Abstract
Objective
Small cell lung cancer (SCLC) is a highly malignant subtype of lung cancer. Cancer stem cell (CSC)-like cells have been implicated in chemoresistance and recurrence. Although previous studies have demonstrated the significance of WASP-interacting protein (WIPF1) in malignant tumors, its underlying molecular mechanism in SCLC is not well known. Here, we demonstrate that WIPF1 can regulate tumorigenesis and its underlying molecular mechanism in SCLC.
Methods
Sphere-formation culture effectively enriches CSC-like cells, such as tumor stem cell-like cells. RNA-seq was used to identify differentially expressed genes between enriched CSCs (3D cultures) and differentiated cells. Next, we adopted RNA interference techniques to investigate the effects of WIPF1 on colony and sphere-formation capacity, as well as cisplatin sensitivity in SCLC cells. Furthermore, we performed western blot analysis to analyze protein expression and employed the STRING database to identify potential signaling pathways.
Results
WIPF1 was significantly upregulated in sphere-formed SCLC cells, relative to differentiated ones under adherent growth conditions (2D cultures). The gene was involved in the regulation of colony formation, sphere-formation capacity, and cisplatin sensitivity in SCLC cell line model. Knocking down of WIPF1 significantly suppressed the proliferation of cancer cells via the YAP/TAZ protein.
Conclusions
Sphere-formation and chemoresistance represent indispensable characteristics of CSC-like cells. Notably, sphere-formation culture is a more effective approach for enriching of CSCs-like cells than traditional adherent culture. Upregulation of WIPF1 in sphere-formed cells, relative to differentiated ones, indicated that it plays an important role in the tumorigenesis of SCLC. Moreover, this process is mediated by the YAP/TAZ pathway, suggesting that it may be a potential therapeutic target for SCLC.
Introduction
Malignant tumor is one of the most leading causes of death in humans. At present, lung cancer has the highest incidence rate and mortality among all types of malignancies, imposing a heavy burden on the society.
1
The WASP-interacting protein (WIPF1) binds to WASP, to activate formation of podosomes and neuron dendrites, and promote cell proliferation and migration by regulating cytoskeletal proteins.21,22 Target genes of p53 highly overlapped with those co-expressed with WIPF1, while the upregulation of WIPF1 promoted tumorigenesis and tumor progression. Eike Staub and colleagues
23
found that c-myc and ESR1, downstream target genes of p53, were co-expressed with WIPF1, and upregulated WIPF1 expression, a phenomenon that was associated with poor prognosis of patients with different malignancies. Reduced expression of WIPF1 in glioblastoma or breast cancer cells significantly downregulated CSC markers and suppressed tumor growth in vivo.
24
Abnormal expression of WIPF1 enhances gastric cancer progression.25,26 Besides, WIPF1 has been confirmed to be a major immune infiltration-related molecule in head and neck cancer.
27
WIPF1 has also been implicated in abnormal cell proliferation, although its role in SCLC remains unknown. Previous reports have indicated that CSCs promote tumor initiation, resistance, and recurrence, thus targeting them may be a feasible strategy for management of SCLC.28,29 On the other hand, although the specific markers for identifying of CSCs,
30
In this study, we investigated the function of WIPF1 in the regulation of cisplatin sensitivity and the underlying molecular mechanism in SCLC. Specifically, we performed RNA-Seq analysis and found that WIPF1 was markedly upregulated in sphere-formed cells relative to attached ones in NCI-H446. Next, we used small interfering RNA (siRNA) technology to knock down WIPF1. Since WIPF1 has been shown to regulate autonomous cell growth via the YAP/TAZ. 32 Therefore, we also analyzed expression of YAP/TAZ proteins and their associations with WIPF1in SCLC.
Materials and methods
Cell lines
Human SCLC cell lines used in the article include NCI-H446 (HTB-171), NCI- NCI-H209 (HTB172), and H526 (CRL-5811). We bought SCLC cell lines from FuHeng Cell Center, Shanghai, China, and the cell lines were cultured in RPMI 1640 medium added with 10% fetal bovine serum (Gibco Technologies, Grand Island, USA). The cell lines were digested with trypsin-EDTA buffer (Invitrogen) when passaged, then placed in incubator.
Cell transfection with WIPF1-siRNAs
WIPF1-siRNAs, synthesized by GenePharma (Shanghai, China), for use in this study are listed in Table 1. Cancer cells were plated in RPMI-1640 medium for 12 h or more, then transfected with 100 nM of WIPF1-siRNAs or control siRNAs that had been diluted in Opti-MEM (Gibco Technologies, Grand Island, USA) mixing well with lipofectamine 2000 (Life technologies, Carlsbad, USA) as described in our previous work. 33
Sequences of WIPF1-siRNAs and control siRNA used in this study.
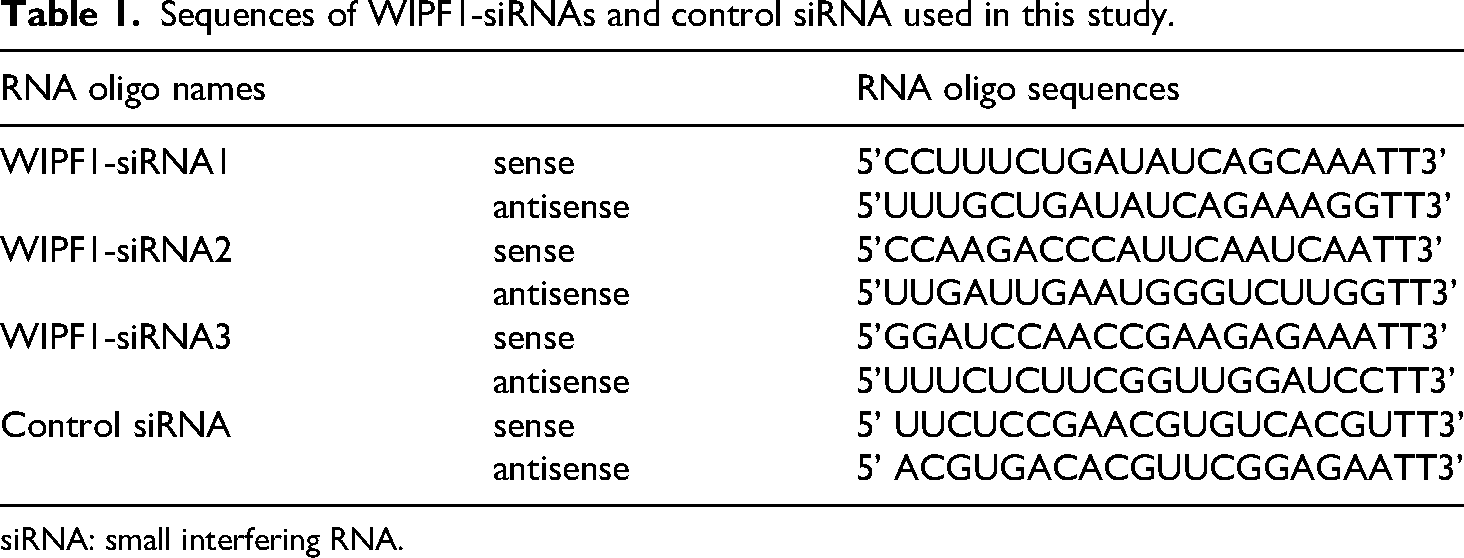

siRNA: small interfering RNA.
Quantitative real-time PCR
The aforementioned SCLC cell lines were treated with WIPF1-siRNAs for 24 h, then total RNA isolated using commercial RNApure Kit (CWbio biotech Co., Ltd, Jiangsu, China). Nanodrop 2000 spectrophotometer (Thermo Fisher Scientific, USA) was used to determine, then equal RNA concentrations used to synthesize cDNA using HiScript II 1st Strand cDNA Synthesis Kit (+gDNA wiper) (Vazyme biotech Co., Ltd, Nangjing, China). Taq Pro Universal SYBR qPCR Master Mix(Vazyme biotech Co., Ltd, Nangjing, China) was then applied on a 7500 Fluorescent quantitative PCR System (Applied Biosystems, USA), targeting genes whose primers are outlined in Table 2.
Sequences of primers used for qRT-PCR.

qrt-PCR: quantitative real-time PCR.
Cell viability assay
SCLC cell line cells were seeded in a microplate (Corning life sciences, USA), at a density of 5000 cells per well, and incubated with WIPF1- or control siRNAs. Cells viability was analyzed through the CCK8 assay kit. Subsequently, in 3 consecutive days, 10 μl of the CCK-8 reagent (Dojin Laboratories, Kumamoto, Japan) was added to each well, followed by determination of absorbance on a Microplate reader (Thermo Fisher Scientific, USA) at a wavelength of 450 nm.
Sphere-formation assay
SCLC cells were placed in ultra-low attachment microplates, and cultured for 14 days in DMEM/F12 medium (Gibco Technologies, Grand Island, USA), with EGF (20 ng/mL, Proteintech, Wuhan, China), bFGF (FGF-basic, 20 ng/mL, Invitrogen, USA), and B27 (20 μl/mL, Gibco Technologies, Grand Island, USA). Thereafter, the number of cell spheres (diameter larger than 100μm) was calculated under a phase-contrast microscope (Nikon Corporation Japan).
Colony formation assay
SCLC cells were placed in conventional microplates and cultured for 14 days in DMEM/F12 medium (Gibco Technologies, Grand Island, USA), with EGF (20 ng/mL, Proteintech, Wuhan, China), bFGF (FGF-basic, 20 ng/mL, Invitrogen, USA), and B27 (20 μl/mL, Gibco Technologies, Grand Island, USA). Thereafter, the Giemsa staining was conducted, and the number of cell colony (diameter larger than 100μm) was calculated under a phase-contrast microscope (Nikon Corporation Japan).
Western blot assay
NCI-H446 cells were harvested, 48 h after treatment with WIPF1-siRNAs. The proteins were quantified via the BCA method, and 45μg separated via SDS-PAGE. The membrane was incubated for 10 h with primary antibodies, namely WIPF1, TAZ, and YAP antibodies (Table 3) at 4 °C, then with peroxidase-conjugated IgG secondary antibody, then analyzed on the Chemi-Doc XRS system. Protein expression was performed using the Image J software.
Information on primary antibodies used for western blot analysis.
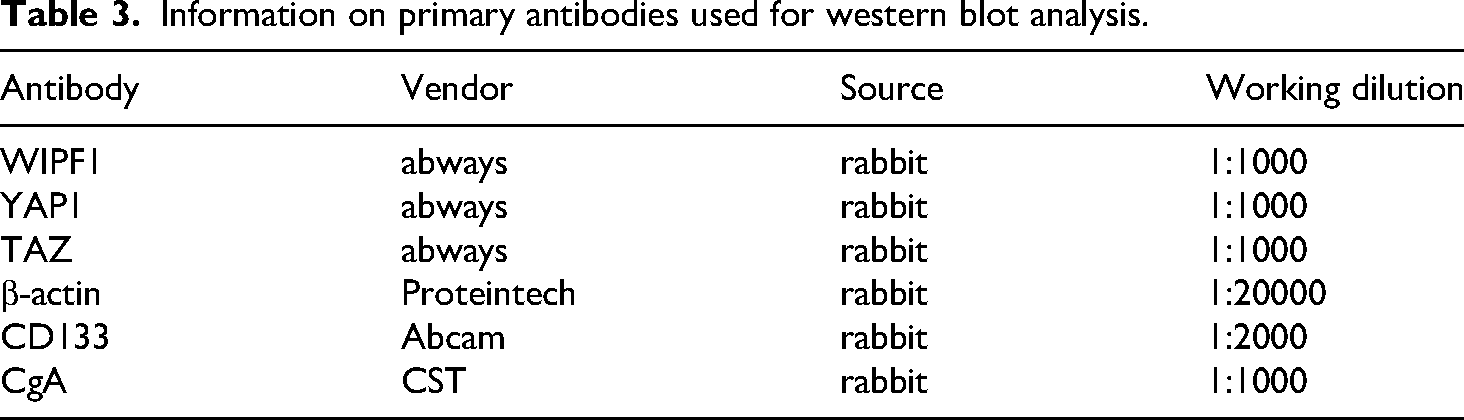

Flow cytometry
The average expression intensity of CD133 in sphere-formed cells and attached cells were determined using Alexa FluorR488labeled primary antibody. Human SCLC lung cancer cells (NCI-H446) were stained with Alexa FluorR488 conjugated CD133 primary antibody (Abcam, USA), Rabbit normal IgG served as the control, facilitating the determination of the average expression intensity in SCLC cells.
Transcriptomic analysis
Transcriptome analysis was performed by the Personal Biotechnology Company Limited (Nanjing, China). Briefly, Total RNA was isolated from sphere-formed cells relative to attached ones in NCI-H446, and mRNAs enriched using oligo dT beads. The number of reads falling in the exons of this meta-transcript were counted and normalized by the size of the meta-transcript and by the size of the library. Differentially expressed genes (DEGs) were identified using NOISeq, as previously described.33,34
Statistical analysis
All experimental data were analyzed using GraphPad Prism Version 5.0. t-test was used to analyze differences between groups; p < 0.05 was considered statistically significant.
Results
WIPF1 was upregulated in sphere-formed relative to attached cells
Although CSC markers have not been clearly identified in SCLC, CD133, uPAR, and CACNA2D1 have been associated with maintenance of stemness and drug resistance.35,36 Our results showed that sphere-formation culture effectively enriched CSC-like cells, such as TSCLCs. Analysis of expression of CD133, CACNA2D1, and uPAR mRNAs revealed that they were upregulated under sphere-formation culture relative to control conditions (supplementary figure1A). Besides, the results implied that the average fluorescence of sphere-formed cells was significantly increased (supplementary figure1B). The results indicated that dissociated spheres can reform secondary spheres in (supplementary figure 1C). RNA-Seq analysis revealed that 296 and 336 genes were upregulated and downregulated, respectively (

WIPF1 was upregulated in sphere-formed relative to attached cells. Scatter plot (A) and hierarchical clustering plot (B) displaying differentially expressed genes (DEGs) between attached and sphere-formed SCLC cells. Red and green dots in (A and B) represent upregulated and downregulated genes, respectively. Note: the selection criteria for the differentially expressed genes were log2|fold change| > 3 and p < 0.05. (C) qRT-PCR results showing changes in expression of TRIM29, WIPF1, EYA4, ALDH3A, LOX, AKR1C1, LZTS1, CRACR2A, and HEYL mRNAs in the sphere-formed relative to attached cells. *Statistically significant difference (p < 0.05) between sphere-formed and attached cells through t-test and at least three repetitions were conducted. SCLC: small cell lung cancer; qrt-PCR: quantitative real-time PCR.
Differentially expressed genes between attached and sphere-formed SCLC cells.

SCLC: small cell lung cancer.
WIPF1-siRNAs downregulate WIPF1 thereby inhibiting proliferation capacity and promoting sensitivity to cisplatin of NCI-H446 and NCI-H209 cells
To suppress WIPF1 expression, SCLC cell lines were transfected with siRNAs designed to specifically target WIPF1, while non-targeting siRNAs served as negative controls. qRT-PCR results (

Knockdown of WIPF1 significantly suppressed the proliferation activity and enhanced sensitivity to cisplatin in small cell lung cancer cells. (A) qRT-PCR results showing changes in WIPF1 expression in WIPF1-siRNA1, 2, and 3 groups of SCLC cell line NCI-H446. (B, C) Western blots showing levels of WIPF1 proteins in WIPF1-siRNA1, 2, and 3 groups of NCI-H446 cells, β-actin was used as a loading control through t-test and 2 repetitions were conducted. (D) CCK-8 assay results showing proliferation of NCI-H446 after transfection with WIPF1-siRNA 1 and 2. (E) CCK-8 assay results showing proliferation of NCI-H446 after transfection with WIPF1-siRNA2 and cisplatin treatment (3.2 µg/ml). *Statistically significant differences (p < 0.05) between WIPF1-siRNA-treated cells and control siRNA (A, C-E) through t-test and at least three repetitions were conducted. SCLC: small cell lung cancer; siRNA: small interfering RNA; qrt-PCR: quantitative real-time PCR.

WIPF1 CCK-8 assay results showing proliferation (A) and cisplatin treatment (3.2 µg/ml) (B) of NCI-H209 cells after transfection with WIPF1-siRNA2. *Statistically significant difference (p < 0.05) between sphere-formed and attached cells through t-test and at least three repetitions were conducted. siRNA: small interfering RNA.
WIPF1 knockdown significantly suppressed colony-forming capacity and sphere formation of SCLC cancer cells
Next, we performed colony formation and sphere-formation assays, on NCI-H446 cell line, to evaluate the function of WIPF1 in SCLC. Results showed a significant reduction in the number of both spheres formation (Figure 4A and 4B) and colonies formation (Figure 4C and 4D) in SCLC cells, after WIPF1-siRNA treatment, proving that this gene might perform a crucial role in tumor development thus could reflect the malignancy and metastatic capacity of cancer cells. 39

WIPF1 knockdown significantly suppressed colony formation and sphere-formation capacity of small cell lung cancer cells. WIPF1-siRNA2 significantly reduced the number of cell spheres (A, B) and the number of cell colonies (C, D). *Statistically significant differences (p < 0.05) between sphere-formed and attached cells through t-test and at least three repetitions were conducted, Scale bar is 100μm. siRNA: small interfering RNA.
WIPF1 regulates SCLC through the YAP/TAZ signaling pathway
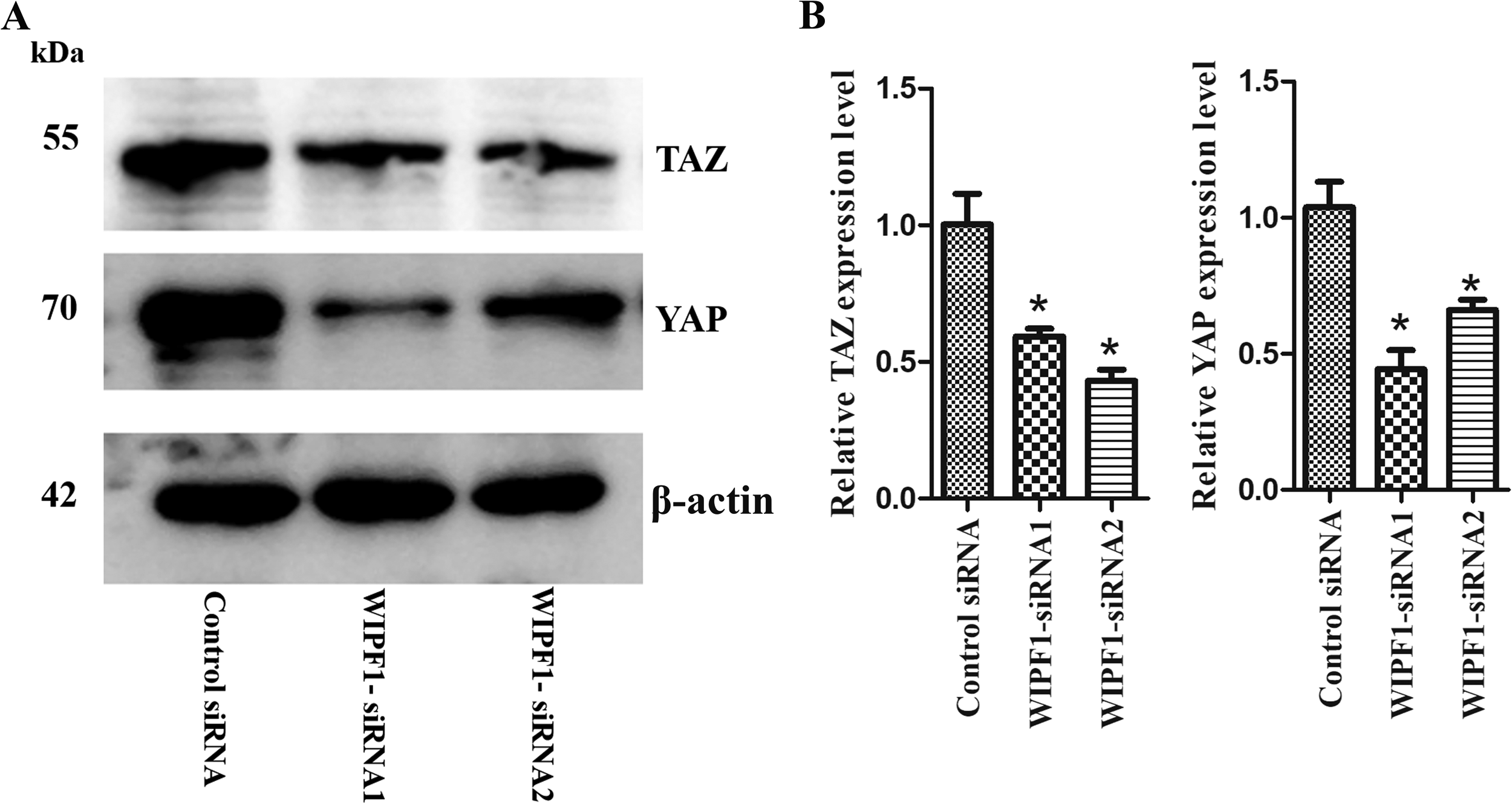
WIPF1 regulated small cell lung cancer cells through the YAP/TAZ signaling pathway. Western blots (A) and relative gray value (B) showed that WIPF1 knockdown significantly downregulated YAP, and YAZ expression in NCI-H446. β-actin served as the loading control. *Statistically significant difference (p < 0.05) between sphere-formed and attached cells through t-test and two repetitions were conducted.
Considering that CD133 is widely recognized CSC-like cells molecular marker and Chromogranin A (CgA) is useful NE marker in SCLC. Western blots were used to observe their expression after treatment with WIPF1-siRNA2. The preliminary results indicated that CD133 and CgA were downregulated after WIPF1-siRNA2 treatment (supplementary figure 5).
Protein–protein interaction network of WIPF1
To understand WIPF1's relationship with other proteins in the pathway, we used the STRING Version: 11.0 (https://www.string-db.org/) to construct a protein–protein interaction network and reveal the protein's direct physical binding and indirect functional connections. Results showed that WIPF1 interacted with four protein kinases, namely BTK, SYK, BRK, and NCK, which might affect the related signaling pathways (

Protein–protein interaction network showing that WIPF1 interacts with four protein kinases (BTK, SYK, BRK, and NCK) through STRING website (https://cn.string-db.org/).
Discussion
Numerous studies have suggested that tumors might possess a stem cell-like subpopulation called CSCs.29,41
Abundance of CSCs in SCLC has been associated with rapid emergence of chemoresistance and spheroidal growth in vitro.45,46 Results of the present study showed that sphere-formation capacity and chemoresistance play crucial roles in growth of CSC-like cells, while sphere-formation cultures represent a more effective approach for acquisition of enriched CSCs-like cells compared to traditional adherent culture.31,47 Transcriptomic analysis has revealed that WIPF1 was upregulated in sphere-formed cells comparing to differentiated ones under adherent growth conditions (2D cultures) in SCLC cell lines. Results from siRNA-mediated silencing of WIPF1 suggested that reduction of this gene suppressed the cell viability and sphere-formation capacity of SCLC cells.
Accumulating evidence has demonstrated the essential role of WIPF1 in tumorigenesis and cancer progression. For example, Lamont and colleagues
48
analyzed methylation profiles in keloids and found that WIPF1 was one of the top 10 hypomethylated genes. While Escoll et al.
24
found that decreased WIPF1 expression was associated with the p53 mutation and significantly downregulated CSC marker expression and suppressed of CSC cell viability, thus restraining tumor growth in vivo through the YAP/TAZ signaling pathway.
Protein–protein interaction network revealed that WIPF1 interacted with four protein kinases, namely BTK, SYK, BRK, and NCK, which might affect related signaling pathways. Taken together, the result of this study indicated that WIPF1 functions in pathogenesis and resistance of SCLC, suggesting its potential as a therapeutic target for this type of cancer.
The present study has several limitations. First, the precise molecular mechanism by which WIPF1 regulates the YAP/TAZ signaling pathway remains incompletely characterized, as we did not perform extended validation through animal models or human tumor specimen analyses. Considering the aggressive nature of SCLC and the scarcity of effective treatment options, further exploration of WIPF1's contribution to SCLC pathogenesis is both timely and warranted. Our subsequent research will focus on these key directions: (1) development of a WIPF1 overexpression plasmid system to proceed with the rescue experiments; (2) systematic screening for cellular proteins interacting with WIPF1 using co-immunoprecipitation coupled with mass spectrometry; and (3) collection of biopsy specimens from SCLC patients for further research. Additionally, we aim to investigate the potential involvement of WIPF1 in regulating novel cell death modalities in SCLC, particularly focusing on its possible connection with pyroptosis pathways. These proposed investigations may provide new insights into therapeutic target development for this recalcitrant malignancy.
Conclusion
In summary, WIPF1 regulates colony formation and sphere-forming ability in SCLC cells, as well as modulates their sensitivity to cisplatin. Knocking down WIPF1 significantly suppresses proliferation capacity and cisplatin sensitivity of SCLC cells via the YAP/TAZ signaling pathway. Protein–protein interaction networks revealed that WIPF1 interacts with four protein kinases (BTK, SYK, BRK, and NCK), which might affect related signaling pathways. Therefore, WIPF1 plays a crucial role in development and progression of SCLC.
Supplemental Material
sj-docx-1-sci-10.1177_00368504251345012 - Supplemental material for WIPF1 regulates stemness in small cell lung cancer
Supplemental material, sj-docx-1-sci-10.1177_00368504251345012 for WIPF1 regulates stemness in small cell lung cancer by Hongyong Fu, Mingji Quan, Qianqian Qiu and Fanjie Jin in Science Progress
Footnotes
Acknowledgement
We are grateful to all peer reviewers and editors for their opinions and suggestions, and our family and colleagues for supporting our work.
Author contributions
H.F. performed the experiments, wrote the manuscript, and was responsible for the conception, and final approval of the manuscript. Q.Q. and M.Q. assisted with the experiments and helped with data analysis. F.J. assisted with the experiments and ordered reagents. We agreed on the journal to which the article has been submitted.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by grants from Medical Science and Technology Research Program of Henan Province (Grant No. LHGJ20190636).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability statement
All relevant data are within the manuscript and its additional files.
Informed consent statement
There are no human participants in this article and informed consent is not required.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
