Abstract
The depletion of the pericellular and territorial matrices in articular cartilage is considered to be one of the earliest events in pathobiology of osteoarthritis (OA). A newly discovered family of proteins with a disintegrin-like and metalloproteinase-like domain (ADAM) may be involved in matrix degradation as well as in cell-cell and cell-matrix interactions. The purpose of this study was to determine by in situ hybridization whether human articular chondrocytes from newborn, normal adult, and OA cartilages express messenger RNA for ADAM-10, one of the members of this family, and by semiquantitative RTPCR to compare the levels of this expression. The results confirmed the expression of ADAM-10 by human articular chondrocytes and revealed the highest levels of expression in the continuously remodeling cartilage of newborns and the most fibrillated areas of OA cartilage, especially the regions of cell clusters. Importantly, ADAM-10 mRNA expression was evident in tissues with the greatest loss of Safranin O staining from the territorial and interterritorial matrix of the chondrocytes. Messenger RNA was upregulated in OA tissue compared to the age-matched normal cartilage, as detected by RT-PCR. Upregulated levels of ADAM-10 mRNA expression appear to be related to the degree of cartilage damage and/ or degradation, which suggests a potential role for at least one member of this new family in the cartilage matrix destruction accompanying OA.
T
These membrane-type metalloproteinases (MT-MMPs) are known to be a component of the cell membrane. One of them, MT1-MMP, could act as a receptor for tissue inhibitor of matrix metalloproteinase-2 (TIMP-2) (Strongin et al. 1995), forming a complex that activates proge15 atinase A (MMP-2) and MMP-13 (Pendas et al. 1997). However, it is unlikely that MT1-MMP plays an initiating role in chondrocytic chondrolysis because we have shown (Büttner et al. 1997) that its message is constitutively expressed in newborn, normal, and OA cartilage, as visualized by in situ hybridization. In our previous study (Chubinskaya et al. 1996) we were able to mimic the earlier cartilage matrix changes by culturing normal human adult cartilage in the presence of IL-1. Depletion of Safranin O staining selectively from the territorial matrix and upregulation of mRNA for MMP-3 and MMP-8 were observed.
Recently, a novel family of metalloproteinase genes with A Disintegrin And Metalloproteinase (ADAM) domain in their structure have been described (Wolfsberg et al. 1995; Howard et al. 1996; Kuno et al. 1997; McKie et al. 1997). To date, 15 full-length members of this family have been cloned. These proteins have a significant analogy with snake venom proteinases. According to Wolfsberg et al. (1995), sequence analysis indicates possible proteolytic activity for some of the ADAMs (numbers 1, 8, 9, and 10). Howard et al. (1996) showed that bovine ADAM-10 has the potential to express metalloproteinase activity. However, no such activity has been demonstrated with the isolated or reconstituted enzyme.
Because members of the ADAM family may have a dual function, proteolysis and adhesion, we decided to test whether these ADAMs could be detected in human articular cartilage with regard to their potential implication in the extracellular matrix remodeling that takes place in normal and pathological conditions. Recently, McKie et al. (1997) have shown the presence of message for ADAM-10, but in isolated cultured human chondrocytes. It is noteworthy, however, that when chondrocytes are isolated from cartilage, the message of several proteins can be up- or downregulated, thus not necessarily reflecting “the state of expression” in the intact tissue. To our knowledge, no studies have been done on chondrocytes within the cartilage tissues and comparing the patterns of expression of ADAM-10 in uncultured human cartilage from organ (tissue) donors and OA patients.
The purpose of this study was to determine whether human articular chondrocytes express the mRNA for ADAM-10 and to correlate the level of this expression with the histopathological appearance of human newborn, normal adult, and OA tissue. Six upstream and downstream primers on the basis of the human sequence for ADAM-10 have been designed in our laboratory and applied for both in situ hybridization and comparative reverse transcriptase-polymerase chain reaction (RT-PCR). Histological evaluation of human cartilages was done using our modification (submitted for publication) of the Mankin score (Mankin et al. 1971). In this article we report the expression of ADAM-10 in human articular cartilage and the upregulation of ADAM-10 message in severely damaged OA cartilage compared to age-matched normal tissues.
Materials and Methods
Reagents
Chemicals, either reagent or molecular biology grade, were purchased from Sigma (St Louis, MO) unless otherwise noted.
Tissue Acquisition
Donor Cartilage. Full-thickness articular cartilages were dissected within 24 hr of death from the load-bearing regions of the femoral condyles of 20 donors with no history of joint disease. Tissues (both male and female, ranging from newborn to 84 years of age) were obtained through the Regional Organ Bank of Illinois. After opening of the joint, the surface of the cartilage was grossly examined and graded by the scale of Collins (1949) as modified by Muehleman et al. (1997). The presence and location of any macroscopically visible fibrillations and full-thickness defects were recorded.
Osteoarthritic Cartilage. Human OA cartilage was obtained as surgical specimens from 18 patients (aged 59-88 years of age, both men and women) undergoing total knee arthroplasty due to advanced OA. Within 3 hr after surgery, full-thickness noncalcified cartilage was dissected from the femoral condyle. Adjacent pieces of the cartilage from both patients and donors were either processed for histology and in situ hybridization or used for total RNA extraction in RT-PCR.
Histological Processing
The cartilage samples were fixed in 4% paraformaldehyde, dehydrated, and embedded in paraffin. Sections (6-μm) were processed for histology and in situ hybridization. Histological sections were stained with Safranin O (Rosenberg, 1971). A histopathological score of 0-13 was assigned to the cartilage samples using a modification (submitted for publication) of the Mankin score (Mankin et al. 1971) with a total of 13 points possible for the most damaged tissue. The score was adjusted from the original Mankin scale, defined for OA in the hip, because the samples do not contain the calcified zone or the tidemark. The scale was expanded to include different staining patterns observed with Safranin O that may indicate earlier degenerative changes in the cartilage.
Although normal cartilage samples were obtained from human donors with no history of joint disease, not all of them appeared to be normal. They exhibited the entire spectrum of gross morphological and histopathological changes from normal to severely damaged tissue. On the basis of two scores, Collins' and Mankin, we define normal cartilage as donor cartilage with Collins' scores of 0-1 (no visible signs of gross anatomic changes or with only slight, negligible fibrillation in articular cartilage) and Mankin scores of 5 or less. Damaged cartilage was defined as donor cartilage with Collins' scores of 2-4 and Mankin scores of 6-13. Cartilage obtained from OA patients was defined as OA cartilage.
Oligonucleotide Probes and PCR Primers for ADAM-10
Six different anti-sense and sense oligonucleotide probes and primers were designed on the basis of ADAM-10 sequence (accession Z48579) to fulfill requirements of both in situ hybridization and RT-PCR: (a) 22-mer, anti-sense, location 117-138, 5′-GACAGAGTGAAATGGCAGAGTT-3′; (b) 21-mer, sense, location 118-138, 5′-ACTCTGCCATTTCACTCTGTC-3′; (c) 21-mer, sense, location 220-240, 5′-TGTTTACCCTGTGCTCTTCTG-3′; (d) 21-mer, antisense, location 944-964, 5′-TGTGAGAGACTTTGGGAGGTA-3′; (e) 21-mer, sense, location 2037-2057, 5′-GCCCCGAGAGAGTTATCAAAT-3′; and (f) 22-mer, anti-sense, location 2356-2377, 5′-CAGCAACGAAGAACAGGGAACA-3′. The specificity of these probes was compared with sequence data available from PC/GENE on the human DNA database. Sequences were checked using Oligo software program, Version 4.1. All oligonucleotide probes were synthesized and purified by Research Genetics (Huntsville, AL). For in situ hybridization, probes were 3′-end-labeled with 5′ -[α -thiol-35 S]-dATP (NEN Du Pont; Boston, MA) using terminal deoxynucleotidyl transferase (Sigma) and hybridized under conditions of highest stringency corrected for each probe.
In Situ Hybridization
In situ hybridization was performed according to the procedure of Sandell et al. (1991) as modified by our laboratory (Chubinskaya et al. 1996; Cole et al. 1996). Evaluation and/ or quantification of the in situ hybridization results was done on the basis of the following criteria: negative message, no silver grains in the cells; weakly positive, less than 20 grains per cell; positive, more than 20 grains per cell.
In Situ Hybridization Controls
Positive Control. As a positive control, the human monocytic cell line THP-1 was used (a gift from Dr. K. Mikecz). The THP-1 cell line has similar characteristics as described for the human U937 histiocytic lymphoma cell line (Howard et al. 1996).
Negative Controls. Sense primers served as negative controls.
Reverse Transcription-Polymerase Chain Reaction
Total RNA was extracted directly from cartilage tissue with acid-guanidium thiocyanate as previously described (Cs-Szabo et al. 1997). Specific primer pairs were constructed for ADAM-10 (primers e and f, listed above) and glyceraldehyde- 3-phosphate dehydrogenase (GAPDH). These primer pairs were designed to raise PCR products of different size [319 base pairs (
Results
By in situ hybridization we were able to detect mRNA for ADAM-10 in the THP-1 cell line that served as a positive control. All three anti-sense primers (see Materials and Methods) displayed similar positive levels of expression for this protein (Figures 1A and 1B). Sense oligonucleotide probes (or upstream primers) revealed no specific binding to this cell line (data not shown).
In situ hybridization experiments showed the high specificity of the designed primers. Even severely damaged OA cartilage clearly exhibited specific binding of the probes within the chondrocytes without nonspecific binding to the extracellular matrix.
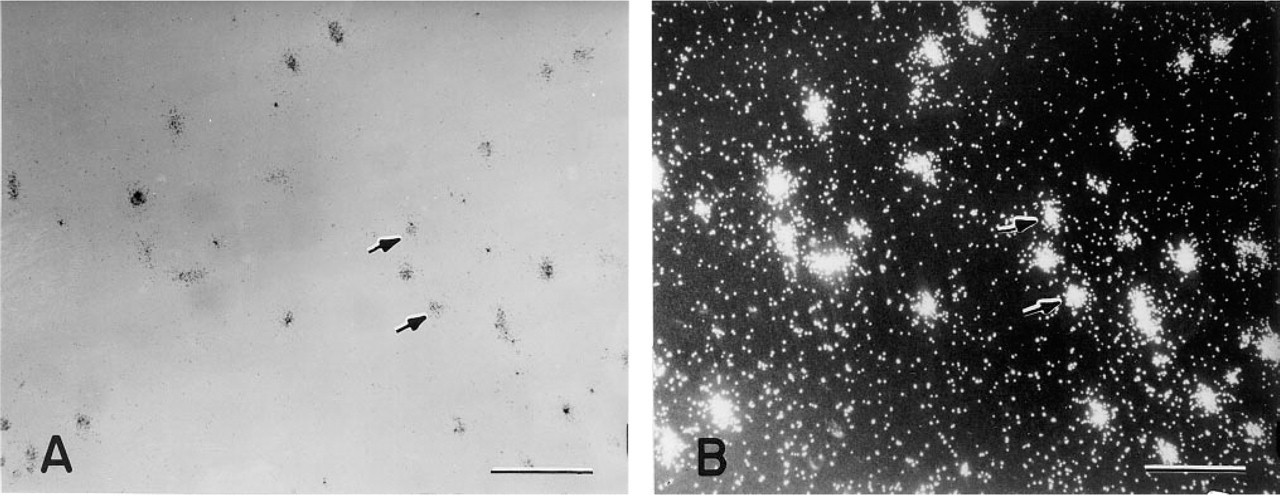
Representative photomicrographs of THP-1 cells hybridized to ADAM-10 anti-sense probes. (A) Brightfield photomicrograph. (B) Darkfield photomicrograph. Bars = 100 μm.
Newborn cartilage had a normal histological appearance, with an intact surface and uniformly distributed Safranin O staining. Normal newborn cartilage does not have the defined cartilage zones that are present in normal adult articular cartilage. This cartilage had clearly detectable message throughout all tissue sections visualized by in situ hybridization (data not shown).
Normal adult articular cartilage used for the current study (Figure 2A) could be described microscopically as cartilage with three distinct zones: (a) an intact superficial layer containing a few rows of flattened chondrocytes and no Safranin O staining; (b) a middle zone with rounded chondrocytes and strongly stained with Safranin O; and (c) a deep zone of cartilage, also with strong Safranin O staining and larger rounded chondrocytes organized in columns (chondrons). Expression of ADAM-10 in this type of cartilage was below the detection limit of the in situ hybridization technique (Figure 2B).
Levels of expression varied among OA cartilage samples. With progression of cartilage damage, ADAM-10 message was upregulated. Cartilages with less damage and lower histopathological scores (Mankin scores 6-9) revealed lower levels of mRNA expression for ADAM-10 (Figures 2C and 2D). Histopathological analysis of cartilages with the most severe degeneration, end-stage OA cartilages with a Mankin score of 10 and greater, showed the absence of the superficial layer, fissures into the middle and deep zones of cartilage, extensive fibrillation of the cartilage, and large clusters of chondrocytes (Figure 2E). These OA cartilages represented the strongest message for ADAM-10 (Figure 2F). The highest expression of this protein was found in the most fibrillated areas as well as in cell clusters. Importantly, ADAM-10 message was evident in cartilage sections in which Safranin O staining was depleted from the territorial (Figure 2C) and interterritorial (Figure 2E) matrices.
Densities of the PCR bands for ADAM-10 and GAPDH in 1 μl of total RNA samples showed a wide variety among the samples (Figure 3). To control these variables, the density of ADAM-10 bands was normalized to the density of the GAPDH-derived PCR bands for each sample. The absolute data of band densities are presented in Table 1. Relative steadystate levels of ADAM-10 mRNA showed age-dependent differences (Figure 3; Table 1). The highest expression was found in newborn cartilage (Figure 3, Lanes 1–3). With the aging of normal adult cartilage, expression was dramatically downregulated (Figure 3, Lanes 4-11) and was very low in older (60–70 years of age) cartilage samples (Figure 3, Lanes 7–11). It is important to note that all cartilage samples from organ donors used for RT-PCR had a normal histological appearance. In OA cartilage samples, mRNA expression for ADAM-10 was upregulated (almost threefold) compared to the expression in the age-matched normal cartilage samples (Figure 3, Lanes 12–15; Table 1). However, there were 10–15% individual differences in the OA samples, reflecting histopathological variations among OA cartilages.
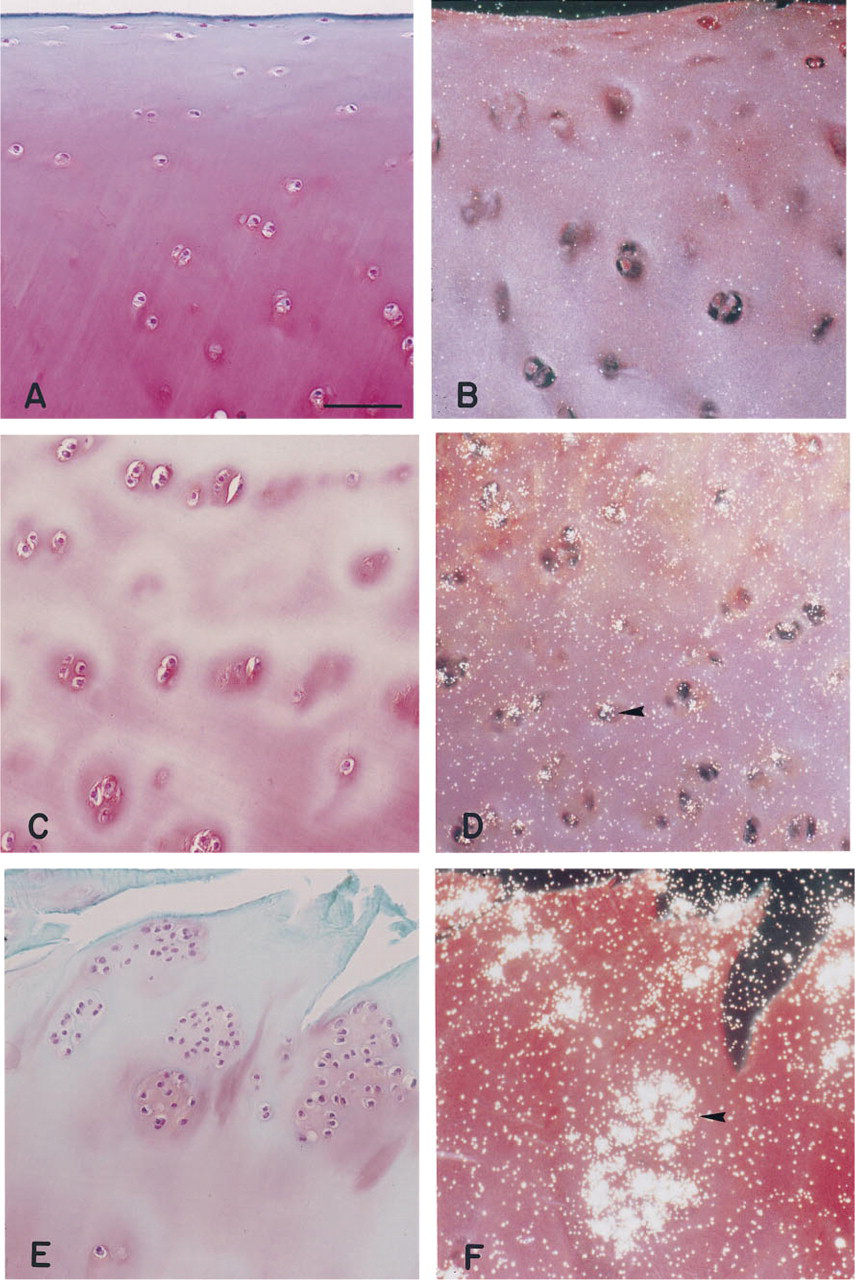
Histological appearance and in situ hybridization of the representative sections from normal adult (34 years of age;
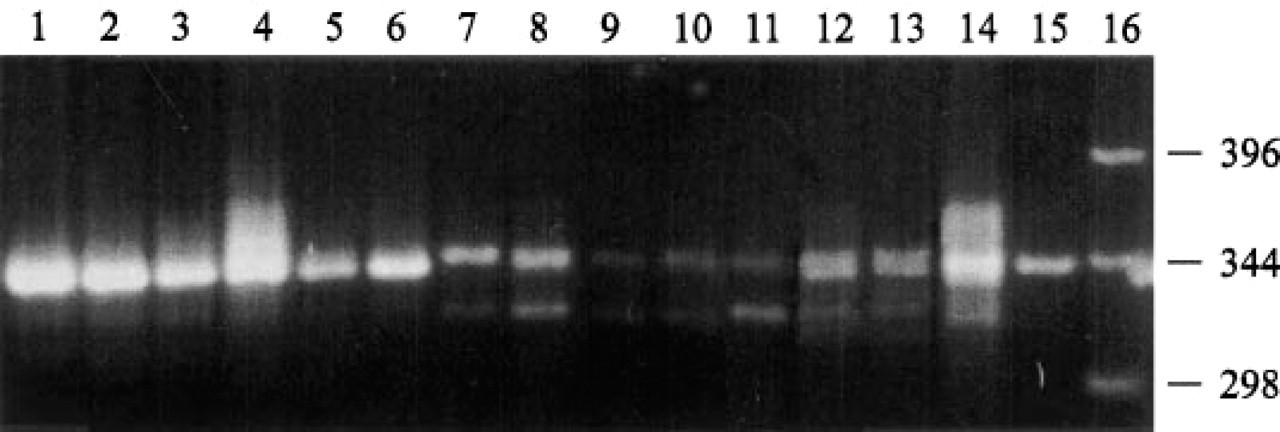
RT-PCR analysis of mRNAs of normal and OA cartilage samples for the presence of ADAM-10 and GAPDH. One μg of total RNAs from cartilage samples of normal donors of different ages (Lane 1, newborn; Lane 2, 14 weeks; Lane 3, 4 years old; Lane 4, 20 years old; Lane 5, 22 years old; Lane 6, 25 years old; Lane 7, 42 years old; Lane 8, 53 years old; Lane 9, 68 years old; Lane 10, 69 years old and Lane 11, 70 years old) and from patients who underwent total knee replacement surgery (Lane 12, 69 years old; Lane 13, 67 years old; Lane 14, 88 years old; Lane 15, 78 years old) were reverse-transcribed and amplified using ADAM-10 (top band) and GAPDH (lower band) primers. Lane 16, 1-KB DNA marker, given in
Sequence analysis of the PCR products from the total RNA of THP-1 cells, newborn, normal adult, and OA cartilage samples showed 99% homology with H. sapiens mRNA for disintegrin-metalloproteinase ADAM-10 only (accession Z48579) (Howard et al. 1996).
Discussion
The present studies clearly demonstrate the expression of mRNA for ADAM-10 in chondrocytes of intact human articular cartilage from organ donors and OA patients undergoing knee arthroplasty. ADAM-10 is a member of a newly discovered family of proteins with A Disintegrin And Metalloproteinase domains in their structure (Wolfsberg et al. 1995; Fox and Bjarnason 1996), which are similar to snake venom or reprolysin proteinases. This protein previously has been identified in different tissues, including brain, heart, kidney, liver, lung, skeletal muscle, spleen, ovary, testis, and a variety of cell lines (Wolfsberg et al. 1995; Herren et al. 1997). However, only recently McKie et al. (1997) demonstrated the expression of three members of ADAM family, ADAM-10, ADAM-12, and ADAM-15, by RT-PCR and Northern blot analysis in resting chondrocytes isolated from human femoral head and in a conditionally immortalized human articular chondrocyte cell line. However, when chondrocytes are removed from tissues and cultured as a monolayer or in alginate beads (our unpublished data), the phenotype of these cells could be changed, showing highly upregulated gene expression for a variety of proteins including collagen Type II, aggrecan, and MMPs, thus not necessarily reflecting “the state of expression” in the fresh (uncultured) tissue (Aigner et al. 1997). To our knowledge, the present article is the first report on the expression of ADAM-10 message in intact human donor and OA cartilage representing a significant number of specimens.
Densities of the PCR-derived bands for ADAM-10 and GAPDH mRNA products a
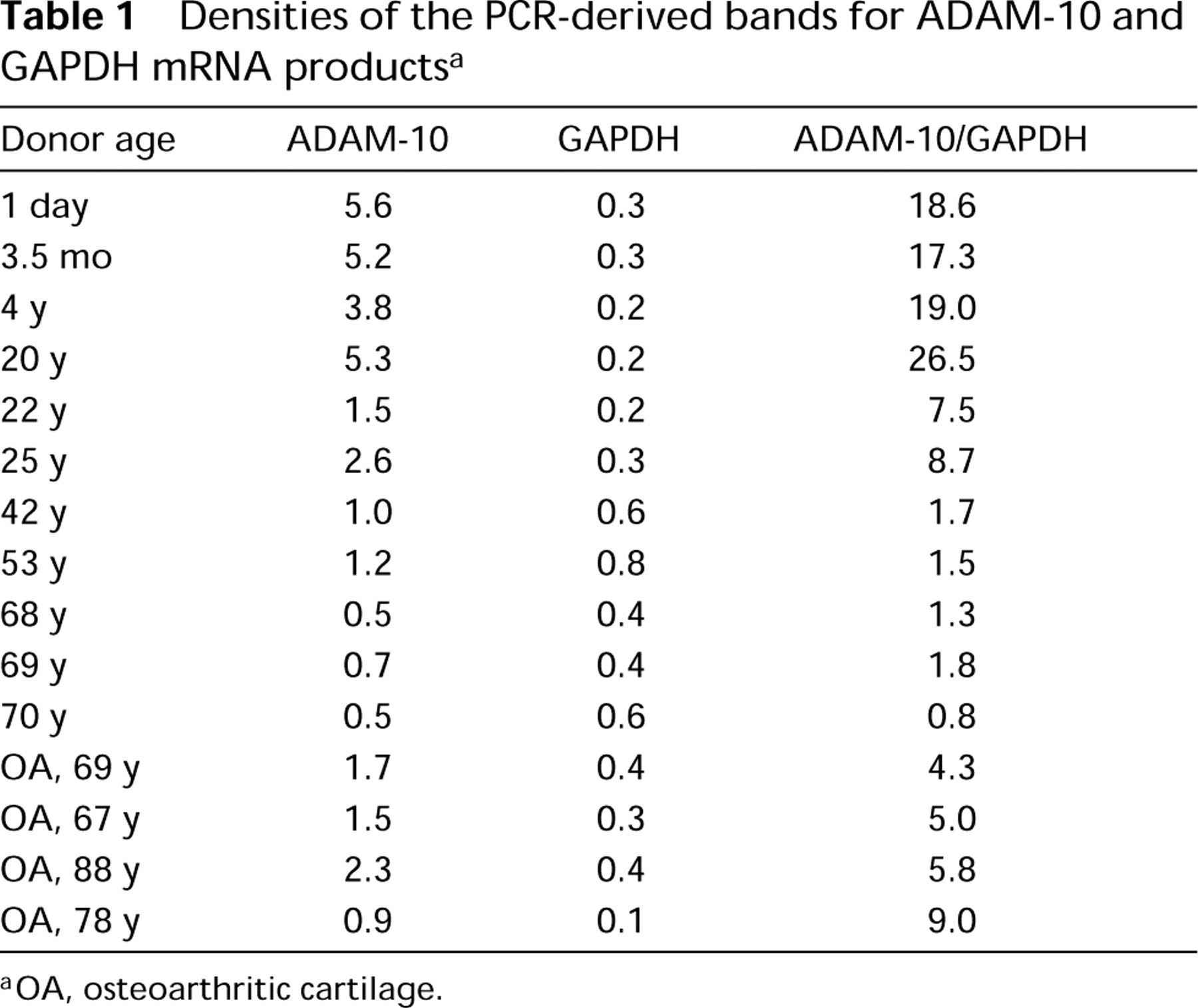

OA, osteoarthritic cartilage.
Utilizing cartilage samples from human donors with no history of joint disease, we document the differences in the levels of expression and message distribution for ADAM-10 with the aging of normal cartilage and compare those to age-matched OA cartilage. Among these donor cartilage specimens, a broad spectrum of histopathological changes can be observed, including loss of the superficial layer, cartilage fibrillation, cloning of chondrocytes, and loss of Safranin O staining. Similar changes are noted for cartilage from OA patients. The strongest signal visualized by in situ hybridization is found in chondrocytes located in the regions with pronounced depletion of sulfated PGs (as indicated by the intensity of Safranin O staining) from the territorial and interterritorial matrices and also in large cell clusters within fibrillated areas of cartilage. Chondrocytes in cartilage samples with a disrupted superficial layer but with otherwise uniform Safranin O staining (regardless of the source of the cartilage) showed no or barely detectable message for ADAM-10. This apparent correlation between localization of ADAM-10 message and loss of PG, together with predicted metalloproteinase activity (Howard et al. 1996), indicates a possible role of ADAM-10 in the turnover of the extracellular matrix.
With RT-PCR, mRNA for ADAM-10 was detectable in all human cartilage samples tested. The highest expression was found in cartilage from newborn donors, a tissue characterized by high rates of matrix turnover during growth and development. In normal adult cartilage, in which the rate of matrix turnover is lower, expression was significantly decreased with increasing age of the donors. However, a threefold upregulation in the levels of ADAM-10 mRNA expression was evident for OA cartilage samples characterized by severe matrix degradation and high turnover, which takes place in an attempted but failing repair process. Densities of the PCR bands for ADAM-10 and GAPDH showed a wide variety among the tissues. This may derive from differences in the isolation and purification procedure of the RNA, the extractibility of the cartilage, cell density and cell numbers in different samples, and/or the metabolic rates of the chondrocytes in this tissue. To control for each of these variables, the densitometric data for ADAM-10 were normalized to the density of the GAPDH-derived PCR band in each sample.
In summary, we have shown the expression of ADAM-10 messenger RNA in intact human articular cartilage from organ donors and OA patients associated with depletion of PGs from the territorial and interterritorial matrices. In addition, the identity in the messenger RNA expression and PCR products between a monocytic cell line and articular chondrocytes suggests some possible common functions of this protein in both types of cells, i.e., the potential enzymatic activity of ADAM-10 against the components of the extracellular matrix. One can speculate that the proteolytic activity of ADAM-10 might be important in the migration of these monocyte type THP-1 cells through the extracellular connective tissue matrix or perhaps in the processing of bioactive proteins (Fox and Bjarnason 1996). Although the function of ADAMs in articular cartilage is unknown, the association of these proteins with either the cell surface or the extracellular matrix suggests a specific role in the pathological and/or physiological processes that lead to remodeling of the extracellular cartilage matrix, and particularly the cell-associated matrix as one of the first targets in early cartilage degeneration.
Footnotes
Acknowledgements
Supported by NIH grant 2-P50-AR-39239.
We thank Dr Gary Davis for brilliant ideas, Dr Ada Cole for critical review of the manuscript, Dr Katalin Mikecz for providing a THP-1 cell line, Drs Joshua J. Jacobs, Klaus Huch, and Holger Koepp for providing cartilage samples, Dr Jeffrey Otto for help in the preparation of ![]() , and Richard Myer, BS, for excellent technical assistance.
, and Richard Myer, BS, for excellent technical assistance.
