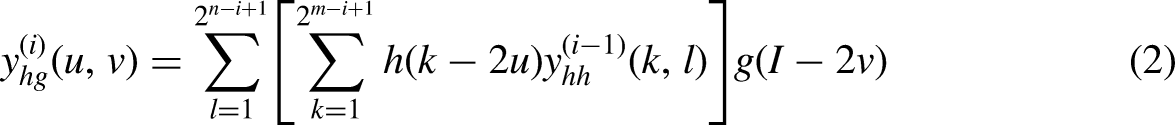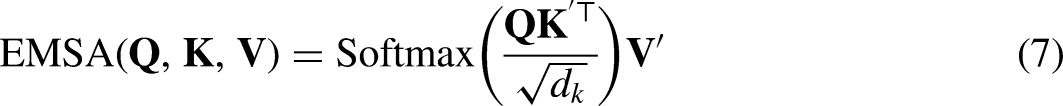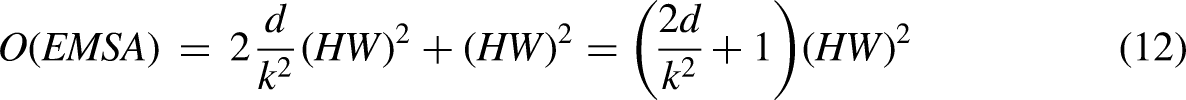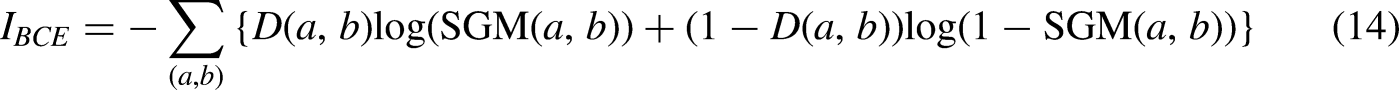Abstract
Nasopharyngeal carcinoma is a malignant tumor that occurs in the epithelium and mucosal glands of the nasopharynx, and its pathological type is mostly poorly differentiated squamous cell carcinoma. Since the nasopharynx is located deep in the head and neck, early diagnosis and timely treatment are critical to patient survival. However, nasopharyngeal carcinoma tumors are small in size and vary widely in shape, and it is also a challenge for experienced doctors to delineate tumor contours. In addition, due to the special location of nasopharyngeal carcinoma, complex treatments such as radiotherapy or surgical resection are often required, so accurate pathological diagnosis is also very important for the selection of treatment options. However, the current deep learning segmentation model faces the problems of inaccurate segmentation and unstable segmentation process, which are mainly limited by the accuracy of data sets, fuzzy boundaries, and complex lines. In order to solve these two challenges, this article proposes a hybrid model WET-UNet based on the UNet network as a powerful alternative for nasopharyngeal cancer image segmentation. On the one hand, wavelet transform is integrated into UNet to enhance the lesion boundary information by using low-frequency components to adjust the encoder at low frequencies and optimize the subsequent computational process of the Transformer to improve the accuracy and robustness of image segmentation. On the other hand, the attention mechanism retains the most valuable pixels in the image for us, captures the remote dependencies, and enables the network to learn more representative features to improve the recognition ability of the model. Comparative experiments show that our network structure outperforms other models for nasopharyngeal cancer image segmentation, and we demonstrate the effectiveness of adding two modules to help tumor segmentation. The total data set of this article is 5000, and the ratio of training and verification is 8:2. In the experiment, accuracy = 85.2% and precision = 84.9% can show that our proposed model has good performance in nasopharyngeal cancer image segmentation.
Keywords
Introduction
Nasopharyngeal carcinoma (NPC), a malignant tumor of the head and neck closely related to Epstein-Barr virus infection, 1 occurs in epithelial cells covering the surface of the nasopharynx and the nasopharyngeal line and mainly presents features such as greyish tissue visible to the naked eye, lamellar arrangement of heterogeneous cells visible by light microscopy, and a pyknotic nucleus, which has shown a yearly increase in the development of the era and is a serious threat to human health. Early detection of NPC is extremely difficult because the early symptoms of NPC are similar to those of daily diseases, and only show symptoms such as nasal congestion, tinnitus, cervical lymph node enlargement, or headache. Studies have shown that the five-year survival rate of early-stage NPC can reach more than 80%, while the mid- to late-stage is below 50%, 2 making the study of early diagnosis of NPC extremely important. 3 The diagnosis of NPC relies on the analysis of medical images for identification, but the need for experienced physicians to manually analyze the images has resulted in patients in medically disadvantaged areas being vulnerable to delays. To solve this problem, a fully automated lesion segmentation method is needed to reduce the workload of physicians and delineate the region of interest (ROI) quickly and accurately.
Up to now, a series of methods have been proposed to accurately segment ROI. Before 2000, they were mainly based on traditional machine learning methods, using means threshold segmentation, region segmentation, edge segmentation, texture features, clustering, etc. for analysis. With the emergence of convolutional neural networks (CNNs), deep learning frameworks began to be widely used. such as the fully convolutional networks model,
4
UNet mode,
5
DeepLab v1&v2
6
,
7
RefineNet,
8
and other model frameworks have shown excellent image processing capabilities in the semantic segmentation process. These methods achieve good segmentation results by weighted training using a large number of T1, T2 and contrast-enhanced T1c shared attributes. However, there are variable lesion regions, different sizes, and irregular boundaries in NPC, so we propose a dual UNet-based segmentation model. First, the input image is trained using the ResNet-50
9
model to initially complete the segmentation task, after which the wavelet transform (WT) is used to extract the low-frequency components of this image and stitch them together in UNet parts, which in turn improves the robustness of the model, after which the transformer is introduced to improve the ROI recognition accuracy. The contributions of this article are as follows:
We segmented the NPC data in the T1 and T2 periods to provide clinicians with sufficient diagnostic reference and improve the reliability of disease analysis. Integrating WT into the image segmentation model, the robustness of the model segmentation is greatly improved by low-frequency adjustment of each part of the improved UNet encoder. Adding a transformer to the end of the encoder of the improved UNet, modeling the input image using the self-attentive mechanism, analyzing the spatial relationship between each pixel, and establishing deep feature correlation, thus improving the training performance of the model and achieving high-precision segmentation of ROI.
The rest of this article is organized as follows: The related work is reviewed in the “Related works” section. The “Methods” section describes in detail the general architecture of the system and the methodological modules. The “Experiments and results” section discusses the experimental setup and the analysis of the results. The “Conclusion” section summarizes our conclusions and future work.
Related works
Traditional image segmentation
Image segmentation, a classical problem in the field of image technology, has attracted many researchers’ enthusiasm and great efforts since the 1970s, and many image segmentation algorithms have been proposed. However, until 2000, traditional image segmentation methods such as thresholding, region, and boundary detection were mostly used. Hao et al. 10 proposed a new local label learning strategy that uses statistical machine-learning techniques to estimate segmentation labels of target images. In particular, we use L1 regularized support vector machine (SVM) and k-nearest neighbor (kNN)-based training sample selection strategy to learn classifiers for each target image voxel from adjacent voxels in the atlas based on image intensity and texture features. Peña et al. 11 combined object-based image analysis and advanced machine learning methods to improve crop identification, evaluating decision trees, logistic regression (LR), SVM, and multilayer perceptron (MLP), and neural network methods to map nine major summer crops from ASTER satellite images captured on two different dates (Arganda-Carreras et al., 2016). 12 We introduced trainable Weka segmentation, which is customized to use user-designed image features or classifiers by providing an unsupervised segmentation learning scheme (clustering) that uses a limited number of manual annotations to train the classifier and automatically segments the remaining data. Prabaharan et al. 13 proposed an adaptive alpha value Havrda–Chavrat entropy method, which segmented the medical microscopic sperm image after Wiener noise removal. A threshold-based segmentation method is implemented and results are obtained on the input image. Khaled et al. 14 proposed a brain image segmentation model based on boundary detection. The boundary segmentation network and boundary information module used to detect and segment the brain tissue of the image were used to distinguish three different boundaries, and a boundary attention gate was added at the encoder output layer to capture more local details. Liu et al. 15 proposed a preprocessing method based on empirical mode decomposition and bilateral filtering. Then, a new clustering method based on gray correlation was extracted from the lung region. Finally, a new lung contour correction technique was adopted to repair the depressed areas caused by pulmonary nodules and blood vessels.
Most of the traditional methods need to rely on manually selecting rules for segmenting images, which makes them unable to deal with complex images, while traditional segmentation methods are sensitive to noise in the image, which may lead to erroneous segmentation or unstable results. Taken together, traditional methods have a series of limitations when dealing with complex medical images and unused application scenarios, including dependence on feature design and lack of adaptivity. These shortcomings are somewhat overcome in modern deep learning methods.
Deep learning for image segmentation
In recent years, due to the success of deep learning models in vision applications, there has been a significant amount of work dedicated to developing image segmentation methods using deep learning models. Cardenas et al. 16 reviewed conventional (non-deep learning) algorithms that are particularly relevant to radiotherapy applications and described in detail the process of pathology image segmentation using deep learning algorithms. Chen et al. 17 aim to solve the limitation that the traditional active contour model can’t fully consider the area and boundary size inside and outside the region of interest in the learning process by developing a new model based on deep learning. The model considers the area and boundary size inside and outside the region of interest in the learning process. Dalca et al. 18 proposed an alternative strategy that combines traditional probabilistic mapping-based segmentation with deep learning, enabling one to train segmentation models for new magnetic resonance imaging (MRI) scans without any manual segmentation of the images. Ren et al. 19 proposed an improved deep fully CNN, called CrackSegNet, for performing dense pixel-level crack segmentation. Chen et al. 20 used deep learning to review more than 100 cardiac image segmentation papers covering common imaging modalities including MRI, computed tomography (CT), and ultrasound as well as the major anatomical structures of interest (ventricles, atria, and vessels). Zhou et al. 21 outlined a deep learning-based approach for multimodal medical image segmentation tasks.
Deep learning methods perform well in spatial modeling and are more suitable when considering local feature tasks, but require large amounts of labeled data for training. This can be a challenge for the field of medical image segmentation, as labeling medical images usually requires expertise and time, and data may be limited. Moreover, deep learning models usually operate as black boxes and lack interpretation of segmentation results, and for the medical image domain, interpretability is important for clinical practice.
Transformer for image segmentation
Transformer is a sequence-to-sequence prediction framework that has a proven track record in machine translation and natural language processing due to its powerful spatial modeling capabilities. Xie et al. 22 proposed a hybrid framework that efficiently connects CNN and transformer (CoTr) for 3D medical image segmentation. CoTr has an encoder–decoder structure. In the encoder, a concise CNN structure is used to extract feature maps, and the transformer is used to capture long-range dependencies. A deformable self-attentive mechanism is introduced in the transformer to focus on only a small fraction of key sampling points, thus greatly reducing the computational and spatial complexity of the transformer. Chen et al. 23 designed TransUNet, a framework that establishes a self-attentive mechanism from a sequence-to-sequence prediction perspective, combining the advantages of transformer and UNet, which, on the one hand, encodes the labeled image blocks in the feature map of the CNN as the input sequence for extracting the global context; on the other hand, the decoder encodes the encoded features are upsampled and then combined with the high-resolution CNN feature map to achieve precise localization. Cao et al. 24 proposed Swin-Unet, which is a pure transformer similar to UNet for medical image segmentation. Lin et al. 25 proposed a novel framework for deep medical image segmentation called dual Swin transformer UNet (DS-TransUNet), which aims to integrate hierarchical Swin transformers into encoders and decoders of standard U-shaped architectures. Gao et al. 26 introduced UTNet, a simple yet powerful hybrid transformer architecture that integrates self-attention into CNNs to enhance medical image segmentation. Valanarasu et al. 27 proposed a gated axial attention model and a local-global training strategy (LoGo) for use in two-dimensional (2D) medical image segmentation to extend the existing architecture by introducing additional control mechanisms in the self-attention module and to train the model efficiently on medical images to further improve medical image segmentation accuracy. The current approach is unreliable and inefficient for heavy medical segmentation tasks, such as predicting a large number of tissue classes or modeling globally interconnected tissue structures. Inspired by the nested hierarchical structure in vision converters, Yu et al. 28 proposed a new three-dimensional (3D) medical image segmentation method (UNesT), which uses a simplified and faster-converging transformer encoder design to achieve local communication between spatially adjacent patch sequences by hierarchical aggregation. Deep learning-based segmentation models usually require large amounts of data, and the models usually have poor generalization due to a lack of training data and inefficient network structures. Tang et al. 29 proposed combining deformable models and medical transformer neural networks for image segmentation tasks to alleviate these problems.
Overall, transformer and deep learning models have their own strengths in different domains and tasks. Transformer's core strength is its excellent sequence modeling capability, which is useful for global contextual understanding in images due to the introduction of self-attention mechanisms that allow the model to better understand the relationships between different locations in the input data. However, transformer models usually contain a large number of parameters, which leads to high computational complexity. When dealing with large-scale images or large datasets, training, and inference may require a lot of computational resources and time. So we try to combine deep learning with transformer and introduce the WET-UNet model.
WT for image segmentation
WT is a transform analysis method that decomposes medical images with unclear features into two parts: a high-frequency module with high temporal resolution and low-frequency resolution, and a low-frequency module with high temporal resolution and low-frequency resolution. Therefore, the image contour is reasonably divided into the low-frequency module, and the high-frequency component of the image can be adjusted to obtain clearer contour information and improve the segmentation accuracy of the subsequent CNN. WT is an ideal tool for image quality enhancement and a traditional image processing method, while deep reinforcement learning combines the advantages of deep learning feature extraction and reinforcement learning strategy learning, so the image can be processed by using deep learning and WT. Guo et al. 30 proposed a deep CNN to predict the “missing details” of wavelet coefficients of low-resolution images, combining the complementary nature of wavelet domain information (divided into low-frequency and high-frequency subbands) with a deep CNN, which also provides structural information of the image through the sparsity of wavelets. Based on the wavelet prediction network, the sparsity boosting property of the residual network is used to fit the wavelet coefficients and further enhance them by residual inference, which contributes to stable training and robust convergence. Huang et al. 31 proposed a face recognition method for wavelets, using the HR wavelet coefficients corresponding to LR to complete the reconstruction of HR images while using a flexible and scalable CNN to obtain higher global perceptual field and local texture features to complete higher accuracy high-resolution face recognition. Liu et al. 32 proposed a multi-stage wavelet CNN (MWCNN) by improving the UNet network by replacing the downsampling and upsampling processes with DWT and IWT while combining convolutional layers to reduce the size of feature mapping and new avenue feature images in the shrinkage network, and then reconstructing the high-resolution feature map using wavelet inverse transform. Since the WT is reversible, it is guaranteed that there is no information loss after the WT. However, a large number of WT operations can over-decompose the feature mapping and lead to a large number of redundant channels, which may be detrimental to the gradient back-propagation during training.
Methods
In this article, we introduce a hybrid network that takes advantage of the WT and transformer. The general architecture of WET-UNet is shown in Figure 1. The network consists of a backbone network and two enhancement modules. Firstly, we feed the original nasopharyngeal cancer images into the backbone network after processing to extract feature maps at different levels; secondly, the feature maps are fed into the WT module and the encoding stage of UNet for low-frequency attribute adjustment to improve the training accuracy of the shallow network for extracting feature models, which in turn ensures the stability and robustness of high-precision ROI region delineation; the output of this stage is fed into the deep network, which uses the transformer encoding block (TEB) module to correlate all shallow features, which is better for capturing global information and long dependencies of the image with better generalization ability and helps to improve the accuracy of image segmentation; the model is used for image reconstruction with the decoded output of the UNet network to extract the lesion region and output. The details of the backbone network and the two modules are described in the following subsections.

The architecture of WETNet. WETNet consists of a backbone network and two information enhancement modules. The backbone network is the commonly used ResNet-50, which is initialized using pre-trained weights learned on the ImageNet dataset. WT and EMSA are, respectively, the wavelet transform module and efficient multi-head self-attention module introduced later.
Backbone network
In computer vision, pre-trained models are often used to speed up training and improve model performance, and the commonly used backbone is ResNet-50. ResNet-50 is a very deep neural network model with very powerful feature extraction capabilities, and we randomly initialize the other parameters of the model. Finally, we use our own nasopharyngeal cancer dataset to fine-tune the whole model by migration learning.
Wavelet transformer
In this article, we perform WT and splicing on the output results of ResNet-50, splice the low-frequency data into the CNN model based on multi-resolution analysis, and complete the fast training of the improved UNet model based on nasopharyngeal cancer data without considering the displacement sensitivity and phase information, and then solve such problems. In our experiments, we use the 2D Haar transform to perform low-pass and high-pass filtering from both the horizontal and vertical directions. Assuming that for a given image of size M × N = 2m × 2n, the decomposition output equation is shown in equations (1) to (4).
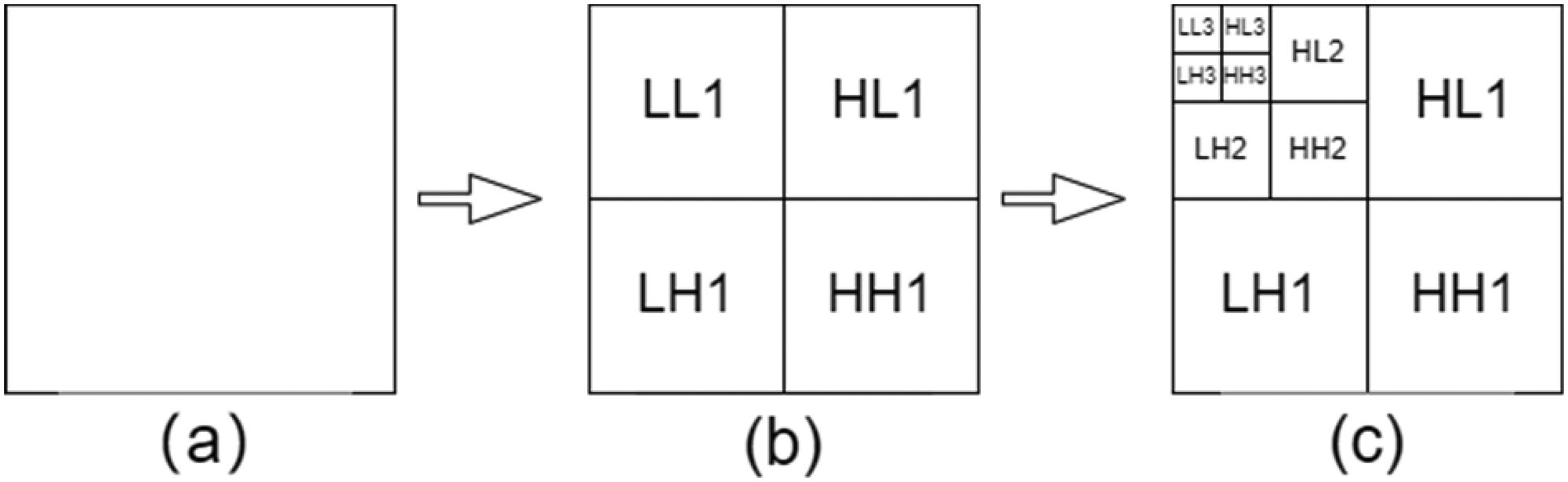
Results of the three-stage Haar wavelet transform, (a) for the original image, (b) for the first-stage wavelet transform, and (c) for the third-stage wavelet transform, where LL1 is the low-frequency information, HL1 is the horizontal high-frequency information, LH1 is the vertical high-frequency information, and HH1 is the diagonal high-frequency information.

Segmentation results of medical images after one to three Haar wavelet transforms.
The core idea of the wavelet-UNet improvement model is to connect the WT low-frequency components with the CNN layer. The four features LL, HL, LH, and HH were obtained by decomposing these wavelet features using fixed parameters without significantly increasing the computational complexity. In Figure 3, we can easily see that the boundary information of the image information is mainly concentrated in the low-frequency component of the WT, so we retain the LL region of the WT and enhance the boundary information of the lesion to improve the subsequent segmentation accuracy. Subsequently, the LL is convolutionally stitched with UNet once to achieve the same convolutional feature connection and ensure the integrity of the next convolution process. Afterwards, similar operations are repeated for LL to ensure the smooth WT, image stitching, and convolution in the UNet downsampling process, and to realize the organic combination of WT and UNet network.
In this module, the image stitching and convolution of WT are performed. Since there is a trade-off between the computational volume of the convolution operation and the target detection capability, to reduce the computational volume, we mainly use its low-frequency components in the WT process to concatenate with the convolution results, ensuring a relatively small computational volume under the condition of higher resolution feature maps, which in turn achieves a trade-off between the computational volume and the target detection capability, ensuring the subsequent medical lesion recognition and improve the recognition capability of lesions.
Transformer encoding block
In using the UNet network coding part for shallow feature extraction, feature mapping from image to high-dimensional space is achieved while keeping the spatial feature distribution unchanged. When the traditional CNN model performs feature extraction, the deep-level information is obtained by downsampling and convolution operations, which leads to the loss of valuable information in the structure of the deep-level network and ignores the overall and local relevance. Therefore, TEB is proposed to replace the deep encoding part of the UNet network, which consists of linear projection (LP), efficient multi-head self-attention (EMSA), and residual (feed-forward) (RF).
Linear projection
To meet the input requirements of the transformer block, we use linear mapping to flatten the feature map output from the last encoded part of the UNet network to change its feature dimension from HWC→ (HW)C while matching the trainable linear mapping to the desired dimension d. In a traditional transformer, in order to obtain each inter-patch correlation information, the choice of imposing positional encoding to complete the association between each token ignores the local relationship with the structural information within each patch, to reduce the impact of this limitation, we use DW Conv and residual structure for linear mapping, defined as shown in equation (5).
Efficient multi-head self-attention
The traditional self-attentive mechanism maps image X into a
To further improve operational efficiency and reduce computational overhead, we propose EMSA. Before performing the self-attentive mechanism operation, we utilize k × k DW Conv with s = 2 to reduce the space size of k and v.

The structure of efficient multi-head self-attention (EMSA).
Finally, we adopt EMSA with h multiple heads, where h is decided by us artificially. Each head outputs a sequence of size
Residual feed-forward
For forward propagation in the transformer block, the dimensionality is usually scaled using MLP for generalization and feature enhancement. Layer normalization (LN) normalizes the feature information before and after MSA to avoid optimization problems. For better performance, we replace the conventional convolution with DW convolution, which is used to extract local information and has a negligible additional computational cost. The output of this part is shown in equations (8) to (10).
We then take the output of the transformer module
Complexity analysis
We analyzed the complexity between the traditional vision transformer (VIT) and our transformer block. The traditional VIT consists of a multi-headed attention mechanism (MSA) and a feed-forward neural network. The softmax layer increases the computation but does not increase the complexity, so it is not included in this comparison. The computational complexity of the traditional VIT model is shown in equation (11).
Joint loss function
In this article, in order to obtain higher accuracy segmentation, we use a hybrid loss function for reducing the difference degree between labeled and segmented data. We define this joint loss function as shown in equation (13)
The BCE loss function is mainly used for fast evaluation and optimization of binary classification tasks, which can effectively capture the errors of the classifier and adjust the weights according to the number and contribution of errors, and is the most widely used loss function in binary classification and segmentation. The expression is shown in equation (14).
Intersection over union (IOU) loss function is used to solve the common loss function of image semantic segmentation, used as a standard evaluation of target detection and image segmentation, which can accurately identify different objects in the image and can overcome the noise and misalignment in the error, used to reflect the gap between the initial segmented image and the doctor labeled information, its expression is shown below:
Experiments and results
We demonstrate the effectiveness and performance of our model through experiments. We first present the sources and relevant details of the dataset, then compare the proposed approach with five networks and finally conduct ablation experiments to evaluate the usefulness and effectiveness of our proposed two modules.
Datasets
In order to validate the effectiveness of our method, we conducted a training test using medical image data from 100 patients with stage T1 and T2 NPC aged between 30 and 50 years old. The training test validated a cut-off ratio of 8 to 1 to 1, totaling 5000 image data. All image data were obtained from the MRI of Hainan Provincial People's Hospital. The data size is in IMA format, and the size of the data available for processing after excluding personal information is 1647*931, which contains a large range of tissue structures from head to neck, while the ROI of nasopharyngeal cancer only accounts for a small portion of the image data, and most of them are black invalid areas and sensitive information such as the hospital where the patient is located, his name and age. Therefore, we first filtered the acquired data to remove image data that did not contain lesion areas; second, we cropped the acquired data. Through close communication with doctors and observing the characteristics of the sample set, we found that the top left corner of the image is “SIEMENS, EX, Se, IM” medical record information, while the top right corner is the name and age of the current patient in Hainan Provincial People's Hospital and other sensitive information, and the bottom left corner is the time at that time, etc. In this regard, we uniformly decimated the data. In this regard, we unified data desensitization and data cropping, and finally manually calibrated these data to ensure that there is no bias in the data. The obtained data are normalized, which reduces the complexity of calculation and improves the accuracy of segmentation. In order to provide a sufficient amount of data, all cropped images are used as original input images during the training and testing phases. To overcome the problem of an insufficient number of samples during training, adaptive random inversion, random cropping, and Gaussian noise33,34 were performed to achieve sample expansion.
Implementation details
For all experiments, we performed simple data expansions using Python, such as random cropping and random flipping, to avoid overfitting problems due to the homogeneity of the data. The main architecture of WET-UNet is a modified UNet, and we used an efficient transformer in the encoder and decoder connection part of the UNet network. All experiments were trained and tested on the PyTorch platform and a single Nvidia GTX3060 GPU. All images were uniformly resized to a resolution of 512 × 512 pixels. The network was trained in an end-to-end fashion. The initial learning rate was set to 0.01, the momentum to 0.9, the weight decay to 1 × 10−4, and the default batch size to 2.
Evaluation metric
During the experiments, we evaluated the ROI segmentation accuracy using the Dice similarity coefficient (Dice), IOU, accuracy, precision, recall, and specificity, using the lesion region labels with the truthfulness of the actual predicted values used to set the parameters and to express the above metrics. When the label is true and the prediction is also true, it is expressed by true positive (TP); when the label is true and the prediction is false, it is expressed by false negative (FN); when the label is false and the prediction is true, it is expressed by false positive (FP); when the label is false and the prediction is also false, it is expressed by true negative (TN). Dice, IOU, precision, accuracy, recall, and specificity are expressed by the above parameters, which are defined as shown in equations (16) to (21), respectively.
Compare experiments
To demonstrate the reliability and superiority of our experiments, we compared several models in processing medical image segmentation, and the specific quantitative results are shown in Figure 5. In the figure, we compare several current widely used models, including the CNN-based methods nnUNet, 35 UNet++, 36 TransUNet, VIT, and Swin-UNet, different models on the nasopharyngeal cancer dataset segmentation results.

Comparison of visualization results of different methods on representative MR images. From left to right, original image, Truth Label, nnUNet, UNet++, TransUNet, VIT, Swin-UNet. Each row contains segmentation results for a particular sample obtained by a different model. The corresponding results predicted by the model are marked with red areas, respectively.
In terms of network architecture, Swin-UNet outperforms the well-designed CNN-based model nnUNet. however, the network architecture itself is not the only determinant of performance. For example, the well-designed CNN-based model nnUNet in the table significantly outperforms the transformer-based model TransUNet. We can see that by directly employing UNet++ for medical image segmentation, UNet++ has many similar, blank feature mappings, while WET-UNet can take full advantage of all the feature mappings. Since the EMSA module guarantees the variation of the received domain, our ET-UNet can learn more diverse feature maps. Our model is based on UNet and a modified transformer of the traditional CNN, which has a lower number of parameters and time complexity compared with TransUNet and Swin-UNet. According to the results, our model achieves good results in this comparison. In addition, we observe that by comparing the original labels, our model predicts no false predictions, also known as false positives, which indicates that the model is more advantageous in noise prediction than other methods after adding the WT to the image processing.
To compare the model performance in detail, we used Dice, IOU, precision, accuracy, recall, and specificity as the quantitative analysis metrics, and the experimental results are shown in Table 1. The accuracy of WET-UNet using ResNet-50 as the backbone reached 0.849, which is higher compared to other models. Using the same training schedule, our proposed WET-UNet significantly outperforms this baseline, achieving a Dice of 0.834 and an IOU of 0.85. The WT module is used in shallow features to enhance the stability and robustness of the model and learn feature textures; the TEB module is used in deep feature extraction, which makes full use of all feature mappings and prefers to learn edges, and thus can obtain better performance. We can clearly see from the figure that the result of each model segmentation will have a part of the error, which we call the error rate. In order to ensure the reliability of the experiment, we mainly compare the error rate with the popular segmentation model in the past two years, and it can be seen that our model has a lower error rate and higher accuracy.
Comparison results of parameters of different models, from left to right, nnUNet, UNet++, TransUNet, VIT, and Swin-UNet.

IOU: intersection over union; VIT: vision transformer.
In extracting deep features, a pure transformer without convolution is better for learning global and local semantic information interaction and obtaining better results. While using the traditional pure transformer also leads to an increase in the number of parameters and time complexity, the proposed efficient multi-head self-attention, compared to the simple multi-head attention mechanism, uses DWConv depth separable convolution to scale k, v, thus reducing the computational overhead. It is proven to be effective through experiments. Based on our experiments, we give the corresponding metrics, which are the floating point number of the model, the parameters, and the average inference time, as shown in Figure 6. By comparison, it is found that ET-UNet is a small to medium-sized network that outperforms other.

Comparison of the number of params, FLOPs, and average inference time of each model in the experiment networks for moderate model complexity, while the average inference time is second only to vision transformer (VIT).
Ablation study
This section focuses on the validation of the two modules mentioned above, and the results are shown in the figure (comparison of metrics). For each module, we independently validate its validity. We adjusted the ratio of training and validation to 8:2, based on the original dataset, for the ablation experiments.
In order to clearly see what each module does and has achieved its function, we evaluate the performance of the base model and the two independent modules using DSC and precision, and also visualize the results of each segmentation. As shown in Table 2, with only one base network, the DSC and precision coefficients are low, but as a base network, it has a small number of parameters and a short inference time. After adding the WT module, we can clearly find that the float of all hard metrics is smaller than other models, which indicates that WT can significantly improve the robustness and stability of the network; adding the TEM module to the base significantly improves the accuracy, but with it comes a part of parameter increase. WET-UNet learns the noise and semantic features through the WT module and TEB module structure. The interaction between and feature correlation further improves the segmentation results. The visualization of the segmentation is specified in Figure 7.

Comparison of segmentation results for different combinations of input modalities.
Results of the ablation experiment.

WT: wavelet transform; DSC: Dice similarity coefficient.
Conclusion
In this study, we propose an innovative WET-UNet network model designed for fast segmentation of nasopharyngeal cancer images. We integrated the UNet architecture, WT, and Transformer module in this model to improve the segmentation performance. Through rigorous experimental validation on nasopharyngeal cancer images collected from Hainan People's Hospital, we draw the following conclusions:
Superior segmentation: Our WET-UNet network performs well in the task of nasopharyngeal cancer image segmentation. Compared with other networks, our method achieves significant improvements in segmentation accuracy and backbone structure segmentation. Improved performance metrics: Our method exhibits higher values in several performance metrics (e.g. Dice coefficient, Jaccard index, and recall), demonstrating its superior performance in nasopharyngeal cancer image segmentation. Moderate parameter complexity: Although we introduced the WT and transformer modules, the parameter complexity of the network remains manageable, making it suitable for practical applications.
Despite the satisfactory results of our study, we would like to honestly admit that there are some limitations. First, our study focused only on nasopharyngeal cancer image segmentation, thus limiting its applicability to this area. Second, the size of the dataset is relatively small, and a broader dataset may offer more possibilities. Finally, the performance of our method is limited by hardware and computational resources, and further computational resources may improve training and inference speed.
Based on the results of this study, our future research direction is to validate the effectiveness of our method in image segmentation for other cancer types to further demonstrate its generalization. At the same time, we will work on optimizing the network structure of WET-UNet to improve performance and reduce parameter complexity. Finally, we plan to apply our method to real medical diagnosis and treatment to verify its utility in clinical practice.
Footnotes
Abbreviations
Acknowledgements
Not applicable.
Authors’ contributions
YZ conceived the study, performed the experiments, obtained the data, and drafted the first manuscript. JL, ZZ, and WL edited the manuscript and organized the data. Ph Z and Sd S analyzed the data and revised the manuscript, and CS confirmed the authenticity of all the original data. All authors have read and approved the final version of the manuscript.
Availability of data and materials
The data sets used and/or analyzed in this study are available from the corresponding authors upon reasonable request. However, the data came from the Hainan Provincial People's Hospital and only a small portion of the data can be disclosed after communication; please contact the corresponding author for more details.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethics approval
Ethical approval to report this case was obtained from the Medical Ethics Committee of Hainan Provincial People's Hospital.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Hainan Province Science and Technology Special Fund (grant number ZDKJ2021042).
Informed consent
Verbal informed consent was obtained from a legally authorized representative(s) for anonymized patient information to be published in this article.
Patient consent for publication
Not applicable.
Author biographies
Yan Zeng is a Ph.D. from Hainan University. Her research area is medical big data, with an emphasis on medical image processing.
Jun Li is a master's student at Hainan University. His research field is medical big data, focusing on medical image processing.
Zhe Zhao is a master's student at Hainan University. His research field is medical big data, focusing on medical image processing.
Wei Liang is a master's student at Hainan University. His research field is medical big data, focusing on medical image processing.
Penghui Zeng is a master's student at Hainan University. His research field is medical big data, focusing on medical image processing.
Shaodong Shen is a master's student at Hainan University. His research field is medical big data, focusing on medical image processing.
Kun Zhang is a Ph.D. from Hainan Normal University. His research area is computer engineering with an emphasis on computing.
Chon Shen is a professor at Hainan University. His research area is medical big data, with an emphasis on image processing.