Abstract
Objective
The prediction of early response in locally advanced nasopharyngeal carcinoma (LA-NPC) after concurrent chemoradiotherapy (CCRT) is important for determining the need for timely consolidation therapy. We developed a radiomic analysis of multi-sequences MR based on machine learning (ML) to assess early response in LA-NPC after CCRT.
Methods
This study retrospectively enrolled 104 LA-NPC patients, randomly divided into training (70%) and test (30%) cohorts. Radiomic features were extracted from five MR sequences (T1, T1C, T2, DWI, and ADC). Feature selection was performed using Pearson's correlation coefficient and LASSO regression to reduce redundancy. ML algorithms were compared to develop models, with suboptimal sequences excluded from the multi-sequence MR fusion model. A combined model integrating the fusion and clinical model was developed using logistic regression, and its diagnostic effectiveness was evaluated using receiver operating characteristic (ROC) analysis.
Results
In the mono-sequence MR analysis, T1 demonstrated the lowest discriminative capacity (AUC = 0.505), followed by T2 (AUC = 0.738). Consequently, we developed a fusion model incorporating ADC, DWI, and T1C features while excluding T1 and T2. In the test cohort, the combined model outperformed both the clinical (AUC = 0.852) and fusion (AUC = 0.886) models, achieving superior effectiveness (AUC = 0.900). Shapley Additive Explanations (SHAP) analysis identified lbp_GrayLevelVariance_ADC as the most influential predictive feature.
Conclusions
A combined model, which merges clinical and multi-sequences MR radiomics model, showed good performance for predicting early response of LA-NPC after CCRT.
Keywords
Introduction
Nasopharyngeal carcinoma (NPC) is a malignant epithelial neoplasm that arises in the nasopharynx and exhibits a significant prevalence in East and Southeast Asia. 1 The diagnosis of NPC is often made at a locally advanced stage, with estimates indicating that this is the case in 75 to 90% of instances due to its deep localization and vague clinical presentation.2,3
Concurrent chemoradiation (CCRT) is the backbone treatment in locally advanced nasopharyngeal carcinoma (LA-NPC). 4 Following CCRT, 58%-89% of patients diagnosed with LA-NPC attain a clinical complete response (CR) within a few months.5–7 Early tumor regression has been identified as an independent prognostic indicator for overall survival (OS) and progression-free survival (PFS) in individuals diagnosed with NPC.8,9 Besides, research conducted by Huang et al. revealed that early intervention in NPC patients with radiographic residual disease resulted in improved short-term efficacy and long-term survival outcomes. 10 Therefore, it is essential to predict individuals with LA-NPC who still have residual lesions after undergoing systematic treatment as early as possible. This enables the implementation of timely consolidation or salvage therapy. A prior investigation conducted in 2023 established a magnetic resonance imaging (MRI)-based clinical radiomics nomogram aimed at forecasting the early response to tumor treatment. 11 However, in order to enhance the predictive performance, this study intends to augment the sample size and integrate novel machine learning techniques that leverage multi-sequence MR data.
Radiomics refers to the process of extracting high-dimensional data from medical images, making them amenable for analysis and mining purposes. It has facilitated the evolution of medical imaging from a tool primarily used for diagnostic purposes to one that serves as a clinical decision support system within the framework of personalized medicine. 12 Although radiomics studies have shown encouraging outcomes in forecasting treatment response in various tumor types, it is worth noting that the majority of conventional radiomics approaches predominantly concentrate on mono-sequence (e.g. MRI contrast-enhanced T1-weighted imaging in Bramen et al.; CT imaging in Jiang et al.)13–15 Multi-sequences MR imaging, in contrast to traditional radiomics, integrates two or more sequences into a unified system. This integration is fundamentally motivated by the complementary nature of MR sequences — while mono-sequence MR may excel in specific tasks (e.g. T1-weighted for anatomical structure, T2-weighted for edema detection), it inherently lacks the capacity to capture multifaceted biological information required for complex clinical diagnoses. By synergizing sequences, multi-sequence MRI overcomes individual limitations and provides a holistic characterization of tissues through high spatial resolution, improved soft tissue contrast, and the capacity to deliver molecular-level biological information with high sensitivity. 16
This systemic superiority has been consistently validated in recent studies, where multi-sequence models demonstrated significantly enhanced performance compared to mono-sequence approaches in predicting rectal cancer response to chemoradiotherapy, predicting disease-free survival in early-stage squamous cervical cancer, improving preoperative assessment of bladder cancer, and enabling precise evaluation of rectal cancer preoperative immunescore.17–22
We searched for relevant multi-sequence MR studies in the field of NPC. In the studies conducted by Cai and Shi, they integrated all radiomic features from T1-weighted images (T1-WI), contrast-enhanced T1-weighted images (T1-C), and T2-weighted images (T2-WI) sequences to predict the survival outcome or early treatment response of NPC patients, but they ignored the role of ADC and DWI.23,24 Zhang's study, compared the radiomic features obtained from individual T1-C, T2, and combined T1-C and T2 images, concluding that the combined images had better predictive efficacy. 25 However, the study did not consider other imaging sequences and lacked an in-depth discussion on the selection and integration of the T1-C and T2 sequences. Lu et al.'s research utilized PET/CT alongside integrated T1, T1-C, and T2 radiomic models to forecast the survival outcomes of patients diagnosed with NPC. 26 Similarly, the study did not provide a discussion on the rationale behind the selection of these specific sequences. Previous researches have shown that radiomic models utilizing Apparent Diffusion Coefficient (ADC) and Diffusion Weighted Imaging (DWI) MRI sequences possess the capability to accurately forecast the individual prognoses of different malignancies, including malignant bone marrow lesions and prostate cancer.27,28 Nonetheless, the effectiveness of radiomic models utilizing DWI and ADC sequences in assessing the early treatment response of NPC remains uncertain. Unfortunately, while prior studies predominantly focused on structural sequences (T1/T2/T1C), these studies above did not thoroughly examine the significance of DWI and ADC sequences in predicting outcomes. Additionally, they lacked in-depth discussion regarding the rationale behind selecting specific combinations of T1, T2, and T1-C sequences. Further study is warranted to explore the potential contributions of DWI and ADC sequences in outcome prediction and to provide a comprehensive understanding of the selection criteria for specific imaging sequence combinations.
This study unveils a combined model that fuses clinical data with multi-sequence MR radiomics, powered by machine learning, to forecast the early response of patients with LA-NPC after undergoing CCRT.
Materials and methods
Study patients
We declare that this study was conducted in strict compliance with the ethical principles outlined in the Declaration of Helsinki (revised in 2024).
From January 2020 to November 2023, we conducted a retrospective study of consecutive patients with pathologically confirmed NPC, involving a dataset of 219 patients. The stage of the disease was classified using the tumor node metastasis (TNM) staging, as delineated in the eighth edition of the American Joint Committee on Cancer (AJCC) guidelines.
Individuals who satisfied the specified criteria were incorporated into this study 29 :
Inclusion criteria: (a) Nasopharyngeal squamous cell carcinoma was confirmed through pathological examination; (b) Classified as stage III to IVA based on pre-CCRT MRI; (c) MRI images of the nasopharyngeal neck region were obtained both prior to and following treatment, encompassing T1-WI, T1-C, T2-WI, DWI, and ADC; (d) All patients underwent radical CCRT; (e) Complete clinical data.
Exclusion criteria included: (a) Inadequate MRI quality resulting from issues such as motion artifacts, blurring, or discontinuities in the images; (b) Background of past treatments for NPC; (c) History of prior malignancy; (d) The presence of concurrent immune system disorders or a background of prolonged hormone medication use; (e) Disease progression after induction chemotherapy (IC) or receiving treatment elsewhere after IC.
In total, 104 patients diagnosed with LA-NPC were taken into this study. We have de-identified all patient details. The patient selection process is illustrated in Figure 1.

Flowchart for the inclusion of patients.
Treatment protocols
The IC regimen employed in this study consisted of a taxane and platinum combination (TP), with intravenous administration of docetaxel at a dosage of 75 mg per square meter or paclitaxel at a dosage of 135–175 mg per square meter, in conjunction with nedaplatin or cisplatin at a dosage of 80 mg per square meter on day1, given once every 3 weeks for a total of 2–3 cycles. Following the conclusion of IC, it was advised that CCRT be conducted within a timeframe of three to four weeks. Radiation therapy (RT) in this study was administered using intensity-modulated radiotherapy (IMRT) mode, utilizing 6 MV photon irradiation. The recommended doses for the planning target volumes (PTVs) derived from the gross tumor volume of the nasopharyngeal carcinoma (GTVnx), gross tumor volume of cervix node (GTVnd), clinical target volume 1 (CTV1), and clinical target volume 2 (CTV2) were 66–70 Gy, 64–70 Gy, 60–62 Gy, and 54–56 Gy, respectively, in 30–33 fractions. Concurrent with RT, chemotherapy was administered intravenously with either cisplatin or nedaplatin at a dosage of 80 mg per square meter on the first day, repeated every 3 weeks. The number of chemotherapy cycles was tailored according to specific physical condition of each patient. It is advised that an MRI examination be conducted within one to three months following the completion of CCRT.
Clinical characteristics acquisition
Clinical characteristics prior to treatment were gathered from the health information system (HIS). These characteristics included age, sex, weight, height, smoking status, drinking habits, family history, EBV DNA level, white blood cell count (WBC), platelet count (PLT), neutrophil count, lymphocyte count, monocyte count, neutrophil to lymphocyte ratio (NLR), platelet to lymphocyte ratio (PLR), lymphocyte to monocyte ratio (LMR), serum albumin (Alb), alkaline phosphatase (ALP), lactate dehydrogenase (LDH), and d-dimer levels. Furthermore, the volume of the nasopharyngeal tumor, maximal axial diameter of positive lymph node (N-axial-length), maximum coronal length of positive lymph node (N-cor-length), and the total volume of lymph nodes (N-total-volume) were obtained by tracing the region of interest (ROI) using ITK-SNAP, an open-source software (www.itk-snap.org).
Tumor response criteria
Tumor response was conducted by comparing MRI images obtained prior to treatment with those obtained one to three months after the completion of CCRT. This evaluation adhered to the Response Evaluation Criteria in Solid Tumors 1.1 (RECIST 1.1) guidelines and was performed by two radiologists respectively. 30 Any discrepancies identified by the two radiologists were figured out through thorough discussion and consensus.
The Cohen's Kappa coefficient(κ) is a metric used to evaluate inter-rater agreement in categorical tasks, particularly by accounting for agreement due to chance. In medical imaging, it quantifies consistency among clinicians in tasks such as tumor segmentation or diagnostic classification, ensuring data reliability.
31
The metric ranges from 0 to 1, with higher values indicating stronger consistency. Cohen's Kappa was used for inter-user agreement of observers.
The responses were categorized into four groups: CR, partial response (PR), stable disease (SD), or progressive disease (PD). Compared with individuals who achieved CR after CCRT, the prognosis of patients with residual tumor is worse.9,32 Subsequently, the patients were categorized into the CR group, and non-CR group encompassing patients with PR, SD, and PD.
MRI image acquisition
Image acquisition was performed using GE 750 W 3.0 T MR or Philips Ingenia 3.0 T. Axial T1WI, T2WI, ADC, T1C and DWI pictures for each patient were obtained through PACS (Carestream, Ontario, Canada) in DICOM format. The manual segmentations of ROI were carried out utilizing ITK-SNAP by one radiation oncologists specializing in NPC radiotherapy with 3 years of experience. The accuracy of the segmentations was further validated by a senior radiation oncologist, who resolved any discrepancies through discussion. Cohen's Kappa was used for inter-user agreement of observers. In our study, we focused on segmenting the primary nasopharyngeal tumor region, while metastatic lymph nodes were not included.
Due to the potential variation in machine parameters, all our data underwent sampling at a 1 mm ×1 mm ×1 mm resolution, thus achieving a consistent spacing across all datasets. Figure 2 presents a visual representation of the radiomics process flowchart.
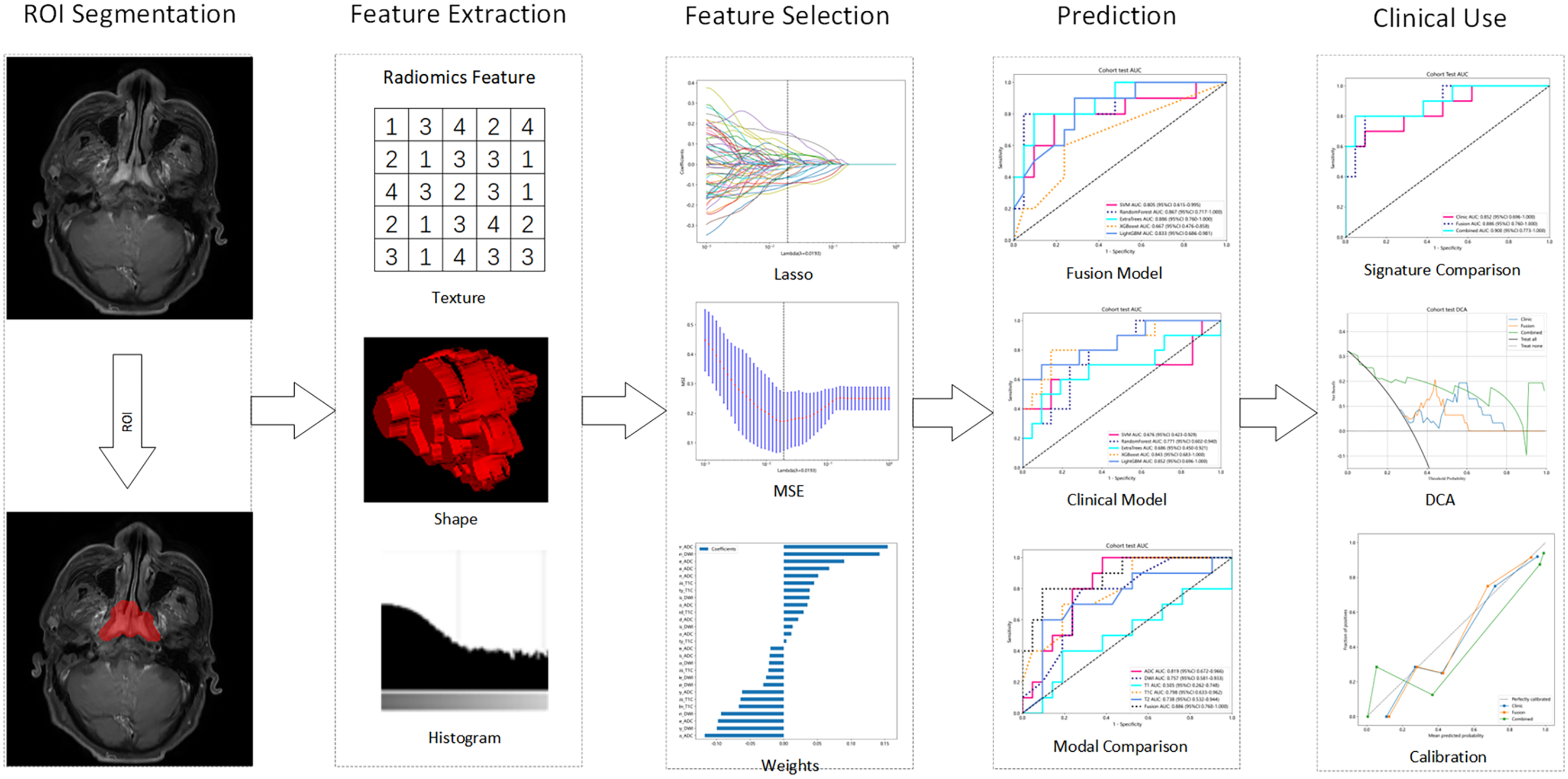
The flow chart of the process radiomics ROI, region of interest; Lasso, least absolute shrinkage and selection operator; T1-C, contrast-enhanced T1-weighted images; ADC, apparent diffusion coefficient; DWI, diffusion weighted imaging; MSE, mean square error; ROC, receiver operating characteristic; DCA, decision curve analysis; AUC, area under the curve.
Radiomics model
For increasing the reliability and reproducibility of the results, we added radiomic quality score (RQS) report as a relevant evaluation method. 33 (Supplement Table 1)
Feature extraction
In our analysis, we meticulously extracted radiomic features that corresponded to mono-sequences (T1, T1C, T2, ADC, and DWI). Our process involved categorizing the handcrafted features into three primary types: (I) Geometry, encompassing the three-dimensional shape characteristics of tumor; (II) Intensity, involving the first-order statistical distribution of voxel intensities in the tumor; and (III) Texture, detailing the patterns or higher-order spatial distributions of intensities. To enhance feature representation, we also applied image transformations such as Laplacian of Gaussian (LoG) and Wavelet to the original data. Various techniques were utilized for extracting texture features, such as the gray-level co-occurrence matrix (GLCM), gray-level run length matrix (GLRLM), gray-level size zone matrix (GLSZM), and neighborhood gray-tone difference matrix (NGTDM) were employed. The extraction of all features was conducted utilizing the pyradiomics tool (version 3.0.1), adhering to the standards established by the Imaging Biomarker Standardization Initiative (IBSI).
Feature selection
In our methodology, we initially standardized all extracted features using the Z-score normalization technique, ensuring a uniform distribution for subsequent analysis. This was followed by a t-test statistical evaluation of these normalized features, wherein we selectively retained those exhibiting a p-value less than 0.05, indicative of statistical significance.
To further refine our feature set and reduce redundancy, we employed Pearson's correlation coefficient to assess collinearity among the features. In instances where a pair of features demonstrated high linear correlation (similarity coefficient greater than 0.9), we strategically retained only one of the two. This step was crucial in minimizing redundancy while preserving the integrity and diversity of the feature set.
The model is instrumental in feature selection as it applies penalties to the absolute values of the regression coefficients. To optimize this model, we determined the best regularization parameter, λ, through a rigorous 10-fold cross-validation process. This approach involved evaluating various λ values to identify the one that minimized the cross-validation error, thereby ensuring the most effective feature selection. The final model included only those features with non-zero coefficients, representing the most relevant and informative predictors for our study. This comprehensive approach to feature selection and model optimization underscores the robustness and reliability of our findings.
Model building
Radiomics model
In our study, we employed the LASSO technique for feature selection and developed a risk model utilizing a range of machine learning algorithms, specifically SVM, ExtraTrees, XGBoost, Random Forest, and LightGBM. Comparative performance assessments were carried out among each mono-sequence. Feature-level fusion, also referred to as early fusion, entails consolidating all features from various modalities into a single feature vector. 34 This process integrates multi-modal features into a unified feature vector prior to model training, creating a comprehensive representation that preserves inter-modal relationships. It allows the model to simultaneously learn from all available imaging data while maintaining the inherent correlations between different sequences. In this study, we utilized the early fusion method to perform feature fusion. Feature fusion was performed on multi-sequences MR features, and the fused features were used to assess the effectiveness of multi-sequences versus mono-sequence approaches. Model robustness was ensured through 5-fold cross-validation in the training dataset, with hyperparameters optimized via Grid-Search. To evaluate the effectiveness of our multi-instance learning approach, we compared two ensemble methods: maximum and average values.
Clinical model
Clinical characteristics were quantitatively represented and subjected to analysis through machine learning algorithms, including SVM, ExtraTrees, XGBoost, Random Forest, and LightGBM, similar to the Radiomics Model.
Combined model
We incorporated radiomics into the clinical model to improve its clinical applicability. The diagnostic performance of the model was evaluated using the area under receiver operating characteristic curves (AUC-ROC) as an evaluative metric. We first selected the best-performing machine learning algorithm based on AUC and used its predicted probabilities as the Rad-Signature. The Rad-Signature was then combined with selected clinical features to form a combined model. Finally, we employed Logistic Regression (LR) to integrate the Rad-Signature and clinical features, leveraging their complementary strengths to build a robust and clinically applicable model. Decision curve analysis (DCA) was employed to evaluate the clinical utility of our predictive models, while calibration curves and the Hosmer-Lemeshow test were utilized to assess the calibration of the models.
SHAP visualization
The Shapley Additive Explanation (SHAP) method quantifies each feature's contribution to the model, ensuring a fair and consistent assessment of feature importance. In this study, SHAP analysis was employed to interpret the model's feature contribution.
Statistical analysis
The dataset was split randomly, with 70% allocated for training and the remaining 30% for testing. To evaluate patient clinical characteristics, we performed statistical analyses using the independent sample t-test, Mann-Whitney U test, and chi-squared test for categorical variables. As detailed in Supplement Table 2, the analysis showed no significant statistical differences between the training and testing groups because almost all features had p-values greater than 0.05. This confirms an unbiased distribution of data across the two datasets. Significant clinical features were discerned utilizing both univariate and stepwise multiple regression analyses. The analyses were conducted utilizing Python version 3.7.12 in conjunction with the statsmodels library version 0.13.2. Additionally, the construction of machine learning models was conducted utilizing the scikit-learn version 1.0.2 interface. Scikit-learn provided a comprehensive and user-friendly framework for implementing various machine learning algorithms and conducting model training, evaluation, and prediction tasks.
Results
Patients
Patients were categorized into the CR group (n = 63, 60.6%), and non-CR group (n = 41, 39.4%). The dataset was divided into training and test sets with a ratio of 7:3. In the training set, there were 73 patients, consisting of 42 CR cases (57.5%) and 31 non-CR cases (42.5%). The test set comprised 31 patients, with 21CR cases (67.8%) and 10 non-CR cases (32.3%). A previous study reported a 65.57% CR rate in LA-NPC patients treated with the TP regimen, which closely aligns with our current findings. 35
Cohen's kappa value was calculated to analyze the observers’ agreement for tumor response evaluation (k = 0.92 > 0.8) and handcraft tumor segmentation consistency (k = 0.85 > 0.8), indicating almost perfect agreement. This strong inter-observer agreement further validates the reliability and reproducibility of our segmentation methodology.
Feature statistics
In this study, we conducted an extensive extraction of radiomic features, yielding a detailed collection of 1834 manually crafted features derived from multiple sequences. The aforementioned features were subsequently categorized into three distinct groups: shape, comprising 14 features; first-order, encompassing 360 features; and a diverse array of texture features. After evaluating the performance of different sequences, we noted that T1 and T2 were less effective. Consequently, we focused on the remaining three sequences, aggregating a total of 5502 features. The extraction was performed using a custom-developed program in Pyradiomics, further details of which are available at http://pyradiomics.readthedocs.io. Supplement Figure 1 visually displays the distribution of these handcrafted features across the various categories.
Evaluation of rad model
Feature selection
In the final analysis stage, we used a LASSO logistic regression model to select features with nonzero coefficients, ensuring inclusion of only the most relevant features. The range of λ values explored in this research extended from 0.001 to 1. Ultimately, the final λ value selected during the 10-fold cross-validation process was λ = 0.0193. Figure 3 presents the coefficients and mean standard error (MSE) obtained from 10-fold validation, offering valuable insights into the model's performance and the significance of the features.

Radiomic feature selection based on LASSO algorithm. Ten-fold cross-validated coefficients (A) and MSE (B) and the histogram of the Rad score based on the selected features (C).
Metrics in combined modal
During the training phase, we applied 5-fold cross-validation and Grid Search to enhance the optimization of the model's hyperparameters, selecting the best parameters based on test dataset performance. The model was then trained on the entire training dataset for improved robustness and accuracy, with specific hyperparameters listed in Supplement Table 3.
In our fusion model comparison, the ExtraTrees classifier outperformed others, showing the highest AUC. (Table 1) It attained an AUC of 0.972 in the training dataset and 0.886 in the test dataset, indicating superior accuracy and generalization capabilities compared to SVM, RandomForest, XGBoost, and LightGBM. (Figure 4) The high AUC values, particularly in the test dataset, underscore the clinical potential of the ExtraTrees-based fusion model, thereby supporting personalized clinical decision-making. Additionally, the robust performance across both training and test datasets suggests that the model is reliable and applicable in real-world clinical settings, where generalizability is critical. The model effectively discriminates high-risk residual disease patients, enabling timely treatment intensification and improving prognosis. Specific metrics for each modality are shown in the Supplement Figure 2.
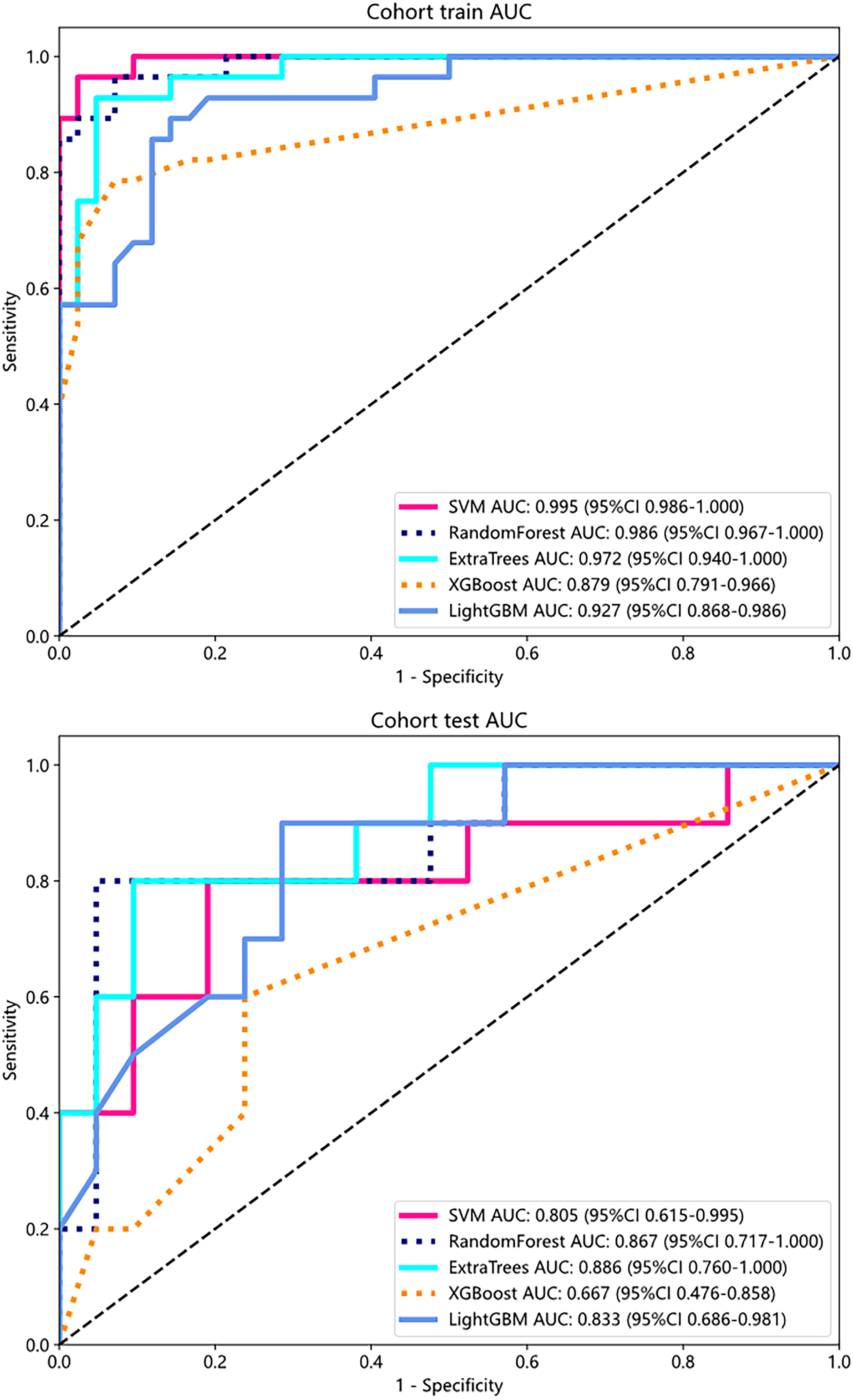
ROC curves for training and test sets of fusion model.
Comparative performance metrics of machine learning models for fusion model.
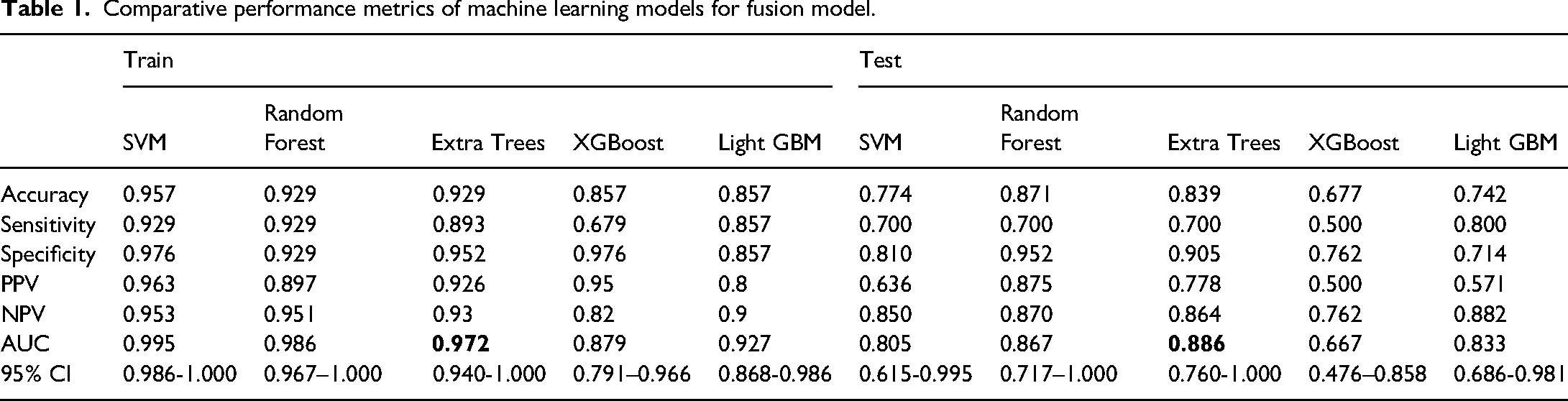
Comparison of radiomics different sequences
In the comparison of different imaging sequences, ADC was proven to be the most effective, exhibiting the highest AUC values. In the training dataset, ADC reached an AUC of 0.951, significantly outperforming T1 (AUC: 0.952), T1C (AUC: 0.767), T2 (AUC: 0.863), and DWI (AUC: 0.868). Notably, in the test dataset, T1's performance drastically declined (AUC: 0.505), while ADC maintained the highest AUC of 0.819 among individual sequences. (Table 2)
Task specific prediction performance of different models.

Based on these results, T1 and T2 features were excluded in the fusion model. The fusion model, integrating features from the selected sequences, demonstrated improved performance, achieving an AUC of 0.972 in the training dataset and 0.886 in the test dataset, demonstrating its superiority over mono-sequence approaches and validating the decision to omit T1 and T2 features. (Figure 5)
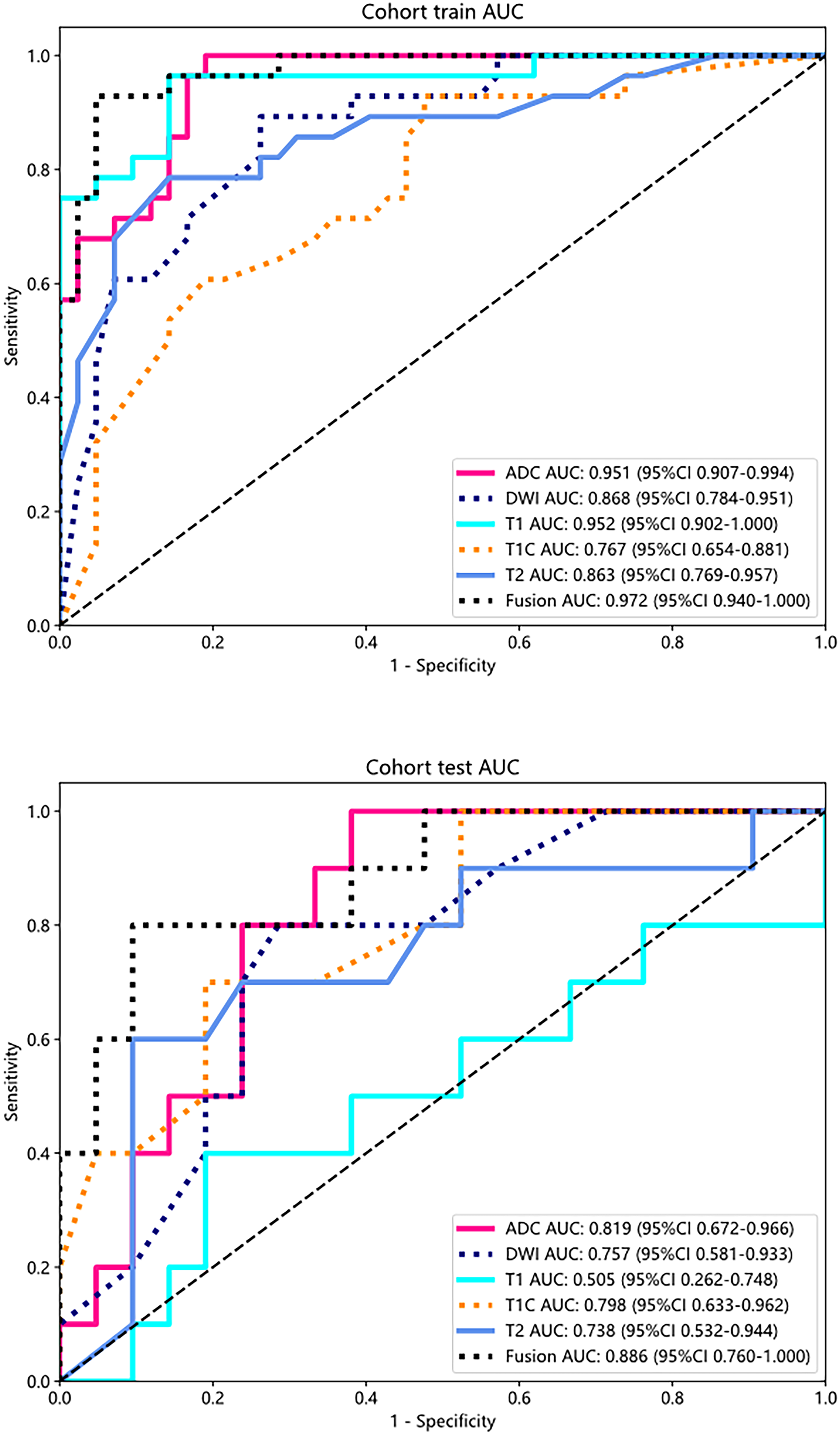
ROC curve of different sequences in train and test cohort. The fusion model, integrating features from the selected sequences(T1C, ADC and DWI), showed enhanced performance with an AUC of 0.972 in the training cohort and 0.886 in the test cohort.
The Delong's test demonstrated that the fusion model achieved statistically significant performance improvement over the T1 model in the test cohorts (p < 0.05). These results confirm the clinical value of multimodal integration in our combined model, underscoring its superior diagnostic performance compared to single-modality approaches. (Supplement Figure 3)
Model comparison
The Combined model, which merges clinical and multi-sequences MR radiomics model, exhibited higher accuracy, sensitivity, specificity, PPV, and NPV in both the training (0.957, 0.939, 0.976, 0.963, 0.953) and test (0.871, 0.7, 0.923, 0.875, 0.87) datasets, outperforming the Clinic and Fusion models. This demonstrates the model's reliability in accurately classifying cases, maintaining robust performance in identifying both positive and negative cases across datasets.
In the training cohort, the clinical, fusion, and combined models demonstrated false positive rates (FPR) of 21.4%, 4.8%, and 2.4%, and false negative rates (FNR) of 17.9%, 10.7%, and 7.1%, respectively. Corresponding values in the test cohort were 9.5%, 9.5%, and 4.8% for FPR, and 40%, 30%, and 30% for FNR. The combined model consistently reduced both FPR and FNR across training and test cohorts, thereby optimizing the trade-off between overtreatment and missed diagnoses - a critical balance for clinical deployment.
Moreover, the Combined model, demonstrated superior performance in terms of AUC. In the training dataset, while the Clinic model achieved an AUC of 0.892 and the Fusion model reached 0.972, the Combined model excelled with an AUC of 0.990, indicating a substantial improvement. Similarly, in the test dataset, the Combined model outperformed both the Clinic (AUC: 0.852) and Fusion models (AUC: 0.886), achieving a remarkable AUC of 0.900. (Supplement Table 4)
These metrics collectively underscore the robustness of the Combined model, highlighting its superior performance not only in terms of AUC but also in other critical clinical performance indicators. This comprehensive evaluation further validates the decision to integrate clinical and multi-sequence MR radiomics features, as it enhances the model's overall diagnostic accuracy and reliability. (Figure 6)

Different models’ ROC on train and test cohort.
Calibration
The Hosmer-Lemeshow (HL) test evaluates the fit between predicted probabilities and actual outcomes, where lower values indicate better calibration, contrary to initial assumptions. In our research, the Combined model demonstrated superior calibration, as indicated by the HL test values of 0.883 and 0.605 for the training and test datasets, respectively. The observed low values indicate a significant alignment between the predicted results and the actual outcomes, thereby underscoring the model's reliability. (Supplement Figure 4)
Model explanation and visualization
SHAP provided a quantitative explanation for the fusion model. The beeswarm plot (Figure 7A) reveals feature contributions by displaying the distribution of SHAP values across samples, where features are vertically ranked by global importance. Red and blue colors indicate positive or negative effects of features on prediction outcomes, with point density clusters showing common effect ranges. Notably, lbp_GrayLevelVariance_ADC emerges as the most influential predictor for CR/nonCR classification and the coloring showed that the model's output increased with decreasing value of this feature. The feature importance analysis (Figure 7B) quantifies these relationships through mean absolute SHAP values, identifying lbp_GrayLevelVariance_ADC, original_maximum3Ddiameter_ADC, and gradient_Skewness_ADC as key predictive features. Together, these visualizations provide clinicians with transparent insights into the model's decision-making process by quantifying individual feature contributions and revealing effect directionality, thereby aligning with current best practices for interpretable machine learning in medical applications.

SHAP beeswarm plot of fusion model. The plot illustrated feature attributions to the model's predictive performance.

SHAP feature importance bar chart. The plot illustrates the relative importance ranking of features.
Figure 8 presents a heatmap illustrating the distribution of 27 selected radiomic features across CR and non-CR patient groups. The color intensity in the heatmap corresponds to feature value magnitudes, with darker shades typically indicating higher feature values. This visualization enables intuitive comparison of radiomic feature differences between patient groups, facilitating identification of features potentially associated with treatment response.

Heatmap visualization of 27 selected radiomic features in the fusion model.
Clinical use
Supplement Figure 5 presents the DCA curves for both the training and test datasets. The analysis indicates that our combined model offers substantial advantages regarding predicted probabilities. Furthermore, the analysis demonstrates a higher potential for net benefit compared to other models, as evidenced in the DCA results.
Discussion
There is a lack of validation for a radiomic analysis of multi-sequences MR using machine learning to predict early response in LA-NPC after CCRT. In this research, a combined model merging clinical and multi-sequences MR radiomics model was constructed to make it come true. This approach demonstrated good performance and proved beneficial for clinicians in assessing the early response in time.
Indeed, numerous studies have examined the efficacy of conventional radiomics models in forecasting early responses or disease progression in patients diagnosed with LA-NPC.11,24,36,37 However, it is crucial to note that traditional radiomics models typically rely on a single or a limited number of sequences for feature extraction and analysis. In contrast, our approach stands out due to its comprehensive utilization of multiple sequences.
With the deepening of cancer radiomics research, the current cancer radiomics based on mono-sequence has exposed its shortcomings. Mono-sequence imaging can only partially reflect the tumor information, and inevitably miss some tumor information. Multi-sequences MR radiomic involves the extraction of comprehensive information from each image sequence, followed by the integration of these data for the purpose of model development. 38 Consequently, the risk of information loss is mitigated. The segmentation of subregions through the integration of multi-sequence MR facilitates the retention of tumor heterogeneity in the radiomic features derived from these subregions, thereby enhancing the overall efficacy of radiomic analysis. 39 Thus, multi-sequences MR models have demonstrated superior performance compared to mono-sequence models. 40
It is worth noting that our multi-sequences MR model construction process involved a selection-based approach. Through evaluating the performance of different sequences, we observed that T1 (AUC: 0.505) and T2 (AUC: 0.738) sequences were less effective in predicting the outcome. Conversely, ADC (AUC: 0.819) emerged as the most effective sequences in the test dataset. By integrating features from the selected sequences (TIC, DWI, and ADC), the fusion model exhibited enhanced performance with an AUC of 0.886 in the test dataset. This result demonstrates the superiority of fusion model over mono-sequence approaches and validates the decision to exclude T1 and T2 features from the model construction.
Notably, the performance of T1 in the test dataset demonstrated a rapid decline, which is consistent with findings from previous studies. In a study conducted by Shi et al., similar outcome was observed where T1 and T2 sequences showed inferior predictive performance compared to T1C sequences in the test dataset. Among the sequences, T1 exhibited the poorest predictive performance. 24 This observation aligns with the clinical perspective, as the T1C sequence provides better information regarding tumor contrast and blood supply due to the enhancement provided by the contrast agent. This enables physicians to more accurately detect and localize tumors and discern differences between tumor and surrounding normal tissue. Due to the absence of contrast agents, T1 sequences alone may not provide sufficient information for determining the location and extent of nasopharyngeal tumors. Therefore, in clinical practice, T1 sequences are often combined with T1C imaging to accurately assess the position and involvement range of nasopharyngeal lesions. Additionally, T2 images exhibit high sensitivity in detecting effusion or edema. Consequently, T2 sequences facilitate improved visualization of lymph nodes, thereby allowing for a more accurate assessment of lymph node metastasis and its specific localization. Recent research has validated the potential clinical application of T2 sequences in differentiating between benign and metastatic lymph nodes within the retropharyngeal region in cases of NPC.41,42 As our study primarily revolved around feature extraction and the establishment of radiomics models focusing on the ROI of nasopharyngeal lesions, the contribution of T2 and T1 sequences to this particular radiomics analysis was relatively minor compared to other sequences.
Hu et al. conducted a study where they found the use of a machine learning model that utilized features extracted from DWI maps as a potential prognosis detection tool for NPC patients. 43 In malignant tumors, water molecule diffusion is often restricted by high cell density, resulting in higher signal intensity on DWI and lower values on ADC. DWI technology provides both quantitative and qualitative information, thereby improving the specificity of disease diagnosis. 43 This study provides evidence that incorporating DWI and ADC sequences in radiomics has the potential to complement or potentially replace the conventional sequences currently used (T2, T1, and T1C). By doing so, it can offer additional high-specificity information and support for decision-making in the field of radiology.
Radiomics clinical visualization methods are diverse, ranging from nomograms to complex interactive tools.44–47 These methods not only enhance the interpretability of radiomics data but also provide intuitive support for clinical decision-making. A nomogram is an interpretability tool used to visualize predictive models, graphically displaying the contribution of each feature to the prediction outcome. In a study by shen et al., a nomogram was constructed to visualize a radiological model for identifying early hematoma expansion in spontaneous intracerebral hemorrhage. 48 Heatmaps are commonly used to display the distribution or patterns of high-dimensional data. A heatmap was utilized to study the association between radiomic features and clinical parameters in patients with early-stage cervical cancer. 47 SHAP is a feature importance interpretation method that demonstrates the contribution of each feature to the model's prediction. The SHAP tool identified two features of the venous phase as the most significant, effectively differentiating between G1 and G2/3 of pancreatic neuroendocrine tumors (pNETs), demonstrating favorable interpretability. 46 Other methods include generating attention maps from deep learning models and developing interactive tools, such as web-based visualization platforms, 45 to enable clinicians to explore radiomics data. With the continuous advancement of technology, radiomics visualization is poised to play an increasingly significant role in precision medicine.
The translation of ML-based medical tools into clinical practice requires careful consideration of regulatory requirements, such as FDA approval or CE marking. Regulatory bodies emphasize the need for robust validation across diverse datasets, transparency in model interpretability, and adherence to data privacy standards. Furthermore, demonstrating clinical utility through prospective trials and ensuring generalizability across different populations are critical steps in the approval process. Addressing these regulatory hurdles is essential to ensure the safety, efficacy, and reliability of AI-driven tools in real-world healthcare settings.
Our predictive model demonstrates encouraging accuracy (AUC = 0.90), suggesting the ability to effectively identify patients at high risk of residual disease. First of all, clinicians can leverage the risk stratification provided by the model to customize treatment plans. For these high-risk patients, treatment intensification should be considered through combination with targeted therapies, immunotherapy, or enhanced maintenance therapy following radiotherapy. Conversely, for low-risk residual disease patients - particularly those with T4 or N3 classification who have undergone induction chemotherapy and concurrent chemoradiotherapy - the use of metronomic chemotherapy may potentially be omitted. Further non-inferiority studies based on these findings will be required to validate the safety of reducing unnecessary interventions in selected patient populations. Secondly, the model's outputs facilitate meaningful discussions between clinicians and patients regarding treatment options, using visual aids to clarify risks and recommendations. Thirdly, accurate predictions allow healthcare facilities to optimize resources, such as scheduling radiotherapy and follow-up care for high-risk patients.
Limitations
However, it is crucial to acknowledge and address the limitations of this study. Firstly, it is important to acknowledge that our study was conducted with a relatively modest sample size and was limited to a single institution, which may affect the generalizability of our findings. The absence of external validation further highlights the need for caution when interpreting the results. In future work, we aim to collaborate with multi-institutional cohorts to validate our findings across diverse populations and settings. Secondly, while standard MRI sequences were used, DCE-MRI and PET-CT offer additional insights in reflecting changes in tissue microcirculation and metabolic activity, providing more accurate diagnostic and treatment recommendations. Although the integration of multi-sequences MR is valuable for advancing radiomics, there are indeed challenges in the implementation process. Variances in spatial structures can be observed among images obtained from different sequences due to factors like temporal differences, patient positioning, device field of view, and the number of slices captured. 38 We are optimistic that future advancements in image registration and alignment tools will facilitate smoother integration of multi-sequence data. We remain committed to exploring these advancements in our ongoing efforts to improve the accuracy and reliability of radiological analyses. Thirdly, the biological interpretation of radiomic features remains challenging. In future studies, we will conduct radiogenomic analyses to correlate radiomic features with molecular characteristics, thereby providing biological explanations for currently ambiguous radiomic signatures.
Conclusion
In conclusion, we have successfully developed and validated a combined model that integrated clinical features with multi-sequence MR radiomics model. This combined model demonstrates exceptional predictive performance, achieves an impressive AUC of 0.900, which significantly outperforms models based on clinical or radiomics features alone. These results underscore the effectiveness of radiomic analysis of multi-sequences MR, supported by machine learning, in enhancing the early response prediction for LA-NPC patients following standard treatment. The superior performance of the combined model highlights its potential as a robust tool for personalized treatment planning and clinical decision-making in LA-NPC management.
Supplemental Material
sj-docx-1-sci-10.1177_00368504251338930 - Supplemental material for Radiomic analysis based on machine learning of multi-sequences MR to assess early treatment response in locally advanced nasopharyngeal carcinoma
Supplemental material, sj-docx-1-sci-10.1177_00368504251338930 for Radiomic analysis based on machine learning of multi-sequences MR to assess early treatment response in locally advanced nasopharyngeal carcinoma by Lei Qiu, Yinjiao Fei, Yuchen Zhu, Jinling Yuan, Kexin Shi, Mengxing Wu, Gefei Jiang, Xingjian Sun, Jinyan Luo, Yurong Li, Weilin Xu, Yuandong Cao and Shu Zhou in Science Progress
Footnotes
Acknowledgements
Thanks to our colleagues and staff at the Department of Radiation Oncology.
Ethical considerations
The study received full ethical approval under the identifier 2023-SRFA-552 from The First Affiliated Hospital with Nanjing Medical University Institutional Review Board. The study process was in accordance with the Declaration of Helsinki Ethics statement.
Authors contributions
LQ, YF and YZ contributed equally to this article and share first authorship.
Data management, Lei Qiu, Yinjiao Fei, Mengxing Wu and Yurong Li; Formal analysis, Kexin Shi, Mengxing Wu and Weilin Xu; Funding, Shu Zhou; Survey, Yuchen Zhu, Kexin Shi, Gefei Jiang and Weilin Xu; Methodology, Lei Qiu; Project Management, Shu Zhou; Resources, Yuandong Cao; Software, Yuchen Zhu, Jinling Yuan, Gefei Jiang and Xingjian Sun; Supervision, Yuandong Cao; Validation, Xingjian Sun; Visualization, Yinjiao Fei, Jinling Yuan and Jinyan Luo; Writing - Original draft, Lei Qiu, Yinjiao Fei and Yuchen Zhu; Writing-review and editing, Weilin Xu, Yuandong Cao and Shu Zhou.
All authors contributed to the article and approved the submitted version.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Thanks to the support of Jiangsu Province Entrepreneurship and Innovation Doctoral Talent Program (2019303073386ER19) and Jiangsu Province People's Hospital Clinical Capability Enhancement Project (JSPH-MC-2021-17).
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability statement
The datasets used and/or analysed during the current study are available from the corresponding author on reasonable request.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.

