Abstract
Digoxigenin-labeled riboprobes of six groups of human mucins were evaluated for sensitivity in archival tissue, using protease XXIV or proteinase K during in situ hybridization.
Keywords
D
For these studies, we used samples representing the tissues from which each mucin gene was cloned originally. Tissue blocks were identified by random search of the archival files of the Department of Pathology, University of California, Davis Medical Center and screened only to select for proper orientation. Five blocks were chosen for each mucin gene.
All tissues had been fixed in 10% neutral buffered formalin and embedded in paraffin using standard techniques. However, the duration that the tissues remained unfixed after their surgical removal and the ultimate duration of fixation was unknown. Adjacent sections from each block were run under identical conditions except for protease treatment. Tissue sections were deparaffinized, hydrated, quenched for endogenous phosphatases, and treated with either protease XXIV (Sigma, St Louis, MO; P-8038, 0.1 mg/ml) or proteinase K (Sigma; P-2308, 0.01 mg/ml) for 10 min at 37C. For hybridization, the tissue sections were placed in a humidification chamber at concentrations specific for each probe and tissue type for 6–8 hr. The slides were reacted with RNase A and RNase T1 for 30 min to hydrolyze remaining unhybridized mucin riboprobe. Subsequent stringency washes were specific for each probe. Slides were incubated with a monoclonal antibody to digoxigenin conjugated to alkaline phosphatase and visualized with NBT/BCIP. All slides were developed for the same amount of time.
Reactivity of ISH product was quantified by image analysis using an AutoCyte workstation (Burlington, NC) and software (Image Pro; Media Cybernetics, Rockville, MD). An area of interest was selected on a slide (either protease XXIV or proteinase K treated) and an image captured. The corresponding field on the adjacent section treated with the other proteinase was located and its image captured. Using the image analysis software, the percentage of the area that contained reaction product was determined. At least five fields per slide were compared. Results were analyzed using a paired t-test.
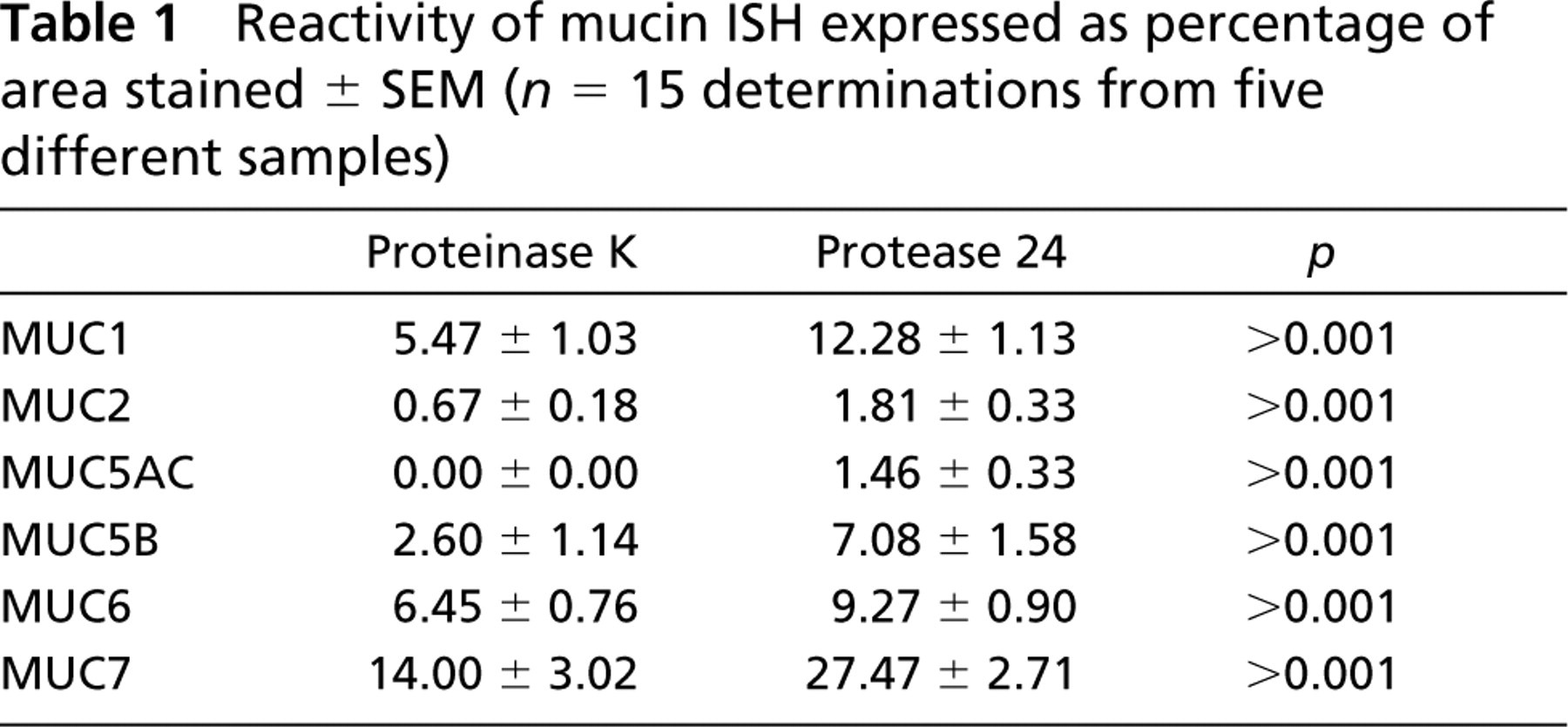
For each mucin, the amount of mucin mRNA as detected with ISH was increased with protease XXIV. With MUC5AC, ISH with proteinase K was not sensitive enough to detect any target mRNA. For the remaining mucin genes studied, the increased sensitivity when protease XXIV was used ranged from 1.5 to 2.7 times greater than when proteinase K was employed (Table 1).
Tissue samples derived from medical procedures and processed routinely for pathological diagnosis are not ideal for use in scientific studies requiring ISH investigations. This results primarily from autolysis or lack of standardized fixation. The size and density of the tissue used may also contribute to variable fixation. The majority of clinical laboratories use neutral buffered formalin for tissue preservation. Formaldehyde crosslinks amino groups preventing loss of cellular mRNA (Basyuk et al. 2000). Underfixed tissue may lose cellular mRNA during autolytic cell breakdown or during tissue processing through extraction of cell components during alcohol dehydration. Over-fixed tissue can presumably lose signal by excessive crosslinkage of protein peptide bonds (Wilcox 1993).
Proteolytic digestion of formaldehyde-fixed tissue is necessary to break crosslinked peptide bonds and allow access to target nucleotides by ISH probes. To a certain extent, protease digestion can compensate for less than ideal fixation. However, protease digestion should be optimized to maximize signal through the unmasking of target nucleotides while still preserving morphology (Naoumov et al. 1988; Unger et al. 1989). Optimal protease digestion is highly dependent on both the duration of fixation and the type of tissue being digested (Fleming et al. 1992).
Although proteinase K is the standard enzyme used in ISH, other enzymes, including pepsin (Unger et al. 1989; Bromley et al. 1994), pronase (Unger et al. 1989; Weiss and Chen 1991), protease VIII (Fleming et al. 1992), protease XIV (Infantolino et al. 1989; Basyuk et al. 2000), and protease XXIV (Naoumov et al. 1988), have also been successfully incorporated into ISH protocols. Proteinase K (P-8038) isolated from the fungus Tritrachium album degrades many proteins even in their native state. Its predominant site of cleavage is the peptide bonds adjacent to the carboxyl group of aliphatic and aromatic amino acids with blocked α-amino groups. It is commonly used for its broad specificity.
Reactivity of mucin ISH expressed as percentage of area stained ± SEM (n = 15 determinations from five different samples)
Protease VIII and protease XXIV are essentially the same enzyme (Enzyme Commission No. 3.4.21.62), both isolated from Bacillus subtilis. These are serine proteases not specific for a given peptide bond but demonstrating preference for hydrolyzing at the carboxyl side of large uncharged amino acid residues (tyrosine, asparagine, and glutamine).
Fleming et al. (1992) found that protease VIII exhibited a wider range of optimal digestion than proteinase K when used for ISH. However, these studies did not compare the effects of these enzymes for use with clinical specimens that had been variably fixed and processed. In our studies, we found protease XXIV to be the enzyme of choice for protease digestion of archival tissue when used for detection of human mucin gene expression. We expect that protease XXIV digestion will aid in the detection of other mRNA species when used in ISH investigation of archival tissues.
Footnotes
Acknowledgements
Approved by the University of California Human Subjects Review Committee and supported by the Cigarette and Tobacco Surtax Fund of the State of California through the Tobacco Related Disease Program, grant 9RT-0214.
