Abstract
Cetaceans inhabit oceans throughout the world. Four specific odontocetes, namely Cuvier's beaked whale (Ziphius cavirostris), Indo Pacific finless porpoise (Neophocaena phocaenoides), pygmy sperm whale (Kogia breviceps), and dwarf sperm whale (Kogia sima), have occasionally been found stranded along Thailand's coastal waters (the Andaman Sea and the Gulf of Thailand). Although shared haplotypes of each species for many locations have been found, and some species have revealed genetic structure through haplotype networks, cetaceans in Thai waters have never been investigated in terms of comparing haplotypes to those that have existed before. Herein, we have illustrated the matrilineally phylogeographic relationships among worldwide populations through Bayesian Phylogenetic tree computations using Markov Chain Monte Carlo (MCMC) and Median-Joining Networks (MJNs). Unique haplotypes of the control region mitochondrial DNA of Thai odontocetes were found for all species. Moreover, a high degree of worldwide haplotype diversity (hd) above 0.8 among the four species was detected, while the lowest degree of nucleotide diversity (π) was observed in the Indo Pacific finless porpoise (1.12% ± 0.184%). An expansion of the effective female population size worldwide of three odontocete species was detected using Bayesian Skyline Reconstruction, but this did not include the Indo Pacific finless porpoise. Because Thai seas are located within the Indo Polynesian province, where this biodiversity hotspot exists, we speculate that these odontocetes may also inhabit specific habitats within the Malay Peninsula and Thailand's territorial waters. Therefore, closer attention and monitoring of these cetacean populations will be necessary for future conservation efforts.
Introduction
Cetacean populations have been in decline globally due to both natural and human causes, including habitat degradation resulting from coastal and river development, depletion of food resources through competition with fisheries, bycatch of the commercial fishing industry, predation by other species, disease outbreaks, and climate change. 1 Whales in particular have been killed for oil and meat for millennia.2–4 Four odontocete species of toothed whales, dolphins, and porpoises were included in this study, namely Cuvier's beaked whale (Ziphius cavirostris), Indo Pacific finless porpoise (Neophocaena phocaenoides), pygmy sperm whale (Kogia breviceps), and dwarf sperm whale (Kogia sima). At present, these species are being negatively impacted by human actions.5–9 Many international organizations now aim to protect these species through the implementation of international agreements, guidelines, and laws. Two examples of these organizations include the Convention on International Trade in Endangered Species of Wild Fauna and Flora (CITES) and the International Union for Conservation of Nature's Red List of Threatened Species (IUCN Red lists). In Thailand, the Ministry of Agriculture and Cooperatives prohibited dolphin fishing in 1990. 10 In the past decade, the Wild Animal Reservation and Protection Act (WARPA) of Thailand were announced in 2019 to protect marine mammals in Thai territorial waters. 11 The four odontocete species included in this study have also been listed as protected species. Many whales in the Mysticetes family (baleen whale) that were severely depleted in the past have now shown signs of recovery, such as increasing numbers of the blue whale (Balaenoptera musculus) in the Antarctic and the humpback whale (Megaptera novaeangliae) in the North Atlantic Ocean after the end of commercial hunts.12–14 In contrast, other odontocetes (toothed whales, dolphins, and porpoises) have displayed a lack of resilience for a number of reasons, including a trend towards a later age for first reproduction, low calving rates, and other social and behavioral traits. 12
Cuvier's beaked whale (goose-beaked whale) is a cosmopolitan species found in temperate, subtropical, and tropical oceans around the world, particularly in deep waters near the continental slope.15–19 During the years between 1982 and 2001, reported numbers of this species were only 11–95 individual whales in the North Atlantic Ocean and 1621–90,725 individuals in the Pacific Ocean. 20 The highest population densities were found in the southern Gulf of California at about 38 animals per 1000 km2. 20 High population densities have also been reported in the waters of Hawaii and California. 21 This species has also been found in the Mediterranean Sea, particularly in the Alborán and Ligurian Seas. 21 More specifically, there are an estimated 429 individual whales inhabiting the Alborán Sea, which connects to the Atlantic Ocean through the Strait of Gibraltar. 21 There have also been reports of this species in the Malay Peninsula and other nearby seas. For example, there has been one sighting of this species in the East Indian Ocean 22 and three strandings along the coasts of northern Java,23,24 Sabah, Malaysia,25,26 and in the coastal waters of the Andaman Sea, Satun Province, Thailand. 27
The Finless porpoise is a coastal marine odontocete that includes two species: the Indo Pacific porpoise (Neophocaena phocaenoides) and the narrow-ridged finless porpoise (Neophocaena asiaeorientalis). 28 The differences between these species have been studied in relation to the morphology of the external skull, the dorsal structure, and their molecular genetics.29,30 Our study focuses on the Indo Pacific species that lives primarily in a narrow band of shallow waters along the coastlines of tropical and temperate Asia. This area starts from the Persian Gulf and extends east to the south of China, while including sections of the East China Sea, the western Indo-Malayan Archipelago and Java.5,31,32 Around the Malay Peninsula, sightings and strandings of Indo Pacific finless porpoise have been reported in the Gulf of Phuket along the coast of Malaysia including in the Malacca Strait and in the Johor estuary that extends from the South China Sea into the Western and Northeastern Gulf of Thailand along to the Cambodian Coast.18,25,26,33,34 Population numbers have been estimated for the Bay of Bengal (Bangladesh, n = 1382 individuals, coefficient of variation (CV) = 54.8%), Hong Kong waters (n = 176 individuals, CV = 26%), and Sarawak (Malaysia, n = 135 individuals, CV = 31%).26,35–37 In the central-western Gulf of Thailand, only 13 groups of finless porpoises were observed to be living in the core habitat at Don Sak, Surat Thani Province 38 ; however, numbers in the Andaman Sea, particularly along the Thai coast, are as of yet unknown.
Pygmy and dwarf sperm whales (genus Kogia) are also commonly found in deep waters, especially over and near the continental slope, but within slightly different habitats. Pygmy sperm whales reside in both tropical and warm temperate waters, while dwarf sperm whales prefer tropical waters. 5 Rarely sighted, there are little data on the abundance of these two species in the wild. Combining both Kogia spp., 395 individuals had been estimated to be in existence in the western Atlantic ocean, while only three have been found stranded in the Mediterranean Sea.39,40 Along the Malay Peninsula, only a few specimens of each species were recorded in Vietnam and in the Indonesian archipelago.24,41 By contrast, both of these species have commonly been found stranded along the southeastern United States and South African coasts. This would suggest that they may be more plentiful in those areas than previously thought.42–44 In Thailand, six dead pygmy sperm whales, as well as three dead and two live dwarf sperm whales, have been reported to be stranded along the coasts of these territorial waters since 1994.45–48
Both the Thai Andaman Sea and the Gulf of Thailand are located in Indo Polynesian province and are recognized as biodiversity hotspots. 49 An overlap in the habitual range of certain distinct species from both the Indian and Pacific Ocean has contributed to the species richness in these areas. Moreover, the overlapping range of some cetacean species has likely resulted from some degree of flexibility in the choice of prey amongst predators and niche partitioning.49–52 However, resource specialization, site fidelity to natal breeding grounds, and social structure are also believed to be contributing factors to the genetic structure of odontocetes.53,54 The degree of conformity between population genetic structure and resource specialization has been proven for many odontocete species such as Risso's dolphin and the killer whale throughout the Northern Pacific region.55,56
Currently, mitochondrial DNA is considered a marker of particular significance for population, biogeographic, and phylogenetic studies. 57 Mitochondrial DNA is comprised of extrachromosomal and small genomes that are generally 16 kb in size and represent maternal inheritance. The ratio of mitrochondrial DNA to nuclear genome copies is also high leading to increased success in the isolation of DNA from decomposed tissue in stranded cetaceans.58,59 A control region representing the noncoding segment of the mitochondrial DNA that is highly variable in many vertebrates has frequently been used to estimate population divergence and gene flow, effective population size, and population trends in wild populations. 60 Extensive numbers of control region sequences obtained from different marine mammal species are available in the GenBank repository. This has led to even greater convenience in data retrieval and for comparisons of the sequences in many cetacean genetic studies. 55 A previous genetic study of Cuvier's beaked whales was performed using a 290-bp mitochondrial DNA control region obtained from 87 individuals in three ocean basins, including those of the North Atlantic, North Pacific, South Pacific, and Indian Oceans. 61 In these waters, haplotype diversity was found to be relatively high with significant genetic differences observed among regions, while two unique haplotypes were identified in whales living in the Mediterranean Sea. A study of genetic variation of Kogia spp. in the Atlantic and Indo Pacific Oceans using a combined 406-bp mitochondrial DNA control region and 398-bp cytochrome b revealed 74 haplotypes of pygmy sperm whales and 27 haplotypes of dwarf sperm whales, with a higher nucleotide diversity observed in the dwarf sperm whales. 62 In that study, only five specimens collected from the Indian Ocean were examined and there were none from oceans in or near the Malay Peninsula. A comparison of genetic diversity of the Indo Pacific finless porpoise in the South China Sea revealed a degree of haplotype diversity that was higher in the Pear River Estuary when compared to the Taiwan Strait, with restricted haplotypes found only in this estuary area.63,64
In Thai Seas, genetic studies on cetacean populations have been limited. Only one out of 27 species found to be inhabiting Thai seas has been studied, the Irrawaddy dolphin (Orcaella brevirostris).45,65 Due to the limited number of cetacean samples in Thai Seas, little is known about how these marine mammals are related to each other in adjacent waters and the oceans worldwide. In the present study, eight mitochondrial control region DNA sequences obtained from the stranded odontocete samples of four species were used to investigate any relevant phylogeographic relationships and to compare these specimens with those acquired from other parts of the world. These comparisons were based on sequences retrieved from the National Center for Biotechnology Information (NCBI) Genbank. In this study, we aimed to investigate whether the unique haplotype identified in the four species found in Thai Seas was in common with the others inhabiting elsewhere and the nearby biodiversity hotspot. We also studied the worldwide maternal genetic diversity of these four species and investigated any changes in the world populations over time.
Methods
Samples
The Phuket Marine Biological Center, Phuket 83000, Thailand provided skin tissue samples from eight deceased cetaceans (Figure 1, Table 1) that were stranded on Thai coasts between 1990–2019. The tissue samples were collected and preserved in 95% ethanol and stored at −20°C. This study was approved of by the Animal Use Committee of the Faculty of Veterinary Medicine, Chiang Mai University, Thailand in 2020 (S14/2563).

Map shows locations (provinces) of 8 stranded samples on Thai coasts (Gulf of Thailand and Andaman Sea) between 1990-2019. The stranded samples are labeled on the map with black symbols, while the locations are indicated by colored areas.
Information on eight tissue samples collected from four species of whales within the waters of Thailand.

F = Female; M = Male; ADM = Andaman Sea; GOT = Gulf of Thailand.
DNA extraction
The skin samples were extracted using a DNeasy Blood & Tissue Kit (QIAGEN, Germany) according to the method previously described66,67 at the Faculty of Veterinary Medicine, Chiang Mai University, Thailand. Extracted DNA samples were prepared for purity evaluation using 2% agarose gel electrophoresis. DNA quantity was determined using a spectrophotometer to measure the 260–280 nm absorbance ratio. Diluted samples (10 ng/μL) were stored at −20°C for subsequent analyzes.
Mitochondrial DNA control region amplification
The mitochondrial control regions of eight samples were amplified using PCR primers: forward, 5′- CAT ATT ACA ACG GTC TTG TAA ACC −3′; and reverse, 5′- GTC ATA AGT CCA TCG AGA TGT C-3′. 68 PCR reactions were conducted in 25 μL reaction volumes using Platinum Taq DNA polymerase (Invitrogen) consisting of 1X reaction buffer, 2 mM MgCl2, 0.4 mg/mL bovine serum albumin, 0.25 mM dNTPs, 0.4 μM of both forward and reverse primers, and 2 μL of the DNA template. The PCR conditions were performed as follows: 95°C for 5 min, 40 cycles of 95°C for 30 s, 50°C for 45 s, 72°C for 1 min, and 72°C for 10 min. All eight PCR products obtained from the amplification were sequenced by ATGC CO., LTD, Pathum Thani, Thailand. The mitochondrial control region sequences obtained from this study, including one from Cuvier's beaked whale (Ziphius cavirostris), two from Indo Pacific finless porpoises (Neophocaena phocaenoides), three from pygmy sperm whales (Kogia breviceps), and two from dwarf sperm whales (Kogia sima), were visually prepared and edited manually using the MEGA-X program version 10.2.2. 69 Subsequently, maximum control region sequence lengths of 466 bp for Cuvier's beaked whale, 480 bp for Indo Pacific finless porpoises, 433 bp for pygmy sperm whales, and 476 bp for dwarf sperm whales were obtained and deposited in GenBank under the accession numbers MZ401334—MZ401341 (Table 1).
Dataset
The mitochondrial control region sequences of the four odontocete species available in GenBank were obtained for subsequent analysis using keyword searches for the following: “mitochondrion”/“control region mtDNA”/“d-loop” + “species name”. Mitogenomes, both full length and partial length of the mitochondrial control regions, were obtained through keyword searches, although only the D-loop Feature presented in the “Graphics” section was exported as a FASTA record. To ensure the correctness of the sequences and species, all were rechecked from the original research articles, and the percent identity was determined using the Basic Local Alignment Search Tool (BLAST). The final datasets of each species obtained from GenBank, including our new sequences, were of different lengths; however, for comparison with all available sequences in each species, the length of each dataset was reduced and aligned with the consensus fragments of 238 bp for Cuvier's beaked whale (63 sequences + 1 sequence of this study), 281 bp for Indo Pacific finless porpoises (100 sequences + 2 sequences of this study), 396 bp for pygmy sperm whales (75 sequences + 3 sequences of this study), and 399 bp for dwarf sperm whales (28 sequences + 2 sequences of this study) using Clustal W program implemented in MEGA X version 10.2.2.69,70 For accession numbers of all sequences used see Table S1.
Phylogenetic tree construction
Phylogenetic trees of the mitochondrial control region DNA sequences for each odontocete species were constructed to observe the phylogenetic relationships of our samples with other samples worldwide using Bayesian Analysis implemented in the MrBayes program version 3.2.7. 71 HKY + I, GTR + G, GTR + I + G, and HKY + I were used as the best tree evolutionary models for Cuvier's beaked whale, Indo Pacific finless porpoise, pygmy sperm whale, and dwarf sperm whale, respectively, which were determined by J model test version 2.1.10.72,73 The run length of Markov Chain Monte Carlo (MCMC) sampling at 10,000,000 iterations for each tree was performed using the convergence diagnostics with the average standard deviation of split frequencies below 0.01. The first 1,000,000 iterations were discarded in the burn-in step. The mitochondrial control region DNA sequences of closely related species were used to serve as an outgroup for each phylogenetic tree: Shepherd's beaked whale (Tasmacetus shepherdi, Accession number AF036227) for Cuvier's beaked whale tree; Cuvier's beaked whale (Ziphius cavirostris, Accession number NC_021435) for Indo Pacific finless porpoise tree; and sperm whale (Physeter catodon, Accession number KU891394) for both pygmy and dwarf sperm whale trees. The phylogenetic trees were then visualized by iTOL version 6.1.1. 74
Haplotypes and genetic diversity
The haplotypes of our samples were investigated and these were compared to the worldwide haplotypes using the alignment of the dataset of each species mentioned above. This comparison was implemented in DnaSP program version 6.12.3. 75 Furthermore, the worldwide genetic diversity of the control region mitochondrial DNA of each species was determined and presented as the number of haplotypes (H), the number of variable sites (S), haplotype diversity (hd), and nucleotide diversity (π) using the same program.76,77 To examine the regional genetic structure, the Median-Joining Networks (MJNs) of each haplotype were constructed using PopART program version 1.7. 78
Demographic analysis
To observe any possible population changes over time in each species, we performed Bayesian skyline plot (BSPs) analysis using the dataset mentioned above. The BSPs were estimated using BEAUti and BEAST program version 2.6.3. 79 The substitution model and a strict clock model were used for each dataset over 10,000,000 iterations with the first 1,000,000 iterations used as a burn-in step. Mutation rates for each species were used for molecular clock calibrations as follows. “The mtDNA control region mutation rate of Cuvier's beaked whale was determined by Dalebout, Robertson, 61 wherein the specific mutation rate at 1.2 × 10−8 bp per site per year was established. For the Indo Pacific finless porpoise, we used a mutation rate of 0.94 × 10−8 bp per site per year established by Lin, Frere. 80 This rate was used to estimate the expansion of the population of Neophocaena spp. in the northwestern Pacific region. As for the pygmy and dwarf sperm whales, the specific mutation rates for these species were not available. Thus, we used the mutation rates reported by Hoelzel, Hancock, 81 which proposed a mutation rate of the control region mitochondrial DNA for cetaceans at 0.5 × 10−9 bp per site per year.
Results
Phylogenetic trees
The phylogenetic tree of Cuvier's beaked whale revealed the monophyletic group for this species (Figure 2). Samples obtained from different oceans were mixed; however, subclades for the Atlantic Ocean and the Mediterranean Sea were detected without any samples from the Indo Pacific Ocean (posterior probability [PP] = 0.98) (haplotypes 5, 6, 7, 11, 12, and 25). Accordingly, our Cuvier's beaked whale sample (END145, haplotype 8) was clustered with a sample from New Zealand (PP = 0.53).
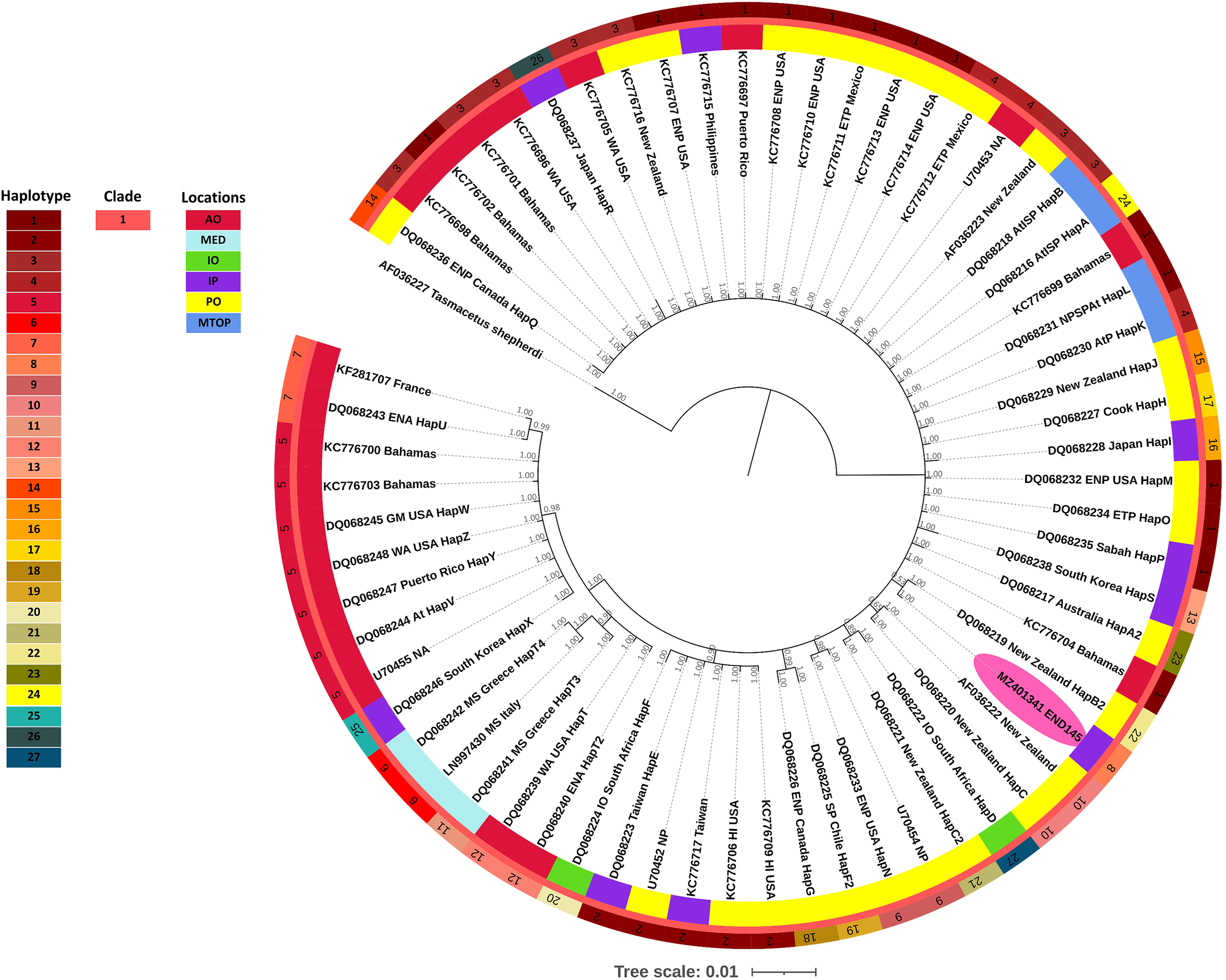
Phylogenetic tree of 64 Cuvier's beaked whales. The 63 control region mitochondrial DNA sequences were taken from the NCBI database and included one (MZ401341 END145) sequence from this study. Shepherd's beaked whale was used as an outgroup. The phylogenetic tree, 239 bp alignment, illustrates the unity clade of Cuvier's beaked whale from around the world with the presence of a red band in the middle layer. The color bands around the tree indicate the haplotypes—outer layer; clade—middle layer; and location—inner layer. Location abbreviations: AO, Atlantic Ocean; MED, Mediterranean; IO, Indian Ocean; IP, Indo Pacific Ocean; PO, Pacific Ocean and MTOP, more than one place.
Two main clades of the Indo Pacific finless porpoise have been presented in the phylogenetic tree (Figure 3). There were only five samples clustered in clade 1 including four from an unknown location and one from Taiwan (PP = 1.00, Figure 3). Our samples, END665 and END086 (haplotype 1), formed a small cluster that was grouped with the other samples in clade 2.

Phylogenetic tree of 102 Indo Pacific finless porpoises. The 100 control region mitochondrial DNA sequences were taken from the NCBI database and included two samples (MZ401340 END665 and MZ401339 END086) from this study. Cuvier's beaked whale was utilized as an outgroup. The phylogenetic tree, 296 bp alignment, illustrated two clades of the Indo Pacific finless porpoise identified from around the world as the red band (clade 1) and the light-yellow band (clade 2) in the middle layer. The color bands circling the tree indicate a haplotype—outer layer; a clade—middle layer; and a location—inner layer. Abbreviations of locations are as follows: IP, Indo Pacific Ocean; IO, Indian Ocean and Unk, Unknown.
According to the phylogeny of the pygmy sperm whale, three clades were detected (Figure 4(a)). Our two samples, END158 and END457 (haplotype 2), obtained from the Gulf of Thailand were the only samples located in clade 1 (PP = 0.99), while clades 2 and 3 were comprised of a mix of samples from different locations. Another sample, END2046 (haplotype 1) is presented in clade 2 and has been clustered with other samples from the Indo Pacific Ocean. Dwarf sperm whales were divided into two distinct clades of related geographical areas: the Indo Pacific Ocean (clade 1); and the Atlantic Ocean (clade 2) (Figure 4(b)). Two samples obtained in our study, END511 (haplotype 2) and END643 (haplotype 1), were included in the Indo Pacific Ocean clade.

Phylogenetic tree of 78 pygmy sperm whales and 30 dwarf sperm whales. (a) The 75 control region mitochondrial DNA sequences were taken from the NCBI database and three were taken from this study (MZ401336 END158, MZ401337 END457 and MZ401338 END2046). The sperm whale was utilized as an outgroup. The phylogenetic tree, 400 bp alignment, illustrated three clades of pygmy sperm whales from around the world as the red band (clade 1), the light yellow band (clade 2), and the light blue band (clade 3) in the middle layer. (b) The 28 control region mitochondrial DNA sequences were taken from the NCBI database and two were included in this study (MZ401334 END511 and END401335 END643). Sperm whale samples were utilized as an outgroup. The phylogenetic tree, 405 bp alignment, indicated 2 clades of the dwarf sperm whale from the Indo Pacific Ocean as the red band (clade 1) and from the Atlantic Ocean as the light yellow band (clade 2) in the middle layer. The color bands circling the tree indicate the haplotypes—outer layer; clade- middle layer; and location- inner layer. Abbreviations of locations are as follows: AIP, Atlantic and Indo Pacific Ocean; AO, Atlantic Ocean; IP, Indo Pacific Ocean; PO, Pacific Ocean and Unk, Unknown.
Restricted haplotype, haplotype network, and geographic distribution
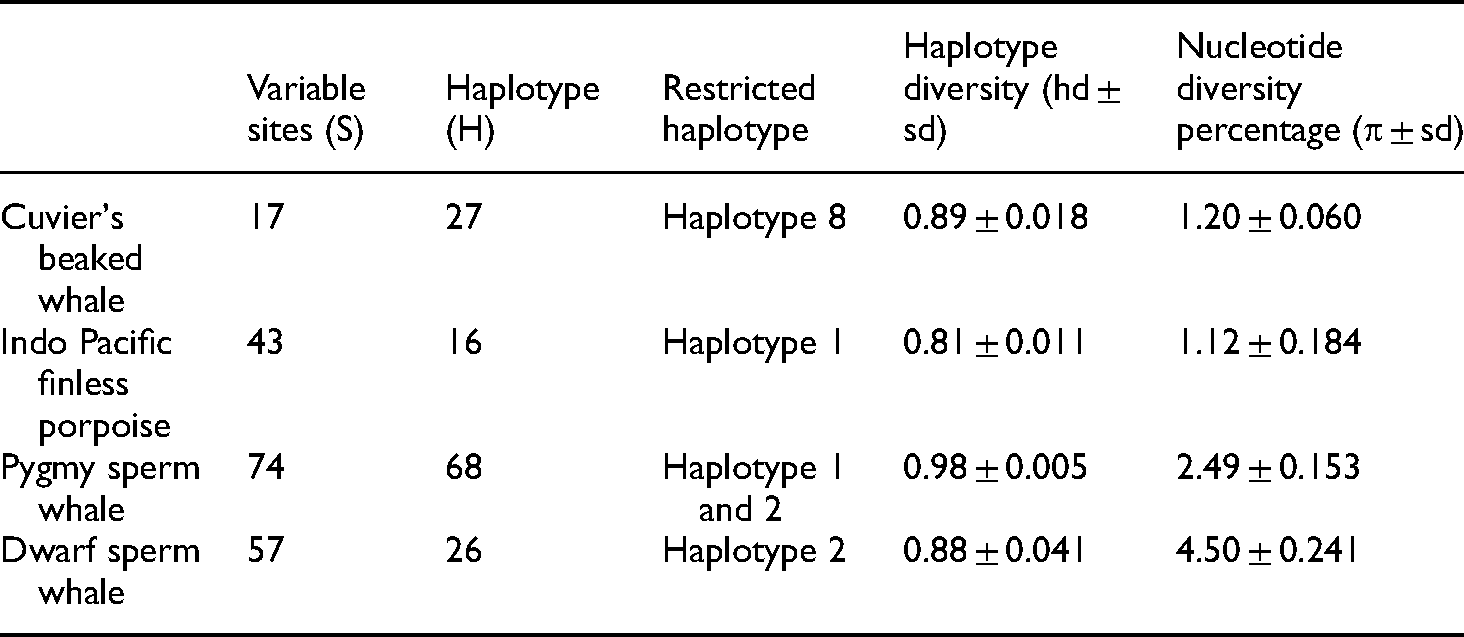
From an investigation of the 238 bp mitochondrial DNA control region of Cuvier's beaked whales, we identified 17 variable sites and 27 haplotypes. The worldwide haplotype diversity and nucleotide diversity values were high for this species (hd = 0.89 ± 0.018 and π = 1.20% ± 0.060%; Table 2). There were 18 haplotypes of Cuvier's beaked whale found to be restricted to a specific geographic area (Haplotype 8, 10, 11, and 13–27; (Figure S1A, Figure 5). The greatest number of Cuvier's beaked whale samples were found in more than one place (MTOP), a nonspecific location designation within the Atlantic, Indian, and Pacific Oceans, and included haplotypes 1, 3, 4, and 24. The major haplotype with the highest frequency for this species was haplotype 1. It was found in many locations but was mainly predominant in the Pacific Ocean (Figure S1A, Figure 5). Unique haplotypes that occurred only in the Atlantic Ocean were 5, 6, 7, 11, 12, and 25. For haplotype 8, restricted haplotypes for the Thai Andaman Sea, were found to be closely related to haplotype 22 from New Zealand and differed by only one nucleotide substitution.

Median-Joining Networks (MJNs) of Cuvier's beaked whale haplotype. MJNs were generated from 238 base pairs of Cuvier's beaked whale control region mitochondrial DNA sequences (115 samples) represented by 27 haplotypes found in 28 locations. Each haplotype is colored according to geographic region or location, and the small black circle represents the inferred haplotype (not sampled). The size of each circle is proportional to its haplotype frequency, while nucleotide substitutions are shown on the branches as a small transverse bar. Abbreviations of locations and haplotypes found at each location are as follows: AO; Atlantic Ocean (5), AU; Australia (23), BC; British Columbia (14, 18), BS; Bahamas (1, 3, 5), CK; Cook Island (17), CL; Chile (19), ETP; Eastern Tropical Pacific (1), GOM; Gulf of Mexico (5), GR; Greece (6, 11), HI; Hawaiian Islands (2), IE; Ireland (12), IT; Italy (6), JP; Japan (16, 26), KR; South Korea (13, 25), MTOP; more than one place (1, 3, 4, 24), MX; Mexico (1, 4), NA; North Atlantic Ocean (4, 5), NP; North Pacific Ocean (2, 9), NWF; North West of France (7), NZ; New Zealand (3, 10, 15, 21, 22), PH; Philippines (1), PR; Puerto Rico (1, 5), SBH; Sabah (1), SC; Scotland (7), TH_ADM; Thailand Andaman Sea (8), TW; Taiwan (2), USA_ENP; United States of America Eastern North Pacific Ocean (1, 9), USA_WA; United States of America West Atlantic Ocean (3, 5, 12) and ZA; South Africa (20, 27).
Restricted haplotype and mitochondrial DNA control region diversity parameters of four species of odontocetes.
The mitochondrial DNA control region diversity of the Indo Pacific finless porpoise was successfully obtained from 265 sample sequences worldwide (Table 2). We found 16 haplotypes and 43 variable sites on 281 bp in this species (Table 2). Four haplotypes (4, 5, 9, and 10), mostly from the eastern South China Sea including the waters of the Taiwan Straits and Japan, contained a greater number of samples than the other regions (Figure S1B, Figure 6). Our two samples (END665 and END086) collected from the Thai Andaman Sea and the Gulf of Thailand had a restricted haplotype (haplotype 1) that was closely related to haplotype 4 from Southeast China (including the Taiwan Straits), Hong Kong, and the Sea of Japan (Figure 6). We also found that haplotype 7 contained four sample sequences obtained from an unknown location and one from Taiwan (Accession numbers MT948057 and MT948059-062), which differed from most of the other locations (Figure 6).

Median-Joining Networks (MJNs) of Indo Pacific finless porpoise haplotype. MJNs were generated from 281 base pairs of Indo Pacific finless porpoise control region mitochondrial DNA sequences (265 samples) represented by 16 haplotypes found in 8 locations (including an unknown location). Each haplotype is colored according to a geographic region or location, and the small black circles represent inferred haplotypes (not sampled). The size of each circle is proportional to its haplotype frequency, while nucleotide substitutions are shown on the branches as a small transverse bar. Abbreviations of locations are as follows: CN; China (2-4, 9), HK; Hongkong (4), IN_ABS; India Arabian Sea (15, 16), JP; Japan (4, 5, 8-14), TH_ADM; Thailand Andaman Sea (1), TH_GOT; Thailand Gulf of Thailand (1), TW; Taiwan (7), Unk; Unknown (3, 6, 7).
Pygmy sperm whales had the largest number of haplotypes; 68 haplotypes and 74 variable sites were obtained from 119 sample sequences (396 bp) with the highest range of haplotype diversity observed among the four odontocete species in our study (hd = 0.98 ± 0.005 and π = 2.49% ± 0.153%; Table 2). For this species, we detected unique haplotypes from our three samples; haplotype 1 from the Andaman Sea (END2046) and haplotype 2, which contained two samples from the Gulf of Thailand (END158 and END457) (Figure S1C, Figure 7). Haplotype 2 had the highest nucleotide substitution numbers, which distinguished it from the others. However, haplotype 1 (END2046) was still closely related to the other haplotypes identified from the Atlantic and Indo Pacific oceans. Although this species exhibited the greatest number of variations in terms of haplotypes, a major haplotype within the larger population was not detected.

Median-Joining Networks (MJNs) of pygmy sperm whale haplotype. MJNs were generated from 396 base pairs of pygmy sperm whale control region mitochondrial DNA sequence (119 Samples) represented by 68 haplotypes found in 8 locations. Each haplotype is colored according to a geographic region or location, and the small black circles represent inferred haplotypes (not sampled). The size of each circle is proportional to its haplotype frequency, while nucleotide substitutions are shown on the branches as a small transverse bar. Abbreviations of locations and haplotypes found in each location are as follows: AIP; Atlantic and Indo Pacific Ocean (32, 35, 38), AO; Atlantic Ocean (6, 7, 9, 10, 12, 15-17, 19-22, 24-27, 30, 31, 34, 43, 50-54, 57-59), ID; Indonesia (3), IP; Indo-Pacific Ocean (3, 5, 11, 13, 14, 18, 23, 28, 29, 35-37, 39-49, 55, 56, 60-64), JP; Japan (4, 67), KI; Kiribati(68), TH_ADM; Thailand Andaman Sea (1), TH_GOT; Thailand Gulf of Thailand (2), Unk; Unknown (8, 33, 48, 65, 66).
While the dwarf sperm whale is a close relative of the pygmy sperm whale, fewer haplotypes were observed; 57 variable sites and 26 haplotypes. The worldwide haplotype diversity of this species (hd = 0.88 ± 0.041) was slightly less than that of the pygmy sperm whale; however, the percentage of nucleotide diversity was determined to be the highest (π = 4.50 ± 0.241) when compared to the other three odontocete species in our study (Table 2). There was a separation of two haplogroups in the Atlantic Ocean and in the Indo-Pacific Ocean represented by 28 substitution nucleotides (Figure S1D, Figure 8). Two samples collected from the Thai Andaman Sea (END511 and END643) were found in the Indo Pacific Ocean haplogroup, along with samples from China and Japan. END511 from Thailand was a restricted haplotype (haplotype 2), while another sample (END643) shared a haplotype with the Indo Pacific Ocean sample (haplotype 1).

Median-Joining Networks (MJNs) of dwarf sperm whale haplotype. MJNs were generated from 399 base pairs of dwarf sperm whale control region mitochondrial DNA sequence (52 Samples) represented by 26 haplotypes found in 6 locations (including an Unknown location). Each haplotype is colored according to a geographic region or location, and the small black circles represent inferred haplotypes (not sampled). The size of each circle is proportional to its haplotype frequency, while nucleotide substitutions are shown on the branches as a small transverse bar. Abbreviations of locations and haplotypes found in each location are as follows: AO; Atlantic Ocean (16-25), CN; China (26), IP; Indo Pacific Ocean (1, 4-15), JP; Japan (3, 4), TH_ADM; Thailand Andaman Sea (1, 2), Unk; Unknown (15).
Estimated effective population size over time
The BSPs analyzes of all four odontocete species are shown in Figure S2. Cuvier's beaked whale, the pygmy sperm whale, and the dwarf sperm whale exhibited a similar trajectory in terms of expansion of the female population. The effective female population size (Nef) for Cuvier's beaked whale had been continuously expanding over the last 350,000 years (Figure S2A). For the Indo Pacific finless porpoise, Nef had been in a stationary phase for 800,000 years without expansion or decline in the female population (Figure S2B). For the pygmy and dwarf sperm whales, Nef had been constant until around 75,000–100,000 years ago when the population expanded (Figures S2C and D).
Discussion
This is the first study to have examined the phylogeographic relationships and genetic information on these four odontocetes originating from Thai Seas, the important boundaries which connect it to the Indian Ocean and the Pacific Ocean. An important highlight of our study is that the restricted haplotypes of these four odontocete species occurred in Thai Seas, although one shared haplotype was found for the dwarf sperm whale, which was the Indo Pacific haplotype. A recalculation of worldwide haplotype diversity values has revealed a degree of conformity to the results of previous studies.61–63 We also detected an expansion of the female populations of Cuvier's beaked whale and the pygmy and dwarf sperm whales, which had not occurred for the Indo Pacific finless porpoise female population.
Cuvier's beaked whale
The worldwide diversity of Cuvier's beaked whale was first studied in 2005 61 using 290 bp of mitochondrial DNA control region segments. This study obtained 33 haplotypes from 87 sequence segments and revealed worldwide haplotype (0.926 ± 0.015) and nucleotide (1.27% ± 0.723%) diversity at high levels; however, the number of haplotypes, as well as the levels of haplotype and nucleotide diversity in our study, were all found to be lower. Using the larger sample size in our study, shorther lengths of consensus sequences were generated. This has led to lower diversity parameters; for example, a smaller number of haplotypes occurred when using only the control region mitochondrial DNA as compared to a combination of the control region and cytochrome b. 62 Most of the haplotypes (e.g. 1, 3, 4 and 24) were distributed among several locations and in all oceans, which may infer that they shared female ancestors in the past. This outcome was similar to that reported for this species in previous studies. 61 However, we found that haplotypes 5, 6, 7, 11, and 12 were also closely related to each other and were found only in the Atlantic Ocean and the Mediterranean Sea. These could form a specific Atlantic Ocean haplogroup or subclade, suggesting that the specific oceanographic and ecological conditions in these areas were involved in shaping the distinct maternal lineage as has occurred in other odontocetes, such as the Risso's dolphin population in the North Pacific and North Atlantic Oceans. 55 Similarly, the unique haplotype that was found for our Cuvier's beaked whale sample from the Thai Andaman Sea may have occurred for the same reason. Because this area is located in a biodiversity hotspot, 49 prey species richness is also likely to promote the differentiation of cetacean populations.51,52 Noticeably, haplotype 25 from a whale meat market located in South Korea (Pacific Ocean) also showed a close relationship with this Atlantic Ocean group. Accordingly, Dalebout et al. 61 suggest that more samples from the Pacific Ocean should be investigated and that agreements banning international trade may have been violated. The mixing of two oceanic (Pacific Ocean and Southern Ocean) Cuvier's beaked whale samples in one clade 82 also produced similar outcomes in our study.
While the estimation trend of Cuvier's beaked whale in the California Current has shown a stable trend from 1993 to 1996, 83 the findings of a recent study also indicate that population trends in the San Nicolas Basin, North Pacific Ocean, were stable or decreased slowly by about 0.8% per year from 2007 to 2018. 84 The stable population trend in this area may indicate a moderate efficiency of conservation management strategies to protect the species, such as using acoustic pingers to reduce bycatch in California drift fishery. 85
Indo Pacific finless porpoise
The number of haplotypes and haplotype diversity of the Indo Pacific finless porpoise in this study was similar to those of previous studies.63,80 The restricted haplotype 1 found in both sites of Thailand (Andaman Sea and the Gulf of Thailand) could infer that this coastal cetacean species has a related ancestor to animals in haplotype 4, which were found in the southern East China Sea and in waters of the Taiwan Straits, Hong Kong, and Japan. There have been records of sightings and strandings of the finless porpoise in Malaysian waters, including Peninsular Malaysia, Sarawak, and Sabah since 1931, 26 but none of those samples have been studied or had sequences deposited in the NCBI database. Consequently, our results provide more genetic evidence that this species may live and migrate around the Andaman Sea coast, southeast to the Malacca Strait-Singapore and north to the Gulf of Thailand. Thus, more samples obtained from Malaysian waters should be examined to expand the biological database for this species.
The phylogenetic tree of the Indo Pacific finless porpoise shows a united clade, except for five samples collected from Chehida's study, 86 which were separated from the first clade. These samples in haplotype 7 were determined to be different from the others by 29 substitution nucleotides. This would suggest that the separation of clade 1 in our study might be due to the misidentification of samples because Neophocaena spp has a sympatric overlap zone in the Taiwan Straits where all three of these species have been found. 87 Thus, it is possible that all samples in clade 1 contain the other Neophocaena spp., but not N. phocaenoides.
As for the population trend of this species, a recent study conducted by Lin et al. (2019) found an increasing trend in the population for this species at 0.76% annually from 2006 to 2014. However, in Hong Kong waters, the Indo Pacific finless porpoise population was found to be fairly stable for a 23-year period from 1996 to 2019. 35 However, the BSPs could not detect changes in the female population size of this species on a global scale.
Pygmy and dwarf sperm whales
In the NCBI database, most of the control region mitochondrial DNA sequences for these species were published by Chivers et al. 62 ; thus, the haplotype numbers and diversity parameters were similar to our results. That study reported the existence of 64 and 24 haplotypes using the control region mitochondrial DNA of pygmy sperm whales and dwarf sperm whales, respectively. 62 In our study, we included 11 and 5 more sequence samples than the previous study for pygmy and dwarf sperm whales, respectively. The new samples in our study were obtained from China, Indonesia, Japan, Kiribati, and the territorial waters of Thailand.88–91 The new worldwide control region mitochondrial DNA haplotypes that we generated increased to 68 haplotypes for pygmy sperm whales and 26 haplotypes for dwarf sperm whales when compared to the outcomes of a previous study. 62 As for the phylogenetic tree of the dwarf sperm whale, our study produced similar results to those of an earlier study 62 with two clades of the Atlantic and Indo Pacific dwarf sperm whales. Both samples from our study (END511 and END643) were located systematically in clade 1 (the clade of the Indo Pacific dwarf sperm whale). Although we inputted five new samples of the dwarf sperm whale into our analyzes, they all originated from the Indo Pacific Ocean suggesting that more samples from the Atlantic Ocean should be investigated in future studies.
The abundance of these two populations is still unknown due to deep diving and ship avoidance behavior. However, an estimated population size of the pygmy sperm whale in Hawaii was determined to be 7251 individuals. 92 Importantly, the population trends of these two species have not yet been fully studied, with the exception of our estimation of the female effective population size.
In this study, the use of mitochondrial DNA sequences alone should be interpreted with caution because of the limitations of this marker. This marker is only inherited from maternal lineage and is susceptible to genetic drift effects. This can result in a degree of incongruity with other nuclear DNA markers.93,94 Studies involving combinations with other nuclear markers should be performed in the future when more samples are available to reveal further insights.
Implications for conservation
Identification of the unique Thai haplotypes of the four odontocete species found in our study represents just the first step in the further exploration of these marine mammals in both Thai waters and nearby seas. The resulting information could be used to monitor and compare the haplotypes of these four species in the future, particularly in the Malay Peninsula. Because odontocetes often exhibit a different pattern of population structure, even within a short distance, 53 the diverse populations are likely to be found in Thai Seas that are connected to both the Indian and Pacific Oceans. Furthermore, the outcomes of future studies may reveal genetic relatedness among the major habitats of these species.
Conclusion
The worldwide phylogeographic relationship of each odontocete species was successfully revealed from the mitochondrial DNA control region established in the online database, while unique haplotypes of the Thai samples were also detected in all four species. We speculate that they not only travel throughout the Malay Peninsula, but may also share a unique population structure and inhabit nearby seas. This would be particularly true for the cosmopolitan oceanic species (Cuvier's beaked whale, pygmy sperm whale and dwarf sperm whale). Although we reported expansions in the female population size of these odontocetes, populations still need to be closely monitored to identify all relevant and emerging trends in the future. This would be particularly true for the Thai cetacean populations. The collection of genetic information from cetaceans inhabiting the seas of Thailand is important for evaluating and developing effective conservation actions and management strategies. In the future, with more samples from these marine mammals, we hope to further explore genetic relationships and expand the biological database of these threatened or endangered species in Thai waters.
Supplemental Material
sj-png-1-sci-10.1177_00368504221103776 - Supplemental material for Relationship of stranded cetaceans in Thai territorial waters to global populations: Mitochondrial DNA diversity of Cuvier's beaked whale, Indo Pacific finless porpoise, pygmy sperm whale, and dwarf sperm whale
Supplemental material, sj-png-1-sci-10.1177_00368504221103776 for Relationship of stranded cetaceans in Thai territorial waters to global populations: Mitochondrial DNA diversity of Cuvier's beaked whale, Indo Pacific finless porpoise, pygmy sperm whale, and dwarf sperm whale by Promporn Piboon, Napat Kriengsakpichit, Anocha Poommouang, Kittisak Buddhachat, Janine L. Brown, Jatupol Kampuansai, Siriwadee Chomdej, Patcharaporn Kaewmong, Kongkiat Kittiwattanawong and Korakot Nganvongpanit in Science Progress
Supplemental Material
sj-png-2-sci-10.1177_00368504221103776 - Supplemental material for Relationship of stranded cetaceans in Thai territorial waters to global populations: Mitochondrial DNA diversity of Cuvier's beaked whale, Indo Pacific finless porpoise, pygmy sperm whale, and dwarf sperm whale
Supplemental material, sj-png-2-sci-10.1177_00368504221103776 for Relationship of stranded cetaceans in Thai territorial waters to global populations: Mitochondrial DNA diversity of Cuvier's beaked whale, Indo Pacific finless porpoise, pygmy sperm whale, and dwarf sperm whale by Promporn Piboon, Napat Kriengsakpichit, Anocha Poommouang, Kittisak Buddhachat, Janine L. Brown, Jatupol Kampuansai, Siriwadee Chomdej, Patcharaporn Kaewmong, Kongkiat Kittiwattanawong and Korakot Nganvongpanit in Science Progress
Supplemental Material
sj-docx-3-sci-10.1177_00368504221103776 - Supplemental material for Relationship of stranded cetaceans in Thai territorial waters to global populations: Mitochondrial DNA diversity of Cuvier's beaked whale, Indo Pacific finless porpoise, pygmy sperm whale, and dwarf sperm whale
Supplemental material, sj-docx-3-sci-10.1177_00368504221103776 for Relationship of stranded cetaceans in Thai territorial waters to global populations: Mitochondrial DNA diversity of Cuvier's beaked whale, Indo Pacific finless porpoise, pygmy sperm whale, and dwarf sperm whale by Promporn Piboon, Napat Kriengsakpichit, Anocha Poommouang, Kittisak Buddhachat, Janine L. Brown, Jatupol Kampuansai, Siriwadee Chomdej, Patcharaporn Kaewmong, Kongkiat Kittiwattanawong and Korakot Nganvongpanit in Science Progress
Footnotes
Acknowledgements
We dedicate the value of this research to the Phuket Marine Biological Center, Phuket, Thailand. To this center, we express our deepest gratitude for providing supported samples, data, and additional information that allowed us to fully analyze the results of this study.
Author contributions
Promporn Piboon conducted the experiment, analyzed the data, and wrote the manuscript. Napat Kriengsakpichit conducted the experiment. Anocha Poommouang offered advise on the laboratory work. Kittisak Buddhachat and Janine L. Brown offered advice on the laboratory work and wrote the manuscript. Jatupol Kampuansai and Siriwadee Chomdej offered advice, provided important scientific detail for the experiment, and also wrote the manuscript. Patcharaporn Kaewmong and Kongkiat Kittiwatanawong provided samples and data. Korakot Nganvongpanit designed the study, analyzed the data, and wrote and finalized the manuscript. All authors read and approved of the final version of this manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The authors are grateful for the research funding provided by the Excellence Center in Veterinary Bioscience, Chiang Mai University, Chiang Mai 50200, Thailand; and the CMU Presidential Scholarship, Chiang Mai University, Thailand.
Data availability
The data that support the findings of this study are available from the corresponding authors upon reasonable request.
Supplemental material
Supplemental material for this article is available online.
Author biographies
Promporn Piboon is a PhD student at the faculty of Veterinary Medicine, Chiang Mai University, Thailand.
Napat Kriengsakpichit is a veterinary student at the faculty of Veterinary Medicine, Chiang Mai University, Thailand.
Anocha Poommouang is a PhD student at the faculty of Veterinary Medicine, Chiang Mai University, Thailand.
Kittisak Buddhachat is an Associate Professor at the Faculty of Science, Naresuan University, Phitsanulok, Thailand.
Janine L. Brown is a senior scientist at the Center for Species Survival, Smithsonian Conservation Biology Institute, Front Royal, Virginia, USA.
Jatupol Kampuansai is an Associate Professor at the Faculty of Science, Chiang Mai University.
Siriwadee Chomdej is an Associate Professor at the Faculty of Science, Chiang Mai University.
Patcharaporn Kaewmong is a scientist at Phuket Marine Biological Center, Phuket, Thailand.
Kongkiat Kittiwattanawong is a scientist at Phuket Marine Biological Center, Phuket, Thailand.
Korakot Nganvongpanit is a Professor at the Faculty of Veterinary Medicine, Chiang Mai University.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
