Abstract
Catalase, the classical peroxisomal marker enzyme, decomposes hydrogen peroxide and is involved in the antioxidant defense mechanisms of mammalian cells. In addition, catalase can oxidize, by means of its peroxidatic activity, a variety of substrates such as methanol and ethanol, producing the corresponding aldehydes. The involvement of brain catalase in the oxidation of ethanol is well established, and severe afflictions of the CNS in hereditary peroxisomal diseases (e.g., Zellweger syndrome) are well known. Whereas the distribution of catalase in the CNS has been investigated by enzyme histochemistry and immunohistochemistry (IHC), very little is known about the exact localization of catalase mRNA in brain. Here we report the application of a tyramine/CARD (catalyzed reporter deposition)-enhanced nonradioactive in situ hybridization (ISH) protocol for detection of catalase mRNA in sections of perfusion-fixed, paraffin-embedded rat brain. Catalase mRNA could be demonstrated in a large number of neurons throughout the rat brain as a distinct cytoplasmic staining signal with excellent morphological resolution. Compared to our standard ISH protocol, the CARD-enhanced protocol for catalase mRNA detection in rat brain showed higher sensitivity and significantly better signal-to-noise ratio. In parallel IHC experiments, using an antigen retrieval method consisting of combined trypsin digestion and microwave treatment of paraffin sections, the catalase antigen was found as distinct cytoplasmic granules in most catalase mRNA-positive neurons. In addition, catalase-positive granules, presumably peroxisomes, were found by confocal laser scanning microscopy in glial cells, which were identified by double labeling immunofluorescence for GFAP and CNPase for astroglial cells and oligodentrocytes, respectively. The excellent preservation of morphology and sensitive detection of both mRNA and protein in our preparations warrant the application of the protocols described here for systematic studies of catalase and other peroxisomal proteins in diverse pathological conditions such as Alzheimer's disease and aging.
Keywords
C
Whereas cultured neurons and astroglial cells contain both catalase and the glutathione system for peroxide detoxification (Dringen et al. 1999; Cimini et al. 2000), their importance varies in the different cell types. Catalase is much more important for clearance of H2O2 in neurons, whereas the glutathione system appears to play a more significant role in glial cells (Dringen et al. 1999). The importance of catalase for the overall function of neurons is further supported by the recent discovery of the role of H2O2 in regulation of dopamine release (Chen et al. 2001).
Catalase has been visualized by the alkaline diaminobenzidine (DAB) reaction (Fahimi 1969) in peroxisomes of both neuronal and glial cells (Holtzman et al. 1973; Holtzman 1982). Because of their small size, they were referred to as microperoxisomes (McKenna et al. 1976; Arnold and Holtzman 1978), emphasizing that their distinction in regular EM preparations from synaptic vesicles was quite difficult (Holtzman et al. 1973). Catalase antigen and other peroxisomal proteins have also been localized by IHC in the CNS (Houdou et al. 1991,1993; Imamura et al. 1994). Using the more sensitive catalyzed reporter deposition (CARD) technique, Moreno et al. (1995) described the mapping of catalase in different brain regions, which corresponds to the distribution of peroxisomal acyl-CoA oxidase recently reported by the same group (Cristiano et al. 2001; Farioli-Vecchioli et al. 2001).
The mRNA encoding for catalase has been demonstrated by Northern blotting in different rat brain regions (Tsay et al. 2000), and the alterations of mRNAs for peroxisomal β-oxidation enzymes have been investigated in developing mouse brain by RT-PCR (Knoll et al. 2000). Reimer and Singh (1990) studied the distribution of catalase mRNA in rat embryos by radioactive ISH and mentioned briefly the presence of catalase transcripts in CNS, without any further details as to the brain regions or cell types.
We recently described a nonradioactive ISH protocol for detection of mRNAs encoding peroxisomal matrix and membrane proteins (Schad et al. 1996; Baumgart et al. 1997,2001). In preliminary attempts to use that protocol for detection of catalase transcripts in rat brain, we noted that, because of the low expression of catalase in brain, the signal-to-noise ratio was rather low and amplification of the signal was necessary. We now report the distribution of catalase mRNA in the CNS of adult rats using a nonradioactive ISH protocol in conjunction with signal amplification by the CARD technique (Adams 1992; Speel et al. 1999). In parallel IHC experiments we have used confocal laser scanning microscopy and an antigen retrieval method (Grabenbauer et al. 2001), and detected the catalase antigen as distinct cytoplasmic granules, presumably peroxisomes, in most catalase mRNA-positive neurons. In addtition, catalase signal was found in astroglial cells and in oligodendrocytes identified by double immunofluorescence for GFAP (glial fibrillary acidic protein) and CNPase (2',3'-cyclic nucleotide 3'-phosphohydrolase), respectively (Kennedy et al. 1980; Sprinkle 1989).
Materials and Methods
Tissue Preparation
Adult male Sprague–Dawley rats weighing 220–250 g were perfused under chloral hydrate anesthesia via the abdominal aorta. After a 30-sec rinse with physiological saline, a fixative containing 0.25% glutaraldehyde, 2% sucrose in 0.1M PIPES buffer (pH 7.4) was perfused for 5 min (for the IHC experiments, 4% freshly prepared paraformaldehyde in the same buffer was used instead of glutaraldehyde). The brains were cut into 1–2-mm frontal or sagittal slices and were embedded in paraffin using an automated vacuum tissue processor (Shandon; Frankfurt, Germany). For experiments, 4-μm sections were cut and mounted on Super Frost Plus slides (Menzel; Braunschweig, Germany).
Preparation of Digoxigenin-labeled Riboprobes
Digoxigenin-labeled riboprobes were produced by in vitro transcription of specific DNA fragments obtained by PCR amplification of plasmid vectors containing the cDNA for rat catalase in a pGEM 7Zf (-) vector and GAPDH in a pBluescript II vector (Schad et al. 1996). For PCR amplification, primer pairs were used that were directed against the vector sequences flanking both oppositely located RNA polymerase promoter sites to amplify the insert with both promoters for transcription of both strands using the same DNA preparation (pBSK-s, 5'-GGAAACAGCTATGACCATGATTACGC-3'; pBSK-r, 5'-TGTAAAACGACGGCCAGTGAGCGC-3'; pGEM-r, 5'-ATTTCACACAGGAAACAGCTATGACC-3'; and pGEM-s, 5'-AGTCACGACGTTGTAAAACGACGGC-3').
In vitro transcription was performed using a digoxigenin labeling kit from Roche Diagnostics (Mannheim, Germany) and the resulting sense and antisense transcripts were cleaved to an average length of 200 bases by alkaline hydrolysis before their use in the ISH experiments.
In Situ Hybridization
Dewaxed and rehydrated paraffin sections were subjected to prehybridization steps including deproteination with 0.1 M HCl for 10 min followed by two PBS (137 mM NaCl, 2.7 mM KCl, 10 mM phosphate buffer, pH 7.4) rinses for 5 min each. Proteinase K digestion was carried out at 37C for 30 min at a concentration of 10–30 μg/ml in a TE buffer (100 mM Tris, 50 mM EDTA, pH 8.0), followed by postfixation in 4% freshly depolymerized paraformaldehyde in PBS. After two PBS rinses the sections were acetylated for 20 min with 0.25% (v|v) acetic anhydride in a TEA buffer (0.1 M triethanolamine, pH 8.0), rinsed in PBS, dehydrated, and air-dried. Before the hybridization the sections were prehybridized for 2 hr at 45C with a mixture consisting of 50% formamide, 50 mM Tris, 25 mM EDTA, 20 mM NaCl, 250 μg/ml yeast tRNA, and 2.5 × Denhardt's solution. Hybridization was carried out overnight at 45C with 5 ng/μl riboprobe in a solution containing 50% formamide, 20 mM Tris, 1 mM EDTA, 0.33 M NaCl, 0.5 μg/μl tRNA, 0.1 μg/μl poly-A RNA, 10% dextran sulfate, and 1 × Denhardt's solution. For each section, 20 μl of this hybridization mix, covered with “Hybri-slips” (Sigma; Deisenhofen, Germany), was used.
The posthybridization washes included a 30-min rinse at 53C in 2 × SSC, followed by 1 hr at 53C with 1 × SSC and 50% formamide. Then the sections were passed through two changes of 0.5 × SSC and 0.2 × SSC for 10 min each at room temperature. After a 10-min rinse in TBS (150 mM NaCl, 100 mM Tris, pH 7.5), nonspecific protein-binding sites were blocked by a 30-min incubation with a blocking buffer containing 1% (w|v) blocking reagent (Roche Diagnostics) and 0.5% (w|v) BSA in TBS buffer.
For detection of the hybridized probe, the sections were incubated overnight with peroxidase-labeled Fab fragments directed against digoxigenin diluted 1:100 in TNB buffer [0.5% (w|v) blocking medium (NEN-Life Science; Cologne, Germany) in TBS]. After two 5-min washing steps in TNT buffer (0.05% Tween-20 in TBS), a 30-min period in TNB buffer, and another two 5-min changes in TNT buffer, the biotinylated tyramine was applied to the sections as a 1:50 dilution in amplificant diluent (NEN-Life Science) for 20 min. After three 5-min changes of TNT buffer the sections were incubated with a 1:100 dilution of streptavidin–HRP conjugates in TNB buffer for 30 min. The color reaction for visualization of the peroxidase activity was performed after three 5-min rinses with TNT buffer using AEC as the substrate for peroxidase for 5 min. Afterwards the sections were rinsed in distilled water, counterstained with hematoxylin, and mounted with glycerol–gelatin. Negative controls were prepared using the corresponding sense probes.
Some parallel sections, after hybridization and posthybridization as described, were incubated with anti-digoxigenin Fab fragments conjugated with alkaline phosphatase instead of peroxidase to obtain a direct NBT/BCIP color reaction without amplification, as described previously (Schad et al. 1996). The originally reported NBT/BCIP incubation time of 4–24 hr had to be extended to 24–48 hr to obtain a minimal signal.
Immunohistochemistry
IHC for detection of catalase was performed on paraffin sections of rat brain samples fixed by perfusion with a fixative containing 4% freshly prepared paraformaldehyde in 0.1 M PIPES buffer (pH 7.4). For antigen retrieval, the protocol recently described by Grabenbauer et al. (2001) was used. Briefly, the deparaffinized and rehydrated sections were subjected to a 5-min digestion with 0.01% trypsin (Sigma) in PBS and an additional microwave treatment in 10 mM citrate buffer (pH 6.0) for 15 min at 720 W, followed by 20-min cooling in the same buffer. For inhibition of endogenous peroxidase activity, the sections were treated with 3% H2O2 for 5 min. Nonspecific binding sites were blocked with 0.5% blocking reagent (NEN-Life Science). The sections were incubated overnight at 4C with a polyclonal anti-catalase antibody diluted 1:500 in the blocking buffer. The antibody has been characterized previously by Beier et al. (1988). After incubation with a biotinylated secondary antibody, the antigenic sites were visualized using a peroxidase-conjugated biotin–streptavidin system (extravidin; Sigma) in conjunction with the Nova Red (Vector Laboratories; Burlingame, CA) peroxidase substrate, which was applied to the sections for 4 min. Afterwards the sections were counterstained with hematoxylin for 1 min, dehydrated, and mounted with Eukitt (Kindler; Freiburg, Germany).
Double-labeling Immunofluorescence
For simultaneous detection of catalase antigen with the glial marker GFAP for astrocytes or CNPase for oligodendrocytes, additional double-labeling immunofluorescence experiments were performed. At first the paraformaldehyde-fixed, paraffin-embedded sections were processed for catalase detection as described above. Then the anti-catalase antibody was detected with an Alexa-488-labeled goat anti-rabbit antibody (Molecular Probes, Eugene, OR; diluted 1:350). Then the sections were incubated overnight with mouse monoclonal antibodies against GFAP (Sigma, St Louis, MO; clone G-A-5, diluted 1:1000) or CNPase (Abcam, Cambridge, UK; clone ab3619, diluted 1:1000). The monoclonal antibodies were detected with an Alexa-594-labeled donkey anti-mouse antibody (Molecular Probes; diluted 1:500). After nuclear counterstaining with TO-PRO-3 (Molecular Probes), the sections were mounted in Mowiol 4-88 (Hoechst; Frankfurt/M., Germany) with 0.5%
The sections were analyzed using a Leica TCS SP confocal microscope (Leica Microsystems; Heidelberg, Germany) with sequential detection of the Alexa-488 fluorescence at 497–643 nm (excited at 488 nm, green), the Alexa-594 emission at 600–640 nm (excited at 544 nm, red), and of the TO-PRO-3 emission at 640–761 nm (excited at 633 nm, printed in blue).
Results
Positive staining representing the catalase mRNA was seen in the cytoplasm of neurons at many sites in rat brain with various intensities. In rat hippocampus, the mRNA of glyceraldehyde-3-phosphate dehydrogenase (GAPDH), used as positive control, showed a strong neuronal expression pattern, with prominent cytoplasmic staining and negative nuclei and nucleoli (Figures 1a and 1c). The intensity of staining could not be further enhanced by prolongation of the incubation time. The catalase mRNA, in comparison to GAPDH, showed a somewhat weaker expression, which could be demonstrated only by the use of CARD as a clear cytoplasmic staining (Figures 1b and 1d). In the hippocampus, pyramidal cells were positive for catalase mRNA and, at lower signal intensity, the neurons of the dentate gyrus (Figure 1b). The frequency of strongly positive neurons was particularly high in the brainstem, where neurons of different cranial nerve nuclei, such as the trigeminal motor nucleus, showed strong catalase mRNA expression (Figure 1f). In cerebellum, Purkinje cells were distinctly positive (Figure 2a), with negative sense controls (Figure 2b). The olfactory bulb, together with brainstem and cerebellum, belongs to the regions of rat brain with the highest catalase mRNA expression, showing a strong staining in the mitral cells and of neurons of the stratum plexiforme externum (Figure 2c). The corresponding sense controls were again completely negative (Figure 2d).
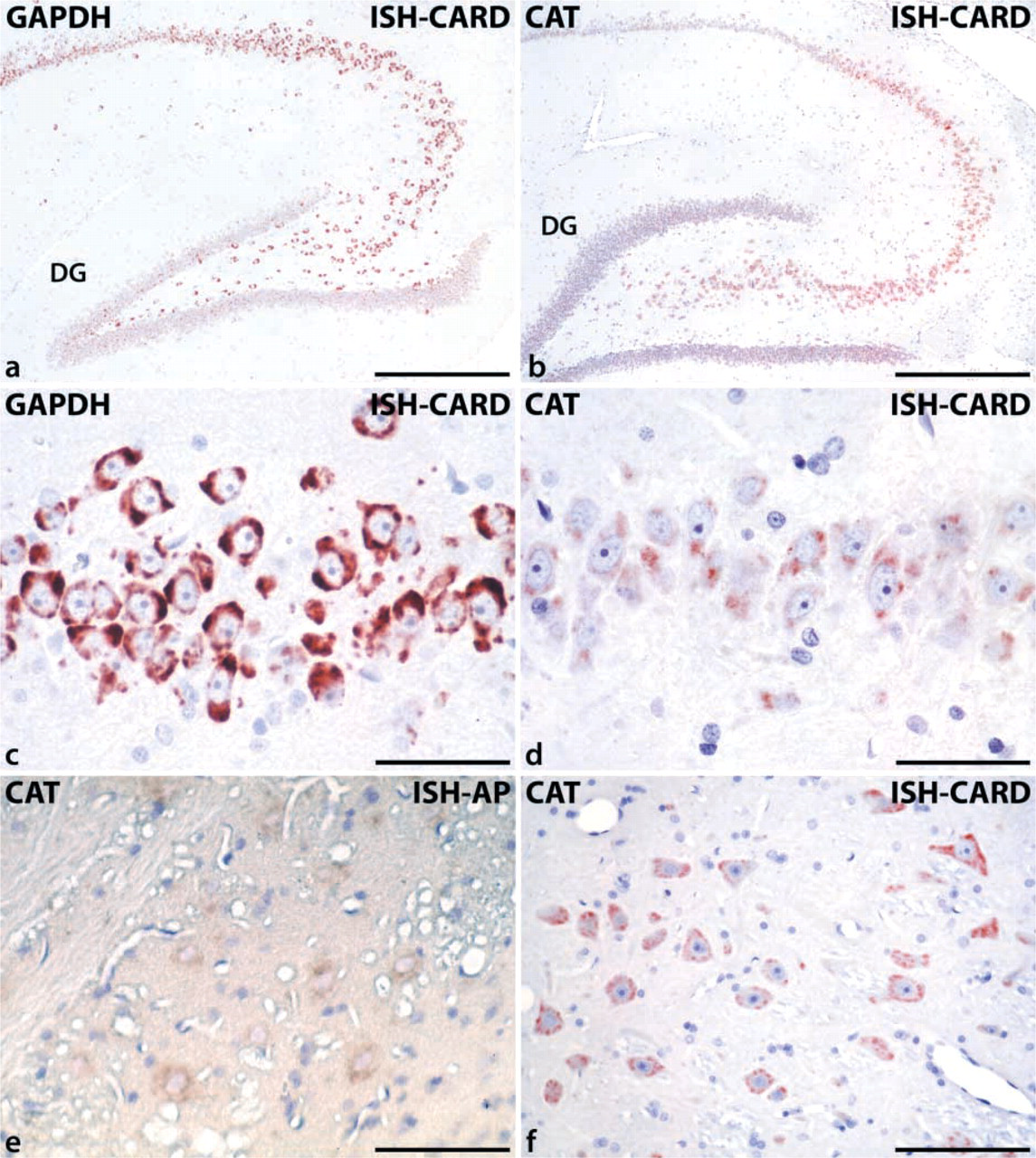
Detection of catalase (CAT) and glyceraldehyde-3-phosphate dehydrogenase (GAPDH) mRNA by CARD-enhanced ISH in perfusion-fixed and paraffin-embedded rat brain. (
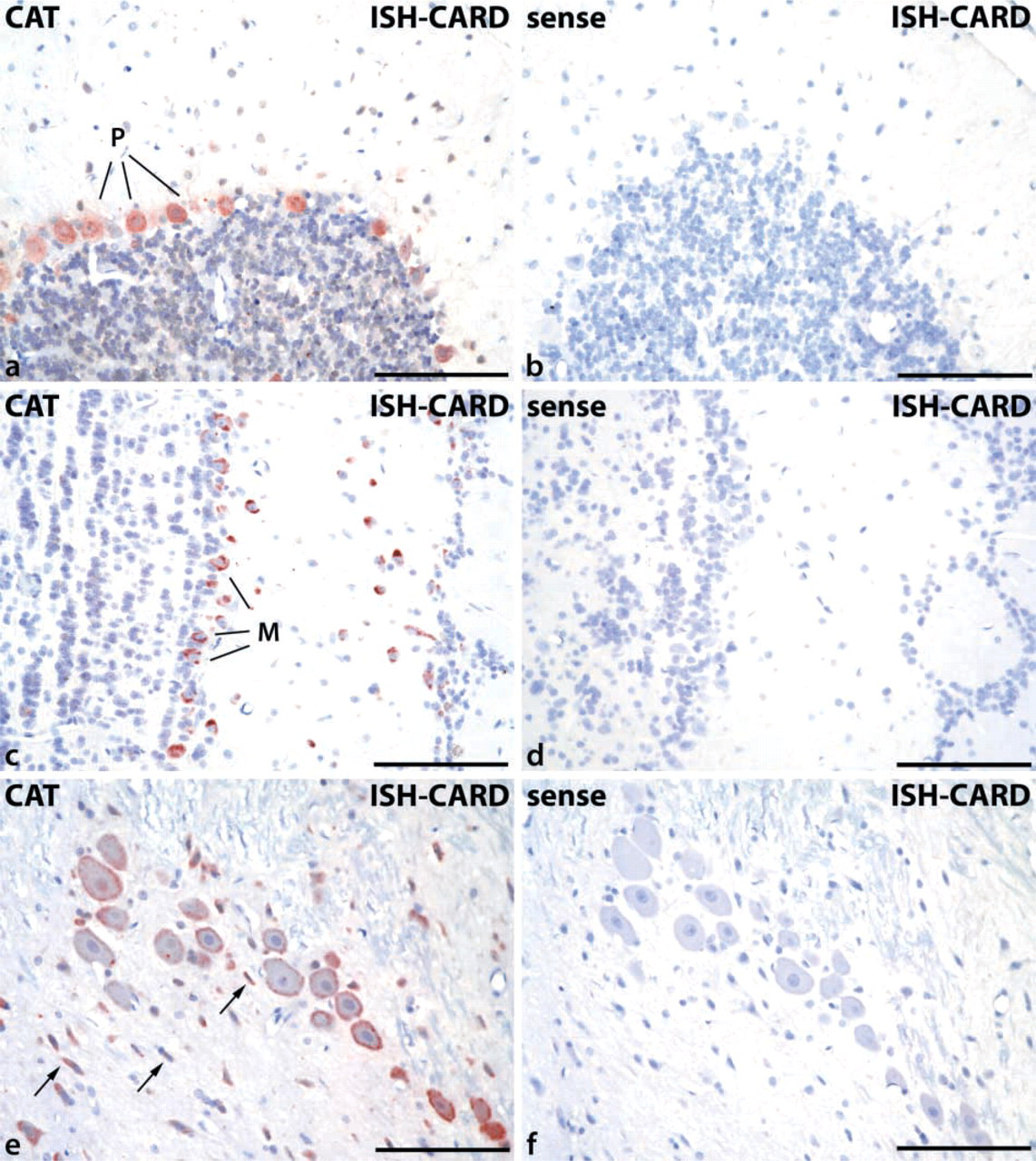
Examples of areas of rat brain with strong catalase mRNA expression, detected by CARD-enhanced ISH. (
In general, the detection of catalase mRNA, in spite of CARD amplification, appeared to be restricted mostly to neurons. The many oligodendrocytes of the corpus callosum, the perivascular astrocytes, and the endothelial cells remained negative. Nevertheless, the staining of catalase mRNA in neurons (e.g., in locus coeruleus; Figure 2e) was often accompanied by positive reaction in smaller cells which, without specific double labeling, could not be definitely ruled out to be of glial origin (arrows in Figure 2e).
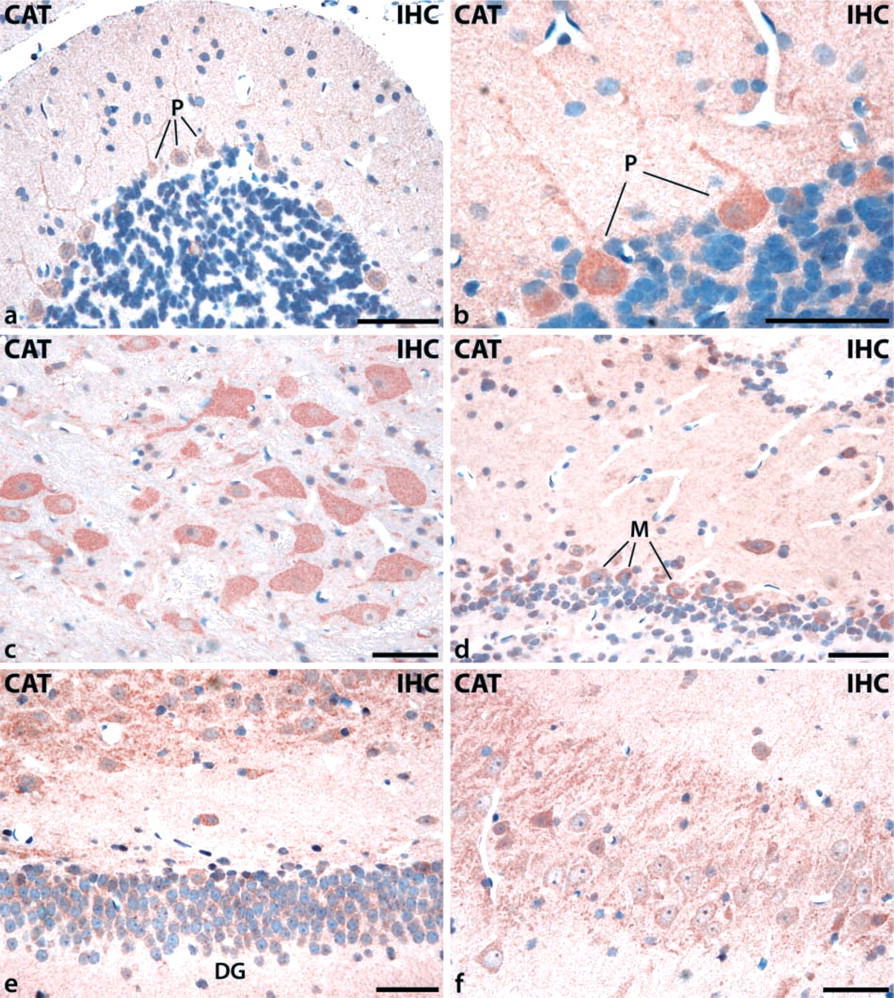
IHC detection of catalase immunoreactivity in different regions of rat brain using perfusion-fixed and paraffin-embedded tissue sections followed by antigen retrieval (Grabenbauer et al. 2001). (
To assess the role of signal amplification by CARD, some sections were hybridized with the same protocol but stained instead with alkaline phosphatase-labeled anti-digoxigenin Fab fragments and NBT/BCIP substrate (Schad et al. 1996). In those sections, only the neurons of the brainstem nuclei, which showed the strongest signal in the CARD-amplified ISH experiments, exhibited, after prolonged incubation time (up to 48 hr), weak cytoplasmic staining with a high degree of nonspecific background (Figure 1e). All control experiments using sense probes remained negative, demonstrating the specificity of the ISH–CARD protocol (Figures 2b, 2d, and 2f).
To assess the validity of our improved ISH protocol, we compared the distribution of catalase mRNA with that of catalase protein using similarly processed paraffin-embedded sections and antigen retrieval (Grabenbauer et al. 2001). The results revealed a finely granular cytoplasmic signal that was also present in extensions of neuronal cells. Corresponding to the mRNA expression pattern, catalase protein could be detected in the same neurons. The number of catalase-positive cells was particularly high in the cerebellum (Figures 3a and 3b), in brainstem (Figure 3c), in the olfactory bulb (Figure 3d), and in the hippocampus (Figures 3e and 3f).
Because of the paucity of catalase mRNA signal in non-neuronal cells (see above), we assessed the presence of catalase immunostaining by using CLSM and double immunofluorescence. Antibodies to GFAP and CNPase were applied as specific markers for astroglial cells and oligodendrocytes, respectively. Finely granular immunofluorescence was observed most prominently as expected in neurons (Figures 4a and 4b) and also in astroglial cells (Figures 4a–4d) and in oligodendrocytes (Figures 4e and 4f). The few catalase-positive granules in the latter cells were mostly localized in the perinuclear cytoplasm, particularly in oligodendrocytes (Figures 4e and 4f).
Discussion
Distribution of Catalase mRNA by ISH Corresponds to That of Catalase Protein
The present study has revealed the presence of catalase mRNA in the cytoplasm of neuronal cells in rat brain, corresponding to the distribution of catalase protein as reported previously by IHC (Moreno et al. 1995). In addition, the application of an improved IHC method to paraffin sections of perfusion-fixed tissues (Grabenbauer et al. 2001) revealed a finely granular staining pattern consistent with small peroxisomes, corroborating previous EM observations (Holtzman et al. 1973). By double immunofluorescence and CLSM, catalase-positive particles were also identified in astroglial cells and oligodendrocytes, although to a lesser extent than in neuronal cells. This appears to correlate with the paucity of mRNA signal for catalase in non-neuronal cells by ISH. The granular staining pattern of catalase in neurons, which was clearly visualized by immunofluorescence (Figure 4), may be of particular significance in studies of peroxisomal biogenesis disorders, in which that enzyme and other matrix proteins of peroxisomes are detected in the cytoplasm (Lazarow and Moser 1995). Our method appears to be superior to those used in previous reports, which showed only a diffuse cytoplasmic staining also in normal animals, although by EM the enzyme was found in peroxisomes (Moreno et al. 1995). The reason for the diffuse cytoplasmic staining seems to be the use of frozen sections and their permeabilization by detergents.
Signal Amplification by CARD
The introduction of tyramine signal amplification has significantly improved the sensitivity of the peroxidase-based color reactions for IHC and ISH techniques. In this study, CARD was applied to enhance the sensitivity of a nonradioactive ISH protocol (Schad et al. 1996) for detection of catalase mRNA in perfusion-fixed and paraffin-embedded rat brain. Compared to the results of ISH experiments obtained with the conventional alkaline phosphatase color reaction, the CARD-enhanced method showed an improved signal-to-noise ratio. The strong signal amplification by CARD enabled us to use short development times for the AEC color reaction, thus leading to clear specific mRNA signals without nonspecific background staining. However, for detection of strongly expressed mRNAs, such as glycerol aldehyde-3-phosphate dehydrogenase (GAPDH), which was used as a positive control in this study, the CARD protocol did not appear to significantly affect the already strong signal.
The high sensitivity of our ISH protocol allowed the use of glutaraldehyde for the perfusion fixation which, compared to paraformaldehyde, led to better tissue preservation. The improved quality of the morphological preparations combined with the high sensitivity of the detection protocol substantially exceeded the resolution of the only previously reported radioactive ISH protocol for catalase in brain (Reimer and Singh 1990).
The Important Role of Catalase and Peroxisomes in Brain
The localization of catalase mRNA by ISH revealed a heterogeneously distributed expression pattern along different subsets of neurons in rat brain. Not only monoaminergic neurons, such as those of the locus ceruleus, but also the GABAergic Purkinje cells, the glutamate-containing mitral cells of the olfactory bulb, and the hippocampal pyramidal cells appear to contain substantial amounts of catalase mRNA. As mentioned above, this correlates well with the immunocytochemical data on distribution of catalase protein in rat brain (Moreno et al. 1995) and with the results of IHC experiments presented in this study (Figures 3 and 4), confirming the important roles of catalase not only in defense against ROS (Dringen et al. 1999) but also in brain metabolism, e.g., of ethanol (Aragon and Amit 1992; Correa et al. 2001).
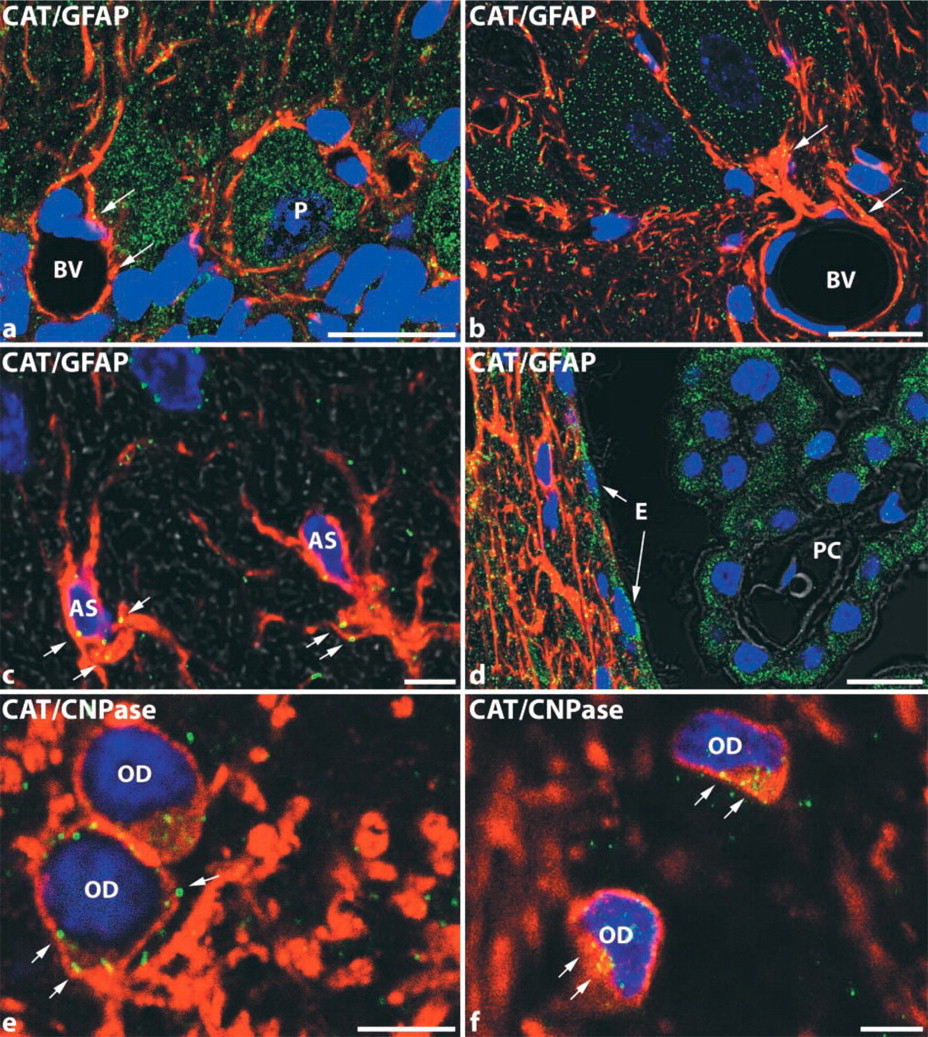
Double-labeling immunofluorescence for detection of catalase in different glial cell types of perfusion-fixed, paraffin-embedded rat brain, analyzed by CLSM. (
As a marker protein for peroxisomes, catalase is subject to severe alterations not only in primary genetic peroxisomal disorders, such as Zellweger syndrome, adrenoleukodystrophy, and infantile Refsum's disease (Wanders et al. 1993; Lazarow and Moser 1995), but also in conjunction with secondary alterations of peroxisomes in tumors (Lauer et al. 1999; Litwin et al. 1999) and inflammation (Beier et al. 1997). For investigation of alterations of catalase expression, either experimentally or under pathological conditions, simultaneous monitoring of catalase mRNA and protein appears to be mandatory. For this task, the presented protocol for the nonradioactive CARD-enhanced in situ detection of mRNA in perfusion-fixed and paraffin-embedded rat brain provides a sensitive technique with high cytological resolution, which should also be useful for investigation of the expression of other peroxisomal proteins.
Footnotes
Acknowledgements
Supported by grants Ba 1155/1–4, Fa 146/1–3, and SFB 601 from the Deutsche Forschungsgemeinschaft and by a fellowship from the Christine-Jung-Stiftung to A.S. by the Medical Faculty of the University of Heidelberg.
The technical assistance of Inge Frommer is gratefully acknowledged.
