Abstract
Genotyping of genetically-engineered mice is necessary for the effective design of breeding strategies and identification of mutant mice. This process relies on the identification of DNA markers introduced into genomic sequences of mice, a task usually performed using the polymerase chain reaction (PCR). Clearly, the limiting step in genotyping is isolating pure genomic DNA. Isolation of mouse DNA for genotyping typically involves painful procedures such as tail snip, digit removal, or ear punch. Although the harvesting of hair has previously been proposed as a source of genomic DNA, there has been a perceived complication and reluctance to use this non-painful technique because of low DNA yields and fear of contamination. In this study we developed a simple, economic, and efficient strategy using Chelex® resins to purify genomic DNA from hair roots of mice which are suitable for genotyping. Upon comparison with standard DNA purification methods using a commercially available kit, we demonstrate that Chelex® efficiently and consistently purifies high-quality DNA from hair roots, minimizing pain, shortening time and reducing costs associated with the determination of accurate genotypes. Therefore, the use of hair roots combined with Chelex® is a reliable and more humane alternative for DNA genotyping.
The identification of a genetic marker introduced in a single modified allele in mice (heterozygous, +/–) is often necessary for designing breeding strategies in an efficient manner. In addition, genotyping becomes necessary when mice carrying null alleles (knockout [KO], –/–) cannot be phenotypically distinguished from wild-type mice (WT, +/+). In a similar fashion, mice harboring LoxP sites in one or both alleles of a given gene (‘flox’ alleles) for Cre-mediated recombination in vivo frequently cannot be distinguished without genotyping.
To determine the genetic constitution of genetically-engineered mice, purified DNA is frequently subjected to the polymerase chain reaction (PCR) using specific primers, such as oligonucleotides, designed to amplify and characterize genomic regions of interest. PCR almost always requires highly purified DNA, a task usually achieved by performing expensive, potentially toxic, and frequently tedious methods (extensively reviewed 1 ). In addition, mouse-genotyping methods rely on DNA purified from ear, tail, toe clippings or other samples usually obtained by painful and potentially harmful procedures. 2 Many cost-effective and reliable DNA purification procedures have been designed to analyze these tissue samples, including the fast hot-alkaline lysis, 3 microwave-based, 4 and guanidinium/silica-based 5 procedures, or the standard time-consuming phenol:chloroform extraction and ethanol precipitation protocol.6,7
Although most available conventional DNA purification methods produce high-quality DNA, some of them are tailored to or work better with specific samples,8,9 whereas others have been adapted for minute samples as diverse as those found in a crime scene. 10 Among the fastest methods of DNA purification, the chelating resin (Chelex®)-based protocol of Walsh and collaborators 11 has gained special attention due to its ability to produce high-quality DNA from virtually any biological sample. Indeed, Chelex® can efficiently purify DNA from bacteria; 12 parasites; 13 insects; 14 arthropods; 15 mammalian cells in culture or blood; 5 sperms; 16 fixed, 17 frozen, 18 fresh and archival 19 specimens; and hair roots from primates or mice.20–22 However, in spite of the evident ample scope of Chelex®-based techniques, very few studies have used this resin for genotyping purposes in mice.16,22–24
The objective of this study was to design and apply a Chelex®-based method of purifying DNA suitable for the routine PCR-based genotyping of laboratory mice. To this end, we used DNA purified from hair roots and other tissues as templates. Our results demonstrate that mice can be accurately genotyped in a fast and reliable manner using hair follicle DNA purified by an easy to follow Chelex®-based approach. This non-invasive and inexpensive methodology may be applied to routine genotyping of mice and to replace traditional methods involving painful collection methods.
Materials and methods
Animals
The studies involving animals have been approved by the Institutional Animal Care and Use Committee (IACUC) of the University. Mice (Mus musculus) with genetic disruption in the somatic Slc12a1 (B6-Slc12a1tm2Haca), Agtr1a (B6.129P2-Agtr1tm1Unc/J), or the sex X-chromosome Ace2 genes were used in our experiments. These mice were provided by Drs Hayo Castrop (University of Regensburg, Regensburg, Germany), 25 Thomas Coffman (Division of Nephrology, Department of Medicine, Duke University, Durham, NC, USA), 26 and Coffman and Gurley (Division of Nephrology, Department of Medicine, Duke O’Brien Center for Kidney Research, Duke University, Durham, NC, USA). 27 Littermates of all possible individual Slc12a1, Agtr1a, and Ace2 allelic constitutions, i.e. male and female Slc12a1+/+, Slc12a1+/−, Slc12a1−/−, Agtr1a+/+, Agtr1a+/−, Agtr1a−/−, male Ace2X/Y (WT) and Ace2−/Y (KO), and female Ace2X/X (WT) and Ace2−/X (carriers), were tested. Animals had ad libitum access to water and chow and were housed under 12 h light and dark cycles. Sentinel mice in the colony were serologically tested for pathogenic agents. Mice were free of pathogenic agents, including ectromelia virus, epizootic diarrhea of infant mice virus, lymphocytic choriomeningitis virus, Mycoplasma pulmonis, mouse adenovirus strains 1 and 2, mouse hepatitis virus, mouse parvovirus, minute virus of mice, polyomavirus, pneumonia virus of mice, reovirus type 3, Theiler’s murine encephalomyelitis virus, sendai virus, endoparasites, and ectoparasites.
Tissue collection
Mice of either gender or genotype, aged 5–10 days, were tail-clipped according to the recommendations in the Guidelines for the genotyping of mice and rats (National Institutes of Health, Bethesda, MD, USA). Briefly, a minimum amount (∼1–3 mm) of tail skin, muscle and bone was excised using sterilized scissors after cleaning and sterilizing the tip of each mouse tail with 70% ethanol. After clipping, gentle pressure was applied to the cut site to stop any potential bleeding followed by application of styptic powder slightly moistened with 70% ethanol. Hair follicles were obtained from 2–4-week-old mice by plucking a small amount of hair (∼10–20 units) from the back of each mouse using small sterilized forceps. 28 To minimize the risk of cross-contamination of hair samples, mice were placed in small and clean individual cages before collecting the hair samples. The scruff of the neck was cleaned and decontaminated with 70% ethanol. Gloves and tweezers were also sprayed with 70% ethanol and dried using clean task wipers (Kimberly Clark® wipes, Kimberly-Clark Ltd, Kent, UK) prior to sampling. The liver was collected post-mortem to assess genotyping performance.
DNA purification
Two widely used and extensively available methods of purifying DNA from the tail clips or hair roots of mice were performed: cation chelating styrene divinylbenzene copolymer resins (Chelex® 100 method) or enzymatic digestion (referred to as the ‘Proteinase K’ method). All solutions used for genotyping were sterilized by autoclaving, filtering (0.22 µm) or prepared under sterile conditions.
Chelex® 100 method
Chelex® suspensions (10% w/v) were prepared under sterile conditions. Aliquots of 200 µL of the resin suspension were placed into sterile 1.5 mL tubes and stored at 4℃ until use. The DNA extraction procedure was based on original protocols published,11,29 and adapted to our needs. Tissue samples were placed in tubes containing 200 µL of 10% Chelex® (Sigma, St Louis, MO, USA) and quickly centrifuged for 30 s at 12,000 × g (Sorvall Legend Micro 21R centrifuge; Thermo Scientific, Waltham, MA, USA), leaving the tissue samples at the bottom of the tubes. The samples were then incubated at 100℃ for 20–40 min with casual swirling every 5–10 min and centrifuged. Accurate aliquots of the supernatants containing DNA were collected (50 µL for hair roots and 100 µL for tail clips) and placed into new tubes, which were then processed or stored at –20℃ until use.
Proteinase K method
Genomic DNA from samples was purified using the DNEasy blood and tissue kit (Qiagen, Germantown, MD, USA) according to the manufacturer’s instructions. DNA elution was performed using 100 µL (tails) or 50 µL (hair roots) of Tris-HCl/EDTA pH 8. Purified DNA samples using this method or Chelex® were assayed for concentration and purity index using a NanoDrop ND-1000 spectrophotometer (Thermo Sci/NanoDrop, Wilmington, DE, USA), and used immediately for genotyping or stored at –20℃ until use. Genomic DNA (50–150 ng) were used in PCRs.
PCR genotyping primers
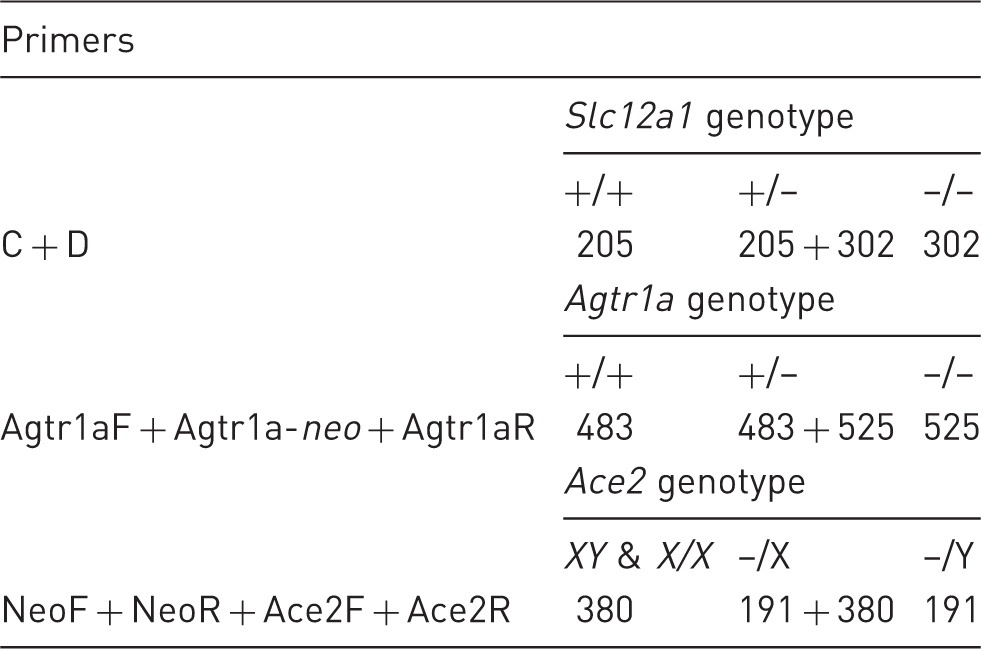
Genotyping primers: oligonucleotide primer names, sequence, and orientation are shown.

PCR design and amplicon analysis
Genotyping amplicons: indicated are primer duplexes or multiplexes and their predicted amplicon sizes in base pairs (bp) for the different genotypes.
Results
DNA purification and genotyping performance
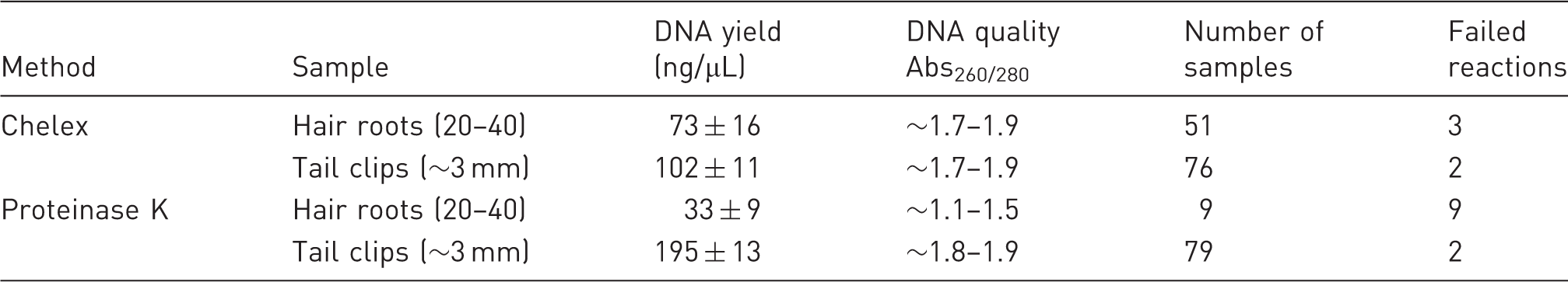
In the present study, two methods of purifying DNA, i.e. Chelex® and Proteinase K, from hair roots or tail clips were initially tested by determining the final concentration of purified DNA followed by assessment of its purity, as estimated by their 260 nm/280 nm absorbance ratio (Abs260/280). As shown in Table 3, both methods produced DNA of an adequate purity from tail clips. However, the Proteinase K digestion-based method of DNA purification produced low amounts of total DNA from hair roots (Figure 1B), which were deemed to be not suitable for genotyping (Abs260/280 < 1.5, Table 3). Instead, the Chelex® method produced enough total DNA from hair roots of the required purity (3.7 ± 0.8 µg, n = 51, Table 3) and integrity for genotyping purposes compared with the Proteinase K-purified DNA from hair roots (Figures 1C and D).
Comparative analysis of hair root and tail clip DNA purified using Chelex and the Proteinase K kit. (a) DNA yields (ng/µL, final volume 100 µL) from tail clips (∼3 mm) using Chelex® resins during the indicated time periods. (b) DNA yields (ng/µL, final volume 50 µL) from hair roots (20–40 units) obtained by using the indicated methods. (c) Representative 0.9% agarose gel electrophoresis of Chelex®-purified genomic DNA obtained from hair roots of Slc12a1 mice. Electrophoresis run was performed at 75 V for 90 min in a mini gel apparatus. (d) Representative 1% agarose gel electrophoresis of purified genomic DNA obtained from hair roots (left panel) and tail clips (right panel) of mice using the DNEasey kit. Performance: concentrations of DNA (ng/µL), quality (Abs260/280) and number of failed polymerase chain reactions of the indicated number of DNA samples obtained from hair roots and tail clips using Chelex®-based and Proteinase K-based methods are shown.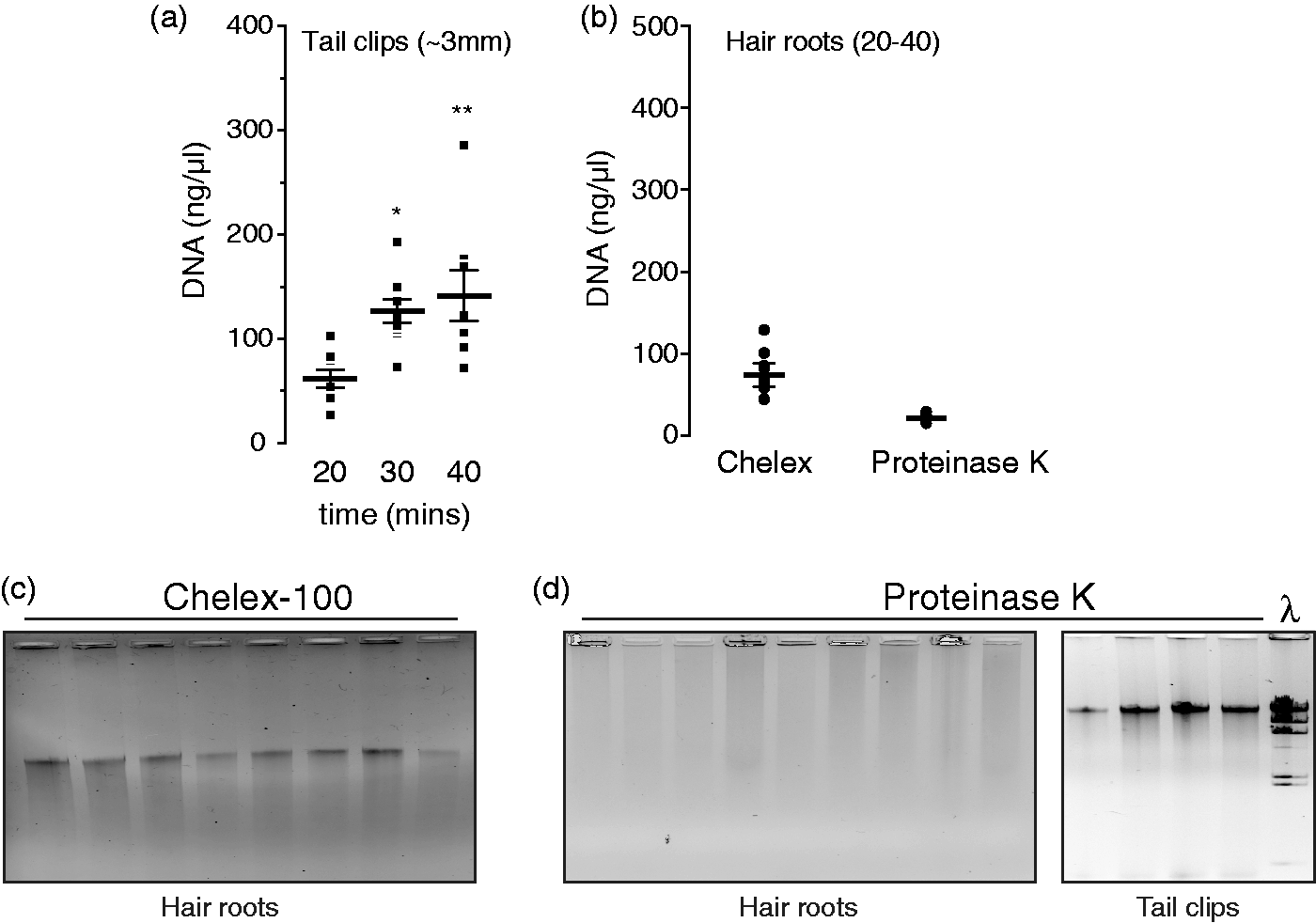
The absolute total yield of DNA, however, was lower when tail clips were subjected to Chelex® action for 20 min instead of overnight as with Proteinase K (∼10 µg versus ∼20 µg, Table 3). Next we determined if longer incubation times with Chelex® resulted in increased DNA yields. To this end, tail clips were incubated in 10% Chelex® at 100℃ for 20, 30 and 40 min, and DNA concentration quantified (total volume: 100 µL). As shown in Figure 1A, the absolute yield of DNA from tail clips incubated with Chelex® for 20 min was 6.2 ± 0.9 µg (n = 8) whereas yields after 30 or 40 min incubation time were 12.7 ± 1.2 µg (n = 9, *P = 0.023) and 14.2 ± 2.4 µg (n = 8, **P = 0.048), respectively. Together, these results indicate that Chelex® has the potential for purifying as much genomic DNA as the conventional Proteinase K method.
Since the quality of Chelex®-purified DNA from tail clips or hair roots was comparable, indicating that both samples were suitable for genotyping, we next determined the potential for producing diagnostic genotypes in Slc12a1 mice using Chelex®-purified DNA from hair roots or tail clips as templates. As shown in Figure 2, Chelex®-purified DNA from hair roots (Figure 2A) or tail clips (Figure 2B) produced comparable genotypes, highlighting the reliability of Chelex®-mediated DNA purification for genotyping and diagnostic purposes. To further validate these results, the genotype of Slc12a1 mice was determined using Chelex®-purified DNA from liver samples (1–2 mm3) obtained post-mortem. Representative results are shown in Figure 2C. There was 100% agreement in genotype results between the liver and hair root samples.
Genotyping validation of Chelex®-purified DNA from hair roots. (a–c) Representative genotyping polymerase chain reactions (PCRs) using Chelex®-purified genomic DNA from: hair roots (a), tail clips (b) or liver (c). As negative controls, Chelex® supernatants in the absence of tissues were used. The Slc12a1 genotypes are defined as homozygous (–/–, 302 bp), wild-type (WT; +/+, 205 bp) or heterozygous (+/–, 302 + 205 bp). (d) PCR genotyping of Ace2 mice from tail clips and hair roots purified using the DNEasy kit (top) and Chelex® (bottom). Shown are the predicted banding patterns corresponding to the following Ace2 genotypes: male knockout (KO; –/Y, 200 bp), female carrier (–/X, 200 + 400 bp) and female WT (X/X, 400 bp). (e–f) Representative genotyping PCR of Agtr1a mice using Chelex®-purified DNA obtained from tail clips (e) and hair roots (f). Shown is the amplicon pattern corresponding to the WT (+/+, 483 bp) and KO (–/–, 525 bp) genotypes.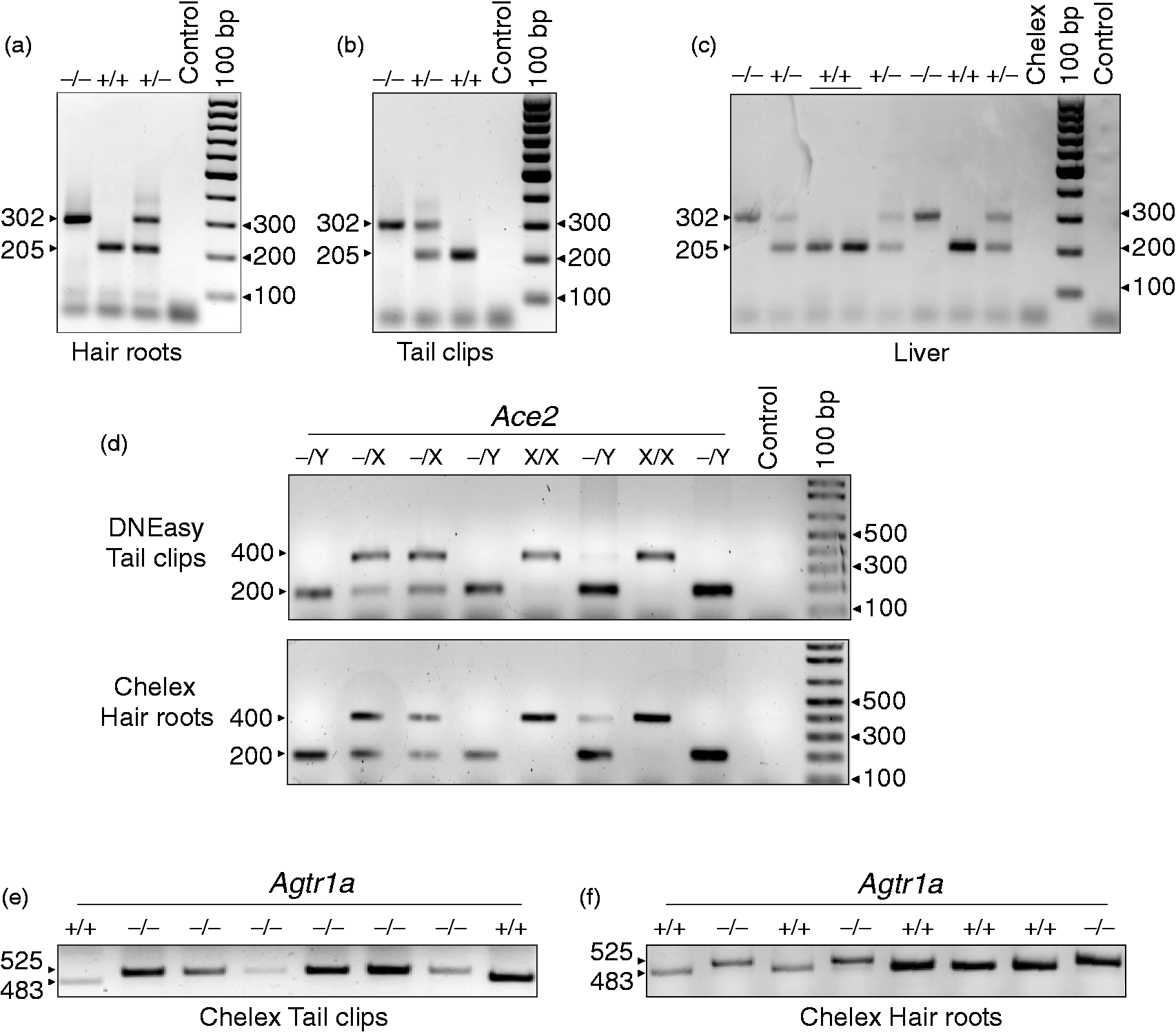
To test the newly developed method on other mouse genetic models wherein the genotyping PCR strategy uses a multiplex of three or four oligonucleotide primers (Table 2), we compared the genotypes of Agtr1a and Ace2 mice using DNA purified from hair roots and tail clips using Chelex® or the DNEasy kit, respectively. As seen in Figures 2D–F, both methods produced comparable results, indicating that Chelex® purification of genomic DNA from hair roots is a reliable, painless, simple, and efficient alternative to commercially available extraction methods.
In the case of PCRs involving hair root DNA purified using Chelex®, almost all of them gave diagnostic results. However, one out of every ∼100 PCRs was also positive for reactions containing no genomic DNA. This was observed typically in the form of a faint band of 205 bp corresponding to the Slc12a1+/+ genotype. Although these false-positives often do not preclude genotype diagnosis, e.g. when a single band of 302 bp is obtained from genomic DNA of Slc12a1 mice (Slc12a1−/−), repeating PCRs eliminated contamination, indicating that the most common source of contamination under our experimental conditions was DNA from the experimenter rather than from cross-contaminated hair follicles. To date, the Chelex®-based method has successfully determined 1272 genotypes from 1297 mice tested (∼98% effective, including tail clips). Notably, Chelex®-extracted DNA stored at –20℃ for a month, remained suitable for a second round of genotyping PCRs. In addition, >30 PCR genotyping reactions were performed using Chelex®-purified DNA from the hair roots of Agtr1 and Ace2 mice, demonstrating that this technique was not isolated to a particular genotype analysis and could handle different PCR genotyping protocols. By contrast, the DNEasy kit could not reliably produce DNA from hair roots that were suitable for PCR.
Discussion
Our results demonstrated that Chelex® resins may be used for purifying genomic DNA from hair roots, tail clips, and tissues for general genotyping using PCRs. As in any other PCR-based strategy, de novo genotyping designs may require optimization depending on the targets, locus selection, and other allelic characteristics, including guaninecytosine (GC) content, repetitive sequences, or melting temperature. Indeed, oligonucleotide design is particularly sensitive to these variables as well as others inherent to the oligonucleotide itself, including specificity, annealing temperature, secondary structure, or stability. 30 We designed several oligonucleotide primers based on widely used algorithms to amplify Slc12a1 alleles by targeting short (<350 bp) stretches of genomic sequences, and by using select sets that produced reliable and diagnostic results.
We successfully genotyped Slc12a1 mice from a wide variety of DNA samples, including feces and toe nails further demonstrating the sensitivity of the Chelex® assay (data not shown). Although Chelex® produced consistently low yields of total DNA from hair roots relative to tail clips (Table 3), as expected, the hair root genomic DNA was of an acceptable purity and therefore suitable for diagnostic genotyping (Figure 2). However, using the DNEasy kit to purify DNA from hair roots could not reproduce the results obtained with Chelex®. Although widely used for purifying DNA from a variety of biological samples, 1 including mouse hair roots, in our hands and in line with recent results, 31 the DNEasy kit produced purified hair root DNA of a low quality (Abs260/280 < 1.5), yield (∼3 µg) and concentration (<50 ng/µL) (Table 3, Figures 1B and D). Accordingly, PCR performance of our primer sets was compromised when using DNA isolated with DNEasy from hair roots. In fact, under our experimental conditions, most of the Slc12a1 PCR genotyping reactions performed using kit-purified DNA from hair roots did not produce reliable results (data not shown), indicating the need for a new strategy in order to keep costs at a minimum.
Clearly, the described DNA purification method using Chelex® may be extended to several tissue samples to produce DNA of a high purity. In fact, clean genotyping results can be obtained within 3 h with significant cost reductions. Indeed, when compared against commercially available enzymatic methods, which are lengthy (usually ∼16 h) and expensive, this is a very economic alternative. Thus, the high sensitivity of the Chelex® extraction of producing hair root DNA, which is more humane than collecting from a tail or ear section, coupled with the clear affordability and welfare benefits of Chelex®, makes this a suitable alternative to commercially available DNA extraction kits. Although several reliable affordable DNA extraction methods exist, besides DNEasy, no other technique has been demonstrated to work consistently with hair follicles. This study does not preclude those other systems from potentially working, in fact it highlights that any sensitive assay may potentially work on hair follicles. Since there is a higher risk of cross-contamination using hair samples, simple precautions should be taken, such as placing mice in small and clean individual cages before collecting hair samples, and cleaning and decontaminating the site of collection prior to sampling. Research laboratories using genetically-engineered mice should consider adapting and using this minimally invasive and less painful technique of collecting tissue for DNA genotyping as this would enhance animal welfare. This technique is particularly beneficial when PCR identification is not required before two weeks of age. Further, due to the non-invasive nature of the procedure, re-confirmation of questionable genotypes can now be easily determined in adult live mice. This eliminates the need for local or general anesthesia which is usually required prior to tail biopsy. Finally these procedures, alone or in combination, can alter the overall well-being of mice,32,33 and are also widely considered to be painful.34–37
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: We are grateful to the STREAMS program, funded by NHLBI (National Heart, Lung, and Blood Institute, NIH) (VO-R), NIH-NIGMS BioSTAR program R25-GM090122, Minority Undergraduate Internship Grant #1-12-MUI-03 (AB), GRAD-PREP program R25 GM086257 (LKM), and NIH 1P30DK096493-01 (SBG). Research was supported in part with funds provided by the following grants: NIH-NIDDK F32DK093226 (NG) and WSU-BSoM Research Incentive Fund 226132 (MDiF). We do not have any affiliation with Chelex®.
