Abstract
Paraffin embedding of tissue is generally perceived to dramatically reduce RNA detectability. As a consequence, in situ hybridization on paraffin-embedded tissue is largely confined to detection of high-copy RNA species (e.g., viral RNA) and/or to detection using typically more sensitive cDNA probes or riboprobes. In this study, several procedures for in situ hybridization on paraffin-embedded rat tissue using oligonucleotide probes complementary to cellular transcripts were developed and quantitatively compared. Certain pretreatments showed marked increases in sensitivity compared to untreated sections. Furthermore, through quantitative assessment using image analysis, sensitivity of optimal pretreatments was equal to that of routinely used fresh-frozen, postfixed tissue sections. The development of such techniques permitting in situ hybridization to be carried out on paraffin-embedded tissue allows a comparison of protein and mRNA distribution to be made in adjacent sections and provides the potential for double labeling by in situ hybridization and immunohistochemistry which may not be possible on post-fixed frozen sections.
S
Materials and Methods
Preparation of Tissue
Male Sprague-Dawley rats of 230 g were terminally anesthetized with sodium pentobarbital and transcardially perfused with 500 ml 0.9% saline containing 25 U/ml heparin. Rats were subsequently perfused with the same volume of 10% formal saline. Brains were removed and stored in fixative for 72 hr before embedding in paraffin. Sagittal sections were then cut at 6 μm using a rotary microtome (Anglia Scientific; Cambridge, UK) and mounted on Fisher Superfrost/ plus slides (BDH; Poole, UK). Sections were stored in dust-free conditions at room temperature (RT).
Slide Pretreatments
All slides were deparaffinized in xylene (15 min), hydrated in graded ethanols [prepared with diethyl pyrocarbonate (DEPC)-treated water], and washed twice in DEPC-treated water. From this stage, the treatments differed. For autoclaving, sections were placed in 10 mM citrate buffer, pH 6.0, and autoclaved in a standard 130C/2 bar/40 min cycle (Astell Scientific; Sidcup, UK). Sections were allowed to cool to 70C, washed for 15 min in DEPC-treated water, and dehydrated through graded ethanols. For microwaving, sections were also immersed in 10 mM citrate buffer, pH 6.0, in a clean polyethylene chamber, covered with a lid, and micro-waved at full power in an 800-W microwave [Panasonic (Matsushita Electric); Uxbridge, UK] for three 5-min periods. After each 5-min period the solution was topped up to compensate for any evaporation. Likewise, sections were washed for 15 min in DEPC-treated water and dehydrated in graded ethanols. For simple heating, slides were placed in DEPC-treated water for 120 min at 90C, before dehydration in graded ethanols. For pretreatment with proteinase K, sections were treated sequentially with 0.2 M HCl (10 min), 1% Triton X-100 (90 sec), 8 μg/ml proteinase K (Boehringer; Mannheim, Germany) in 20 mM Tris, 2 mM CaCl2, pH 7.0 [for 20 min, 40 min, or 90 min, at 37C, based on optimal conditions as previously described (Oliver et al. 1997)]. The digestion was subsequently stopped in PBS containing 2 mg/ml glycine and 5 mM EDTA before dehydration in graded ethanols.
For comparison, sections from fresh-frozen rat brains were probed for proenkephalin mRNA using a typical, well-characterized in situ hybridization protocol (Sirinathsinghji et al. 1995; Rigby et al. 1996). Briefly, the brains of rats were rapidly removed after decapitation and frozen in isopentane chilled to −40C. Sections were then cut in a cryostat (Reichert-Jung; Nussloch, Germany) at 10 μm and allowed to dry onto Fisher Superfrost/plus slides (BDH) before fixation in chilled 4% paraformaldehyde in 0.01 M PBS, pH 7.2 (5 min). Slides were then washed twice in PBS and processed through graded ethanols to 95% ethanol, in which they were stored at 4C until use, at which point they were removed from the ethanol and allowed to air-dry.
Design and Synthesis of Oligonucleotide Probes
An oligonucleotide probe complementary to nucleotides 388–432 of rat proenkephalin (Yoshikawa et al. 1984) was synthesized on an Applied Biosystems DNA synthesizer (model 394; Perkin Elmer, Foster City, CA) and purified by 8 M urea/8 M polyacrylamide preparative sequencing gel electrophoresis.
In Situ Hybridization
The oligonucleotide probe was labeled in a reaction mixture containing 0.4 pmol oligonucleotide, 1 x reaction buffer (Boehringer), 2.5 mM cobalt chloride, 25 U terminal deoxy-nucleotidyl transferase (TdT; Boehringer), and 19 μCi [35S]-deoxyadenosine 5′(α-thiotriphosphate) (NEN; Hounslow, UK). Specific activity was 1250 Ci/mmol (12.5 mCi/ml) for 15 min at 37C. On cessation of the labeling reaction, dithiothreitol was added to a final concentration of 40 mM. The radiolabeled probe was subsequently purified from unincorporated nucleotides using Sephadex G-50 spin columns. Hybridization was carried out as described previously (Sirinathsinghji et al. 1995; Rigby et al. 1996). Briefly, 3 × 105 cpm of 35S-labeled probe in 100 μl of hybridization buffer [50% deionized formamide, 4 x standard saline citrate (SSC), 5 x Denhardt's, 200 μg/ml acid-alkali-cleaved salmon sperm DNA, 100 μg/ml long-chain polyadenylic acid, 25 mM sodium phosphate, 10% dextran sulfate, and 40 mM dithiothreitol] was placed on each slide and coverslipped with a Parafilm (Sigma; Poole, UK) coverslip. Hybridization was carried out overnight in a humid environment at 37C. To define nonspecific hybridization, adjacent sections were incubated in either labeled sense probe or a mixture of labeled anti-sense probe with excess (100 x) unlabeled anti-sense probe. As a further control, the sense probe was also assessed using excess unlabeled sense probe. After hybridization, the sections were washed for 1 hr in 1 x SSC (containing 1% sodium thiosulfate) at 55C, followed by 1 hr in the same buffer at RT. Slides were then rinsed in 0.1 x SSC, dehydrated in graded ethanols, air-dried, and exposed to Hyperfilm βmax (Amersham; Little Chalfont, UK) for 48 hr.
For qualitative histological assessment of the effect of the various pretreatments on tissue morphology, sections that had undergone in situ hybridization after the pretreatment procedures outlined above were stained with hematoxylin and eosin and examined under a Leitz DMRD microscope (Leica; Nussloch, Germany).
Evaluation of In Situ Hybridization Results
The sensitivity of the pretreatments was calculated by densitometric analysis of in situ hybridization signal on autoradiographs. All hybridizations of all sections were carried out using the same probe mixture and same washes at the same time to ensure optimal comparability. Furthermore, reference standards were used to allow interfilm comparison where required. Image analysis was carried out using a MCID computerized image analysis system (Imaging Research; St Catharines, Ontario, Canada).
Quantitative analysis was carried out on at least three sections from each brain. The hybridization signals of the sections were correlated to a standard curve obtained from exposing a series of radioactive 35S standards (obtained by serial dilution of an aliquot of the 35S-labeled proenkephalin probe) to the same film as the sections. The optical density measurements of the standards (and therefore the sections) were translated into absolute amounts of radioactive nucleotide and proenkephalin mRNA detected in attomol/mm2.
Results
Effect of Pretreatment Regimens on Sensitization of In Situ Hybridization Signal
In situ hybridization signal for proenkephalin was observed predominantly in the caudate putamen (Figure 1), as previously described (Harlan et al. 1987; Beckstead 1995). Densitometric data are displayed in Figures 1 and Figures 2. The in situ signal in postfixed frozen sections and paraffin sections pretreated by microwaving for 15 min in citrate buffer were the most intense of all sections tested. There was no significant difference between these two groups (Figures 1A, 1D, and 2). A lower amount of signal was observed in all other paraffin-embedded sections, irrespective of pretreatment.
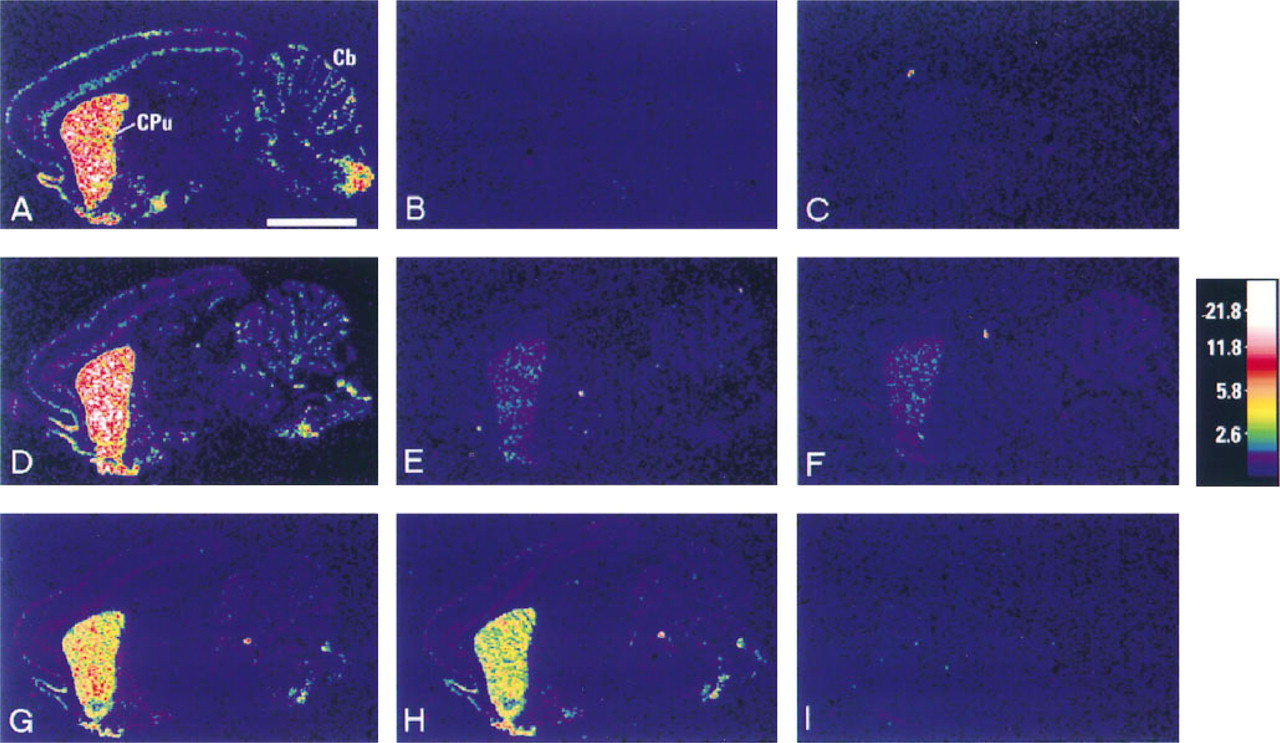
Sensitivity of various in situ hybridization retrieval methods as shown by computer generated images. (
The optimal pretreatment in terms of sensitization was microwaving for 15 min in citrate buffer, and proteinase K digestion for 20 minutes was the second most effective (Figure 1G). There was little difference in terms of intensity between sections that had undergone proteolytic digestion for 20 min and those that had undergone 40 min (Figure 1H), but considerable difference was observed in those digested for 90 min (Figure 1I). No positive signal for proenkephalin in the brain was detectable over background in sections subjected to 90 min of proteinase K digestion. Only weak signal was observed after autoclaving of paraffin sections (Figure 1E). The least sensitive method of pretreatment [second to not pretreating paraffin-embedded sections at all (Figure 1C) or using the harsh treatments of autoclaving or proteinase K digestion for 90 min] was that of heating the sections in water at 90C (Figure 1F). Only weak signal was detectable, with little difference noted between this group and those autoclaved (see Figure 2). All control sections probed with a mixture of anti-sense probe and 100 x excess of unlabeled anti-sense probe (Figure 1B), those probed with sense probe, or with sense probe with 100 x excess unlabeled sense probe gave results not significantly higher than background.
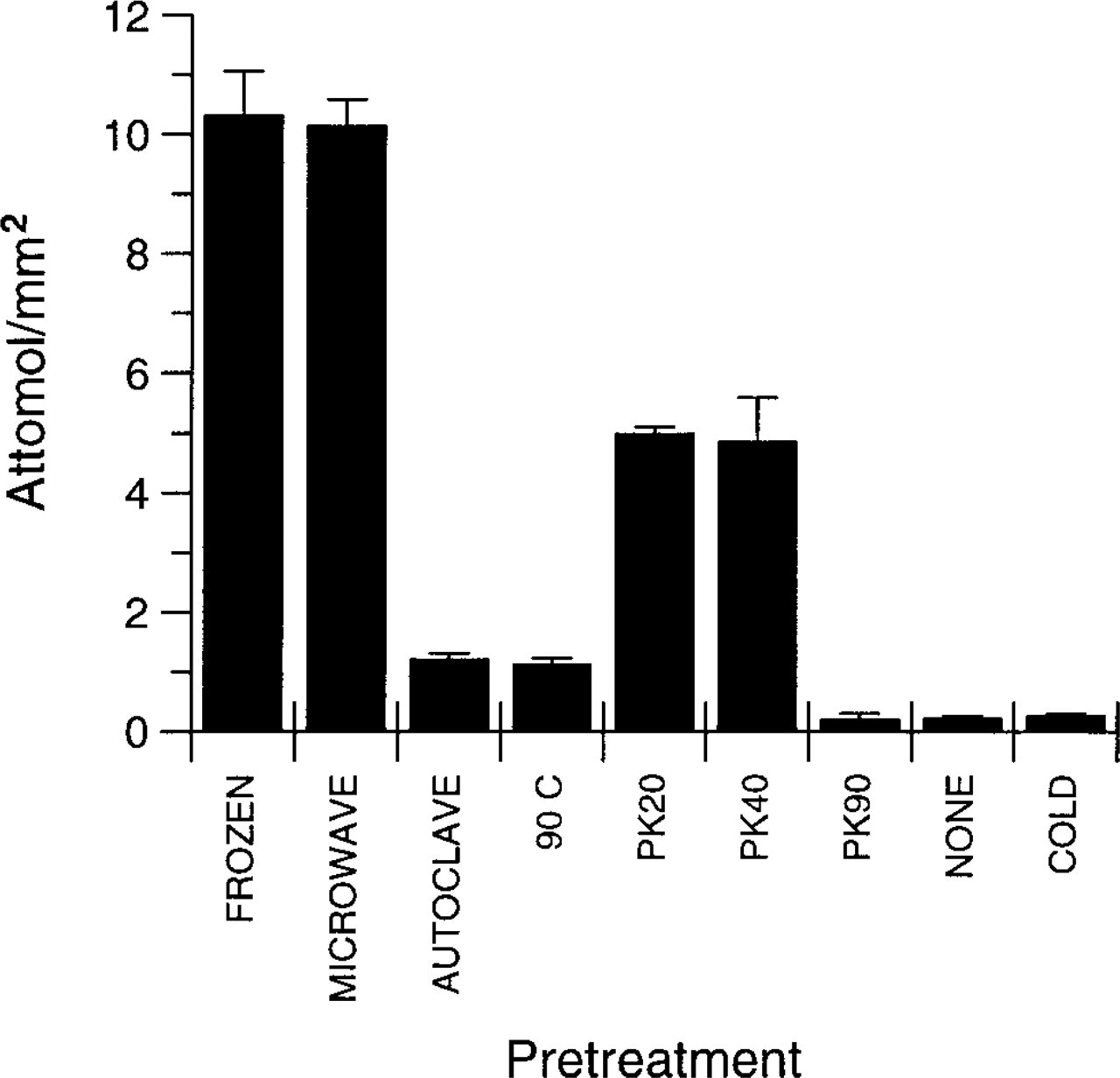
Graph illustrating comparison of pretreatments. All data were obtained from the same autoradiographic film, using sections from one individual animal, probed using the same probe and washed in the same wash solutions to ensure optimal comparability. Data were consistently reproducible when pretreatments were repeated on series of sections from different animals. Note that there is no difference between the frozen postfixed sections and the paraffin sections that were pretreated by microwaving. Sections treated by autoclaving and 120 min in distilled water at 90C (90 C) gave less intense retrieval and gave similar results. Proteinase K treatment for 20 (PK20) or 40 (PK40) min were also indistinguishable from one another. Sections pretreated with 90-min proteinase K digestion (PK90), those not pretreated with any retrieval conditions (NONE), and those probed with a mixture of labeled and 100 x excess unlabeled anti-sense probe (COLD) were all comparable and were visually indistinguishable from background (see Figure 1). Data are mean ± SEM; n = 6 (FROZEN and COLD groups) or n = 3 (all other groups).
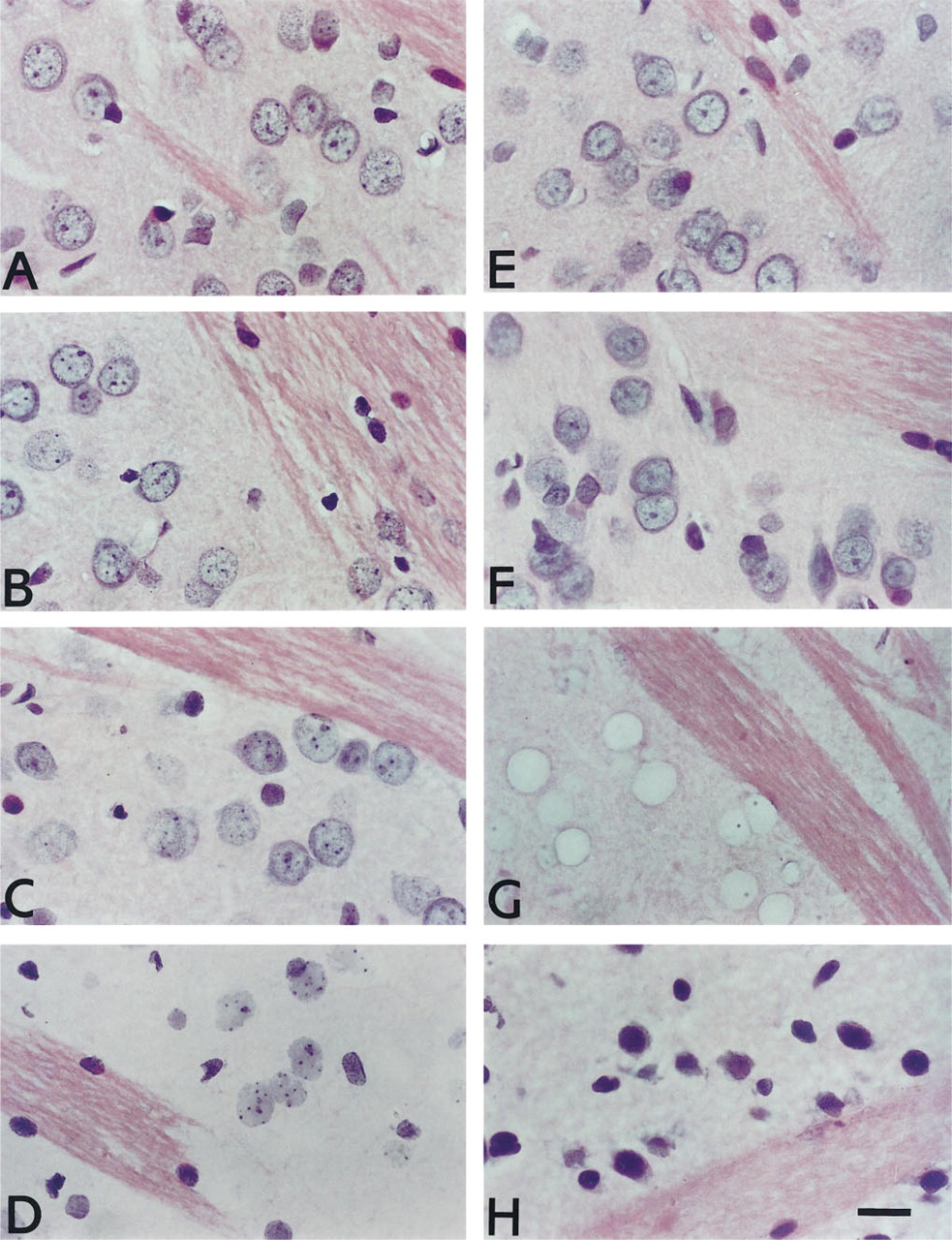
Effect on cell morphology of various pretreatments. (
Effect of Pretreatment Regimens on Tissue Morphology
Histological assessment was carried out on hematoxylin and eosin-stained sections that had undergone in situ hybridization for proenkephalin. Sections that had not undergone any pretreatment apart from basic deparaffinization and dehydration were regarded as the benchmark and showed characteristically intense hematoxylin staining primarily of chromatin and, to a lesser extent, the somal cytoplasm, with glial cells (e.g., oligodendrocytes and astrocytes) staining more intensely. Eosin stained the cytoplasm and myelin of white matter tracts, the latter being the most intense (Figure 3A). Sections pretreated with microwaving, heating at 90C or proteinase K treatment for 20 min had negligible effects on cell morphology (Figures 3B, 3E, and 3F). Chromatin structure was marginally impaired compared to controls in sections pretreated for 40 min in proteinase K but was still perfectly acceptable (Figure 3C). Very poor preservation of cell morphology was observed in sections pretreated by autoclaving or 90 min of proteinase K digestion (Figures 3D, and 3G). Sections that had been autoclaved showed minimal eosin staining of the neuropil, although strong white matter fiber staining was preserved. All neuronal nuclei appeared to have disappeared from the section, although some glial nuclei remained, often in a damaged state. In some cases, the nucleoli appeared to be spared and occupied the appropriate position in the void in which the nucleus had been (Figure 3G). Sections that had been digested for 90 min in proteinase K also demonstrated absence of eosin-staining in the neuropil, leaving strong eosinstained white matter fibers (Figure 3D). Cytoplasmic hematoxylin staining was also affected and gave cells the appearance of homogenous blue nuclei in which the nuclear chromatin structure was poorly preserved (Figure 3G). The nucleoli in some cells were still apparent. Frozen postfixed sections had relatively poor morphology compared to all paraffin sections except those pretreated with 90 min of proteinase K or autoclaving (Figure 3H). Hematoxylin staining of the nucleus was very intense, and it was difficult to distinguish neurons from glia. Chromatin structure was indistinguishable. Eosin staining overall was relatively weak, and was marginally stronger in the white matter tracts. Individual fibers were much less clearly distinguishable than in any of the paraffin sections, even those subjected to severe pretreatments, such as autoclaving.
Discussion
In this study we have compared an array of pretreatment regimens using quantitative densitometric techniques to determine the optimal conditions for sensitization of in situ hybridization signal on paraffin-embedded brain sections. Many methods have been documented for retrieving antigenicity in archival paraffin sections (Shi et al. 1991,1993; Umemura et al. 1995) and for staining of cells positive for TdT-mediated end-labeling (TUNEL) (Strater et al. 1995), but no comparative analysis has previously been made with regard to sensitivity for in situ hybridization using oligonucleotide probes in brain tissue.
It is generally believed that the greatest hybridization signal is achieved using fresh tissue, cut frozen and lightly postfixed in 4% paraformaldehyde in PBS. In this study we achieved a high degree of sensitivity using this method, although the signal intensity in paraffin sections pretreated by microwaving was equally high. Assuming that the oligonucleotide probe is small enough to penetrate into the depth of the section completely and that all β-particles reach the overlying film, it can be concluded that microwave pretreatment of paraffin tissue allows more sensitive detection of mRNA than in fresh tissue, because in this experiment we have compared routinely used 10-μm frozen post-fixed sections with 6-μm paraffin sections. The reasons for enhanced sensitivity may involve several factors. First, microwaving may allow greater penetration of the probe because of its ability to disrupt crosslinked proteins formed as a result of formaldehyde fixation (Fox et al. 1985; Puchtler and Meloan 1985; Morgan et al. 1994). These bonds may exist even in the lightly postfixed frozen tissue, thereby inhibiting probe penetration. A second possibility is that when the rats are perfused with fixative before brain removal and paraffin embedding, there may be less chance of RNase-mediated mRNA degradation than in animals decapitated before fresh tissue removal, owing to instantaneous RNase inhibition. The greater sensitivity of in situ hybridization after microwaving than after proteinase K treatment for 20 min was demonstrated by stronger signal (Figure 2). Interestingly, 40 min of proteolytic digestion had no major effect on signal intensity compared to 20-min digestion. However, increasing digestion time to 90 min caused a total loss of signal, presumably because of decreased retention of the target mRNA through digestion of crosslinked cellular components that maintain intracellular morphology. The reverse situation was observed in paraffin sections that received no pretreatment (apart from deparaffinization), in which no specific signal was observed. This was presumably due to lack of permeabilization. Weak signal was observed in sections pretreated with heating in distilled water at 90C for 120 min. This combination of soaking solution, temperature, and duration of incubation was an optimal combination for the retrieval of bcl-2 immunoreactivity (Umemura et al. 1995). However, only weak in situ signal was evident in this group, suggesting that although this pretreatment may be suitable for bcl-2 immunostaining, it is unsuitable for mRNA retrieval. It has been observed by others that for antigen retrieval to take place, the buffer must have an ionic content, e.g., as in citrate buffer (Shi et al. 1991; Morgan et al. 1994).
Although the microwave procedure appears to optimally sensitize the in situ signal, these data must be discussed in the light of the morphological analysis if sections are to be examined by light microscopy in addition to preliminary autoradiographic analysis. It was clear from the hematoxylin and eosin-stained sections that optimal cell morphology was maintained in sections heated at 90C in distilled water or those pretreated with 20 min of proteinase K digestion. However, these had significantly weaker signal intensities than those microwaved. Ultimately to decide on which protocol to use would depend on the absolute need of optimal morphology combined with estimated quantity of target RNA. In most circumstances, the use of the microwaving conditions outlined in this study would be entirely satisfactory. This would have the clear advantage of also allowing the potential for double labeling by immunohistochemistry and in situ hybridization on adjacent sections or on the same section, because microwaving in these conditions has been used by ourselves and others to retrieve antigenicity (Shi et al. 1991,1993). Furthermore, it has recently been shown that microwaving with these conditions is able to sensitize in situ signal using cRNA probes in many different tissues without the need to vary duration of permeabilization incubations according to tissue type, which is often required when proteolytic methods are used (Sibony et al. 1995; Lan et al. 1996).
We conclude that after suitable pretreatment, such as microwaving for 15 min in citrate buffer, RNA detectability by in situ hybridization is greatly enhanced, to a greater degree than in frozen postfixed sections. This method has many advantages, including preservation of superior morphology, enhanced potential for double labeling, and more convenient and stable storage characteristics with increased longevity.
Footnotes
Acknowledgements
We are grateful to Ms A. Jennings for technical assistance and to Dr M. Rigby for expertise in image analysis.
