Abstract
Objective
Today, several new machine learning models are being applied to perform survival analyses; however, due to the variations of the size and characteristics of samples, predictive performance of many models is not ideal. In this work, relevant data were collected from the surveillance, epidemiology, and end results (SEER) database of esophageal cancer patients and a multivariate survival analysis model was established based on DeepSurv model to predict the survival rate of esophageal cancer patients and provide reasonable recommendations for individual treatment of these patients. The application of novel machine learning models in survival analysis has been gaining momentum; however, due to the heterogeneity in sample size and characteristics, the predictive accuracy of many models remains suboptimal. The aim was to forecast the survival rates of esophageal cancer patients and to offer informed recommendations for personalized treatment strategies.
Methods
In this retrospective cohort study, the data of 5276 patients were collected from the SEER database using SEER*Stat software(version 8.4.3). The collected data was divided into training and testing sets with 7:3 ratio and the predictive performance of the model was explored using the concordance index (CI). The efficiencies of different treatment methods were compared. In this study, we developed a survival analysis model based on the DeepSurv model. The data was stratified into training (70%, n = 3693) and testing (30%, n = 1583) sets with a 7:3 ratio. The model's predictive efficacy was assessed using the CI, and the effectiveness of various treatment modalities was comparatively analyzed.
Results
The selected 5276 patients with esophageal cancer were divided into a training set (70%, 3693) and a testing set (30%, 1583). Cox univariate regression analysis results showed that race, diagnosis year, and marital status were risk factors affecting prognosis (p < 0.05). The consistency indexes of training and test sets were 0.84 and 0.83, respectively. These tests indicated that the model had good performance. There is a significant difference in the therapeutic efficacy between the surgical group and the non-surgical group (p < 0.001).
Conclusions
According to DeepSurv model, a survival prediction model was developed for patients with esophageal cancer and reasonable suggestions were provided for individual treatment selection. Utilizing the DeepSurv model, we have developed a predictive survival model tailored for esophageal cancer patients, providing actionable insights for the selection of individualized treatment approaches.
Introduction
Esophageal cancer is a prevalent and aggressive malignancy in clinical oncology. The global incidence and mortality rates of esophageal cancer in malignant tumors are about 3.2% and 5.3%, ranking 7th and 6th in malignant tumors, respectively.1,2 With the advent of innovative therapeutic approaches, the survival rate of patients with esophageal cancer has been significantly improved. Nonetheless, for those afflicted with advanced stages of the disease, the 5-year survival rate remains disconcertingly low, typically falling below 25%. 3 The etiology of esophageal cancer is multifactorial, especially in early stages where the symptoms are subtle and nonspecific. This leads to a delayed diagnosis for the majority of patients, who are typically identified during the intermediate to late stages of the disease, significantly impacting their prognosis. Consequently, it is imperative to proactively identify and understand the risk factors that influence patient outcomes.4–6
There are many factors that affect the prognosis of esophageal cancer, such as age, tumor stage, case type, and treatment modalities. Previous researchers in this domain have predominantly employed conventional statistical techniques such as Cox proportional hazards (CoxPHs) regression and logistic regression to construct predictive models, which were often constrained by limited sample size and features.7,8
CoxPH is the most common semi-parametric model. 9 Despite its utility, the CoxPH model's risk function is based on a linear assumption, which may oversimplify the complex dynamics of disease progression. This limitation has prompted the need for a more sophisticated and robust survival prediction model.
With the popularity of machine learning, many algorithms have been applied to survival analysis, including survival decision tree,10,11 support vector machines,12,13 random forest,14,15 and nonlinear CoxPH methods. Notably, deep learning-based survival analysis models have emerged as a powerful tool, adept at handling nonlinear relationships without the restrictive assumptions inherent in traditional models.16–18 This research applied deep survival analysis models to conduct statistical analyses. We aim to enhance the accuracy of survival predictions and provide a rational framework for patients to make more informed decisions regarding their treatment strategies. 19
Data and analysis methods
Data
All esophageal cancer patients were selected from surveillance, epidemiology, and end results (SEER) database (SEER Research Data 17 Registries, Nov 2020 Sub (2010–2020)), which contained data of cancer patients in the United States. This database included a great amount of patient-related information, including demographic information, tumor data information, treatment methods, and prognosis information. All patient information has been de-identified. This study is a retrospective cohort study, and we have signed a data usage agreement and obtained permission for data use. This study was conducted in accordance with the 1975 Helsinki Declaration revised in 2024 and was waived by an Institutional Review Board (IRB)/Independent Ethics Committee (IEC).
Data acquisition process
In this research, screening process for esophageal cancer patient data is illustrated in Figure 1.
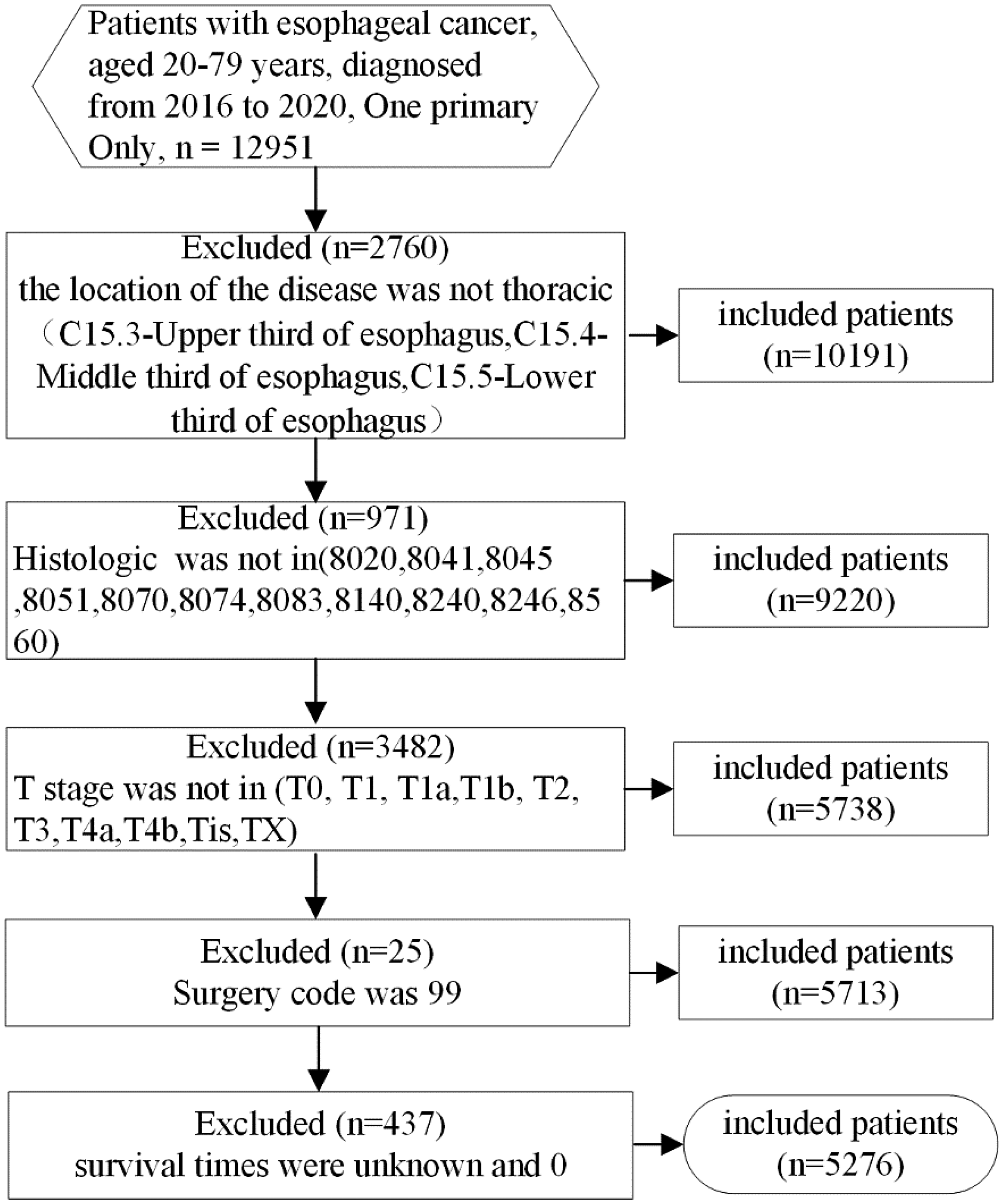
Data screening flowchart for esophageal cancer patients.
Analysis and design of deep learning models
DeepSurv is a deep learning method for predicting the risk of patient death, including an input layer, hidden layer, and output layer. This study is based on the DeepSurv model, which uses deep learning technology to predict the survival status of patients and provide recommendations for treatment methods. The reporting of this study conforms to STROBE guidelines. 20
Model structure
Input layer: Input layer receives clinical characteristics and other relevant information of patients. These characteristics include age, gender, race, diagnosis year, marital status, tumor stage, treatment method, etc. These input features were standardized or normalized to ensure good numerical stability during training process.
Hidden layer: This layer was the core computational part of the model, containing multiple neurons which were applied to extract high-dimensional features from input data. Each hidden layer was transformed using a nonlinear activation function (such as ReLU or SELU, as shown in formulas 1 and 2), which enabled the model to capture complex relationships within the data. The numbers of hidden layers and neurons in each layer were determined by experimental optimization to ensure the accuracy and generalization capability of the model.
Output layer: This layer contained neurons to predict patient's risk function. This layer outputs a continuous value through a linear activation function, representing the survival risk of the patients. To train the model, negative log partial likelihood was used as loss function, which was similar to the objective function of the CoxPH model. The modeling process is shown in Figure 2.

Diagram of modeling process.
Statistical method
In the research, R software (version 4.3.1) was used for data preprocessing, and Python software (version 3.8.0) was used for model training. p-value < 0.05 is statistically significant.
Result
Basic information statistics of esophageal cancer patients
In the research, a total of 5276 patients with esophageal cancer were adopted as study material. No normal distribution was observed among the covariates. The average median age of the patients was 65 [58,71] years, average survival time of the patients was 9 (4,17) months, adenocarcinoma and squamous cell carcinoma account for 72.3% and 25%, respectively. Also, the proportions of surgery and non-surgery were 32.1% and 67.9% respectively, and those for upper, middle, and lower third of esophagus were 5.5%, 16.3%, and 78.1% respectively. Other information is summarized in Table 1.
Basic information statistics of the cohort.

Feature selection
To evaluate the stability of multivariate models in terms of various features, COX univariate analysis was applied to explore important factors affecting prognosis. During model training stage, tests were performed using all factors and some selected factors to explore the predictive performance and stability of the developed model. In training dataset, variables with significant effects on prognosis were identified via COX univariate regression analysis. Analysis results showed that race, diagnosis year, and marital status were significantly affected the prognosis of esophageal cancer patients, with p-values of less than 0.05. The specific analysis results are shown in Table 2. Selection of these variables helped simplify the model and improve its interpretability, while decreasing overfitting risk.
Prognostic risk factors.

CI: concordance index.
Specifically, significant differences were found in prognosis among patients of different ethnic groups. Diagnosis year reflected the development of diagnostic and treatment techniques and prognosis varied depending on diagnosis year. Marital status affected the social support and mental health patients, thereby affecting prognosis.
During model training phase, both all features and a subset of significant features were applied for the training and validation of model. For full-factor model, all available features were applied to train DeepSurv model to maximize data utilization. For partial-factor model, only significant features selected through COX univariate analysis were applied to train the model, simplifying model structure and evaluating its performance.
Development of prediction model
DeepSurv model network settings: The number of input layer nodes was equal to the total number of factors, hidden layer had two layers, the number of neurons was 10, output layer had one node, learning rate was 0.001, 21 momentum was 0.887, 22 dropout was 5.667e-3, 23 and the number of iterations was 1000.
DeepSurv model is a deep learning-based Cox proportional risk model, which used a neural network structure to perform nonlinear transformations on input features and capture complex relationships among different features. In input layer, the number of nodes was equal to the total number of all input features. In this research, input features included multiple factors, such as race, year of diagnosis, marital status, etc., that affect the survival of esophageal cancer patients. Input layer received a variety of clinical and demographic features of patients, which were preprocessed and input into the model in a standardized or normalized manner. Hidden layer was set to two layers, each containing 10 neurons. ReLU (Corrected Linear Unit) was applied to activate the function. In deep learning, ReLU can effectively alleviate the problem of vanishing gradients.24,25 Hidden layer transformed input features via nonlinear activation functions, extracted higher-order features, and captured complex patterns and relationships in the data. Output layer had one node, which estimates the risk function 
Experimental result
After 1000 iterations, loss function was stabilized, as shown in Figure 3.
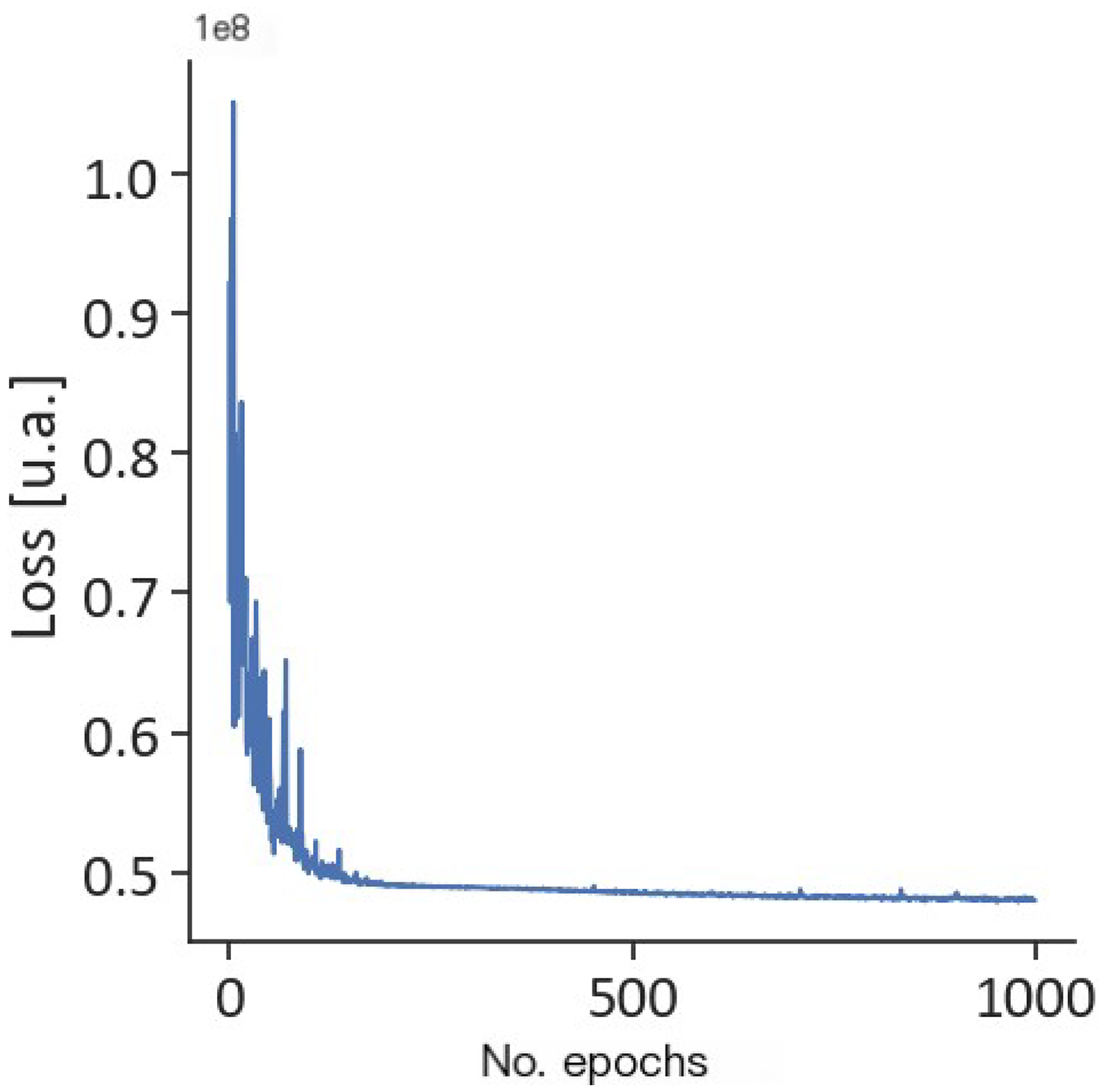
Loss function.
The CI scores on the training set and testing set are 0.84 and 0.83, indicating that the model has good predictive performance. Surgery is a traditional treatment for esophageal cancer. To verify the therapeutic effect of surgery, an experiment was conducted on the surgery group and the non-surgery group. The results showed that surgery is beneficial for patients with esophageal cancer (p < 0.01). Comparison results are illustrated in Figure 4.

Comparison of surgical and non-surgical treatment effects.
To improve prediction result efficiency and accuracy and prevent overfitting and underfitting, three parameters were optimized in the model, with the parameter settings shown in Table 3.
Parameter values.

After optimization, the final parameter values are shown in Table 4.
Optimized parameter values.

After optimization, the iterative calculation process of the loss function is shown in Figure 5.
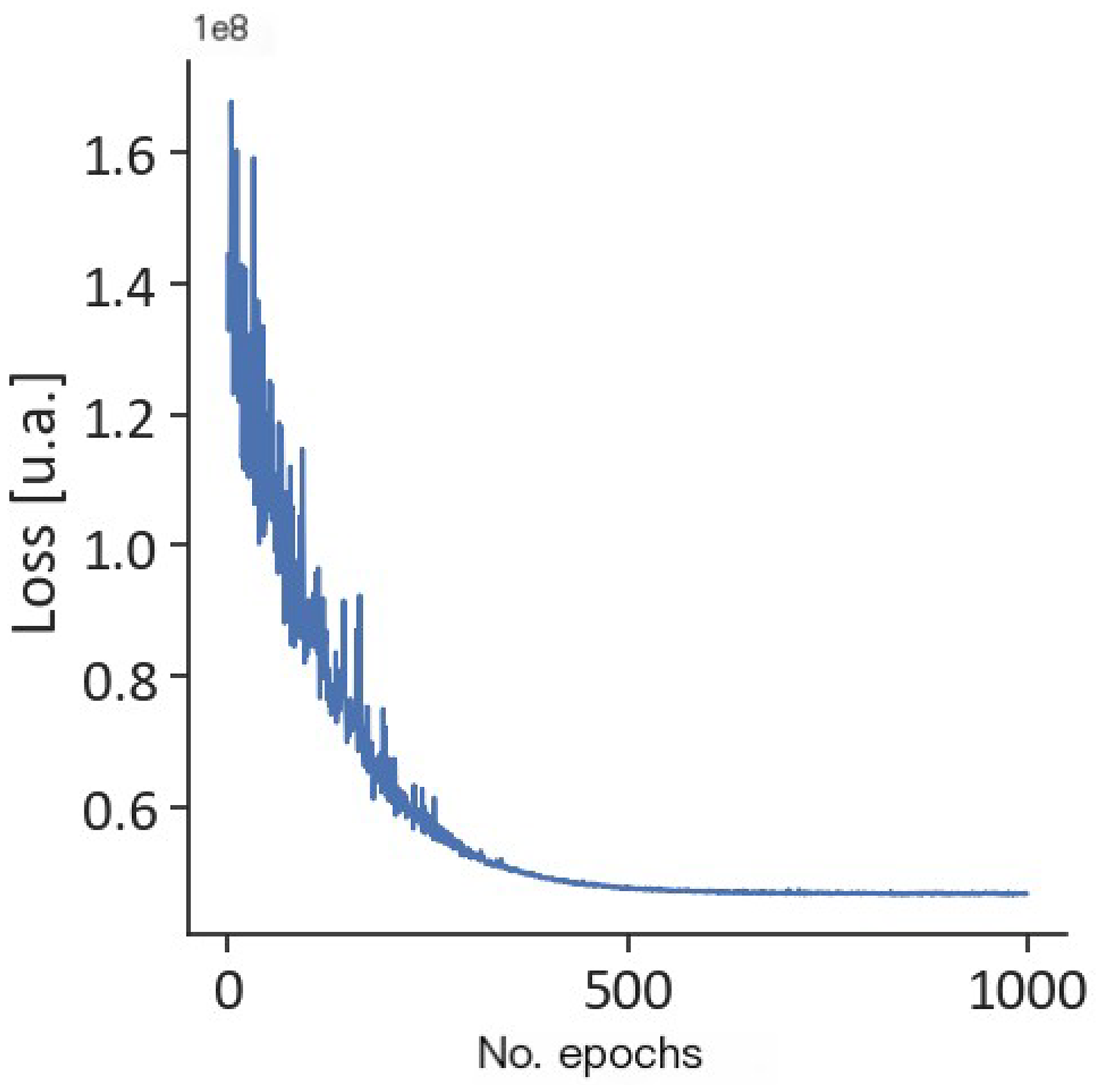
Iterative calculation diagram of loss function after optimization.
Discussion
In terms of treatment options, surgery was beneficial for esophageal cancer patients and could improve their survival time, according to experimental comparisons.
Rahman et al. utilized models generated through elastic net regression, random forest, and extreme gradient boosting to demonstrate that surgery is a significant prognostic factor for esophageal cancer. 26 Similarly, the study confirmed the prognostic significance of surgery for patients with esophageal cancer using multiple machine learning methods. 27 In contrast, another study found that the post-surgical survival rate of esophageal cancer patients was not high through the CoxPH model and Random Survival Forest model. 28
Recently, there have been some models using the SEER database to analyze cancer survival rates. A study developed a staging system for esophageal squamous cell carcinoma based on DeepSurv using SEER data, with a CI of 0.732. 29 Additionally, another study also developed an online prognostic tool for high-grade gliomas using SEER data, with a CI of 0.73–0.74. 30
In this study, data from 5276 esophageal cancer patients were obtained. Initial statistical analysis and univariate regression identified significant prognostic factors. A predictive model for survival analysis was then developed, achieving consistency indexes of 0.84 and 0.83 for the training and testing sets, respectively, which improved to 0.84 for the test set after optimization. This model had relatively stable CI scores under different conditions. As a deep survival analysis model, it could handle more complex situations and had better advantages in dealing with multivariate, large sample, and nonlinear data.
This study also had some limitations. First, the study data were obtained from the SEER database and we could not access some factors that may affect prognosis, such as response to neoadjuvant treatment, protocol of chemotherapy, and sequence and protocol of treatments (neoadjuvant vs surgery only). Second, this study was a retrospective cohort study and further clinical validation is required to confirm the reliability of the findings. Finally, this study may be at risk of overfitting and requires evaluation through methods such as cross-validation or external validation datasets.
Conclusions
Based on Deepsurv model, a predictive model was constructed that forecasts survival periods. Compared to the non-surgical group, the surgical group had a longer survival period.
Footnotes
Acknowledgements
We would like to thank the SEER team for providing the data. We would also like to thank those who participated in SEER for their contribution to the survey.
Author contributions
ZX, WZ, and LW contributed to conception and design of the analysis. ZX, CX, LW, XL, and BC contributed to collection of the data. ZX, CX, and LW contributed to the data and analysis of tools. ZX, CX, LW, and XL performed the analysis. ZX, FG, CX, WZ, LW, XL, QF, YD, XL, and BC contributed to conception of the manuscript, reviewing and editing of the manuscript, and final approval of the version to be published.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by Anhui Province university natural science research key project [No.2023AH040056, No.2023AH050333]; the Natural Science Foundation of Anhui Provincial Department of Education[2024ZK49].
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.


