Abstract
Introduction:
Chronic rhinosinusitis (CRS) is a heterogeneous inflammatory disease with variable treatment outcomes based on endotypes. Perioperative stratification of these patients remains a significant challenge. In the present study, we sought to use machine learning (ML) approaches in conjunction with clinical and inflammatory biomarker data to identify patients presenting for revision surgery as a proxy for recalcitrant disease and better understand which data types may be useful in classifying patients.
Methods:
Six hundred thirty-four CRS patients were included. All features were categorized into 5 sets: histopathologic markers, cytokine and chemokine markers, demographic factors, comorbidities, and medication history. Leave-one-out cross-validation was employed to train random forest (RF) classifiers separately according to each feature group.
Results:
There were 298 patients with CRS without nasal polyps (CRSsNP) and 336 patients with CRS with nasal polyps (CRSwNP). The training results of the RF model revealed that for CRSsNP patients, the combination of cytokine and chemokine markers possessed the highest predictive accuracy, while the classification accuracies of other feature groups were relatively lower. For CRSwNP patients, the combination of cytokine and chemokine markers along with eosinophil and neutrophil-related markers provided the highest classification performance. Considering all patients, the classification model obtained by jointly training multiple feature combinations demonstrated commendable performance.
Conclusion:
Integrating multiple clinical and inflammatory feature sets significantly enhances the predictive modeling of surgical history in CRS patients. These findings highlight the importance of multimodal data integration for improving classification performance and provide valuable insights for developing more accurate and clinically applicable ML models in CRS. The code has been made publicly available through https://github.com/hrlblab/CRS_prediction.
Introduction
Chronic rhinosinusitis (CRS) is a prevalent inflammatory disorder of the sinonasal mucosa, affecting ~5% to 12% of the general population.1,2 It is now widely recognized as a heterogeneous condition with distinct phenotypes and endotypes, as emphasized in the European Position Paper on Rhinosinusitis and Nasal Polyps 2020 (EPOS 2020). 3 It is associated with a significant healthcare burden due to frequent physician visits, medical treatments, and the need for surgical interventions. 4 CRS is typically classified into 2 major phenotypes: CRS with nasal polyps (CRSwNP) and CRS without nasal polyps (CRSsNP), each demonstrating distinct clinical and immunopathological characteristics. 5
Inflammatory biomarkers have emerged as crucial tools for differentiating CRS subtypes, guiding treatment decisions, and predicting disease progression.6,7 Indeed, cytokine profiling and molecular markers can provide key insights into the immune dysregulation underpinning CRS, potentially allowing for more targeted therapeutic strategies. 5 Type 1 inflammation, commonly associated with CRSsNP, is characterized by elevated levels of IFN-γ, IL-1β, IL-6, and IL-8, with a predominance of neutrophils. Conversely, CRSwNP exhibits a type 2 inflammation profile, dominated by IL-4, IL-5, and IL-13, often with eosinophilic infiltration. 8 Additionally, type 3 inflammation, characterized by IL-17A and IL-22, has been implicated in both phenotypes, further complicating treatment approaches. 6 Additionally, endotyping has therefore emerged as an important framework for understanding CRS heterogeneity and guiding future management, including the introduction of biologic therapies for patients with severe type 2-driven disease, as emphasized in recent European guidelines (EPOS 2020; EPOS/EUFOREA 2023).3,9
Surgical intervention, including endoscopic sinus surgery (ESS), is a cornerstone of CRS management when medical therapy fails. 10 However, a subset of patients experiences recurrent disease, necessitating revision surgery.4,11 Understanding the role of biomarkers in predicting surgical outcomes could optimize patient selection for surgery and identify individuals at risk for multiple interventions. 5 Recent studies suggest that cytokine expression patterns correlate with surgical history,12,13 with elevated IL-5 and IL-13 levels linked to increased revision rates in CRSwNP and mixed inflammatory profiles observed in CRSsNP. 6
With advancements in computational methods, machine learning (ML) has emerged as a powerful tool in CRS research, enabling the analysis of complex, nonlinear relationships between biomarkers, surgical history, and disease progression. 14 Several studies have demonstrated the potential of ML models in CRS classification, treatment response prediction, and surgical outcome forecasting. 15
Recent research has developed ML models to predict postoperative outcomes in CRS patients undergoing ESS. 14 By integrating preoperative clinical parameters, biomarker profiles, and imaging data, these models have achieved high accuracy in forecasting disease control, partial control, or relapse after surgery.16,17 Some approaches have incorporated microRNA markers, significantly improving predictive performance. 18 Additionally, ML has been applied to eosinophilic CRS prediction, utilizing a combination of blood biomarkers, inflammatory mediators, and clinical observations to differentiate subtypes with high sensitivity and specificity. 14
In this study, we apply ML techniques to analyze the relationship between biomarker profiles and surgical history in CRS patients. Our objective is to develop predictive models that can stratify patients based on their likelihood of requiring revision surgery, providing insights into the pathophysiology of treatment-refractory CRS. We hypothesize that specific cytokine patterns and inflammatory endotypes are associated with surgical recurrence and that ML-driven analysis can improve prognostic accuracy beyond traditional clinical parameters.
Materials and Methods
Patient Enrollment and Data Curation
Adults presenting to the Vanderbilt Asthma, Sinus, and Allergy Program or the Vanderbilt Bill Wilkerson Center for ESS between September 2015 and September 2024 enrolled in an IRB-approved longitudinal prospective cohort study. The diagnosis of CRS was determined according to the guidelines outlined in the European Position Paper on Rhinosinusitis and Nasal Polyps, as well as the International Consensus Statement on Allergy and Rhinology, 19 and patients were selected for surgery after failing a trial of appropriate medical therapy. Patients were excluded if they had odontogenic rhinosinusitis, suspected mycetoma, current monoclonal antibody therapy, systemic steroid use within 1 month of surgery, cystic fibrosis, or known autoimmune disease. Two classification tasks were defined based on surgical history: a binary classification to predict whether a patient had ever undergone sinus surgery, and a multi-class classification task to predict the number of prior surgeries a patient had undergone.
To ensure data integrity, all samples missing the target variable for either classification task were excluded. For the binary task, this included missing values in the variable indicating prior surgical experience, while for the multi-class task, samples lacking a valid count of previous surgeries were removed. For the remaining features, missing values were imputed using the column-wise mean. All numerical features were then standardized using z-score normalization to ensure consistency in scale across variables.
Cytokine and Cell Count Assessment
Prior to beginning each surgical procedure, a 9 × 24 mm polyurethane sponge was placed into the middle meatus in each side under endoscopic visualization and left in place for 5 minutes. This was then retrieved and centrifuged at 14 000g for 10 minutes to elute mucus. Samples were then gently vortexed and again centrifuged for 5 minutes to remove cellular debris. Supernatants were removed and frozen at −80 °C for subsequent cytokine analysis. Cytokine assays were performed using a standard sensitivity multiplex cytokine bead assay (BD Biosciences, Franklin Lakes, NJ, USA) according to the manufacturer’s protocol; the procedure for assessment has been previously described.20,21 Histopathological evaluation of excised sinonasal ethmoid tissue was performed by a head and neck pathologist in a blinded fashion, and the mean number of eosinophils and neutrophils over 5 randomly selected high-powered fields (HPF) was recorded.
Feature Extraction
Based on clinical relevance and data availability, all features were grouped into 5 functional categories:
Eosinophil and Neutrophil-Related Histopathologic Markers (EO): Eosinophil count per HPF, neutrophil count per HPF, computed tomography score, and eosinophil-based histopathology score.
Cytokine and Chemokine Markers (CY): Mucus biomarkers based on a cytometric bead assay measuring levels of 17 inflammatory cytokines and chemokines (IL-1α, IL-1β, IL-2, IL-3, IL-4, IL-5, IL-6, IL-7, IL-8, IL-9, IL-10, IL-12/IL-23 (p40), IL-13, IL-17A, IL-21, tumor necrosis factor-α, interferon-γ, Eotaxin, RANTES, GM-CSF).
Demographic Factors (DM): Sex, race, BMI, and whether patients are current smoker.
Comorbidities (CM): Presence of other comorbid medical conditions, including asthma, allergic rhinitis, migraine, other headache disorders, fibromyalgia, arthritis, temporomandibular joint disorder, obstructive sleep apnea, gastroesophageal reflux disease, eustachian tube dysfunction, congestive heart failure, chronic obstructive pulmonary disease, diabetes mellitus type 1, diabetes mellitus type 2, and ear pain.
Medication History (MH): Preoperative medication use data, including antihypertensive medications, nasal corticosteroid medications, antidepressant medications, migraine medications, psychiatric medications, gastroesophageal reflux disease medications, and anti-leukotriene medications.
For the binary classification task, the target variable was defined as the presence or absence of any prior sinus surgery. For the multi-class task, the number of previous surgical procedures was discretized into 3 categories: no prior surgery (0), a single surgery (1), and multiple surgeries (>1).
These class labels served as the outcome variables for their respective tasks.
ML Classification
For both tasks, we adopted the leave-one-out cross-validation (LOOCV) approach, and we trained random forest (RF) models with different feature sets. In each iteration of LOOCV, 1 subject is held out as the test case while the remaining subjects are used for training, thereby ensuring that nearly the entire dataset contributes to model fitting and that every patient is evaluated exactly once. This approach reduces the risk of biased estimates that may arise from small training folds in k-fold CV, particularly after stratification into CRSsNP and CRSwNP subgroups. Although LOOCV can be computationally more intensive and may produce higher variance estimates than repeated k-fold CV, our experimental design mitigated this concern by repeating LOOCV across 100 random seeds. This strategy provided a robust and nearly unbiased assessment of model performance given the available data. The mean value across 100 experiments was taken as the final result. Model performance was evaluated using macro-average area under the receiver operating characteristic curve (AUC-Macro), with final results reported as the average across all 100 runs. In addition, we reported the logistic regression (LR) training results as a baseline for comparison.
Statistical Analysis
To compare predictive performance across feature sets, we applied paired 2-sided t-tests on the distribution of AUCs obtained from repeated LOOCV runs (n = 100 seeds). For each subtype (CRSsNP and CRSwNP) and task (binary and multi-class), pairwise tests were performed between the integrated feature sets (eg, EO + CY + DM + CM vs individual groups such as CY alone). A significance threshold of P < .05 was adopted. No correction for multiple testing was applied, given the exploratory nature of the analysis; results of significant comparisons are reported in Figure 1. All analyses were performed in Python (version 3.12.9) using the SciPy Library (version 1.15.2).
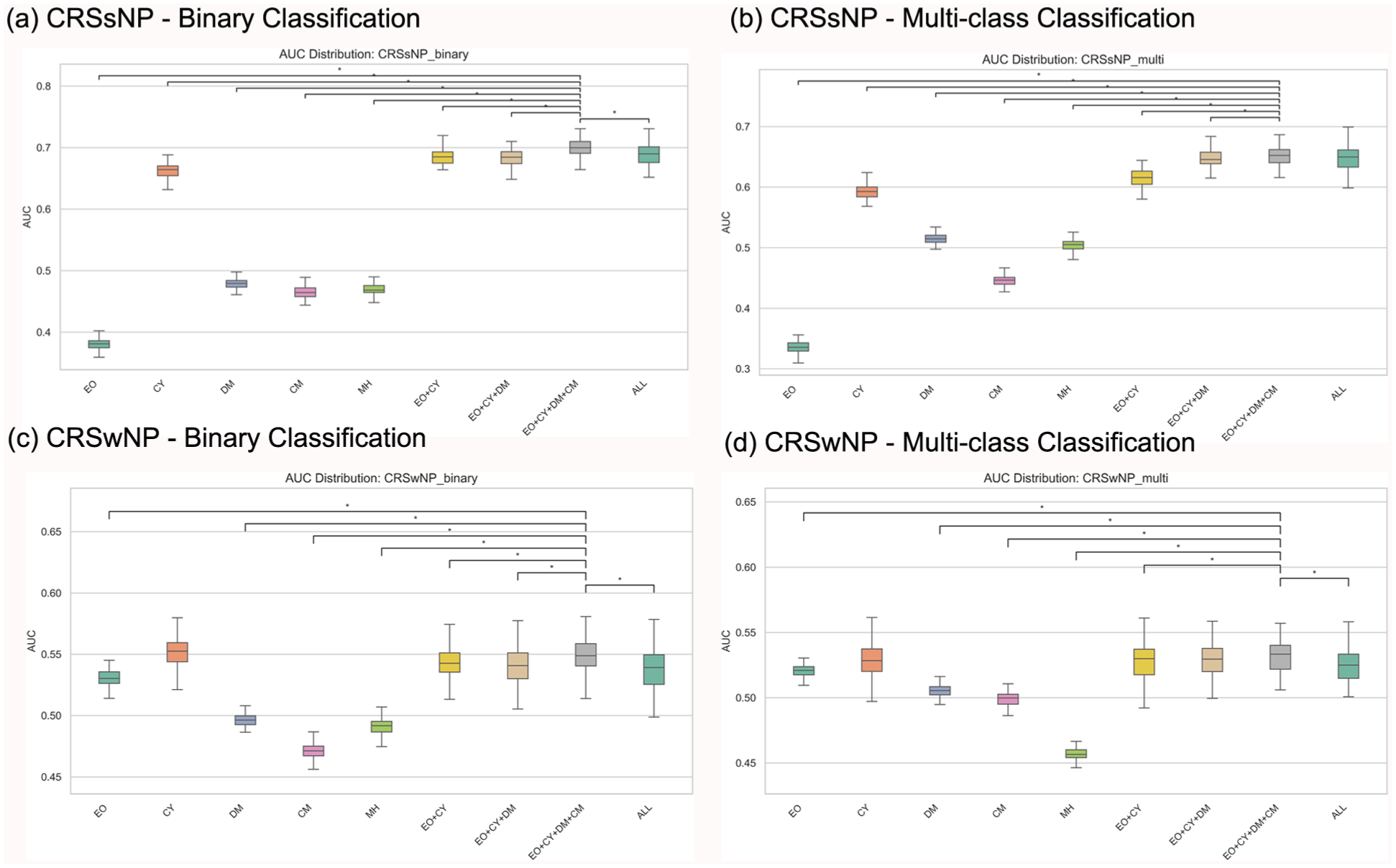
AUC distributions and paired t-test results across different feature sets for 4 classification tasks. Each subfigure shows the AUC distributions for the evaluated feature combinations. Statistical comparisons were performed between the EO + CY + DM + CM combination and all other feature sets.
Results
The classification performance of different feature sets was evaluated using the AUC-Macro metric. All features were categorized into 5 sets: eosinophil and neutrophil-related markers, cytokine and chemokine markers, demographic factors, comorbidities, and MH of the patients. The results demonstrate the predictive capability of these feature sets within the 2 symptom subtypes, CRSsNP and CRSwNP, individually, highlighting variations in classification power when different features or their combinations are used (Table 1).
Clinical and Demographic Characteristics of Enrolled CRS Patients.

Abbreviation: CRS, chronic rhinosinusitis.
Mean (SD; Range); n (%).
CRSsNP Internal Performance Comparison
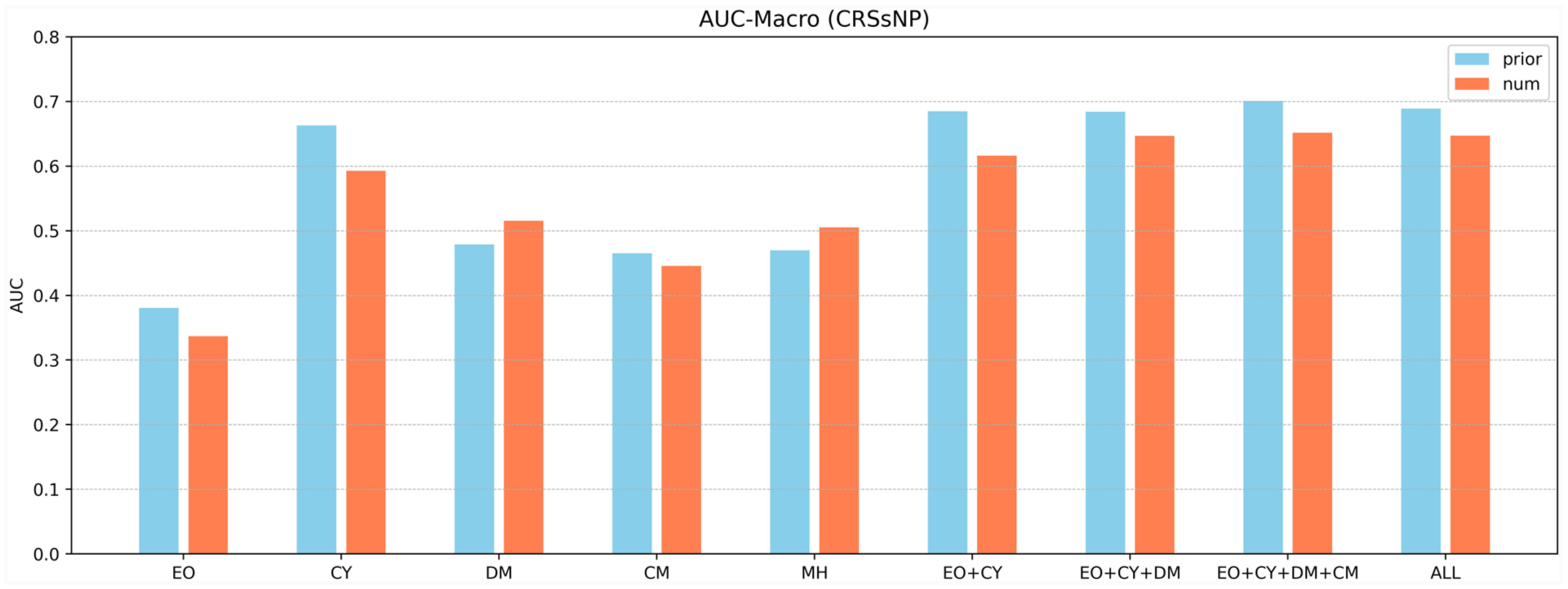
Within the CRSsNP symptom cohort, as shown in Figure 2 and Table 2, RF model with CY achieved the highest AUC-Macro value among single feature groups, with an AUC of 0.6630 in the binary classification task and 0.5927 in the multi-class classification task. EO yielded AUCs of 0.3806 and 0.3367 for the binary and multi-class tasks, respectively. DM achieved AUCs of 0.4787 (binary) and 0.5154 (multi-class), while CM showed AUCs of 0.4650 (binary) and 0.4453 (multi-class). MH feature set yielded AUCs of 0.4697 (binary) and 0.5051 (multi-class).
AUC results for CRSsNP.
AUC Results for CRSsNP.
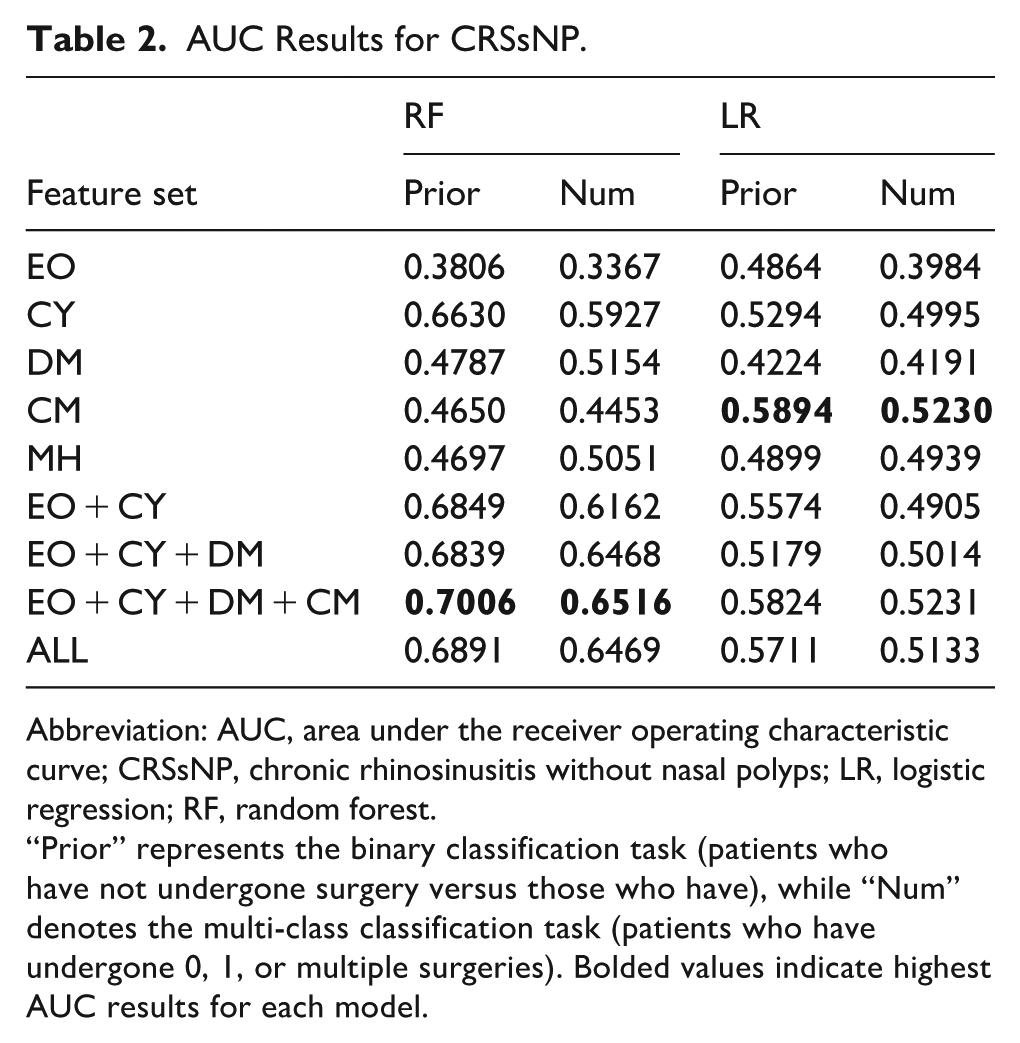
Abbreviation: AUC, area under the receiver operating characteristic curve; CRSsNP, chronic rhinosinusitis without nasal polyps; LR, logistic regression; RF, random forest.
“Prior” represents the binary classification task (patients who have not undergone surgery versus those who have), while “Num” denotes the multi-class classification task (patients who have undergone 0, 1, or multiple surgeries). Bolded values indicate highest AUC results for each model.
When combining feature sets, EO + CY achieved AUCs of 0.6849 (binary) and 0.6162 (multi-class). EO + CY + DM further increased the AUCs to 0.6839 and 0.6468, respectively. The combination of EO + CY + DM + CM yielded the highest overall performance, achieving AUCs of 0.7006 in the binary task and 0.6516 in the multi-class task. Using all available features (ALL) resulted in AUCs of 0.6891 (binary) and 0.6469 (multi-class).
In comparison, the LR model demonstrated consistently lower performance across both tasks. Among the single feature sets, its best results were observed with CY (AUCs of 0.5294 in binary and 0.5096 in multi-class), while EO, DM, CM, and MH all yielded AUCs close to 0.50. For integrated feature sets, EO + CY + DM + CM achieved the highest LR performance with AUCs of 0.5824 (binary) and 0.5231 (multi-class), still notably below the corresponding RF results. These findings indicate that RF provided superior discriminative ability in CRSsNP compared to LR, particularly when multiple feature groups were integrated.
CRSwNP Internal Performance Comparison
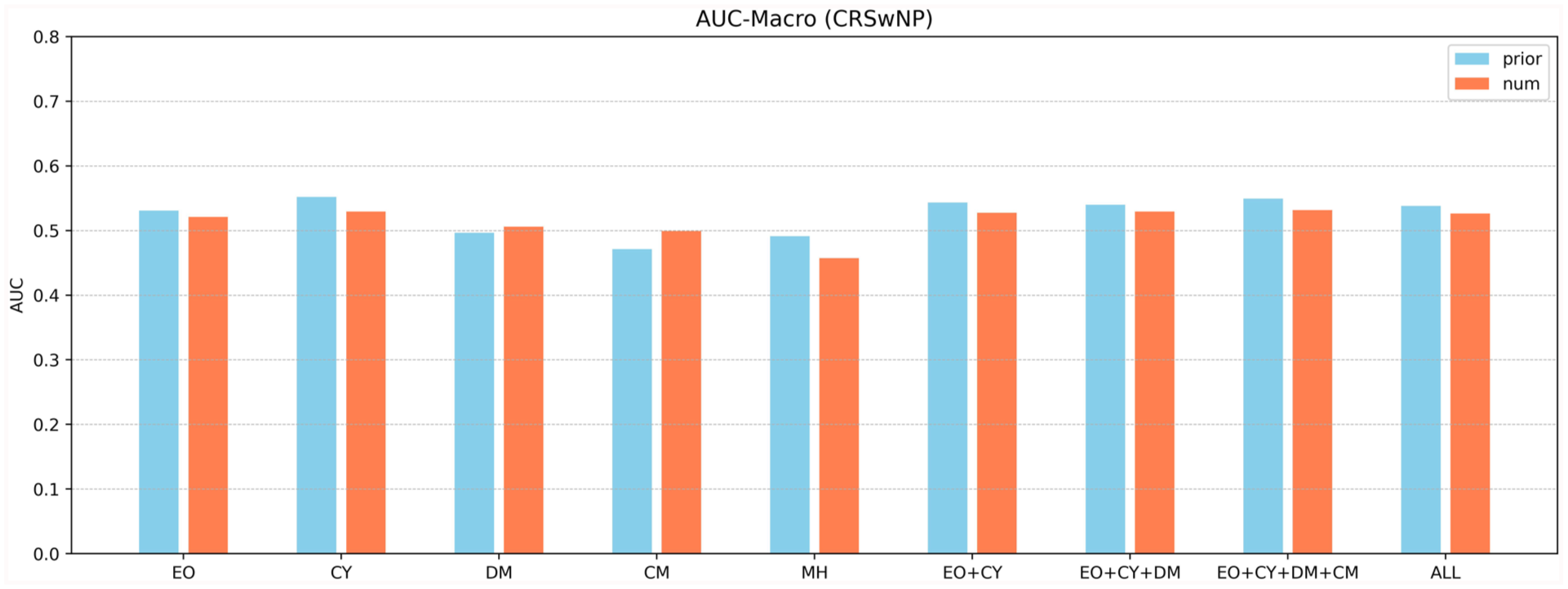
Within the CRSwNP cohort, shown in Figure 3 and Table 3, the RF model with EO achieved an AUC of 0.5306 in the binary classification task and 0.5209 in the multi-class classification task. CY achieved slightly higher AUCs of 0.5519 (binary) and 0.5293 (multi-class).
AUC results for CRSwNP.
AUC Results for CRSwNP.
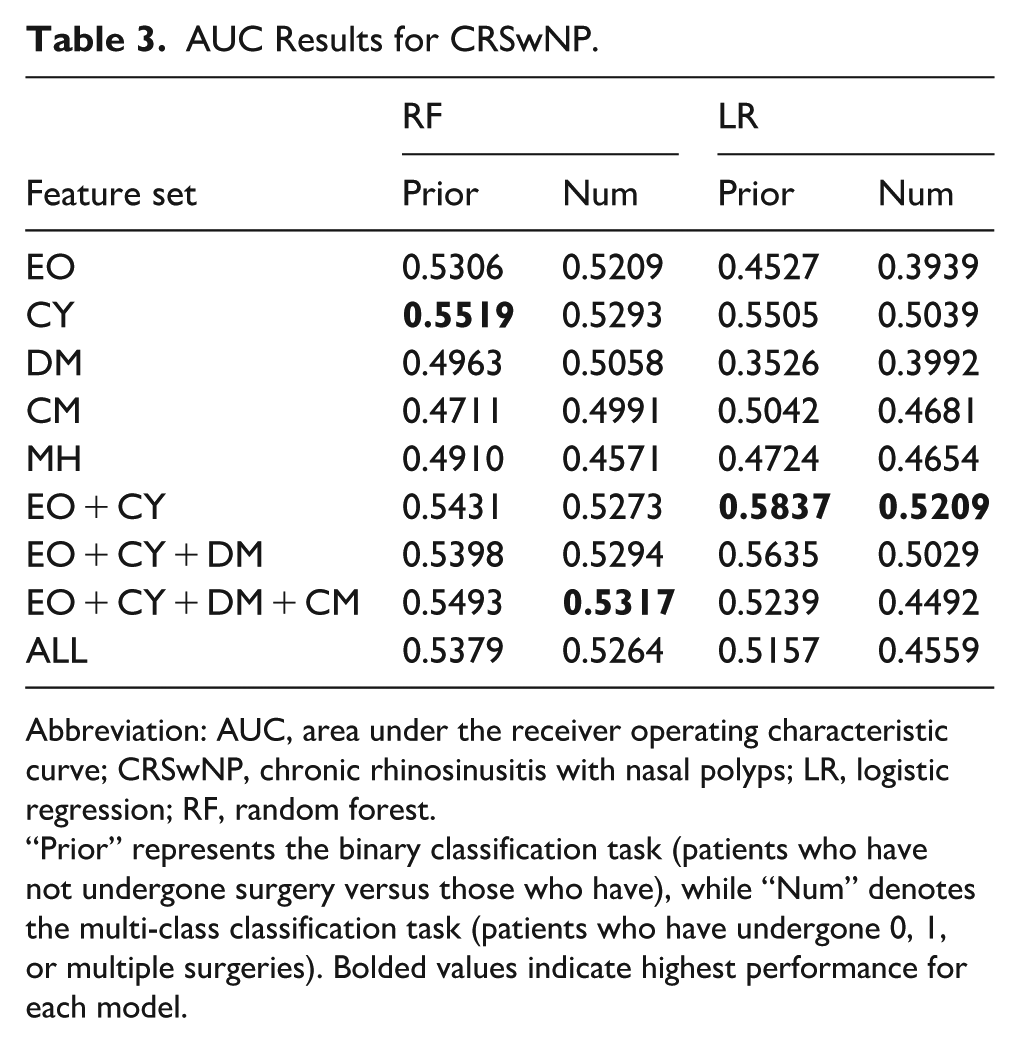
Abbreviation: AUC, area under the receiver operating characteristic curve; CRSwNP, chronic rhinosinusitis with nasal polyps; LR, logistic regression; RF, random forest.
“Prior” represents the binary classification task (patients who have not undergone surgery versus those who have), while “Num” denotes the multi-class classification task (patients who have undergone 0, 1, or multiple surgeries). Bolded values indicate highest performance for each model.
DM yielded AUCs of 0.4963 (binary) and 0.5058 (multi-class), while CM yielded 0.4711 (binary) and 0.4991 (multi-class). MH showed AUCs of 0.4910 (binary) and 0.4571 (multi-class).
For feature combinations, EO + CY achieved AUCs of 0.5431 (binary) and 0.5273 (multi-class). EO + CY + DM resulted in AUCs of 0.5398 (binary) and 0.5294 (multi-class). The combination of EO + CY + DM + CM yielded AUCs of 0.5493 in the binary task and 0.5317 in the multi-class task, representing the highest performance among the tested feature combinations. Using all features (ALL) achieved AUCs of 0.5379 (binary) and 0.5264 (multi-class).
In comparison, the LR model demonstrated similarly constrained performance. Among the single feature sets, CY again yielded the best results with AUCs of 0.5727 (binary) and 0.5051 (multi-class), slightly higher than RF in the binary task. EO and DM produced AUCs close to 0.50, while CM and MH performed near chance level. For integrated feature sets, EO + CY provided the highest LR result in the binary task (0.5837), marginally exceeding RF for the same feature set, whereas EO + CY + DM + CM reached 0.5239 in the multi-class task, which remained lower than the corresponding RF performance. These findings suggest that in CRSwNP, both RF and LR models offered only limited discriminative ability, with occasional cases where LR marginally outperformed RF, but overall results remained close to chance.
Statistical Significance of Feature Group Differences
The distribution of AUC values for different feature combinations across the 4 classification tasks and paired t-test results are shown in Figure 1.
CRSsNP Binary Classification
The EO + CY + DM + CM combination achieved a mean AUC of 0.7006. Paired t-tests between EO + CY + DM + CM and other feature combinations yielded the following P values: EO (P < .001), CY (P < .001), DM (P < .001), CM (P < .001), MH (P < .001), EO + CY (P < .001), EO + CY + DM (P < .001), ALL (P < .001).
CRSsNP Multi-Class Classification
The EO + CY + DM + CM combination achieved a mean AUC of 0.6516. Paired t-tests between EO + CY + DM + CM and other feature combinations yielded the following P values: EO (P < .001), CY (P < .001), DM (P < .001), CM (P < .001), MH (P < .001), EO + CY (P < .001), EO + CY + DM (P = .036), ALL (P = .069).
CRSwNP Binary Classification
The EO + CY + DM + CM combination achieved a mean AUC of 0.5493. Paired t-tests between EO + CY + DM + CM and other feature combinations yielded the following P values: EO (P < .001), CY (P = .186), DM (P < .001), CM (P < .001), MH (P < .001), EO + CY (P = .002), EO + CY + DM (P < .001), ALL (P < .001).
CRSwNP Multi-Class Classification
The EO + CY + DM + CM combination achieved a mean AUC of 0.5317. Paired t-tests between EO + CY + DM + CM and other feature combinations yielded the following P values: EO (P < .001), CY (P = .166), DM (P < .001), CM (P < .001), MH (P < .001), EO + CY (P = .023), EO + CY + DM (P = .254), ALL (P = .015).
Discussion
Decision Curve Analysis
As is shown in Figures 4 and 5, decision curve analysis (DCA) was performed to evaluate the clinical utility of the predictive models. In the CRSsNP cohort, RF models demonstrated modest positive net benefit across a limited range of threshold probabilities (~0.2-0.6), with integrated feature sets such as EO + CY + DM + CM and ALL performing slightly better than single feature groups. In the CRSwNP cohort, DCA revealed that the net benefit of RF models was very limited. No feature set consistently demonstrated a clinically meaningful advantage.
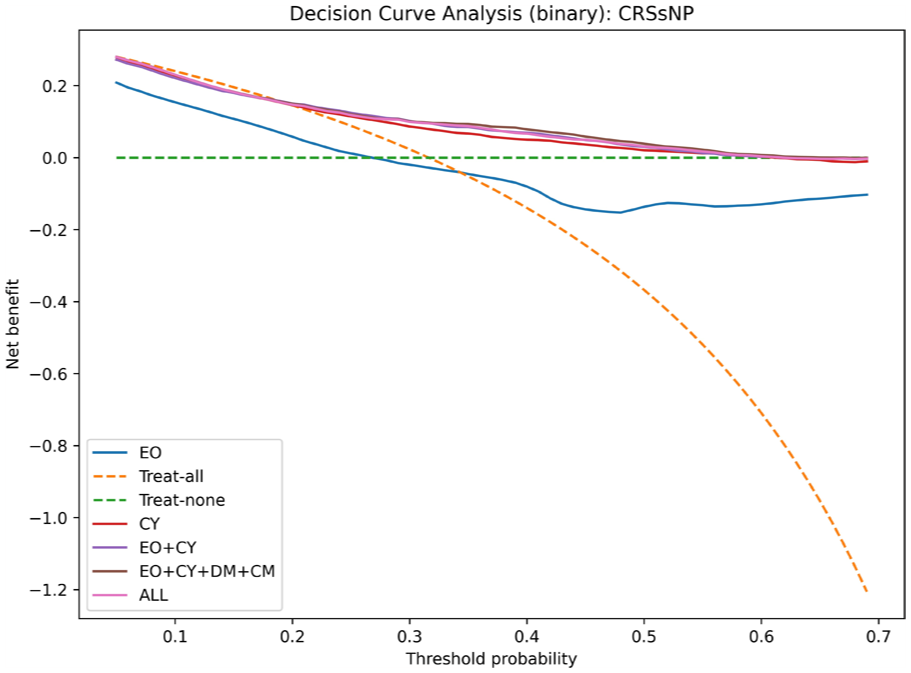
Decision curve analysis for CRSsNP. Threshold probability is restricted to the range of 0.05 to 0.7 to reflect the clinically relevant decision space. CRSwNP, chronic rhinosinusitis with nasal polyps.
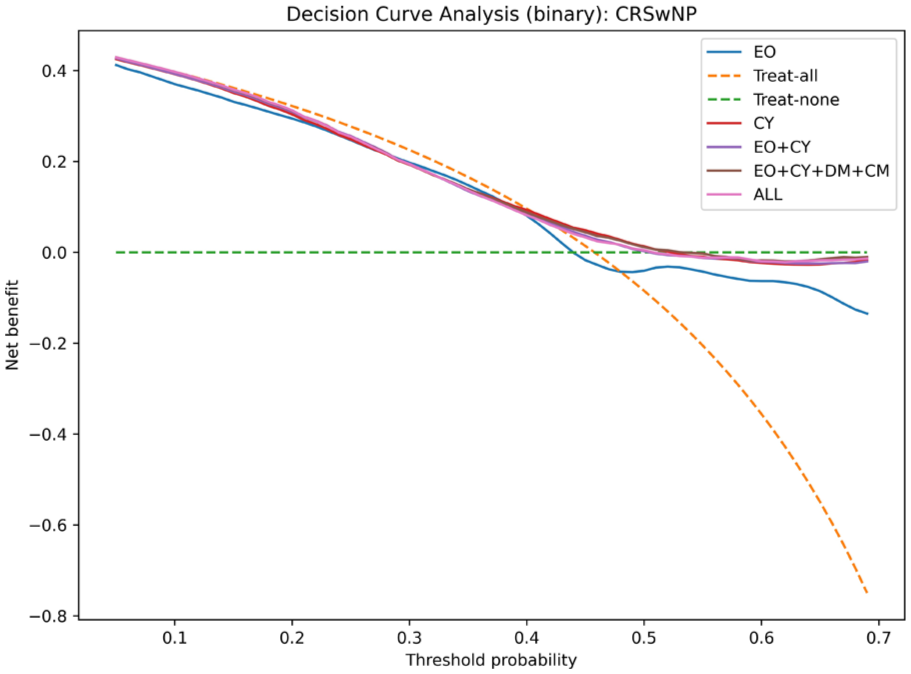
Decision curve analysis for CRSsNP. Threshold probability is restricted to the range of 0.05 to 0.7 to reflect the clinically relevant decision space. CRSNP, chronic rhinosinusitis with nasal polyps.
Summary of Findings
Analysis of Individual Feature Sets
In the CRSsNP cohort, the feature set consisting of cytokine and chemokine markers demonstrated the strongest individual predictive capability, suggesting the central role of inflammatory mediators in distinguishing patients who had undergone surgery. Eosinophil and neutrophil markers also contributed to classification performance but to a lesser extent. Demographic and comorbidity features showed relatively limited discriminatory power when used independently. The inclusion of MH as a feature appeared to negatively affect classification results, potentially due to treatment-related variability among patients. The highest classification performance was achieved by combining multiple feature sets, particularly EO, CY, DM, and CM, while the addition of MH did not yield further improvement.
In the CRSwNP cohort, the predictive performance of individual feature sets was generally lower compared to CRSsNP. Eosinophil and neutrophil markers exhibited relatively strong performance among single feature sets, underscoring the contribution of eosinophilic inflammation in CRSwNP pathophysiology. However, demographic factors, comorbidities, and MH individually contributed little to classification accuracy. Combining eosinophil and cytokine markers led to slight improvements, but the overall classification gains through feature integration were modest. This suggests that CRSwNP classification based on preoperative clinical features remains challenging, likely due to the heterogeneous nature of the disease and the limited signal strength of available markers.
Analysis of Overall Performance
Across all classification tasks and subtypes, models incorporating the feature combination of EO + CY + DM + CM consistently achieved the highest or near-highest AUC values. In the CRSsNP group, the EO + CY + DM + CM combination yielded an AUC of 0.7006 for the binary classification task and 0.6516 for the multi-class classification task. In the CRSwNP group, the EO + CY + DM + CM combination resulted in AUCs of 0.5493 and 0.5317 for the binary and multi-class tasks, respectively.
The paired t-test results demonstrated that, particularly in CRSsNP, EO + CY + DM + CM significantly outperformed most individual feature sets and smaller combinations (eg, EO, CY, DM, CM). In CRSwNP, although the overall classification performance was lower, the EO + CY + DM + CM combination still provided a consistent advantage over individual feature sets.
The analysis of feature combinations revealed that, within CRSsNP, incorporating cytokine and chemokine markers significantly improved classification performance, reinforcing the importance of inflammatory mediators in distinguishing cases. While adding demographic, comorbidity, and medication-related features provided a minor increase in classification accuracy, the overall gain was limited, and the inclusion of MH even reduced classification performance in some cases. In contrast, within CRSwNP, combining eosinophilic and cytokine-related feature sets produced the highest classification performance, confirming the synergistic role of eosinophilic inflammation and cytokine activity. Further incorporation of demographic, comorbidity, and medication-related variables resulted in only marginal improvements in predictive power. When multiple feature sets were combined, the highest AUC-Macro scores were achieved, demonstrating that multimodal feature integration enhances classification performance, although the primary predictive contributions stem from core inflammatory markers.
Limitations and Future Work
This study has several limitations. First, the range of cytokine and chemokine markers included was relatively limited, which may have restricted the ability to fully capture the complex inflammatory endotypes associated with CRS. Important mediators such as IL-17A, IL-22, and thymic stromal lymphopoietin (TSLP) were not assessed, potentially missing key inflammatory signals relevant to disease progression and surgical outcomes.
Second, the patient cohort size was modest, particularly after stratification into CRSsNP and CRSwNP subgroups, which may have limited the generalizability and statistical power of the findings. Small sample sizes increase the risk of overfitting and reduce the robustness of ML models when applied to external populations.
Third, the use of revision surgery as the primary outcome should be interpreted with caution. While repeat surgical intervention can reflect persistent disease, it is not a pure marker of biologically recalcitrant CRS. Decisions regarding revision surgery are also influenced by surgeon preference, patient decision-making, surgical technique, adherence to postoperative care, and access to healthcare services. These non-biological factors may confound the association between revision surgery and true disease intractability, thereby limiting the specificity of our outcome measure.
Fourth, the overall predictive performance of both RF and LR models was modest, with many AUC values hovering around 0.5 and only a few clearly exceeding this threshold. This highlights important limitations in the current modeling approach. Several factors may account for this observation: (i) the relatively small cohort size, particularly after stratification into CRSsNP and CRSwNP, which increases noise and limits statistical power; (ii) the restricted set of cytokine and chemokine markers, which may not fully capture the biological heterogeneity of CRS; and (iii) the intrinsic complexity and multifactorial nature of CRS, where non-biological influences such as surgical technique, adherence, and healthcare access also affect outcomes. Together, these issues likely contributed to the low discriminative ability observed across models.
Future work should aim to expand the panel of cytokine and chemokine measurements to provide a more comprehensive characterization of the immune landscape in CRS. Additionally, larger, multi-center datasets should be collected to enhance the external validity and clinical applicability of predictive models. Advanced modeling approaches, including deep learning architectures and longitudinal modeling techniques, may also be explored to capture temporal patterns and improve predictive performance.
Conclusion
This study provides a systematic evaluation of feature set integration for predicting surgical history in CRS patients, with separate analyses for CRSsNP and CRSwNP subtypes. Across 4 classification tasks, the combination of eosinophil and neutrophil markers, cytokine and chemokine markers, demographic factors, and comorbidities (EO + CY + DM + CM) consistently achieved superior or near-superior predictive performance in the CRSsNP cohort. Paired statistical tests confirmed the significance of this combination over individual feature sets in multiple settings. In contrast, for CRSwNP patients, the predictive performance of both individual and combined feature sets was only marginally above chance, underscoring the substantial heterogeneity of this subgroup and highlighting that current models are not yet clinically useful for this population.
These results offer valuable insights for future research, emphasizing the importance of expanding cytokine panels, leveraging larger multi-center datasets, and adopting advanced modeling techniques to improve the predictive accuracy and clinical applicability of ML models in CRS.
Footnotes
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This project was supported by the Burroughs Wellcome Fund PSIA award (N.I.C.), NIAID L30AI186158 (N.I.C.), and CTSA award UL1TR000445 from the National Center for Advancing Translational Sciences. Its contents are solely the responsibility of the authors and do not necessarily represent official views of the National Center for Advancing Translational Sciences or the National Institutes of Health.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
