Abstract
Introduction
Atherosclerosis serves as the primary etiological factor for coronary artery disease, cerebral infarction, and peripheral vascular disorders. The foundation of atherosclerosis lies in lipid metabolism dysregulation, marked by the accumulation of lipids and complex carbohydrates, alongside hemorrhage, and thrombosis. This process subsequently leads to the proliferation of fibrous tissue and calcification, culminating in the thickening and stiffening of the arterial walls and the narrowing of the vascular lumen.1–3 Atherosclerosis stands as a leading global cause of mortality, presenting an urgent challenge that demands resolution. Nevertheless, the intricate mechanisms underlying the pathogenesis of atherosclerosis remain poorly understood. Crucially, endothelial dysfunction is a pivotal initiating event in atherosclerosis. Endothelial cells (ECs) line the vascular lumen and maintain vascular homeostasis. 4 Dysregulation of their proliferation and migration directly contributes to atherosclerotic progression: aberrant proliferation leads to intimal thickening, while enhanced migration facilitates endothelial-to-mesenchymal transition and neointima formation. 5 These processes promote lipid infiltration, inflammatory cell recruitment, and ultimately, plaque development and instability. Recent studies have established that long non-coding RNAs (lncRNAs) play crucial roles in various biological and pathological processes associated with this condition6,7 particularly in modulating endothelial cell behaviors such as proliferation, migration. The initial step of atherosclerosis is vascular endothelial dysfunction. EC are directly exposed to the bloodstream, and their damage can lead to inflammation, lipid deposition, and plaque formation. Meanwhile, umbilical cord derived human umbilical vein endothelial cells (HUVECs) are easy to isolate, culture, and passage, while retaining endothelial cell characteristics such as expression of markers such as von Willebrand factor vWF and CD31. 8 Therefore, HUVEC can directly simulate the endothelial damage mechanism of atherosclerosis.
Long noncoding RNAs (lncRNAs) are defined as noncoding RNA molecules exceeding 200 nucleotides in length, characterized by the absence of significant open reading frames (ORFs). These lncRNAs predominantly reside in the nucleus and exhibit a relatively low level of sequence conservation, with only approximately 12% of lncRNAs being conserved across species beyond humans. LncRNAs play a crucial role in regulating various biological processes, including cellular proliferation, invasion, and apoptosis. Consequently, lncRNAs are emerging as promising biomarkers and therapeutic targets for a range of human diseases.9,10 lncRNAs have been identified as playing a crucial role in cellular processes such as differentiation, proliferation, and apoptosis. 11 Recent studies have further elucidated their significant involvement in the etiology of cancer.12–14 Additionally, lncRNAs have been implicated in the pathophysiology of cardiovascular diseases (CVDs).15,16 For example, lncRNA Malat1 was validated by Chen et al., which play a vital role in cardiac microvascular endothelial cells via its blocking effects on miR-26b-5p/Mfn1 pathway-mediated mitochondrial dynamics and apoptosis. 17 Long noncoding RNA OIP5-AS1 was reported as a regulator that the oxidized low-density lipoprotein (ox-LDL) promoted HUVECs apoptosis through recruiting EZH2 to GSK-3β promotor region. 18
Nuclease-sensitive element-binding protein 1(YB-1) is a regulator of pre-mRNA alternative splicing. YB-1 regulates the binding and stability of cytoplasmic mRNA, which regulates the transcription of a great deal of genes. The translation was actualized by accommodating the interaction between the mRNA and the initiation factors of eukaryotic. At present, YB-1 has a lot of credible reports, such as, DHX9/YB-1 complex was confirmed involved in phenotypic transformation of vascular smooth muscle cells (VSMC). 19 YB-1/PD-L1 is also verified a function of cancer, which can play an important positive role in chemotherapy and antitumor immunity. 20 In colorectal cancer cells, YB-1 was reported as highly phosphorylated to inhibit the proliferation of CRC, which can play function by Ak strain transforming (protein kinase B) signal channel. 21 They all elaborate on the functions of YB-1. At present, a few studies have confirmed the relationship between YB-1 and atherosclerosis, 22 but its mechanism still needs to be further clarified. In this study, we determined the expression of matrix metalloproteinase 2 antisense RNA 1 (MMP2-AS1) in HUVECs under the condition of transforming growth factor-β1 and explored the mechanistic role of MMP2-AS1 in HUVECs proliferation and migration. More importantly, our study suggests a potential role of the MMP2-AS1/YB-1 axis in the pathogenesis and clinical diagnosis of atherosclerosis, warranting further validation.
Materials and methods
Cell culture
The HUVECs were extracted from Shanghai Institutes for Biological Sciences (Shanghai, China, RRID:CVCL_2959) and immortalization. HUVECs underwent 48 hours culturing in Dulbecco's Modified Eagle Medium Nutrient Mixture F-12 (Gibco, Life Technologies Limited Paisley, PA49RF, UK) containing 10% of fetal bovine serum (Corning, USA). Our study in accordance with the Helsinki Declaration of 1975 as revised in 2024.
Cell transfection
Specific siRNAs targeting MMP2-AS1 (si-MMP2-AS1) were used to knock down MMP2-AS1, and si-NC were used to negative control. The pcDNA3.1 with the full sequence of YB-1 was used to overexpress YB-1, and pcDNA3.1 were negative controls. All plasmids were obtained from Gene Pharma Co., Ltd (Shanghai, China). The transfection of these plasmids and siRNA into HUVECs was performed using Lipofectamine 2000 (Invitrogen; Thermo Fisher Scientific, Inc., USA) according to the manufacturer's guide.
Cell counting kit-8 (CCK-8) assay
Cell viability was measured via CCK8 kit (Dojindo, Japan) per manufacturer's protocol. Briefly, the transfected HUVECs were seeded into 96-well plates at a density of 5 × 104 cells each well, and the HUVECs were inducted at 37°C for 1 hour with the CCK-8 of 10:1 dilution ratio. Then, the cell proliferation of HUVECs was determined by measuring on an Enzyme labeling instrument (BioTek, SynergyTM H1/H1 M, USA) at 450 nm.
Real-time quantitative Polymerase Chain Reaction (qPCR)
Total RNA was extracted from HUVECs which treated with TGF-β, ox-LDL, H2O2, and anoxia using TRIzol Reagent (Sparkjade, China), cDNA was synthesized using HiScript® III RT SuperMix for QPCR (Vazyme Biotech, Nanjing, China) and subjected to QPCR using TB Green® Premix Ex Taq™ II (Tli RNaseH Plus) (Takara, Japan) with Real-Time PCR System (Bio-Rad, C1000 Touch, USA), and it used to glycerAldehyde-3-phosphate de hydrogenase (GAPDH) as a reference gene. The primers were used in this experiment: GAPDH, 5′-TGACAACTCCCTCAAGATTGTCA-3′ and 5′-GGCATGGACTGTGGTCATGA-3′; MMP2-AS1, 5′-CTTGACTTTCTTCCACTGCCCTC-3′ and 5′-AAGTTGAGGCAGGAAGATTGAGC-3′; YB-1, 5′-GGGGACAAGAAGGTCATCGC-3′ and 5′-CGAAGGTACTTCCTGGGGTTA-3′.
Western blot analysis
Cultured HUVECs were lysed in lysis buffer containing radio immuno precipitation assay (Solarbio, Beijing, China), protease inhibitor phenyl methyl sulfonyl fluoride (Solarbio, Beijing, China) and cocktail (Roche, Basel, Switzerland). The BCA Protein Assay Kit (Solarbio, Beijing, China) was used to examine the protein density. Later, total samples were separated with 12% Sodium Dodecyl Sulfate-PolyAcrylamide Gel Electrophoresis and then transferred onto the Poly Vinylidene DiFluoride membrane (Merck Millopore, USA). Primary antibodies applied were anti-YB1 (Abcam, USA, Dilution, ab76149, 1:1000) and anti-β-actin (Cell Signaling Technology, USA, Dilution, 4970, 1:1000). Secondary antibodies applied were Goat Anti-Rabbit (Abcam, USA, Dilution, ab205718, 1:2000). Using ECL kit (Thermo Fisher Scientific, USA) and ChemiDoc system (Bio-Rad, USA) to take photos and observe the result. Using image J 1.52P (Java 1.8.0-172) software to analyze the bands of protein.
Migration assay
Transwell migration assay was used to examine the migration of HUVECs which contains 8μm pore sized membrane (Corning, USA). Briefly, on upper chamber, the treated HUVECs were plated with serum-free D-DEM/F-12 medium at a density of 4 × 105 cells each well, nevertheless, in the lower chamber, D-DEM/F-12 medium with 20% fetal bovine serum was filled. The plated cells were cultured for 24 hours in a cell incubator, and then, the migrated cells of HUVECs were fixed with 4% paraformaldehyde (Solarbio, Beijing, China) and staining with 0.1% crystal violet. The number of migrated HUVECs was analyzed using a light microscope (Nikon, Japan).
The wound healing assay was carried to measure the migration potential of HUVECs. Mechanically, treated The HUVEC was seeded on a 6-well plate, and each well had 2 × 106 cells. After 24 hours incubation, a wound like a straight-line was generated by scratching the bottom of well with a 1 mL pipette tip. Obtaining the pictures at a specific time, and the picture is analyzed by image J software.
Kyoto encyclopedia of genes and genomes (KEGG) analysis differential gene expression
Perform differential analysis of genes between the disease group and the control group using R package limma (Version 3.58.1) from GSE97210 database. Set | logFC |>0 and P-value < 0.05 as the threshold for differentially expressed genes (DEGs). Among them, genes with logFC > 0 and P-value < 0.05 are upregulated differentially expressed genes (upregulated genes), while genes with logFC < 0 and P-value < 0.05 are downregulated differentially expressed genes (downregulated genes).
EdU assay
The experiment was conducted according to the reagent manual of BeyoClick™ EdU-488 (Beyotime, China). Incubate the cells with culture containing 50 μM EdU at 37 °C for 2 hours. Remove the culture medium and fix the cells with 4% paraformaldehyde at room temperature for 15 minutes. Treatment with 0.5% Triton X-100 for 20 minutes increased cell membrane permeability. Add reaction mixture Alexa Fluor 488 azede. React at room temperature in the dark for 30 minutes. Stain the cell nucleus with 4',6-Diamidino-2-Phenylindole (1 μg/mL) for 5 minutes. Observation under fluorescence microscope: EdU+ cells: green fluorescence (labeling proliferating cells); DAPI+ cells: blue fluorescence (labeling all cells).
Oil Red O staining
HUVECs were fixed with 4% paraformaldehyde for 15 to 20 minutes and rinsed 3 times with phosphate buffered saline (PBS). 0.5% ORO (Oil Red O) mother liquor: dissolve 0.5 g ORO powder in 100 mL isopropanol (stored away from light). Prepare working fluid before use: mother liquor: distilled water = 3:2, mix and let stand for 10 minutes, filter with filter paper. Immerse the sample in ORO working solution (Oil Red O staining kit, Beyotime, China) incubate at 37°C for 10 to 15 minutes, wash with 60% isopropanol for 30 seconds, rinse with PBS 3 times (terminate staining). Hematoxylin counterstaining of cell nuclei (1–2 minutes), water washing returns to blue. Observation under an optical microscope (lipid = bright red particles, nucleus = blue).
Conservation analysis
PhyloP algorithm (UCSC Genome Browser: https://genome.ucsc.edu/) revealed evolutionary conservation of MMP2-AS1 across vertebrates.
Statistical analysis
The data of this study was shown as the mean ± standard error. And significance analysis of that data was according to t-test, One-way analysis of variance (ANOVA) and two-way ANOVA, respectively, with the Graph Pad Prism 5 software. Unpaired two-tailed Student's t-test was used for intergroup comparisons. Statistical significance was deemed to the P-value < 0.05. All the data was at least three experiments, and every experiment was independent.
Result
Increased expression of MMP2-AS1 in TGF-β1-treated HUVECs
We first explored MMP2-AS1 expression from GSE97210 database, the result showed that MMP2-AS1 exhibited high expression in atherosclerosis, which is up to three times than healthy (Figure 1(a)). So, we predicted that MMP2-AS1 may play an important role in atherosclerosis. TGF-β1 is a factor that induces atherosclerosis, 23 so we detected that the expression of MMP2-AS1 in HUVECs treated with TGF-β1 is increased (Figure 1(b)). Using KEGG analysis, the expression of MMP2-AS1 in the human disease and healthy groups was analyzed and found to be more enriched in the disease group. Differential genes in the two groups were also analyzed, yielding 1758 upregulated genes and 2323 downregulated genes (Figure 1(c) and (d)). Next, we conducted a conservative analysis of MMP2-AS1 by University of California Santacruz (UCSC), which showed that MMP2-AS1 has highly conservatism (Figure 1(e)). The above results showed that MMP2-AS1 might regulate atherosclerosis.
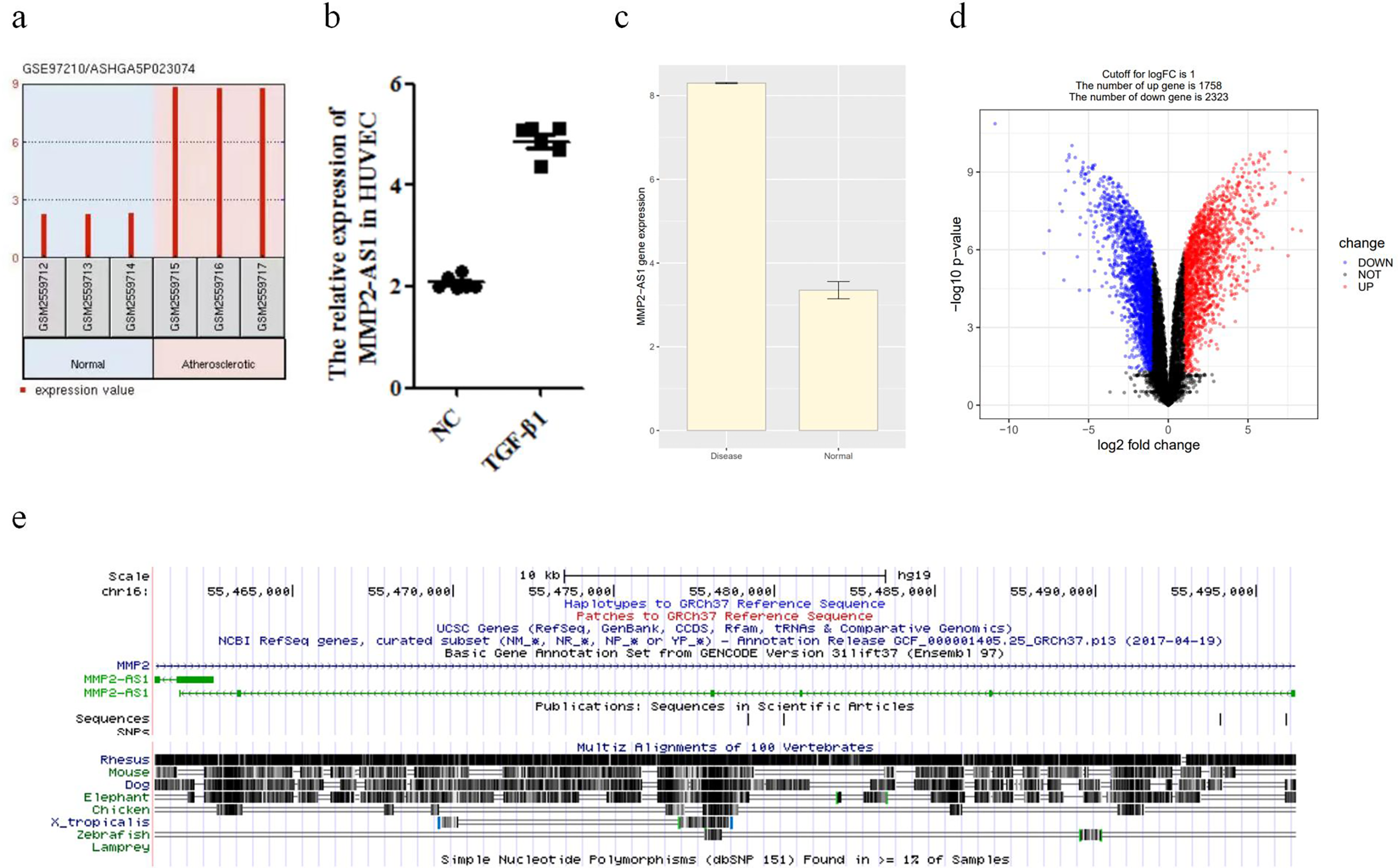
Increased expression of MMP2-AS1 in TGF-β1-treated HUVECs. (a) The expression of MMP2-AS1 in atherosclerosis through the prediction of GSE97210 database. (b) The relative expression of MMP2-AS1 in HUVECs treated with TGF-β1. **P < 0.01. (c) and (d) The relative expression of MMP2-AS1 in normal and AS groups. (e) The conservatism of MMP2-AS1 in different species was predicted using UCSC. HUVEC: human umbilical vein endothelial cells.
The regulation of MMP2-AS1 in HUVECs
To further determine the role of MMP2-AS1 in the pathogenesis of atherosclerosis in vitro, we used ox-LDL, 24 anoxia, 25 hydrogen peroxide 26 to induce the pathological state of HUVECs to simulate the pathogenesis of atherosclerosis in the human body. According to previous study reports, the optimum treatment concentration of ox-LDL is 50 μg/mL in HUVECs. 24 So, we treated it, and the result showed that the expression of MMP2-AS1 increased with the processing time of ox-LDL (Figure 2(a)). Meanwhile, HUVECs were treated with hydrogen peroxide in 100 μM, which concentration had been reported.27,28 The result acts MMP2-AS1 may participate in regulation of HUVECs, and the expression of MMP2-AS1 increased with the processing time of hydrogen peroxide (Figure 2(b)). To investigate the effect of anoxia conditions on the expression of MMP2-AS1 in HUVECs, we used 1% oxygen to treat HUVECs. 29 Similarly, in anoxia treated HUVECs, the expression level of MMP2-AS1 increased in a gradient with increasing anaerobic treatment time (Figure 2(c)).

The regulation of MMP2-AS1 in HUVECs. (a) The relative expression of MMP2-AS1 in HUVECs treated with ox-LDL (50μg/mL) at 0, 12, 24, 36, and 48 hours. ***P < 0.001. (b) The relative expression of MMP2-AS1 in HUVECs treated with hydrogen peroxide (100μM) at 0, 2, 8, 12, and 24 hours. ***P < 0.001. (c) The relative expression of MMP2-AS1 in HUVECs treated with anoxia. The oxygen concentration is 1% and treated 0, 2, 8, 12, and 24 hours. ***P < 0.001. HUVEC: human umbilical vein endothelial cells; ox-LDL: oxidized low-density lipoprotein.
Knockdown MMP 2—as 1 inhibits proliferation and migration of HUVECs
To explore the roles of MMP2-AS1 in atherosclerosis, the function assays of HUVEC were conducted. Effects of MMP2-AS1 on cell proliferation and migration of HUVECs were studied. Inhibition of MMP2-AS1 was achieved using corresponding si-MMP2-AS1, and then, si-NC and si-MMP2-AS1 were transfected into HUVECs, that showed the si-MMP2-AS1 knockdown successfully after transfection (Figure 3(a)). CCK8 assay and EdU assay showed that knockdown si-MMP2-AS1 inhibits the proliferation of HUVECs obviously (Figure 3(b) and (c)) and then, cell wound scratch assay and transwell assay showed that si-MMP2-AS1 attenuates the migration of HUVECs (Figure 3(d) and (e)). Oil red staining experiments showed that knocking down si-MMP-AS1 can inhibit HUVEC lipid droplet formation and reduce lipid deposition (Figure 3(f)). Above functional test demonstrated a conclusion is that knockdown MMP2-AS1 inhibits proliferation, migration and lipid deposition of HUVECs.
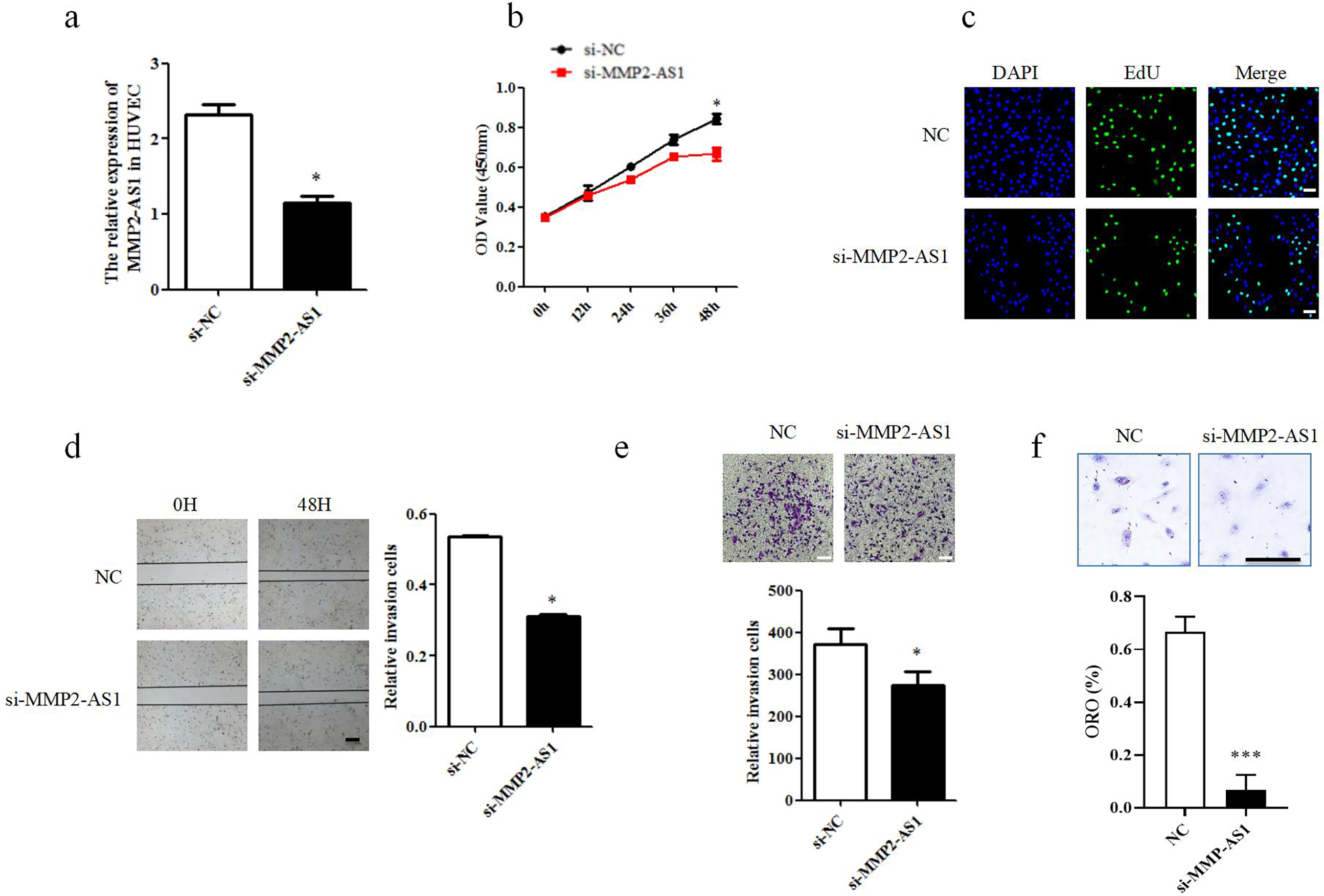
Knockdown MMP2-AS1 inhibits proliferation and migration of HUVECs. (a) The relative expression of MMP2-AS1 in HUVECs. ***P < 0.005. (b) CCK8 assay approved that the proliferation of si-MMP2-AS1. ***P < 0.005. (c) EdU assay approved that the proliferation of si-MMP2-AS1. Scale: 20 mm, *P < 0.05. (d) Scratch wound healing test approved that the migration of si-MMP2-AS1. Scale: 50 mm, *P < 0.05. (e) Transwell assay approved that the migration of si-MMP2-AS1. Scale: 20 mm, *P < 0.05. (f) Oil red staining experiments approved that the lipid deposition of si-MMP2-AS1, Scale: 20 mm, ***P < 0.005. HUVEC: human umbilical vein endothelial cells.
MMP2-AS1 interacts with YB-1 in HUVECs
To further verify the mechanism of MMP2-AS1, we predicted the downstream target of MMP2-AS1 by catRAPID (Figure 4(a)). From the results of the score, YB-1 (YBOX1) has a great combination advantage. Next, we use catRAPID express system to analyze the sequence of YB-1 and MMP2-AS1, which showed that YB-1 and MMP2-AS1 have a strong binding effect (Figure 4(b)). Then we verified YB-1 in a series of tests to see if it worked with MMP2-AS1. QPCR showed the YB-1 was regulated by MMP2-AS1 (Figure 4(c)). The expression of MMP2-AS1 was detected by PCR and agarose gel electrophoresis and found that using YB-1 pull-down RNA can enrich MMP2-AS1 more. RNA immunoprecipitation (RIP) assay shows MMP2-AS1 interacts with YB-1 in HUVECs (Figure4(d)).

MMP2-AS1 interacts with YB-1 in HUVECs. (a) and (b) The prediction of downstream binding proteins of MMP2-AS1 used catRAPID website. (c) The relative expression of YB-1 of HUVECs. ***P < 0.001. (d) Using IgG as negative control, YB-1 can combined with MMP2-AS1. HUVEC: human umbilical vein endothelial cells.
MMP2-AS1 inhibits proliferation and migration of HUVECs by targeting YB-1
Western blot analysis showed increased levels of YB-1 in the plasmid, while MMP2-AS1 siRNA decreased YB-1 (Figure 5(a)). The result showed that MMP2-AS1 can be used as a regulator for YB-1. CCK8 assay, EdU assay and transwell assay approved that si-MMP2-AS1 recoverd the proliferation and migration induced by YB-1(Figure 5(b) to (d)). Thus, si-MMP2-AS1 inhibits proliferation and migration of HUVECs by targeting YB-1. The hypothetical molecular mechanism is shown as Figure 6.
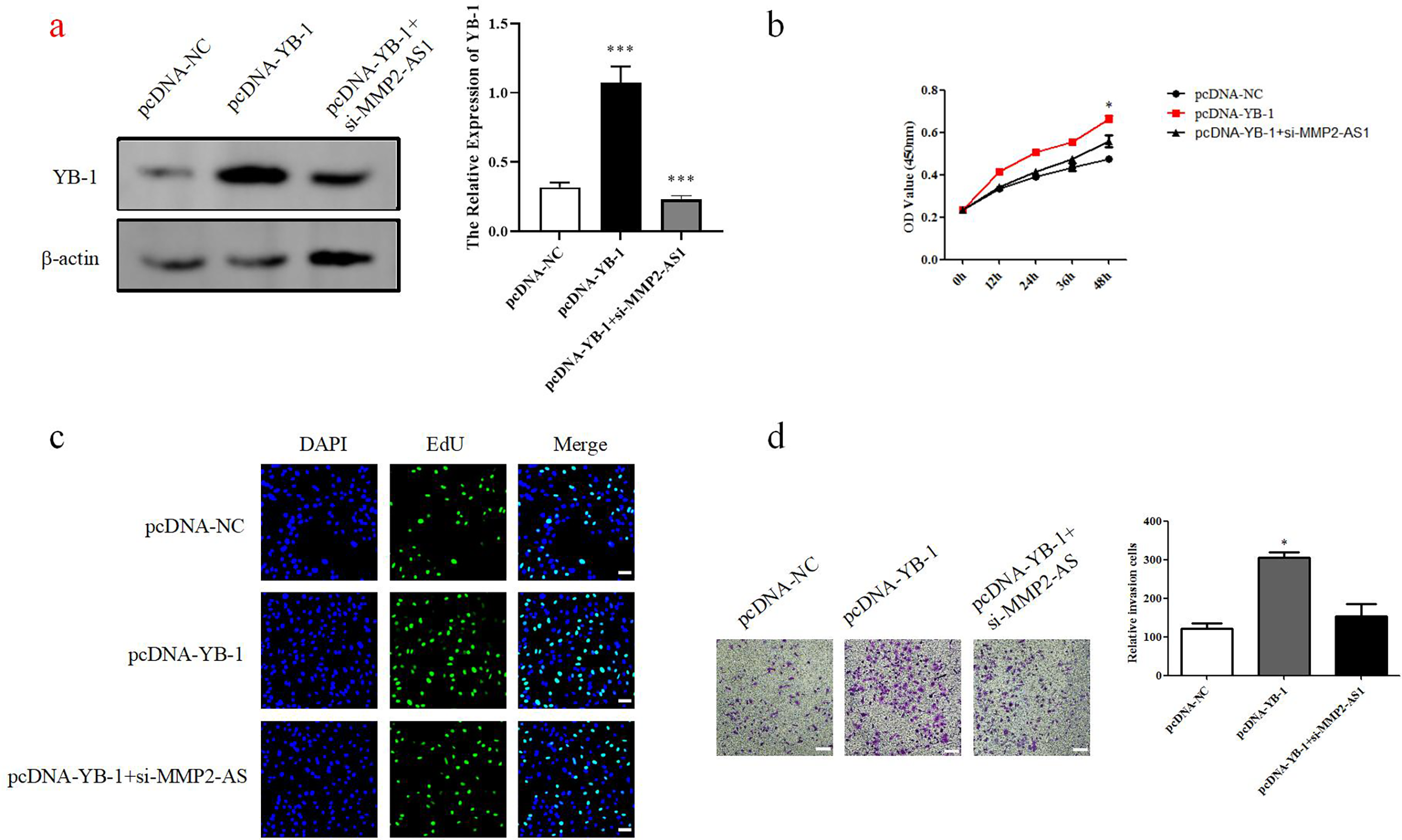
MMP2-AS1 inhibits proliferation and migration of HUVECs by targeting YB-1. (a) The pcDNA3.1, pcDNA-YB-1, and pcDNA-YB-1 + si-MMP2-AS1 were transfected in HUVECs. The protein levels of this four groups were showed. *P < 0.05. (b) CCK8 assay approved that the proliferation of pcDNA3.1, pcDNA-YB-1, and pcDNA-YB-1 + si-MMP2-AS1. ***P < 0.001. (c) EdU assay approved that the proliferation of pcDNA3.1, pcDNA-YB-1, and pcDNA-YB-1 + si-MMP2-AS1. Scale: 20 μm, *P < 0.05. (d) Transwell assay approved that the migration of pcDNA3.1, pcDNA-YB-1, and pcDNA-YB-1 + si-MMP2-AS1. Scale: 20 μm, *P < 0.05. HUVEC: human umbilical vein endothelial cells.

MMP2-AS1 and YB-1 in atherosclerosis. The expression of MMP2-AS1 was upregulated in atherosclerosis and YB-1 was upregulated in atherosclerosis. They performed negative correlation. Silencing MMP2-AS1 inhibited the proliferation and migration of HUVEC through targeting YB-1, and promoted atherosclerosis. HUVEC: human umbilical vein endothelial cells.
Discussion
Cardiovascular diseases include stroke, coronary artery diseases, and hypertension which cause multiple morbidity and mortality worldwide. Atherosclerosis is closely related to inflammatory reaction by endothelial dysfunction, the proliferation and migration of HUVECs, and the remodeling of extracellular matrix (ECM). All of the above causes the atherosclerotic plaque. 30
While lncRNAs have traditionally been viewed as disruptors of genomic transcription, emerging research has highlighted their crucial involvement in a range of biological processes, including neurodegenerative disorders, 31 cancer, 32 inflammation, diabetes, and cardiovascular diseases.11,33 Prior investigations have established that lncRNAs are essential for both genomic regulation and the maintenance of human health, particularly through their effects on cell proliferation, migration, and apoptosis.34,35 The role of lncRNAs in genomic regulation and disease underscores the significance of the lncRNA-mediated regulatory network. For instance, silencing of the lncRNA MMP2-AS1 markedly inhibited cell proliferation and migration, it further suggests that lncRNA MMP2-AS1 may participate in the regulation mechanism of atherosclerosis. Furthermore, basic local alignment search tool alignment analyses revealed that MMP2-AS1 specifically associates with YB-1; inhibiting YB-1 suggests that it mediates the regulatory effects of MMP2-AS1 on the growth and proliferation of HUVECs. 36
In the current investigation, we observed that the expression of lncRNA MMP2-AS1 was significantly elevated in HUVECs subjected to ox-LDL in a time-dependent manner. This cellular model effectively simulated the pathogenesis of atherosclerosis induced by excess ox-LDL, with the heightened expression of MMP2-AS1 suggesting its potential critical involvement in endothelial dysfunction associated with atherosclerosis. Functional assays conducted on HUVECs demonstrated that the silencing of MMP2-AS1 expression resulted in a marked reduction in both proliferation and migration of these cells. Furthermore, mechanistic studies indicated that MMP2-AS1 interacts with YB-1 to enhance its expression. This lncRNA MMP2-AS1/YB-1 axis may play a significant role in the regulatory mechanisms underlying atherosclerosis in HUVECs. 37 This aligns with established evidence that ox-LDL is a pivotal driver of endothelial dysfunction in atherosclerosis. Ox-LDL binds to scavenger receptors (e.g. LOX-1) on HUVECs, triggering intracellular oxidative stress, Nuclear Factor kappa B cells activation, and pro-inflammatory cytokine release (e.g. InterLeukin-6, vascular cell adhesion molecule), thereby promoting monocyte adhesion—a critical initial step in plaque formation. 38 Recent studies further reveal that ox-LDL regulates noncoding RNA networks in HUVECs: it upregulates pro-atherogenic lncRNAs (e.g. OIP5-AS1, MALAT1) while suppressing protective miRNAs (e.g. miR-126-3p), creating a feedforward loop that exacerbates endothelial injury.39–41 Our data add MMP2-AS1 to this regulatory axis, demonstrating its specific induction by ox-LDL and functional role in HUVEC proliferation/migration. This cellular model effectively simulated the pathogenesis of atherosclerosis. This study explored the downstream mechanisms associated with MMP2-AS1 and its role within the context of atherosclerosis. Our findings indicated that MMP2-AS1 interacts with the YB-1 protein, thereby modulating its expression levels.
Although this experiment is innovative, all experiments are carried out at the cellular level, lacking in vivo validation of atherosclerosis animal models (such as ApoE −/− mice). Also, no more representative primary cells were used. The pathological significance and therapeutic potential of MMP2-AS1/YB-1 axis in vivo cannot be confirmed. At the same time, clinical sample analysis was insufficient. Although the database (GSE97210) was cited to show that MMP2-AS1 was highly expressed in atherosclerosis, patient tissue samples were not used to verify the expression level or correlation. Further exploration is needed at a deeper level.
Conclusion
Our study revealed that MMP2-AS1 as a regulatory factor plays an important role in the migration and proliferation of HUVECs via the MMP2-AS1/YB-1 axis.
Footnotes
Ethics approval and consent to participate
Not applicable. Our study is not on human tissue / samples and does not require approval by the ethics / review committee. We have verified the review board and received their exemption.
Authors’ contributions
ZTL was involved in data analysis, figures prepared, and wrote the first draft of manuscript; KW in performed the experiments, and data acquisition and discussion; and YFC and ZLL in conceived the concept, designed the manuscript, coordinated and critically revised manuscript, responsible for its financial support, and corresponding works.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability
The data used to support the findings of this study are available from the corresponding author upon request.
