Abstract
Pearl millet (
Keywords
Introduction
Throughout history, agriculture has faced significant challenges due to different global climates. In particular, arid and semiarid regions are susceptible to various abiotic stresses, such as salinity, drought, heat, and cold, which negatively affect crops. Pearl millet (
Pearl millet is considered the sixth most important cereal in the world, serving as a staple food for approximately 90 million impoverished people. It covers a large agricultural area of 30 million hectares in the arid and semiarid tropical regions of Asia and Africa. To highlight its significance, the FAO Committee on Agriculture (COAG) designated 2023 as the International Year of Millets. 5 One of its notable characteristics is its exceptional tolerance to abiotic stresses, especially in salt-affected regions, which makes it indispensable for cultivation in challenging environments. Further research in this area has great potential to improve pearl millet varieties with enhanced stress tolerance, thus supporting sustainable agriculture in arid and semiarid regions. A previous study on pearl millet examined the genetic diversity of six local pearl millet varieties in Saudi Arabia by analyzing their chloroplast DNA trnL-F regions. Variations have been observed among the varieties, shedding light on their adaptation to the harsh climate of the Arabian Peninsula.6,7 Understanding stress tolerance mechanisms in plants can lead to better strategies for enhancing crop resilience and food security, especially in salty regions.
Nature's adaptability is evident in the ability of plants to withstand abiotic stress. Understanding the mechanisms underlying stress tolerance in pearl millet offers exciting prospects for sustainable agriculture and food security. By exploiting plant resilience, we can address the challenges of the ever-increasing global population and create a sustainable agricultural future. Unraveling stress tolerance mechanisms offers great potential for sustainable agriculture and environmental protection. 8 Leveraging plant resilience is crucial to ensure nutritious food for the world's growing population.
In Dhawi, 9 we investigated how pearl millet responds to salt stress, which is a critical concern in arid and semiarid regions. Fourteen-day-old seedlings were divided into control, 75 mM NaCl, and 150 mM NaCl groups and irrigated daily for 7 days. RNA sequencing of leaves revealed 27.6% differentially expressed genes (DEGs), comprising 3246 upregulated genes and 7408 downregulated genes, compared with the control and 150 mM NaCl groups. Pathway analysis revealed the involvement of upregulated genes in crucial pathways such as phytyl-PP biosynthesis and lysine degradation, whereas downregulated genes were associated with coumarin and cholesterol biosynthesis. Moreover, a prior study by Dhawi, 10 utilizing enrichment analysis to emphasize biological processes crucial for thermal response in pearl millet, revealed that among 36,041 genes, 10 exhibited significant fold changes previously unknown for such responses. Computational tools for exposing conserved amino acid transporter domains. Phylogenetic analysis revealed distinct clustering patterns among genes, indicating evolutionary relationships and potential functional similarities. In addition, molecular function analysis revealed phosphorelay sensor kinase activity and protein kinase activity in the thermally treated leaves, suggesting a complex genetic basis for pearl millet tolerance to multiple environmental stresses. In a recent investigation conducted by Huang et al., 11 seedlings of pearl millet were cultivated for a duration of 13 days, followed by exposure to a heat treatment at 40°C for 14 h. Subsequent analysis of pearl millet seedlings post-heat stress revealed discernible variations. Leaf samples were collected at intervals of 1, 48, and 96 h post-heat stress exposure, corresponding to distinct experimental groups. Transcriptome sequencing was employed to elucidate the molecular responses of pearl millet leaves subjected to varying durations of heat treatment (1, 48, and 96 h). The comprehensive analysis identified a total of 12,451 unique genes exhibiting differential expression. Specifically, 4659 DEGs were observed following 1 h of heat stress exposure, 3981 DEGs after 48 h, and 8286 DEGs after 96 h. This indicates a progressively greater impact of prolonged heat stress on the transcriptional profile of pearl millet leaves. Considering the current trend of climate change, characterized by an annual increase in temperature of 1–1.5°C, 12 our study aims to investigate the effects of elevated temperature, specifically 50°C, for a brief duration of 60 s. This experimental approach is designed to mitigate the initiation of apoptosis or programmed cell death in response to heat stress, while concurrently enabling the examination of gene expression patterns through transcriptional analysis. We present insights from transcriptomic analysis, which is a powerful tool for unraveling the underlying mechanisms of stress response networks in pearl millet. This was achieved by comparing fold changes concerning the control for two primary abiotic stress factors on pearl millet: salt and heat. Continued research on pearl millet opens the possibility of creating more resistant varieties and contributing to sustainable agriculture in arid and semiarid environments.
Materials and methods
Plant materials
Seeds of the pearl millet cultivar (
After 10 days of growth, pearl millet samples were categorized into three groups: a control group devoid of any treatment, a salinity stress group treated with 150 mMol NaCl for 3 days (200 ml of saline each day), and a heat stress group exposed to 50°C for 60 s at 13 days of age. Post-treatment, samples were collected from 13-day-old seedlings, immersed in RNAlater Stabilization Solution following the Lader method,13,14 and stored at 4°C. RNA extraction from stems was conducted using the Trizol method developed by Chomczynski and Mackey. 13 The experimental configuration comprised a control group of 14 seedlings grown under normal conditions and an abiotic stress group consisting of 14 seedlings subjected to both heat and salt stress simultaneously. The analysis comprises three replications of RNA analysis per group.
Transcriptome sequencing data analysis
Genomic DNA and double-stranded cDNA quality assessment involved Qubit dsDNA estimation and agarose gel electrophoresis. Each sample's 500 ng genomic DNA and 200 ng cDNA were combined for library preparation using the NEBNext UltraII DNA library preparation kit for Illumina (Cat no: E7770, New England Biolabs), following the manufacturer's protocol. Sequencing was carried out on the S4 Lowell of NOVASEQ 6000 with 150-bp paired-end chemistry, outsourced to NGB Diagnostics, Noida, India.
DNA samples were shared with an approximate 300–400 bp insert size. Fragmented genomic DNA underwent end repair, A-tailing, indexed adapter ligation, purification, and PCR enrichment to obtain the final genomic DNA library. Library quantification utilized a Qubit Fluorometer (Invitrogen), while the HSDNA kit on Tapestation (Agilent Technologies, USA) assessed the distribution of library fragments. Tagged genomic DNA libraries were pooled in equal ratios, loaded onto the c-bot automated system for cluster generation, and then onto the Illumina S4 Flow Cell of the Illumina Novaseq 6000 Sequencing system. Sequencing utilized 2150 bp paired-end chemistry. Following sequencing, samples were demultiplexed, and indexed adapter sequences were trimmed using CASAVA v1.8.2 software from Illumina Inc. The entire library preparation and sequencing processes were outsourced to commercial service providers, ensuring robust and standardized procedures.
Quality control of sequencing data
In next-generation sequencing (NGS), factors like sequencing platform, reactants, and sample quality impact overall sequencing quality and base error rate. To assess sequencing quality distribution, we analyzed the full length of sequences, identifying positions with low sequencing quality. This analysis ensures the overall reliability of sequencing data.
Sequencing evaluation summary
A total of 93.68 GB of raw data was obtained. Cleaning involved adapter removal and discarding short sequences (<20 bp). After cleaning, 93.06 GB of high-quality data was available.
Mapping and differential analysis
Unigenes were indexed using BWA, and cleaned data were mapped against these unigenes using the BWA MEM algorithm. SAM files were processed using SAMTOOLS.15–17 For differential analysis, DESeq2 was used to identify DEGs based on specific criteria. Volcano plots, violin plots, and a heatmap of top 30 genes were generated for result visualization. Hierarchical clustering was performed to explore relationships among genes.
Read quality check and FastQC analysis:
To assess the quality of the reads, FastQC version 0.11.1 (www.bioinformatics.babraham.ac.uk), 15 was employed. The cleaned data underwent quality checks using FastQC. The parameters examined included base and sequence quality distributions, average base content per read, and GC distribution. Universal Illumina adapters were removed, and sequences shorter than 20 bp were discarded. Preprocessed reads were assembled de novo using Trinity with default settings. CD-HIT was then applied to cluster sequences at 90% similarity, and the longest isoform from each cluster was selected as a unigene. Unigenes were indexed using BWA, and cleaned data were mapped against them using the BWA MEM algorithm.
Quantification and DESeq2 analysis:
Reads mapped to genes were quantified using Samtools. DESeq2 was applied for differential analysis, identifying DEGs based on log2 fold change and adjusted p-value criteria. Unigenes were annotated using BLAST and the UniProt database. KEGG and GO enrichment analyses were performed using clusterProfiler and topGO, respectively.
Data submission to NCBI:
Pearl millet gene assembly v.1.1 reference genome was aligned using HISAT2 (http://daehwankimlab.github.io/hisat2/). 16 The resultant SAM files were then converted to the BAM format using Samtools (http://www.htslib.org). 17 Pearl millet gene assembly v.1.1 reference genome alignment, raw count matrix creation, normalization, and data submission to NCBI as Sequence Read Archive (SRA) were carried out using HISAT2, Samtools, and DESeq2. The RNA sequences are deposited under the accession number PRJNA991076.
Results
Visualizing DEGs in pearl millet under abiotic stress and pathway analysis:
The investigation into RNA sequence analysis of pearl millet shoots under abiotic stress delved into the examination of a substantial pool of 36,041 genes. From this comprehensive dataset, a total of 17,271 genes were successfully identified, each linked to specific gene IDs, KEGG annotations, UniProt information, and their corresponding known functions. Notably, among these, 2388 genes exhibited a distinctive pattern of upregulation in response to the imposition of heat stress, while a separate set of 4327 genes experienced downregulation. In the context of salt stress, a different pattern emerged, with 2013 genes showing an upregulated response and 4221 genes undergoing downregulation, as visually represented in Figure 1.
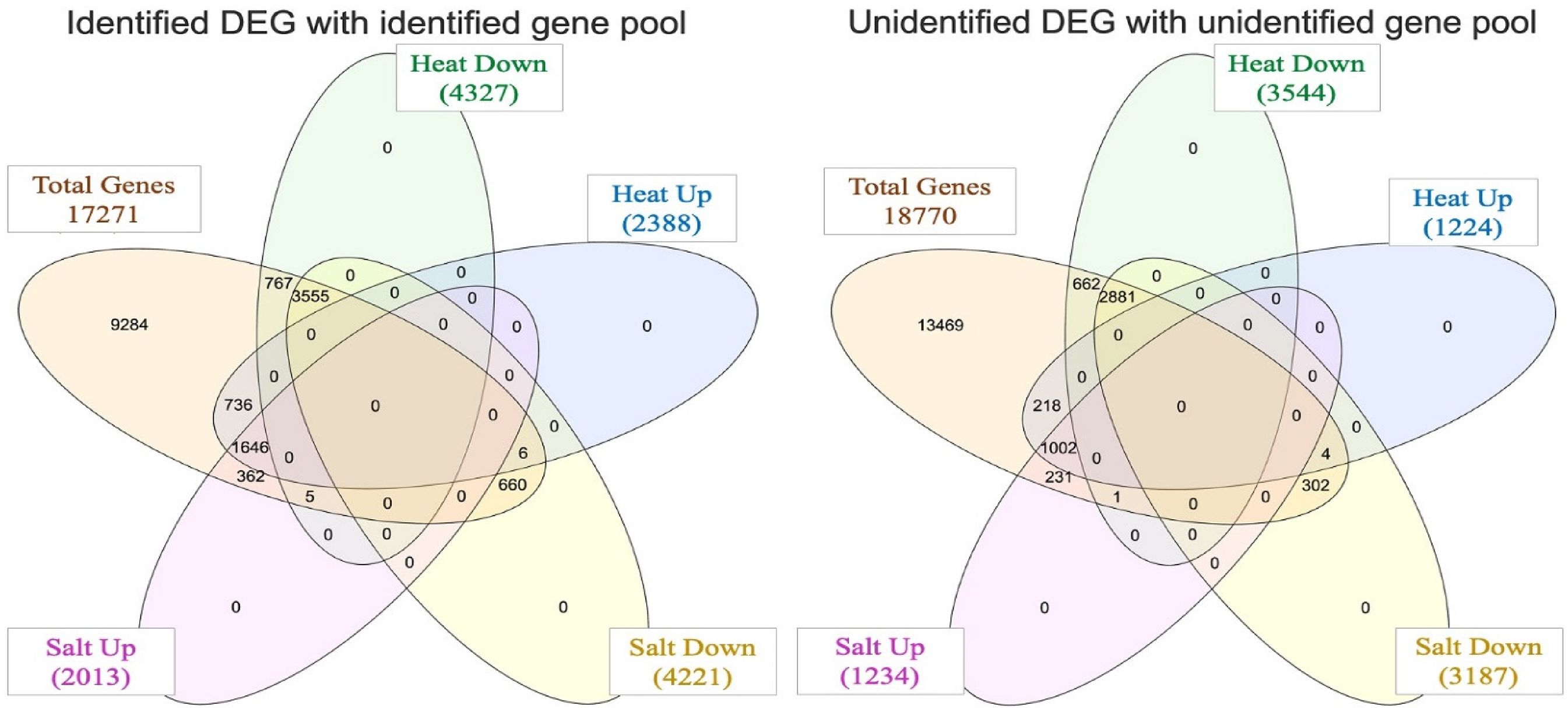
Venn diagram representing the total number of genes, both identified and unidentified, that demonstrate differential expression in pearl millet shoots under heat and salt stress conditions compared with the control group.
Moreover, a meticulous comparative analysis brought to light the intriguing observation that 3555 genes underwent a shared downregulation in both salt and heat stress conditions, hinting at potential common regulatory mechanisms. In contrast, 1646 genes exhibited a coordinated upregulation in pearl millet shoots, indicating shared molecular responses to the stressors, as depicted in Figure 2.

Venn diagram representing the total number of identified differential expressions (upregulated and downregulated) in pearl millet shoots under heat and salt stress conditions compared with the control group.
Within the scope of this study, a comprehensive examination of DEGs was specifically conducted for pearl millet shoots subjected to salt stress. The detailed findings of this analysis are meticulously presented in Supplemental Table 1, providing a statistical and descriptive breakdown of these DEGs. This tabular representation not only offers insights into the distribution of these genes but also sheds light on their characteristic features under the specific conditions of salt stress.
Furthermore, the study expanded its inquiry to unveil the intricate molecular landscapes influenced by both heat and salt stress conditions. Supplemental Table 2 elucidates the KEGG pathways associated with the DEGs in pearl millet shoots. This detailed table not only enumerates the genes participating in each pathway but also provides an insightful narrative on the potential molecular processes and metabolic pathways that bear the influence of these stressors. In essence, the tables serve as valuable tools for researchers, offering a nuanced understanding of the gene expression dynamics and the molecular intricacies governing pearl millet's responses to heat and salt stress.
The assessment of fold changes in gene expression serves as a pivotal metric for discerning alterations in pearl millet shoots under normal conditions (control) as opposed to those subjected to abiotic stress conditions, specifically heat and salt stress. Figure 3 serves as a graphical representation, providing a visual depiction of the magnitude of upregulation or downregulation in gene expression under stress conditions relative to the control group. This figure facilitates a comprehensive understanding of the differential gene expression patterns induced by different stressors.

Fold change in DEGs between shoots in control compared with samples under abiotic stress conditions (heat and salt).
The fold change data play a crucial role in constructing a heat map, a powerful tool that visually portrays the expression levels of numerous genes across various samples of heat-stressed and salt-stressed pearl millet shoots in comparison with the control group. Figure 4 employs color gradients to represent upregulated and downregulated genes, allowing for the identification of distinct gene expression patterns in response to diverse stress conditions. The heat map serves as an insightful visualization, enhancing the interpretation of the intricate gene expression dynamics induced by heat and salt stress.
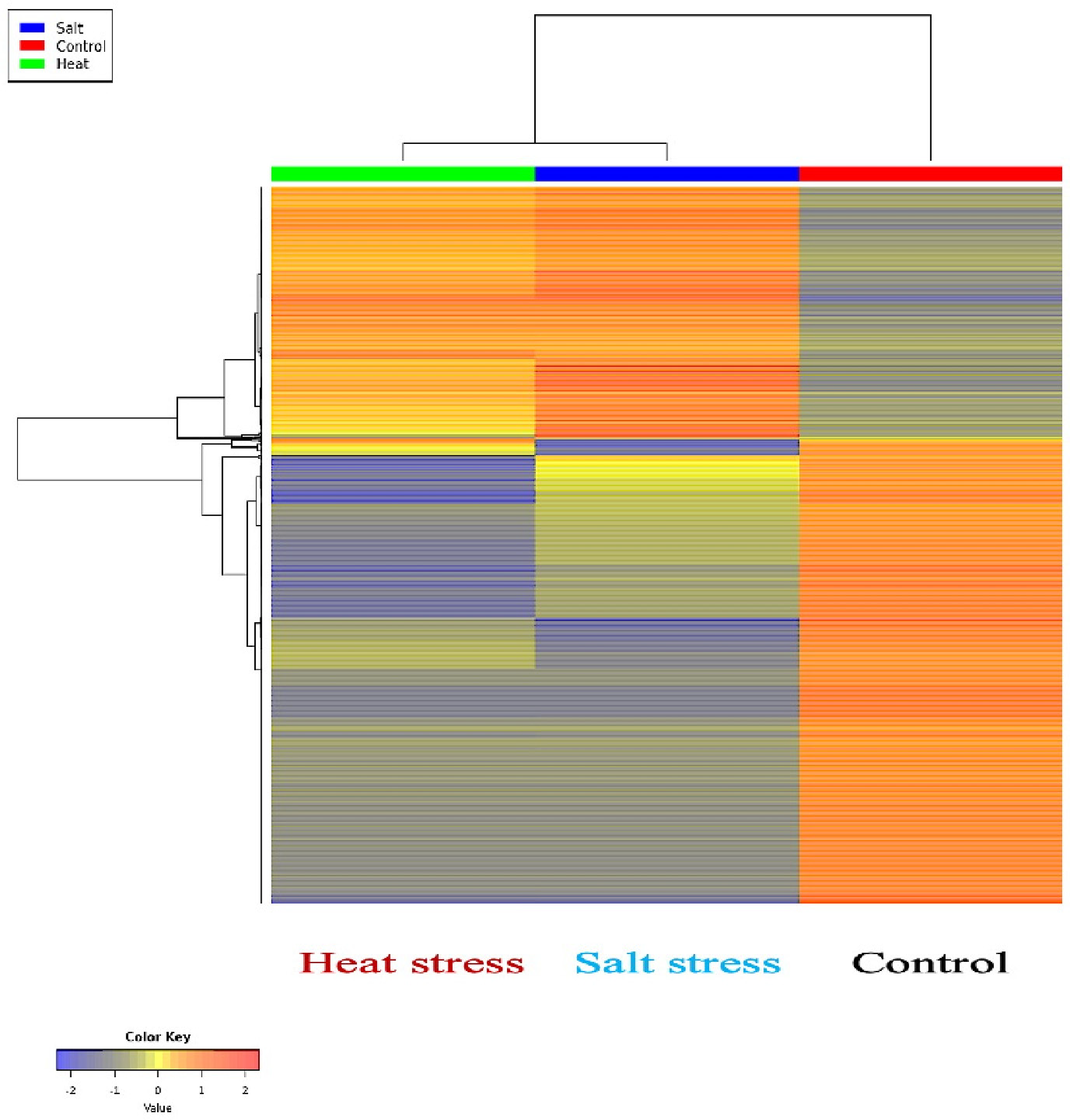
A heat map visually represents the expression levels of multiple genes in various samples of heat-stressed and salt-stressed pearl millet shoots compared with the control group. The map uses color gradients, with red indicating upregulated genes and green representing downregulated genes, to highlight the gene expression patterns within each sample.
Additionally, Principal Component Analysis (PCA) offers a comprehensive representation of DEGs in pearl millet shoots under both control and salt-stressed conditions. Figure 5, the PCA plot, provides a visual insight into the clustering of samples based on their gene expression profiles. This representation aids in understanding how stress influences the overall transcriptomic landscape by illustrating the grouping of samples with similar expression patterns. The PCA plot serves as a valuable analytical tool for discerning the impact of stress on the global gene expression dynamics of pearl millet shoots, offering a nuanced perspective on the transcriptomic responses under different conditions.

Principal component analysis (PCA) representation of pearl millet shoots with DEGs under both control and salt-stressed conditions.
Utilizing R scripts, 18 various expression plots were generated to elucidate the intricate details of gene expression patterns. The MA plot, featured in Figure 6, provides a visual representation of differences in measurements between two samples by transforming data into M (log ratio) and A (mean average) scales. Subfigures a1 and a2 of Figure 6 display the M scale (log ratio) for heat stress and salt stress DEGs, respectively, offering insights into the variation in read counts between samples. The M scale represents the log ratio of measurements between the two samples compared with the control, providing valuable information about the magnitude of changes in gene expression under abiotic stress.
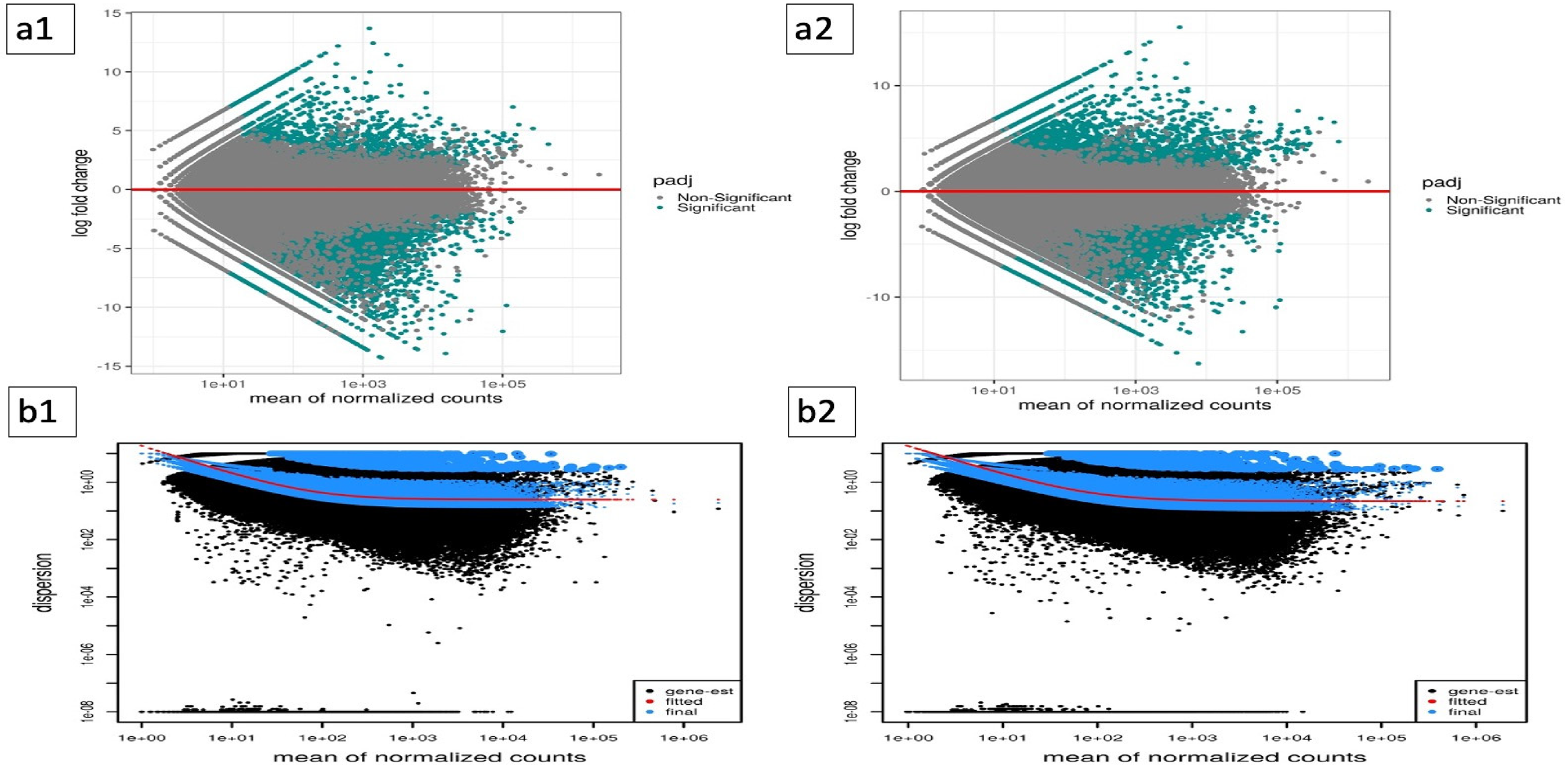
M scale (log ratio), which represents the log ratio of measurements between the two samples compared with the control. (a1) and (a2) display heat stress and salt stress DEGs, respectively, calculated as the logarithm (base 2) of the fold change under abiotic stress (heat or salt). The dispersion plot shows the mean of normalized read counts using the DESeq2 tool for (b1) heat stress DEGs and (b2) salt stress DEGs.
The volcano plot, showcased in Figure 7, is a 2-D scatter plot that visualizes the significance and magnitude of changes in features, such as genes or transcripts, within a specific comparison. Figure 7 consists of Volcano Plots representing different categories of genes based on their fold change and statistical significance. The X-axis represents the fold change between the two groups on a log2 scale, while the Y-axis shows the negative log10 of the p-values from the statistical test of the comparison. Colors are used to distinguish non-DEGs (blue), upregulated genes (red), and downregulated genes (green). This representation aids in identifying significant changes in gene expression between the groups.

The volcano plot is a specialized 2-D scatterplot designed to visualize the significance and magnitude of changes in transcript features within a specific comparison: (a) heat versus control or (a2) salt stress versus control. The x-axis represents the fold change between the two groups and is displayed on a log2 scale. Meanwhile, the y-axis shows the negative log10 of the p-values obtained from the statistical comparison. The colors used in the plot convey different categories: blue represents non-DEGs (abs(log2FC) < 2 and padj > 0.05), red denotes upregulated genes with log2FC ≥ 2 and padj ≤ 0.05, and green indicates downregulated genes with log2FC ≤ −2 and pad ≤ 0.05. (a1) and (a2) display the DEGs under heat stress and salt stress, respectively. The p-value distribution plot is provided for (b1) heat stress DEGs and (b2) salt stress DEGs.
The distribution of normalized counts across samples is effectively portrayed by a violin plot, as seen in Figure 8. Violin plots showcase the expression patterns for heat stress and salt stress DEGs. Figure 8 offers valuable information on the variation in gene expression within each group. Notably, salt stress expression exhibited a one-fold increase compared to heat stress in pearl millet shoots. Additionally, a heatmap generated using million electron-volts software is presented in Figure 8. The heatmap illustrates the expression profiles of the top 30 genes based on the lowest p-value (considered as DEGs). Colors in the heatmap represent downregulated genes in green, normal genes in black, and upregulated genes in red. Thermal stress showed a higher number of upregulated genes compared to salt stress and the control, further delineating the distinct gene expression patterns under different stress conditions.

Violin plot illustrates the distribution of normalized counts across the samples for (a1) heat stress DEGs and (a2) salt stress DEGs. The HeatMap showed a graphical representation where the plot was generated for the top 30 genes, based on the lowest p-value (at abs(log2FC) ≥ 2 and p-value ≤ 0.05 for DEGs), using million electron-volts software. In this representation, green, black, and red represent downregulated, normal, and upregulated genes, respectively. HeatMaps for pearl millet shoots DEGs of (b1) heat stressed and (b2) salt stressed.
The outcomes of sequence-based KEGG annotation are presented in the ensuing section, showcasing the top 10 KEGG distribution plots for DEGs under both heat stress and salt stress conditions. As depicted in Figure 9, this analysis illuminates the specific KEGG pathways that undergo enrichment in response to abiotic stress.

Results of sequence-based KEGG annotation, presenting the top 10 KEGG distribution plots for both (a1) heat stress DEGs and (a2) salt stress DEGs. Top 30 biological process enrichment plots based on KEGG and GO enrichment analysis for pearl millet shoots: (b1) heat stress DEGs and (b2) salt stress DEGs.
Figure 9 further incorporates the top 30 biological process enrichment plots, derived from both KEGG and GO enrichment analyses, specifically tailored for pearl millet shoots under the influence of heat and salt stress conditions. These plots not only elucidate the biological processes enriched with DEGs but also provide a comprehensive understanding of the molecular mechanisms steering the plant's response to stress.
This thorough analysis yields valuable insights into the intricate molecular responses of pearl millet shoots when subjected to salt stress, unraveling key pathways, discernible gene expression patterns, and potential targets for further exploration of stress tolerance and adaptation mechanisms. The collective findings contribute significantly to our comprehension of the underlying molecular dynamics that govern the plant's resilience and adaptation strategies in the face of environmental challenges.
Metabolic pathway diagrams depicting pearl millet shoots’ response to abiotic stress, specifically heat and salinity, were generated using the iPath online tool (https://pathways.embl.de/ipath3.cgi?map = metabolic#, accessed on 30 July 2023).19,20 Both downregulated and upregulated DEGs were annotated using KEGG reference numbers. The visualization of metabolic pathways specifically emphasized common downregulated genes, showcasing the KEGG reference numbers associated with 20 metabolites (Figure 10). Additionally, metabolic pathways linked to KEGG reference numbers of 17 metabolites highlighted common upregulated genes (Figure 11) in pearl millet under the mentioned abiotic stress conditions.

The metabolic pathways associated with the KEGG reference numbers of 20 metabolites, which are commonly downregulated, and DEGs in pearl millet leaves under abiotic stress conditions of heat (50°C for 60 s) and salt (150 mM NaCl), are examined in comparison with the control.

The metabolic pathways associated with KEGG reference numbers of 17 metabolites that are commonly upregulated and DEGs in pearl millet leaves under abiotic stress conditions of heat (50°C for 60 s) and salt (150 mM NaCl) are examined in comparison with the control.
Discussion
Heat and salt stress impact on gene expression and adaptive mechanisms in pearl millet
The repercussions of environmental stresses, specifically heat and salt stress, present a formidable challenge to the growth and development of plants. To enhance plant tolerance and productivity in adverse conditions, a thorough comprehension of the molecular mechanisms governing plant responses to these stresses is imperative. A study by Ouertani et al. 21 noted a significant increase in the number of differentially expressed kinase genes in both leaves and roots after 8 h of salt exposure, underscoring the potential involvement of the kinase signaling pathway in mediating the plant's response to salt stress.
The convergence of heat and salt stress introduces additional complications such as drought and osmotic stress, compromising vital processes including photosynthesis, protein synthesis, energy metabolism, and lipid metabolism. 22 DEG M-Scale analysis reveals distinctive molecular responses in plants to heat stress and salt stress, offering valuable insights into the variation in read counts between samples and facilitating a comprehensive understanding of specific genes and pathways that are activated or suppressed during stress (see Figures 1–11).
Pearl millet, akin to numerous plant species, exhibits intricate molecular responses to environmental stresses, as unveiled by gene expression analyses under heat and salt stress. These findings underscore pearl millet's sophisticated adaptive mechanisms to navigate adverse conditions. However, it is imperative to consider alternative perspectives, recognizing that extensive molecular responses may not consistently correlate with improved tolerance or successful adaptation. 23 Despite the insights gained from differential expression analyses, the activation of certain pathways and upregulation of specific genes do not guarantee enhanced plant survival or productivity under stress conditions 24 Evaluating the physiological and biochemical consequences of these responses is essential to discern whether they contribute to effective stress tolerance in pearl millet and other plant species. 25 Our results indicate a high number of downregulated genes under abiotic stress, either heat or salt; however, these complex molecular responses may signify stress-induced damage and disruption of normal cellular processes rather than efficient adaptive mechanisms. A critical examination of the functional implications of these responses is necessary to assess their effectiveness in promoting plant adaptation and survival under adverse environmental conditions. 26 In the current study, cascading events triggered by heat stress and salt stress in pearl millet, resulting in changes in gene expression (see Figures 1–9) and subsequent biochemical and physiological changes (see Figures 10–11), underscore the ability of pearl millet to perceive and respond to environmental signals. In addition to gene expression, these stresses pose challenges to fundamental processes, compromising the plant's regulatory capacity over photosynthesis, protein synthesis, and energy and lipid metabolism. 27 The unique molecular responses observed in pearl millet emphasize the importance of understanding the underlying mechanisms. Unraveling gene expression patterns and physiological responses represents a crucial step toward developing strategies to enhance plant tolerance and productivity in the face of challenging environmental conditions. 28
Cross tolerance in pearl millet: shared abiotic KEGG pathways for salt and heat stress responses
Pearl millet (
The study results (Figures 9–11) reveal four common abiotic KEGG pathways that play vital roles in pearl millet's response to salt and heat stress: phenylpropanoid biosynthesis, photosynthesis (antenna proteins), photosynthesis, and plant hormone signal transduction. These metabolic pathways are believed to be essential in the mechanism of cross-tolerance. Cross tolerance is an adaptive strategy observed in plants, whereby those exhibiting tolerance to one type of abiotic stress also demonstrate enhanced resilience to other stressors. This phenomenon underscores the interconnectedness of stress response mechanisms and improves plants’ ability to thrive in harsh environments.30,31
Phenylpropanoid biosynthesis is stimulated in pearl millet shoots in response to abiotic stress (heat and salt). Phenylpropanoid biosynthesis is a fundamental pathway in plants that is responsible for the production of phenolic compounds. These compounds act as secondary metabolites with diverse roles, including defense against environmental stressors. In pearl millet, the activation of this pathway under salt and heat stress may lead to the synthesis of various phenolic compounds, such as flavonoids, lignin, and phenolic acids.
Abiotic stress leads to the production of reactive oxygen species (ROS) such as superoxide anions, hydrogen peroxide, and hydroxyl ions. 32 To counteract the propagation of these harmful ROS, plants activate a well-orchestrated and finely-tuned antioxidant system. 33 Phenolic compounds play a crucial role in this process by possessing powerful antioxidant properties and assisting in the scavenging of harmful ROS in abiotic plants. In addition, the phenylpropanoid biosynthetic pathway is stimulated, leading to the production of various phenolic compounds with strong antioxidative potential.34,35 The accumulation of phenolic compounds helps protect pearl millet plants from oxidative stress, thereby enhancing their ability to withstand salt and heat stress.
Our study showed that in response to heat and salt stress, pearl millet shoots expressed genes related to photosynthesis antenna proteins and photosynthesis. Photosynthesis is a crucial process in plants that converts light energy into chemical energy, enabling them to produce carbohydrates and sustain growth. However, under salt and heat stress, photosynthesis can be severely impaired. The photosynthesis–antenna proteins pathway plays a key role in optimizing the efficiency of photosynthetic light harvesting. This allows the plant to balance energy absorption from light and prevents excess energy from causing photodamage. By maintaining a balanced antenna protein system, pearl millet can continue to perform photosynthesis, albeit at a reduced rate, under stressful conditions.
On the other hand, the upregulation of genes related to photosynthesis, specifically those involved in CO2 concentration mechanisms, improves water use efficiency, as the closure of stomata (tiny pores on leaf surfaces) during stress reduces CO2 availability. By enhancing photosynthetic efficiency, pearl millet minimizes carbon loss and maintains a stable supply of carbohydrates for growth and stress tolerance. Similarly, under abiotic stress, pearl millet showed upregulation of plant hormone signal transduction. The signal transduction pathway of plant hormones plays a vital role in the transmission of stress signals and the initiation of appropriate responses. Pearl millet modulates hormone signaling pathways, specifically abscisic acid (ABA) and ethylene, in response to salt and heat stress. The application of ABA aids in the regulation of water balance and the closure of stomata, thus mitigating water loss in the presence of salt and heat stress. In contrast, ethylene plays a crucial role in modulating stress responses and affecting development in unfavorable circumstances.36,37
Pearl millet uses various abiotic KEGG processes, such as phenylpropanoid biosynthesis, Photosynthesis–antenna proteins, Photosynthesis, and plant hormone signal transduction, to augment its resilience to salt and heat-induced stress. The comprehension of these pathways and their functions in stress tolerance can yield significant insights for the development of cross-tolerant breeding strategies aimed at producing resilient pearl millet varieties, thus providing food security in demanding ecological conditions.
Pearl millet heat stress-specific KEGG pathways
Heat stress is a critical factor that profoundly impacts crops and agriculture, and its effects have been exacerbated by climate change and global warming. As global temperatures rise, heatwaves become more frequent and intense, significantly affecting crop growth and productivity. When temperatures surpass the optimal range for crop development, heat stress negatively affects plant physiology and metabolism, leading to reduced photosynthesis, impaired growth, and ultimately lower crop yields. 38 To address the adverse effects of heat stress and climate change on crops, various strategies can be implemented, including breeding and genetic improvement. The development of heat-tolerant crop varieties through selective breeding and genetic engineering can aid in better adaptation to changing climate conditions, thereby ensuring a more resilient agricultural system. 39 In our study, we focused on understanding the influence of heat stress on transcriptomic pathways in pearl millet to foster greater resilience in agriculture. Our analysis revealed specific upregulated KEGG pathways in pearl millet shoots under heat stress, including cytochrome P450, cutin, suberine, and wax biosynthesis, zeatin biosynthesis, and biosynthesis of various plant secondary metabolites, such as crocin biosynthesis, ginsenoside biosynthesis, and saponin. These findings shed light on the molecular responses of pearl millet to heat stress and provide valuable insights for enhancing its heat tolerance and overall performance under changing environmental conditions.
In our study, we observed the association of cytochrome P450 with pearl millet shoots exposed to heat stress, indicating its crucial role in promoting survival under such conditions. The enzyme family Cytochrome P450 s (CYPs) is widely known as the largest group of enzymes responsible for facilitating hydroxylation processes that depend on NADPH and/or O2. These reactions occur across various life domains. In both plants and mammals, cytochrome P450 enzymes (CYPs) are crucial for xenobiotic detoxification. 40 In addition, CYPs exhibit multifunctional properties and play a pivotal role in the synthesis of secondary metabolites, antioxidants, and phytohormones in higher plant organisms. Although the molecular and biochemical processes mediated by CYPs have been extensively studied and documented, the specific correlation between their biochemical processes and their impact on various plant functions is not fully understood.
The advent of NGS has opened up new opportunities to investigate the role of cytochrome P450 enzymes (CYPs) in various plant processes, including responses to environmental stress. Many CYP genes are regulated in response to environmental challenges and are also significantly involved in the interplay between abiotic and biotic stress responses. Considering this, cytochrome P450 enzymes (CYPs) have great potential as promising candidates for genetic modification of agricultural species to enhance their resilience against both biotic and abiotic challenges. 41
Furthermore, transcriptomic analysis revealed that during heat stress, pearl millet activates the cutin, suberine, and wax biosynthesis pathways to enhance the production of these protective compounds. The cuticle, a waxy layer covering the aerial surfaces of plants, plays a crucial role in limiting nonstomatal water loss and providing protection against environmental stresses.
The increased synthesis of cutin, suberine, and wax helps to reduce water loss from the plant surface, thereby improving its water-use efficiency. In addition, these compounds form a physical barrier that shields the plant from external stresses, including heat and drought. Notably, cuticular wax plays a significant role in enhancing plant tolerance to various abiotic and biotic stressors. It serves as a defense mechanism against excessive UV radiation, high temperature, bacterial and fungal diseases, insects, high salt, and low temperature.42,43 Drought, a major form of abiotic stress, poses substantial challenges to global food security and terrestrial ecosystems’ overall health because it adversely affects plant development and crop yield. However, there remains a lack of understanding regarding the molecular and evolutionary mechanisms governing the response of plants to drought stress through cuticular wax. Utilization of cuticular wax to enhance water usage efficiency in agricultural practices holds promise for promoting environmental sustainability. By harnessing the protective properties of cutin, suberine, and wax, we can develop more resilient crop varieties and improve agricultural practices despite climate change and increasing environmental challenges.
Pearl millet shoots under heat stress also showed the expression of zeatin, which is classified as a cytokinin and belongs to a group of plant hormones responsible for regulating various physiological activities, including growth and responses to stress. Zeatin's primary role is to promote cellular division in nonmeristematic tissues. It is produced within the plastids and then transported through the xylem. When faced with heat stress, pearl millet activates zeatin biosynthesis to support growth and development under adverse environmental conditions. Zeatin acts as a crucial plant hormone that promotes cell division and elongation, facilitating the proper development of plant tissues, even in the presence of heat stress. By sustaining growth, pearl millet enhances its overall health and strengthens its ability to withstand the detrimental impacts of high temperatures. 44
Furthermore, our work reveals that pearl millet, when subjected to heat stress, triggers the production of several plant secondary metabolites, such as crocin, ginsenoside, and saponin. The secondary metabolites exhibit several functionalities, including antioxidant defense, allelopathy (the suppression of nearby plant development), and defense against herbivores and infections. Pearl millet exhibits an enhanced capacity to tolerate stress and withstand heat damage through the production of these metabolites.45–47
Heat stress triggers specific KEGG pathways in pearl millet, allowing the plant to adapt and survive under challenging conditions. The activation of pathways related to drug metabolism, xenobiotic metabolism, cuticle biosynthesis, zeatin production, and various plant secondary metabolites collectively contribute to the enhancement of heat stress tolerance in pearl millet. Understanding these pathways provides valuable insights into developing climate-resilient crop varieties and ensuring food security in a changing climate.
Pearl millet salinity stress-specific KEGG pathways
Salinity stress is a major environmental factor that negatively impacts the growth and productivity of crops, including pearl millet (
Salinity stress in pearl millet shoots was linked to specific metabolic pathways, such as carbon fixation during photosynthesis. The phenomenon of salinity stress can interfere with the intricate process of photosynthesis, thus diminishing the accessibility of energy and carbon necessary for the optimal growth and development of plants. To surmount this obstacle, pearl millet employs a mechanism where it modulates its carbon fixing in photosynthetic organisms to maximize the assimilation of carbon dioxide (CO2). In the presence of salinity stress, pearl millet may adopt diverse mechanisms to cope with adverse conditions. One such strategy involves the upregulation of carbonic anhydrase activity, which facilitates the conversion of bicarbonate (HCO3-) into carbon dioxide (CO2). 48 This mechanism aids in the preservation of photosynthetic efficiency, even despite diminished CO2 availability resulting from the closing of the stomata as a reaction to salinity-induced stress. Pearl millet effectively preserves water and sustains carbohydrate production through the process of fine-tuning carbon fixation, thus enabling its survival in saltwater environments. Similarly, an experiment was conducted whereby rice plants were exposed to three levels of drought stress (−10, −25, and −40 kPa) for a duration of 15 days. The results of this study indicate that the imposition of short-term drought stress has a positive impact on the process of photosynthetic carbon fixation. The activities of sucrose synthase and sucrose phosphate synthase were strongly impacted by abiotic stress. The activity of vacuoles invertase exhibited a notable rise, with a positive correlation between the severity of drought and the level of vacuoles invertase activity. 49
A significant alteration was identified in the expression of RNA associated with the MMR pathway in pearl millet shoots. DNA repair pathways in plants affect plant growth and development. The aforementioned pathway is responsible for the correction of mistakes that arise throughout the processes of DNA replication and recombination. Under conditions of salinity stress, pearl millet may undergo DNA damage because of the buildup of reactive oxygen species (ROS) and other variables associated with stress. To preserve the integrity of its genome and inhibit the transmission of genetic changes, pearl millet triggers the activation of the MMR pathway. This method facilitates the detection and correction of discrepancies or minor insertions and deletions within the DNA, thus safeguarding the integrity and functionality of the plant's genetic material, even in the presence of salinity-induced stress. The process of DNA MMR is constant throughout evolutionary history and serves a critical function in preserving the stability of the genome. The initiation of MMR occurs when MutS proteins identify and rectify single base–base mismatches, short loops consisting of unpaired nucleotides, and specific forms of DNA damage. Arabidopsis thaliana, a plant species, possesses MutS protein homologs (MSH) that resemble those observed in other eukaryotic organisms. In addition, Arabidopsis thaliana exhibits a distinct MSH7 polypeptide. The findings from experiments conducted on salt-treated msh7 seedlings indicated a deficiency in G2/M arrest. Consequently, msh7 seedlings demonstrated diminished susceptibility to salt-induced stress, as indicated by a decline in rosette and leaf sizes, stomatal density, overall leaf count, silique length, and seed count per silique. The results of this study indicate that inhibition of MSH7 activity may be a potential strategy for improving plant adaptation responses to salinity stress.49–51
Nitrogen metabolism refers to the biochemical reactions and pathways involved in the utilization, transformation, and elimination of nitrogen compounds in living organisms. The impact of salinity stress on nutrient absorption and use, particularly with regard to nitrogen, which plays a critical role in numerous physiological functions inside plants, is evident. To adapt to the constraints of nitrogen scarcity in saline environments, pearl millet employs regulatory mechanisms to modulate its nitrogen metabolic pathway. In the present investigation, we observed that under conditions of salt stress, pearl millet exhibits the potential to modulate the expression of genes associated with the processes of nitrogen absorption and metabolism. This allows the plant to optimize nitrogen utilization efficiency, ensuring that nitrogen is allocated to critical processes, such as amino acid synthesis and protein production, which are essential for stress adaptation and growth. Consistent with a prior investigation, it has been observed that nitrogen metabolism is linked to the resilience of photosynthesis in rice under water stress conditions, primarily by influencing CO2 diffusion, antioxidant capacity, and osmotic adjustment. 51 Exposure to salinity stress triggers distinct KEGG pathways in pearl millet, facilitating the plant's ability to acclimate and endure in saline habitats. The enhancement of salinity stress tolerance in pearl millet is achieved by activating processes associated with phenylpropanoid production, carbon fixation in photosynthetic organisms, MMR, and nitrogen metabolism. Comprehending these pathways can facilitate the advancement of crop types that are tolerant to salt, thereby guaranteeing the sustainability of agriculture in locations impacted by salinity-induced stress.
Study limitations
The study has some limitations. For instance, the relatively short durations of stress exposure—three days for salt stress and 60 s for heat stress—may not fully capture extended stress responses, potentially resulting in a limited understanding of molecular dynamics. Additionally, the transcriptomic analysis predominantly focused solely on shoots, overlooking potential variations in gene expression across other vital plant tissues. In addition, the reliance on a single timepoint for RNA sequencing analysis, specifically with 13-day-old seedlings, may overlook dynamic changes in gene expression during different developmental stages or timepoints under stress conditions. Furthermore, the experiments conducted in controlled laboratory conditions may not entirely replicate the complex and variable conditions of the field, highlighting the necessity for field studies to offer a more realistic assessment of pearl millet's response to abiotic stress in its natural environment. Lastly, while transcriptomics provides valuable insights into gene expression, the lack of direct reflection on protein abundance or post-translational modifications underscores the importance of integrating transcriptomic data with proteomic analyses for a more comprehensive understanding of the molecular mechanisms involved in stress tolerance.
Conclusion
Pearl millet exhibits cross-tolerance to both salt and heat stress, as evidenced by the activation of shared abiotic KEGG pathways. Phenylpropanoid biosynthesis, photosynthesis–antenna proteins, photosynthesis, and plant hormone signal transduction are crucial pathways involved in the plant's response to these stressors, enhancing its resilience in challenging environmental conditions. The production of phenolic compounds via the phenylpropanoid biosynthesis pathway aids in antioxidant defense and protection against stress-related damage. The fine-tuning of photosynthesis through the antenna protein pathway allows pearl millet to maintain energy absorption and minimize photodamage under heat and salt stress. Plant hormone signal transduction, particularly the regulation of ABA and ethylene, aids in water balance and stress responses. Understanding these pathways can be valuable for developing climate-resilient pearl millet varieties. Moreover, under heat stress, pearl millet specifically activates pathways related to cytochrome P450, cutin, suberine, and wax biosynthesis, zeatin biosynthesis, and various plant secondary metabolites. These pathways enhance heat tolerance by promoting cell division, protecting against stressors, and providing antioxidant defense. In addition, pearl millet responds to salinity stress through specific KEGG pathways, including carbon fixation in photosynthetic organisms, MMR, and nitrogen metabolism. The modulation of carbon fixation facilitates efficient CO2 assimilation, even under conditions of limited CO2 availability due to closed stomata. The MMR pathway plays a vital role in maintaining genomic stability and protecting against DNA damage caused by salinity stress. Pearl millet also optimizes its nitrogen metabolism to allocate nitrogen to critical processes, thus supporting stress adaptation and growth. Overall, understanding these specific KEGG pathways in the response of pearl millet to heat and salinity stress provides valuable insights for breeding stress-tolerant varieties and ensuring food security in challenging ecological conditions. By harnessing plant cross-tolerance and specific stress-responsive pathways, we can develop sustainable agriculture practices and enhance resilience despite climate change and increasing environmental challenges.
Supplemental Material
sj-docx-1-sci-10.1177_00368504241237610 - Supplemental material for Abiotic stress tolerance in pearl millet: Unraveling molecular mechanisms via transcriptomics
Supplemental material, sj-docx-1-sci-10.1177_00368504241237610 for Abiotic stress tolerance in pearl millet: Unraveling molecular mechanisms via transcriptomics by Faten Dhawi in Science Progress
Footnotes
Acknowledgments
I would like to express my thanks to Asma Abdulmohsen Aljogaiman for her assistance in producing the images. Her support and contributions have been invaluable, playing a crucial role in advancing and ensuring the success of this research project.
Authors’ contributions
The author took full responsibility for conceptualizing the manuscript, conducting data analysis, providing explanations, writing, and overseeing the final production.
Data availability
The RNA sequences of the pearl millet shoots in the current study for all samples were submitted to the National Center for Biotechnology Information (NCBI) database under the Sequence Read Archive (SRA) data, with the accession number PRJNA991076.
Declaration of conflicting interests
The author declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author received no financial support for the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
Author biography
Faten Dhawi, also known as Faten Dhawi Almuhanna, is an Associate Professor at King Faisal University, an accomplished author, and a biotechnology scientist with a passion for understanding the molecular and metabolic responses of plants to abiotic stress. Her research aligns with climate change mitigation efforts as part of sustainable development goals at the molecular and genetic levels. Additionally, she has conducted research on plant-microbial interactions and phytoremediation in polluted sites. She holds a PhD in Biochemistry and Molecular Biology from Michigan Technological University, MI, USA.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
