB1B haplotype within a 200-kb interval and a reduced transcript were identified. Therefore, the nucleotide sequence of the 200-kb region was determined in a patient with the syndrome, and 186 known and five unknown SNPs were found. Of these SNPs, only one in an intergenic region 44-kb upstream of a BUB1B transcription start site co-segregated with the disease. This substitution was therefore a strong candidate mutation for the syndrome. If the nucleotide substitution identified was the disease-causing mutation, its introduction into cultured normal human cells should result in a decrease of BUB1B transcripts and increased chromosomal instability. To examine this possibility, a TALEN-mediated two-step single base pair editing strategy was designed (Fig. 2). The first step included TALEN-mediated targeted integration of a selection cassette into the SNP flanking region. The targeting vector contained a neomycin-resistant gene and a herpes simplex virus thymidine kinase (hsvTK) gene, separated by a 2A peptide sequence, allowing expression of the discrete protein products from a single open reading frame. The knock-in clones that were positively selected using neomycin. The second step involved the targeted excision of the selection cassette, and introduction of the single nucleotide substitution. Single-nucleotide-edited clones were selected negatively using ganciclovir treatment. This substitution was introduced bi-allelically into the human colon cancer cell line HCT116. Cell clones showed reduced BUB1B transcripts and chromosomal instability in the form of PCS and MVA, which are cellular hallmarks of the syndrome. These results suggest that the nucleotide substitution identified was the causal mutation of PCS (MVA) syndrome (Ochiai et al., 2014). The single base pair editing technique was therefore useful to investigate nucleotide variants of unknown functional relevance (Urnov, 2014).
Bi-allelic introduction of single nucleotide substitutions into cultured human cells using a transcription activator-like effector nuclease (TALEN)-mediated two-step single base pair editing technique. The first step involved the TALEN-mediated targeted integration of a selection marker into the single nucleotide polymorphism flanking region. The targeting vector contained a neomycin-resistant gene and a herpes simplex virus thymidine kinase gene separated by a 2A peptide. The second step involved the targeted excision of the selection cassette and introduction of the single nucleotide substitution.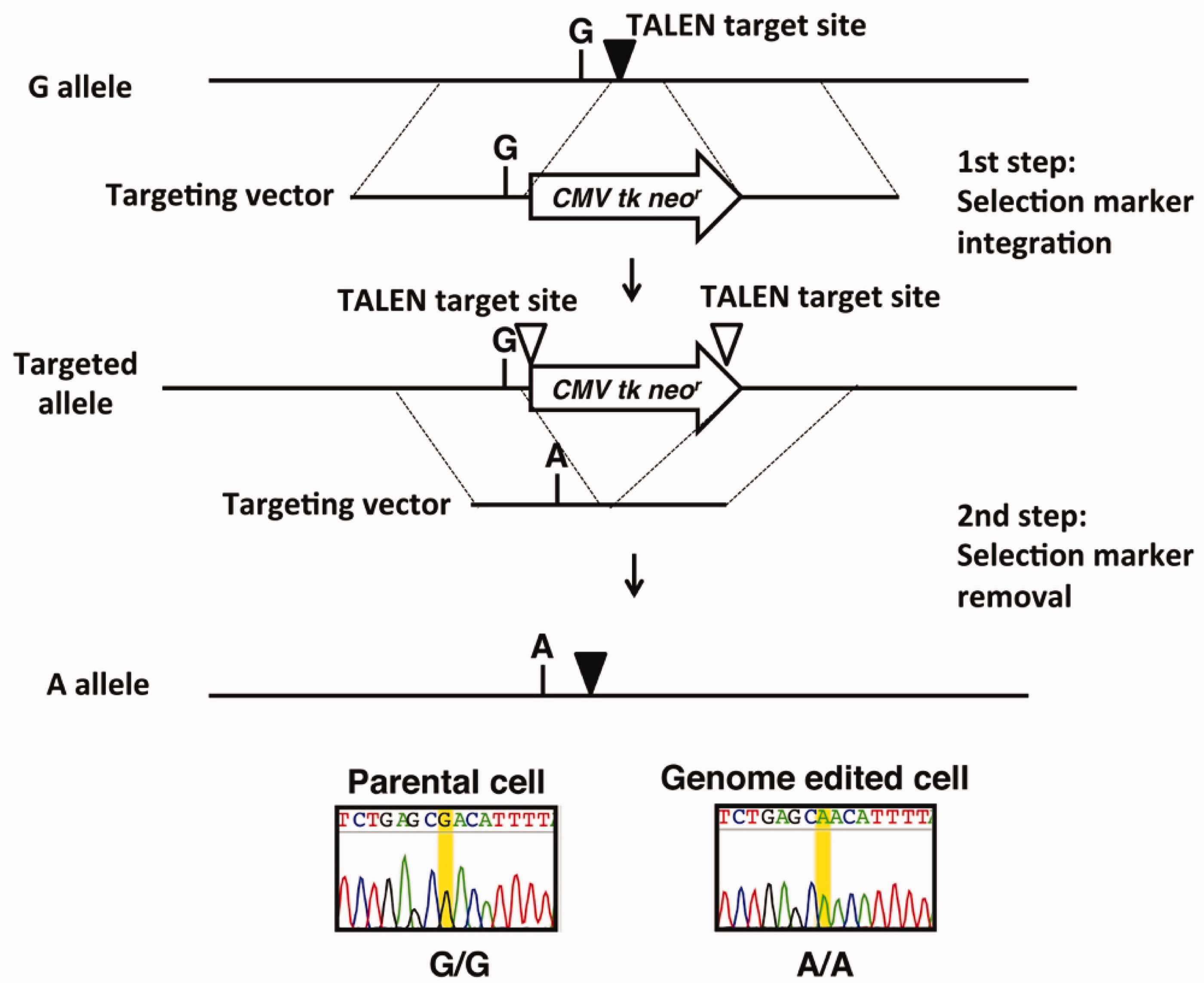
4. CONCLUSIONS
A number of epidemiological studies have demonstrated an increased risk of cancer incidence in normal individuals carrying nucleotide variants of DNA repair genes (Naccarati et al., 2007). This study showed that the radiation-induced MN formation of NBS heterozygous carriers was intermediate between that of patients with NBS and a control individual, as reported previously (Stumm et al., 2001; Neubauer et al., 2002; Dumon-Jones et al., 2003). These results suggest an apparent relationship between heterozygous mutations of familial hyper-radiosensitive diseases, mild radiosensitivity, and cancer incidence. As well as heterozygous mutations, many SNPs in DNA repair genes associated with risk of cancer have been reported. However, the functional evaluation of such variants may prove difficult because their effect sizes may be smaller than those of heterozygous mutations. It is therefore necessary to establish an evaluation system to compare radiosensitivity using model human cells with a uniform genetic background. This strategy will be useful to study the effects of candidate nucleotide variants underlying individual radiosensitivity.
The single base pair editing technique described here is able to introduce single nucleotide substitutions anywhere within the genome, without any footprints besides targeted sites; however, it is labour intensive because it requires multiple sets of TALENs, two separate targeting constructs, and two rounds of gene targeting. Recently, oligonucleotides have been used as donor templates to generate novel allelic variants of endogenous loci in a single step without selection (Chen et al., 2015). Moreover, the CRISPR/Cas9-based RNA-guided DNA endonuclease is currently emerging as a simple and efficient artificial nuclease (Cong et al., 2013). The targeted DNA cleavage by the CRISPR/Cas9 system only requires one short length (16–20 bases) of gRNA sequence for the recognition of target sequence and the common Cas9 enzyme, while TALENs require the design and assembly of two customised nucleases for one target locus. Therefore, it is much easier to engineer the CRISPR/Cas9 system than TALENs. These editing technologies are facilitating the future generation of model human cells carrying nucleotide variants of DNA repair genes for further investigation of radiosensitivity.
