Abstract
We present a specific and sensitive method for simultaneous detection of three mRNA species in individual neurons. The method relies on the use of riboprobes labeled with [35S]-UTP, digoxigenin-UTP, or biotin-UTP. The nonradioactive probes were sequentially revealed by incubation with anti-digoxigenin immunoglobulins or streptavidin conjugated to peroxidase, followed by the use of fluorochrome-labeled tyramides as peroxidase substrates. The radioactive probe was revealed by conventional autoradiography. There was no interaction among the different probes or the various detection systems. We demonstrate the use of this method by illustrating on laser scanning confocal microscopy the co-localization of the mRNAs coding for corticotropin-releasing factor (CRF), arginine vasopressin (AVP), or peptidylglycine α-amidating monooxygenase (PAM) in rat hypothalamic paraventricular nucleus (PVN) and its modulation by endogenous glucocorticoids. Our results suggest that this method could be used not only to study the regulation of the hypothalamo-pituitary-adrenal axis but also in various models in which mRNAs are present at low concentrations.
Keywords
I
Materials and Methods
Chemicals
Restriction enzymes were from Gibco BRL (Eragny, France) or MBI Fermentas (distributed by Euromedex; Strasbourg, France). T3 or T7 RNA polymerase was purchased from New England Biolabs (distributed by Ozyme; Montigny-le Bretonneux, France). RNasin and RNase-free DNase I were from Promega (Charbonnieres, France). ATP, GTP, CTP, UTP, digoxigenin-11-UTP, biotin-16-UTP, RNase A, tRNA, 5(6) carboxytetramethylfluorescein-N-hydroxysuccinimide ester, and anti-digoxigenin immunoglobulins (Fab fragments) coupled to horseradish peroxidase (HRP) were from Boehringer Mannheim (Meylan, France). 5(6) Carboxytetramethylrhodamine-N-hydroxysuccinimide ester was purchased from Sigma (St Quentin Fallavier, France). Tyramide was from Aldrich (L'Isle d'Abeau Chesnes, France). [α35S]-UTP (SA 1300 Ci/mmole) and streptavidin coupled to HRP were purchased from New England Nuclear (Les Ulis, France). The nuclear emulsion (K5) was from Ilford Anitec (St-Priest, France) and the photographic chemicals were from Kodak-Pathé (Paris, France). All other reagents (molecular biology grade, or the highest possible grade) were purchased either from Sigma or Aldrich.
Animals
Male Sprague-Dawley rats (180-200 g bw) were purchased from the Centre d'Elevage R. Janvier (le Genest St. Isle, France). They were housed in our laboratory under controlled temperature (22–24C) and a constant 12-hr light/ dark cycle for at least 3 weeks before the experiment. They had free access to standard rat chow and tapwater. All experimental procedures were performed in accordance with local animal use regulations; studies were approved by the University Committee on the Use and Care of Animals. Bilateral sham operation or adrenalectomy (ADX, n = 4 in each group) was performed under ether anesthesia using a dorsal approach. ADX rats had free access to 0.9% NaCl in tapwater. One week later animals were sacrificed by decapitation between 1000 and 1100 hr. Brains were carefully removed, immediately frozen on dry ice, and stored at -70C until sectioning.
Probes
The CRF probe was a 770-
In Situ Hybridization
Coronal sections (6 μm) through the hypothalamic PVN were cut in a cryostat microtome at-20C. They were thawmounted onto gelatin twice-coated slides, dried on a slide warmer, and kept at -70C. The sections were warmed at room temperature (RT) and fixed in 4% formaldehyde in PBS, pH 7.2, for 5 min. After two washes in PBS, they were placed in 0.25% acetic anhydride in 0.1 M triethanolamine/ 0.9% NaCl, pH 8, for 10 min and delipidated in ethanol and chloroform. They were hybridized for 20 hr at 56C with a buffer containing 10 mM Tris, pH 7.4, 1 mM EDTA, 600 mM NaCl, 50% (v/v) formamide, 10 % (w/v) dextran sulfate, 25 μg/ml yeast tRNA, 1 X Denhardt's solution, 0.1 M DTT, and 1.5 X 107 dpm/ml CRF probe, 20 μl/ml AVP probe, and 40 μl/ml PAM probe under a glass coverslip (these various concentrations of probes have been determined in pilot experiments to saturate the mRNA to be hybridized and to give the lowest possible background). All the subsequent steps were performed at RT unless otherwise specified. Coverslips were removed in 2 X SSC (1 X SSC is 0.15 M NaCl, 0.015 M sodium citrate, pH 7.2). The sections were washed in 2 X SSC for 30 min, treated with RNase A (10 μg/ml in 2 X SSC) for 30 min at 30C, and subsequently washed in 1 X SSC/10 mM β-mercaptoethanol (β-ME) twice for 10 min, 0.5 X SSC/10 mM β-ME for 10 min, 0.1 X SSC/10 mM β-ME for 10 min, 0.1 X SSC/10 mM β-ME twice for 30 min at 65C, and finally in 0.1 X SSC/10 mM β-ME for 10 min. Slides were subsequently incubated for 10 min in Tris-buffered saline (0.1 M Tris, 0.9% NaCl, pH 7.4) containing 0.1% Triton X-100 (TBST) and then for 1 hr in TBST containing 3% normal sheep serum (NSS). Slides were incubated overnight in TBST containing 1% NSS and the anti-digoxigenin immunoglobulins conjugated to HRP, diluted 1:100. Slides were subsequently washed three times for 10 min in TBST and incubated for 10 min in 0.2 M Tris, 10 mM imidazole, 0.01% H2O2, pH 8.8 (TIH) containing 50 μM tyramide coupled to fluorescein (FITC). After three 5-min washes in TBST, the HRP was inactivated by incubation for 15 min in 0.1 M HCl/0.9% NaCl, and slides were further washed three times for 5 min in TBST. Then the slides were incubated for 30 min at 37C in TBS containing 0.5% DuPont blocking reagent (TBSB) and further incubated for 30 min at 37C in TBSB containing the streptavidin conjugated to HRP diluted 1:500. After three 5-min washes in TBST slides were incubated for 10 min in TIH containing 50 μM tyramide coupled to rhodamine (TRITC) and washed three times for 5 min in TBST. Then the slides were rapidly dipped in distilled H2O, followed by 70% ethanol, and dried under a stream of warm air. They were subsequently dipped in nuclear emulsion diluted 1:2 in H2O and exposed for 8 days at 4C. Slides were developed for 4 min in D-19 diluted 1:2, fixed for 5 min in Unifix, washed, and coverslipped with mowiol.
Controls included hybridization with the sense probes, inactivation of the HRP before incubation with the tyramine coupled to fluorochromes, or omission of the anti-digoxigenin immunoglobulins or the streptavidin coupled to HRP.
Confocal Microscopy
Specimens were viewed under a Leica TCS laser scanning confocal microscope coupled to a DMR inverted microscope equipped with planachromatic 20/0.4 and planapochromatic 63/1.4 objectives (Heidelberg, Germany). The argon-krypton laser generated excitation bands at 488 nm for FITC, 568 nm for TRITC, and light for differential interference contrast to detect autoradiographic silver grains. FITC and TRITC signals were recovered through bandpass filters centered at 520 and 600 nm, respectively. Simultaneous threechannel scanning was performed, adjusting the pinhole diameter to make optical sections about 1 μm thick, giving acceptable resolution on the z-axis for both the fluorescent and the autoradiographic focal planes. The power of the laser beam and the gain of each photomultiplier were optimized to minimize the injection of the signal from one channel to another. Leica Scanware software was used to store confocal images, which were transferred to a Power Macintosh computer (Apple Computer) and prepared with Photoshop software (Adobe Systems). The silver grains were inverted to white to improve visualization. Final images were printed on a Codonics NP 1600 printer.
Results
To determine the specificity of our technique, we ran several control experiments using sections obtained from sham-operated rats. First, we tested a possible interaction among the various probes or among the different revelation systems. Serial sections through the hypothalamic supraoptic nucleus [SON, which is known to contain both AVP and PAM mRNAs (Grino et al. 1990), but is devoid under basal conditions of CRF mRNA (Watts 1992)], were hybridized with AVP and/or PAM anti-sense probe and fully processed. PAM mRNA, which appeared as green (FITC) fluorescence was distributed over the entire SON, as previously described using isotopic ISHH (Grino et al. 1990). PAM mRNA was predominantly located in the cytoplasm, whereas the nucleus showed a weaker, punctate signal (Figures 1 and 6). AVP mRNA, which appeared as red (TRITC) fluorescence, was present only in the cytoplasm and was restricted to the ventral portion of the SON, as previously described using isotopic ISHH (Young et al. 1986; Figure 2). When sections were hybridized with both probes, the same distribution pattern of AVP and PAM mRNAs was observed and the co-localization of AVP and PAM mRNAs appeared as yellow to orange fluorescence (Figure 3). Second, we hybridized the sections with both the AVP and the PAM anti-sense probes. Slides were processed as described in Materials and Methods except that the anti-digoxigenin immunoglobulins coupled to HRP or the streptavidin coupled to HRP were omitted. Under this condition, the corresponding FITC or TRITC signals could not be detected (data not shown). Third, slides were hybridized with both the AVP and the PAM anti-sense probes and fully processed, except that the HRP coupled to the anti-digoxigenin immunoglobulins or the HRP coupled to streptavidin was inactivated with HCl before incubation with tyramide coupled to FITC or with tyramide coupled to TRITC, respectively. Under these conditions, the fluorescent signal corresponding to the AVP (Figure 4) or to the PAM probe (Figure 5) was not detected, demonstrating that incubation with HCl was effective in inactivating the HRP without affecting the subsequent steps. When SON sections were hybridized with AVP, PAM, and CRF probes and dipped in nuclear emulsion, fluorescent signals were comparable to those obtained in the previous experiment, and silver grains were sparse and randomly distributed, demonstrating the lack of interaction between the nonradioactive and the isotopic detection of the signals (Figure 6). Finally, PVN sections were hybridized with the AVP, PAM, and CRF sense probes and fully processed. As illustrated in Figure 7, only a faint, scattered green signal could be detected. Concomitantly, silver grains were sparse and randomly distributed all over the field, and most probably represented the background noise of the nuclear emulsion. When PVN sections of shamoperated rats were hybridized with the three antisense probes, four different populations of cell bodies were detected in the medial parvocellular portion (Figures 8 and 10). Cells labeled for AVP and PAM (scattered magnocellular cells), cells labeled for CRF (which appeared covered by grain clusters) and PAM (CRF+/AVP- hypophysiotrophic cells), cells labeled for AVP, CRF, and PAM (CRF+/AVP+ hypophysiotrophic cells), and cells labeled only for PAM (which may synthesize other neuropeptides than CRF and AVP). After ADX, no cell labeled only for CRF and PAM could be detected, and all the cell bodies that contained CRF and PAM mRNAs expressed AVP mRNA (Figures 9 and 11).
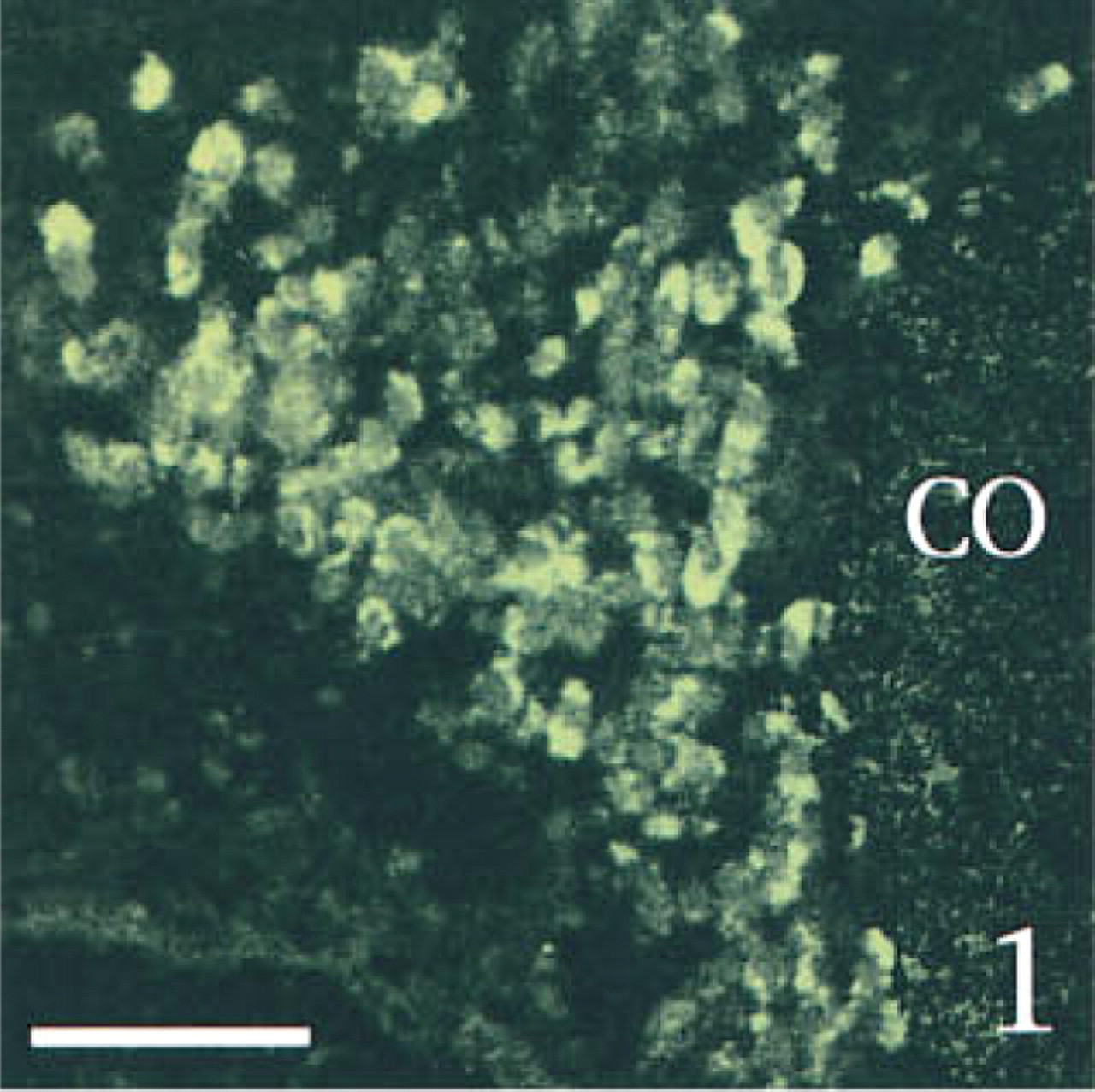
Two-channel confocal scanning of serial sections through the SON hybridized with the PAM anti-sense probe ( Figure 1 , FITC labeling), the AVP anti-sense probe ( Figure 2 , TRITC labeling), or both probes, showing the co-localization of both labelings as a yellow to orange fluorescence ( Figure 3 ). Bars = 100 μm.
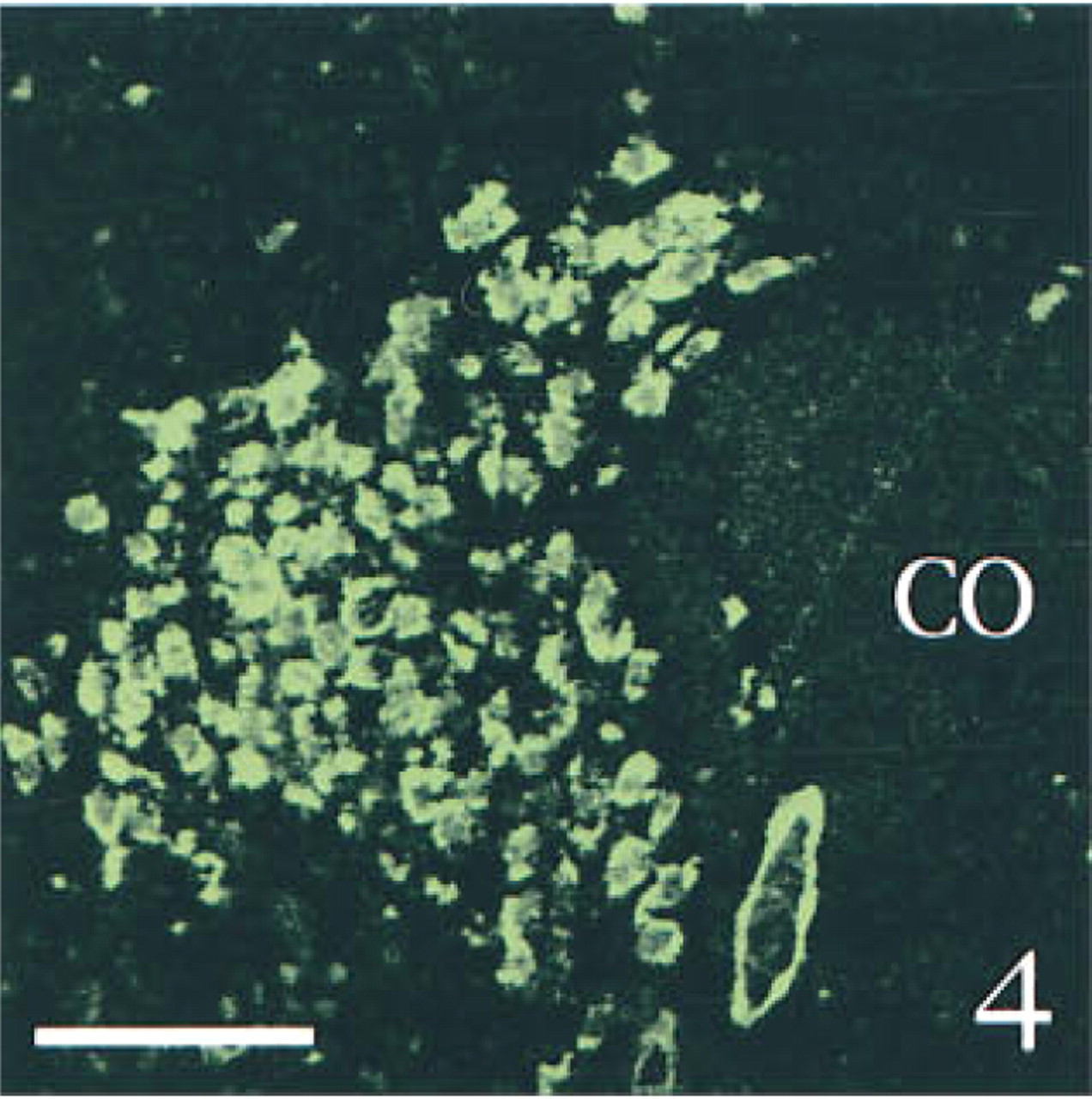
Two-channel confocal scanning of serial sections through the SON, hybridized with both the PAM and the AVP anti-sense probes. ( Figure 4 ) The section was treated with HCl (to inactivate HRP coupled to biotin) before incubation with tyramide coupled to TRITC; consequently, only FITC fluorescence (corresponding to PAM mRNA labeling) is detected. ( Figure 5 ) The section was treated with HCl (to inactivate HRP coupled to the anti-digoxigenin immunoglobulins) before incubation with tyramide coupled to FITC; consequently, only TRITC fluorescence (corresponding to AVP mRNA labeling) is detected. Note that the lower right corner of Figures 4 and 5 shows the autofluorescence in the FITC range of the same venule. Bars = 100 μm.
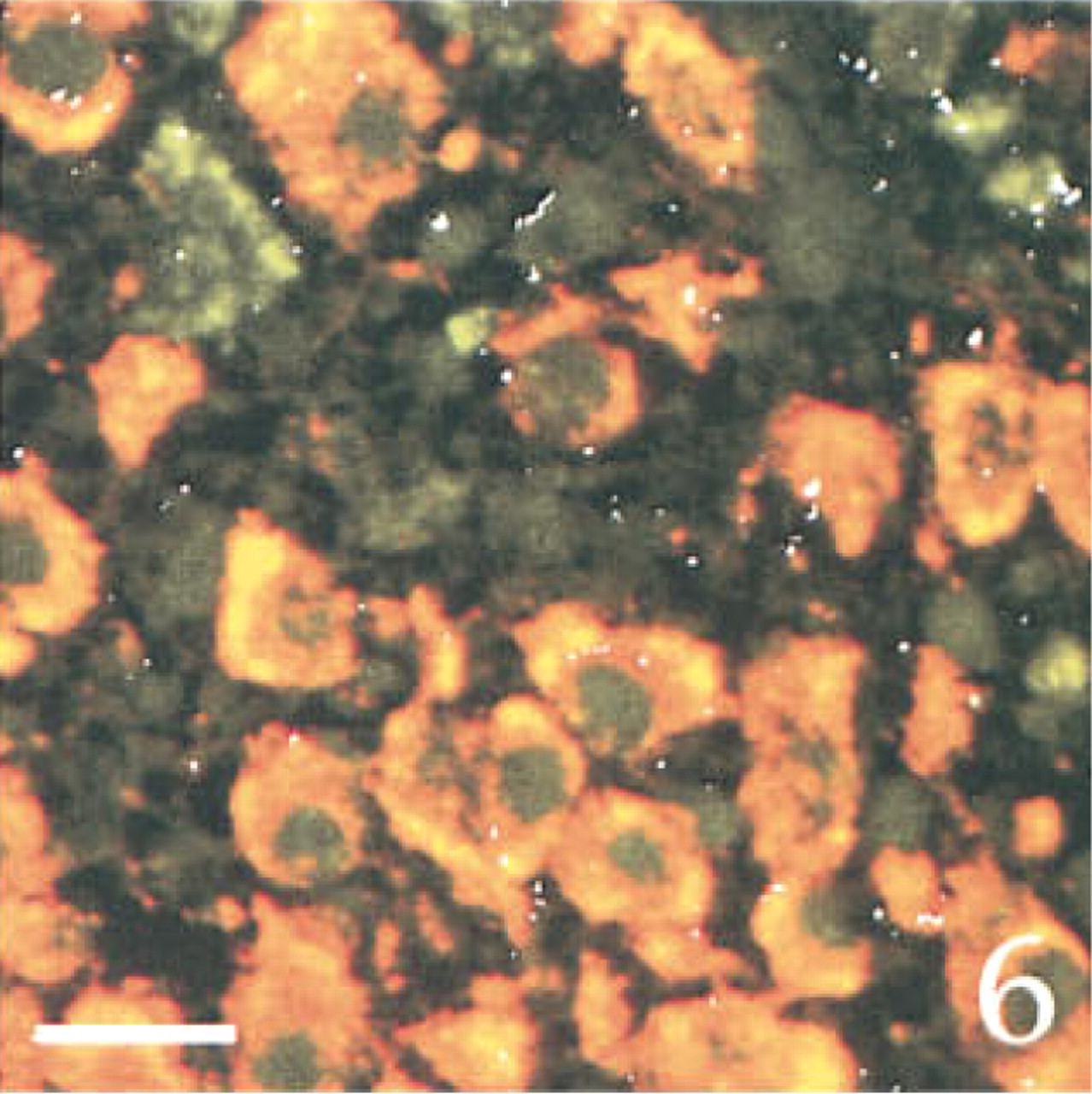

Evidence for a lack of interaction between the various detection systems of the triple in situ hybridization method. The figure shows a three-channel confocal scanning of a section through the ventral region of the SON hybridized with both the PAM (that shows FITC labeling), the AVP (that shows TRITC labeling), and the CRF (that shows autoradiographic grains inverted to white color for better vizualization) anti-sense probes. A large majority of cell bodies co-localized PAM and AVP labeling. The autoradiographic grains were sparse, randomly distributed, and mainly located outside of cells, consistent with the lack of CRF expression in the SON. Note the cell nuclei clearly labeled with the PAM probe. Bar = 10 μm.
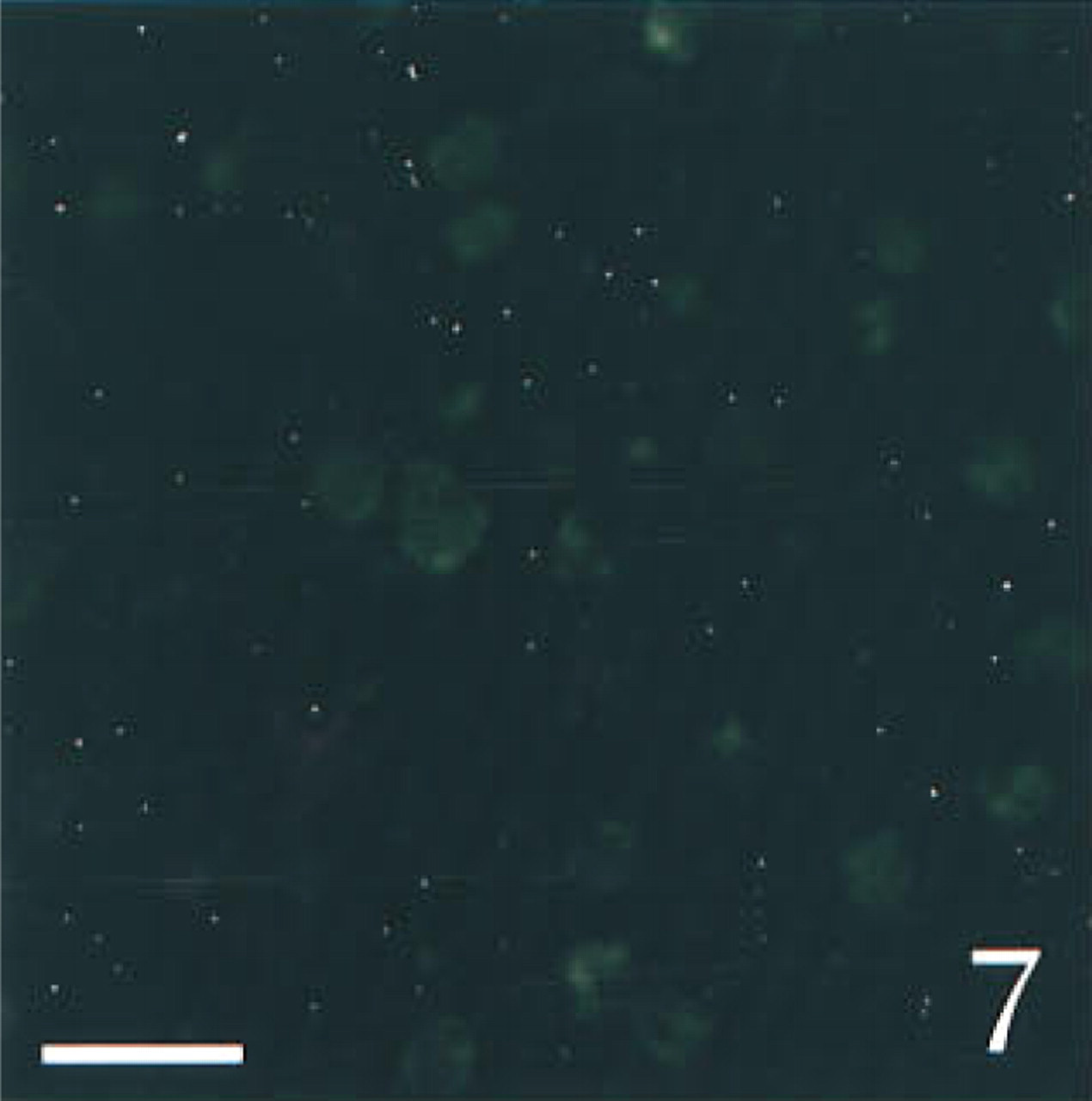
Three-channel confocal scanning of a section through the PVN hybridized with PAM, AVP, and CRF sense probes. Only a faint green fluorescent background is detected. The silver grains (white) are sparse and randomly distributed, demonstrating the specificity of the probes. Bar = 10 μm.
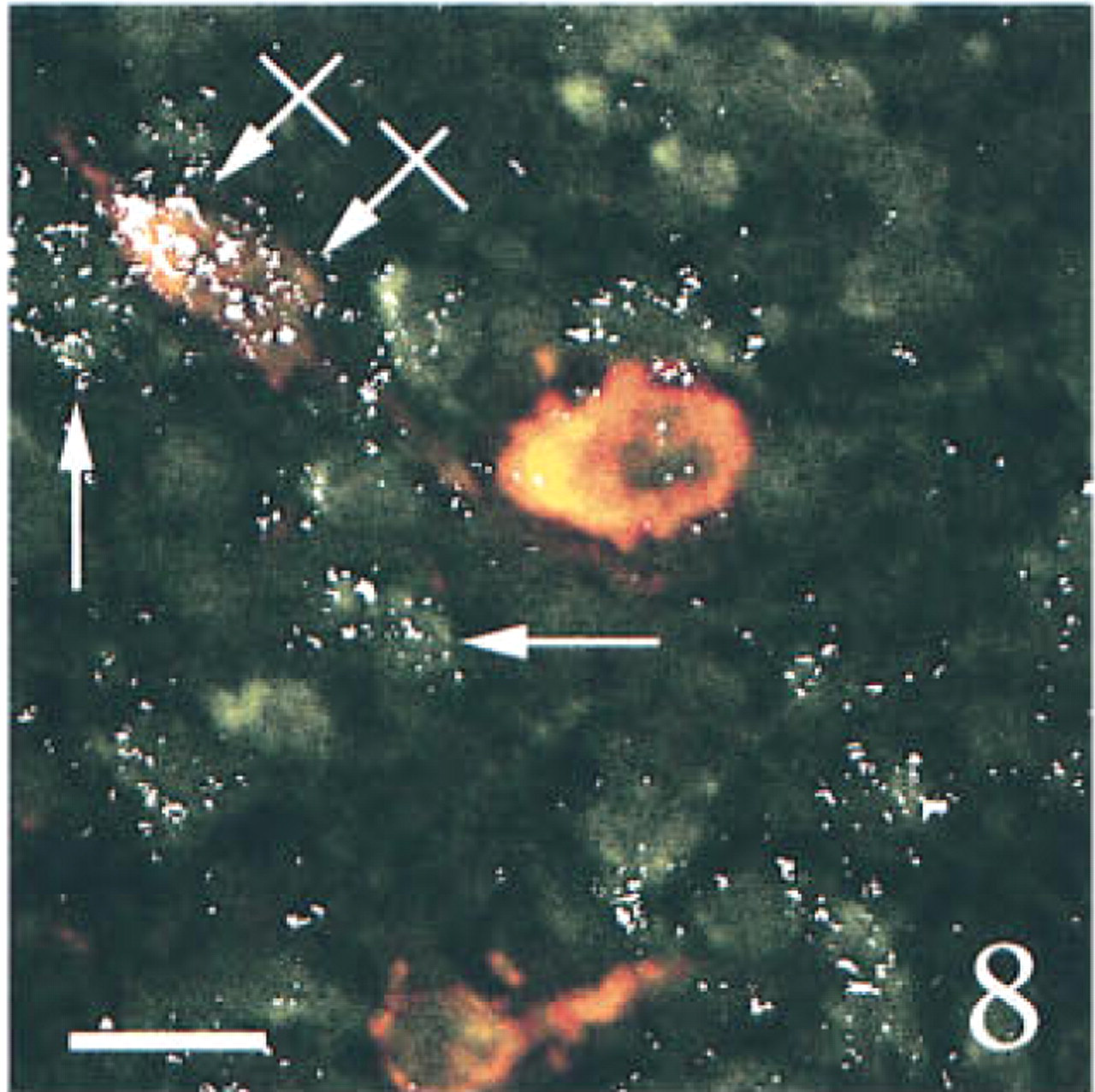
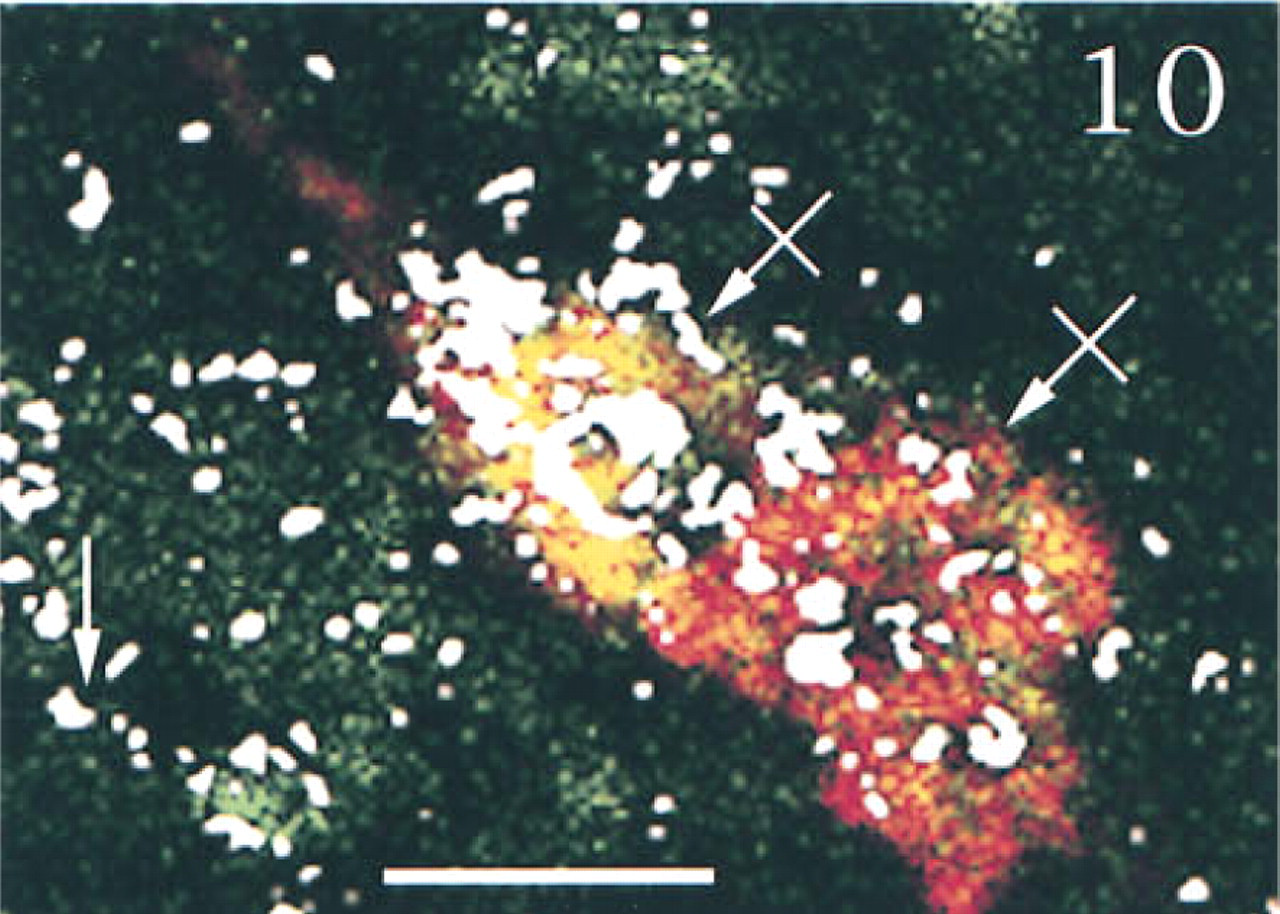
Three-channel confocal scanning of sections through the medial parvocellular portion of the PVN obtained from a sham-operated ( Figure 8 ) or an adrenalectomized ( Figure 9 ) rat, hybridized with PAM, AVP, and CRF anti-sense probes. Parvocellular triple labeled (crossed arrows), double labeled (for CRF and AVP, arrows), or single labeled (for PAM, arrowhead) cells are shown. The scattered magnocellular neurons display only a yellow to orange labeling after the expression of PAM and AVP mRNAs. Note the increased frequency of triple labeled cells after adrenalectomy. Bars = 10 μm.

Discussion
This study describes a new method for labeling three distinct mRNA species within a highly heterogeneous population of neurons. Our results show that this technique is specific and sensitive, and that the different markers are well distinguishable, allowing their co-localization to be easily detected.
The specificity of the probes and the revelation systems was assessed in several ways. Sections hybridized with sense probes did not show any signal above background, demonstrating the specificity of the hybridization step. Such specificity was most probably due to the perfect homology between the probes and the mRNAs to be detected and to the high stringency of the hybridization (50% formamide, 56C) and washing (0.1 X SSC, 65C). There was no interaction between the two steps of fluorescent detection of the nonradioactive probes. Incubation with HCl after revelation of the first (FITC) fluorescent signal was effective to inactivate the HRP coupled to the anti-digoxigenin immunoglobulins. This was demonstrated by the lack of signal when the sections were treated with HCl before incubation with tyramide coupled to fluorescein. In addition, hybridization of SON sections with the AVP or the PAM probe or with both probes revealed the same distribution pattern for AVP and PAM mRNAs. Finally, the lack of silver grain clusters in triple labeled sections through the SON, a nucleus that is, under basal conditions, devoid of CRF mRNA (Watts 1992), demonstrated a lack of interaction between the fluorescent signals and the autoradiographic emulsion.
The high sensitivity of our triple labeling method is illustrated by the relatively intense PAM and AVP hybridization signals in the parvocellular cell bodies of the PVN. These two mRNAs are known to be expressed at low levels in the parvocellular PVN compared with the magnocellular PVN (Grino et al. 1990). Such a level of sensitivity was most probably related to the use of riboprobes. It is known that riboprobes give stronger signals than the commonly used oligonucleotides (Marks et al. 1992). Pilot experiments showed that hybridization with oligonucleotide probes, followed by fluorescent detection, gave much lower signals, PAM mRNA being at the limit of the detection level. However, we found that the intensity of the PAM hybridization signal was increased by reducing the mean average size of the PAM riboprobe from 3800
A triple labeling ISHH of HRP-labeled oligonucleotides to centromeric regions of human chromosomes has been recently described by van Gijlswijk et al. (1997). However, this technique requires several rounds of hybridization and washing. Taking into account the minimal length needed for hybridization (about 16 hr; Cox et al. 1984), washing, and signal detection, such a protocol does not appear suitable for ISHH of mRNAs using riboprobes. One advantage of our experimental protocol is that it allows hybridization of tissue sections in a single round. The validity of our technique to detect three mRNAs in the same cell body is illustrated by the co-localization of AVP, CRF, and PAM mRNAs. It is known that AVP and CRF mRNAs (Wolfson et al. 1985; Paulmyer-Lacroix et al. 1994a,b) and immunoreactivity (Sawchenko et al. 1984; de Goeij et al. 1992) are co-localized in the parvocellular PVN cells and that amidation of their COOH-terminus, which is catalyzed by PAM, is necessary for their biological activity (Eipper et al. 1992), strongly suggesting that the PAM protein is synthesized in CRF- and AVP-containing cell bodies. However, there was thus far no direct proof of this co-localization. Our results clearly show that the mRNAs coding for the abovementioned neuropeptides and the amidating enzyme are present in the same cell bodies, both in control and in adrenalectomized rats.
Finally, in the present study we have used laser confocal microscopy. The simultaneous capture for several signals implies the risk for capturing a part of the signal of one channel through the other. This drawback is well controlled under laser confocal microscopy because both gain and offset of photomultipliers can be accurately adapted to the emission intensity, and high-order bandpass filters can be wedged very near the peak emission frequency of the corresponding fluorochrome. However, conventional fluorescence microscopy with good filters sets and dichroics and a high quality CCD-video camera, together with postacquisition pseudo-colorization and overlays, could provide good results.
In conclusion, this work demonstrates for the first time that two nonradioactive and one radioactive ISHH can be easily associated to allow detection of three mRNA species in the same cell body by laser scanning confocal microscopy. This technique is specific, sensitive, and can be performed in a single round of hybridization. Our results strongly suggest that our method could be used not only to study the regulation of the hypothalamo-pituitary-adrenal axis but also in various models in which mRNAs are present at low concentrations.
