Abstract
Barx2 is a member of the Bar class of homeobox genes and has been shown to regulate specific cell adhesion molecules, L1, Ng-CAM, N-CAM, and cadherin 6. By Northern blotting and in situ hybridization, we show that Barx2 is expressed throughout the gut and is located in epithelial cells of the proliferative and differentiative regions of the stomach, esophagus, and intestine. Barx2 was expressed in muscle cells of the muscularis externa and also showed a graded pattern of expression in intestinal enterocytes, decreasing in a crypt-to-villous direction. We speculate that Barx2 may regulate cell adhesion molecules in epithelial cells of the gut.
Barx2 is a member of the Bar class of homeobox genes that include BarHI and BarH2 from Drosophila (Higashijima et al. 1992) and Cnox from the cniderian Chlorohydra viridissima (Schummer et al. 1992). The homeodomain of Barx2 shares 87% amino acid identity with Barx1 and both genes are expressed in various epithelial tissues undergoing remodeling (Tissier-Seta et al. 1995; Jones et al. 1997; Sander et al. 2000; Herring et al. 2001). In the gut, a number of homeobox genes are expressed in temporally and spatially restricted expression patterns. A number of these, including Cdx-1 and Cdx-2, are expressed in intestinal epithelial cells (Duprey et al. 1988; Walters et al. 1997). Although the functions of these homeobox genes are still being elucidated, it is clear that some of them regulate the expression of cell adhesion molecules (Jones et al. 1997; Sellar et al. 2001; Hinoi et al. 2002). Most recently, Cdx2 has been shown to be a regulator of LI cadherin in the gut (Hinoi et al. 2002). Barx2 has been linked with expression of a number of cadherins, notably L1-cadherin, Ng-cam, N-cam, and cadherin 6 (Jones et al. 1997; Sellar et al. 2001).
Barx2 has been shown to promote myogenic differentiation and to regulate the expression of muscle-specific genes (Herring et al. 2001; Meech et al. 2002). Herring et al. (2001) reported expression in the stomach and the small and large intestine, although they did not define its location in these complex tissues. Here we used in situ hybridization (ISH) analysis to specifically locate Barx2-expressing cells in the adult gastrointestinal tract, and we show that Barx2 is expressed in smooth muscle cells and epithelial cells.
A 254-bp Barx2 3′ coding fragment adjacent to the homeobox was amplified by PCR from adult rat genomic DNA and used to probe for Barx2 expression. This region shares minimal nucleotide sequence identity between Barx1 and Barx2 (33% for mouse Barx1 and Barx2 and 36% for human Barx1 and Barx2: GenBank accession nos. AF277160, L77900, AF213356 and AH008405, respectively), and under our stringent conditions the probe is specific for Barx2. The primers 5′-tgg aca gga agc acc cac aaa ac-3′ (upper) and 5′-tag ctt aat ggt ggg ggt tcc gaa g-3′ (lower) were designed to amplify the 3′ end of Barx2 between the stop codon and the homeobox. The PCR fragment was cloned into pGEM-T easy vector (Barx2-3′ coding), sequenced in both directions and identified as Barx2 by database sequence searches using ANGIS (Australian National Genomic Information Retrieval System). A magnesium concentration of 1.5 mM was used in the PCR with the following parameters: (a) one cycle at 94C (30 sec), 72C (30 sec); (b) while the PCR was held at 72C the TAQ polymerase was added; (c) 35 cycles of 94C (30 sec), 55C (1 min), 72C (1 min); and (d) a final extension step of 72C (7 min). This Barx2-3′ coding fragment was used as a gene-specific probe for the Northern and ISH experiments.
Four-month-old female Sprague-Dawley rats were sacrificed and tissue segments from the stomach (fundic and body regions), esophagus (proximal and distal), duodenum, jejunum, ileum, cecum, and colon (proximal, middle, and distal for each) were removed, fixed, and processed for ISH (Powell and Rogers 1990). Antisense and sense [α33P]-rUTP radiolabeled riboprobes were hybridized to tissue sections (Powell and Rogers 1990). Hybridized sections were washed at a final stringency of 0.1 × SSPE at 65C for 30 min and exposed for 10 days at 4C. Tissues were stained with hematoxylin and images captured under bright-field and darkfield illumination with an Olympus BH2 microscope fitted with an Olympus darkfield condenser (model U-DCW) and a Sony digital video camera (model SSC-DC50P) and analyzed with Image Pro Plus analysis software (Media Cybernetics; Carlsbad, CA).
For Northern blotting analysis, total RNA was isolated from gut tissues and poly A(+) RNA prepared (Poly-A-Tract kit System IV; Promega, Madison, WI). Approximately 3-μg samples of poly A(+) RNA were fractionated in a 1% agarose formaldehyde gel, transferred to Zetaprobe GT membrane (Bio-Rad; Hercules, CA) and probed with the 254-bp Barx2-3′ coding fragment labeled with [α-32P]-dCTP. A rat β-actin RNA probe was synthesized from pTri-βactin-125 (Ambion; Austin, TX) and used to quantitate loading between different tissues. Filters were washed in 2 × SSC/0.1% SDS/65C for 30 min and exposed to a phosphor image screen for 24 hr. All experiments were approved by the Animal Ethics Committee of the Womens and Childrens Hospital, Adelaide, South Australia.
We first detected Barx2 expression in the forestomach, the body compartment of the stomach, duodenum, ileum, jejunum, and colon by Northern blotting analysis (Figure 1). Expression was uniform in the proximal, middle, and distal subregions with the exception of the proximal colon, which showed reduced expression compared to the distal colon. Barx2 expression was elevated in the forestomach compared to the body compartment and may reflect the different anatomic variation between these two regions. We did not detect Barx2 expression in the cecum.
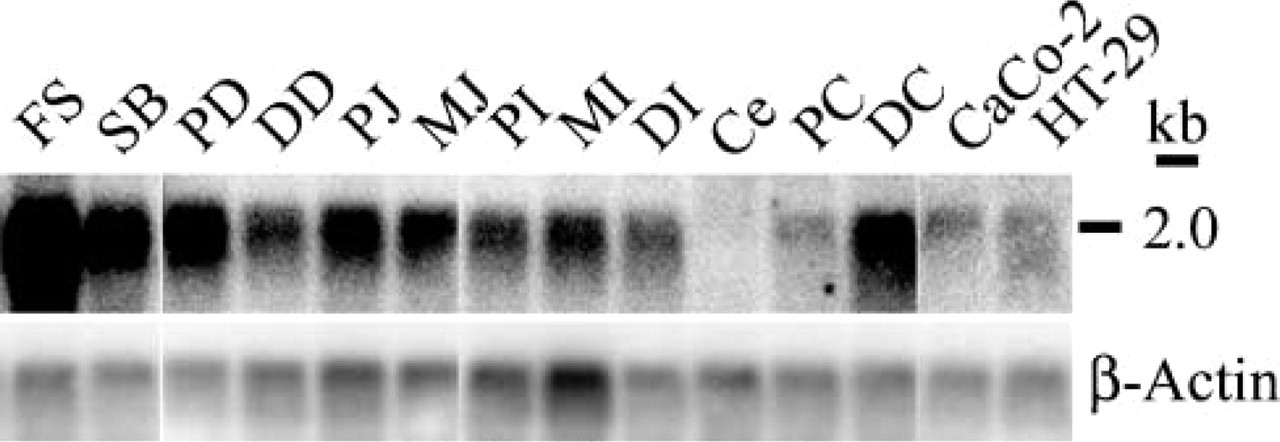
Analysis of Barx2 expression in adult rat gut and colon adenocarcinoma cell lines by Northern blotting. A transcript of 2.0 kb was identified in the gut and in two colon adenocarcinoma cell lines.
ISH analysis revealed that in the esophagus and forestomach Barx2 is expressed in the basal and suprabasal layers, which contain proliferating and differentiating cells, respectively (Figures 2A-2F). In the stomach body compartment, Barx2 was expressed mainly in the region towards the bottom of the glands containing the basophilic chief cells and appeared to decrease towards the upper portion of the glands where the acidophilic parietal cells are located (Figures 2J-2L). In the small and large intestine we found Barx2 expression to be strongest in the crypts and much reduced in the villous enterocytes (Figures 2M-2ZA). We also detected diffuse Barx2 expression in cells in the muscularis externae in the gut (Figures 2H and 2N; and data not shown). These expression data provide morphological evidence that Barx2 is involved in intestinal muscle cell function and provide a link between previous in vitro experiments and whole-tissue analyses (Herring et al. 2001; Meech et al. 2002).
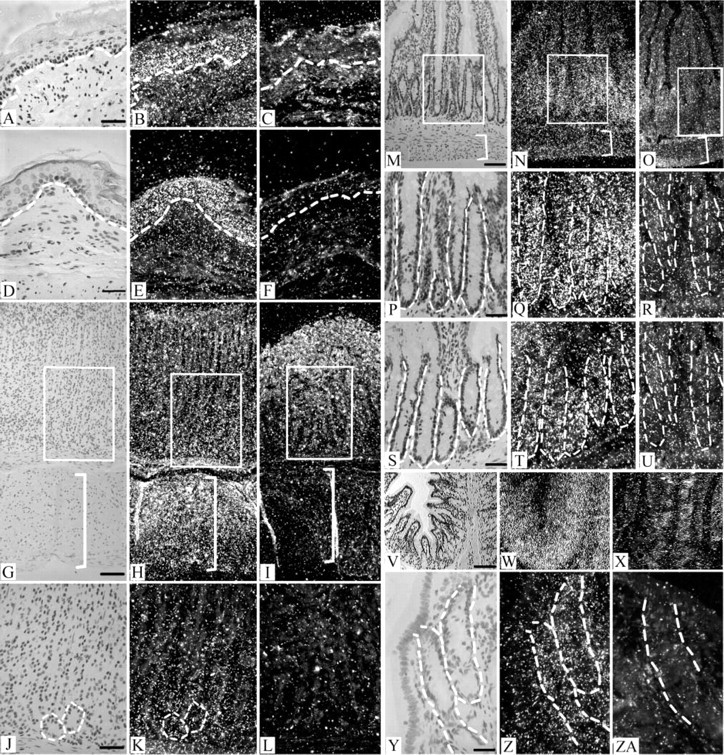
Barx2 detection by ISH in adult rat gut. Barx2 is expressed in the proliferative and differentiative regions of the esophagus, stomach, small intestine, and large intestine. Expression was weakly detected in villous enterocytes (
Mounting evidence suggests that Barx2 regulates the cell adhesion molecules L1 and cadherin 6. L1 is a neural cell adhesion molecule whose promoter can be stimulated or repressed by Barx2, depending on the circumstances (Jones et al. 1997). Barx2 and L1 share common expression locations, with both co-localizing in similar regions during mouse embryo development (Jones et al. 1997). We previously reported that Barx2 is uniformly expressed in the outer root sheath of developing wool follicles and the basal layer of the embryonic ectoderm, regions that are immunoreactive for L1 (Nolte et al. 1999; Sander et al. 2000). Our findings that Barx2 expression is abundant in the crypts and decreases in the villous enterocytes of the rat intestine is very similar to the immunohistological location of L1 previously reported in comparable regions of mouse small intestine (Thor et al. 1987).
We also detected Barx2 expression in two human colon adenocarcinoma cell lines, Caco-2 and HT-29, at a low level compared to expression in normal gut tissue. Recently, BARX2 expression levels have been found to positively correlate with cadherin 6 in ovarian surface epithelium and in ovarian cancer cell lines, and there is speculation that it is a suppressor of cancer progression (Sellar et al. 2001). Taken together with our findings of Barx2 expression in diverse epithelia of the gut, this raises the question of whether dysregulation of Barx2 might be associated with gastrointestinal cancers.
