Abstract
LRP5 is a novel member of the low-density lipoprotein receptor family that is genetically associated with Type 1 diabetes. As a start to defining the normal function of LRP5 and to generate testable hypotheses of its potential role in Type 1 diabetes pathogenesis, we carried out an extensive expression analysis of this gene at the mRNA and protein levels in normal human, monkey, and mouse, as well as in non-obese diabetic (NOD) mice at several stages of diabetes development. In all species, expression of LRP5 was found in four functionally important cell types: the distributed mononuclear phagocyte system, the islets of Langerhans, vitamin A-metabolizing cells, and CNS neurons. Given the critical role of macrophages in the onset and progression of islet cell destruction in Type 1 diabetes and the hypothesized role of retinoids as modifiers of diabetes progression, these findings suggest that LRP5 may confer Type 1 diabetes risk by altering the normal functioning of one or more of these regulatory systems. Specifically, given that the LRP5 polymorphisms associated with diabetes are in the promoter region of the gene, alterations in LRP5 expression may be responsible for diabetes susceptibility and therefore may be potential targets for therapeutic intervention.
T
The lipoprotein receptor family is made up of a rapidly expanding number of structurally related proteins which serve a variety of functions in lipid metabolism and signal transduction (Nakagawa et al. 1998).
The original member of this family, the LDL receptor, mediates uptake of cholesterol from the circulation by receptor-mediated endocytosis of lipoproteins (Hey et al. 1998). However, other members of this family have been shown to have functions unrelated to lipoprotein metabolism (Strickland et al. 1995). LRP1, for example, mediates uptake of chylomicron remnants, but in addition serves as a receptor for the protease inhibitor α2-macroglobulin, and has been shown to bind over 25 other ligands. LRP1 therefore appears to be a general scavenger receptor (Krieger and Herz 1994; Willnow et al. 1999). More recently other members of the family, including megalin, the VLDL receptor, and the apolipoprotein receptor 2, (ApoER2) have been shown to play critical roles in such diverse physiological functions as steroid and retinoid uptake, and neuronal migration during brain development (Christensen et al. 1999; Trommsdorff et al. 1999). The ligands and functions of LRP5 have not been fully elucidated, but it has thus far been proposed to mediate the endocytosis of ApoE-enriched lipoprotein particles (Kim et al. 1998) and to participate in cell proliferation (Dong et al., 1998). Although the extracellular domain of LRP5 consists of the protein motifs present in other members of the LDL receptor family, they are arranged in a unique pattern (Hey et al. 1998), suggesting that its ligand specificity may be different from that of other members of the family.
To begin to understand the biology and function of LRP5, we have examined in detail its expression in non-diabetic human and monkey tissues as well as in non-diabetic mice and in the Type I diabetes model, the non-obese diabetic (NOD) mouse. We show that, in all species, LRP5 is expressed in the distributed phagocytic system, in the pancreatic islets, in retinoid-storing cells, and in neurons of the CNS. These patterns suggest that LRP5 may play a role in several physiologically important cell signaling systems and suggest several potential pathogenic mechanisms by which LRP5 may contribute to Type I diabetes susceptibility.
Materials and Methods
Tissues
Human tissues were obtained from the National Disease Research Interchange (NDRI; Philadelphia, PA). Human thymoma samples were a generous gift of Dr. I. Roxanis (John Radcliffe Hospital; Oxford, UK). African Green monkey (Chlorocebus aethiops) and non-diabetic Swiss-Webster mouse tissues were obtained though Laboratory Animal Resources at Merck Research Laboratories. NOD mice (Podolin et al. 1993), were obtained from Drs L. Wicker and L. Peterson at Merck Research Laboratories.
Antibodies
Anti-human Antibody. Peptide SYRHLRPPPPSPCTDSS from the C-terminus of LRP5 was synthesized by Research Genetics (Huntsville, AL) as an eight-branch multiple antigenic peptide (MAP) with a molecular weight of 16,008. Antibody to the peptide was produced in two rabbits according to established protocols by Cocalico Biologicals (Reamstown, PA). The antibody was used either directly in crude serum or after affinity-purification. Affinity-purification was performed on a 1 ml BioRad Affi-Gel 15 column (Hercules, CA) to which 2 mg of the MAP peptide was bound covalently. The antibody was eluted with 100 mM Glycine-HCl, pH 2.5, and the eluant was neutralized with 1 M phosphate, pH 8.0. The affinity purification was monitored by ELISA.
Anti-mouse Antibody. A polypeptide corresponding to amino acids 281–533 of the extracellular domain of mouse LRP5 was produced as follows: PCR primers 5′-AAGCTCAGCTTCATCCACCG-3′ and 5′-GAGCTCCCGTCTATGTTGATCACCTCG-3′ were used to amplify a 965-nucleotide product using plasmid DNA containing the full-length LRP5 cDNA as the template. The PCR fragment was digested with the restriction endonucleases Nco I and Sac I and subcloned into the NcoI and Sac I sites of pET-32b (Novagen; Madison, WI). The plasmid construct was transformed into AD494(DE3) pLysS (Novagen). The resulting thioredoxin:LRP5 fusion protein was expressed and purified using a His-Bind Resin (Novagen) according to the manufacturer's instructions. Imidizole elution from the His-Bind resin yielded approximately 0.6 mg of purified protein per 100 ml bacterial culture. The purified protein was used to immunize two rabbits from CoVance (Denver, PA).
Recombinant Adenoviruses
Adenoviruses containing the mouse and human LRP5 genes were constructed to provide cells expressing high levels of recombinant protein. The cDNAs for hLRP5 and mRP5 were fused to the human CMV promoter and the bovine growth hormone polyadenylation signal and cloned into the adenovirus shuttle vector pΔE1sp1A (Bett et al. 1994) with the transcription unit directed towards the packaging signal. The complete virus genomes were created by homologous recombination in E. coli BJ 5183 (Chartier et al. 1996) between the shuttle vector and plasmid pE1-E3. The viral DNAs were released from the plasmid by restriction digestion, transfected into 293 cells by calcium phosphate co-precipitation, and plaques were obtained after 9 days. The resulting Ad 5-based viruses carry the LRP5 gene in E1 and have the E3-deletion of BHG10 (Bett et al. 1994). Viruses were amplified, and large-scale stocks were grown in NUNC cell factories and purified by double Cs-banding. For expression studies, 293 cells and COS cells were infected with 200 and 3000 particles/cell, respectively, and harvested 48 hr and 72 hr after infection.
Western Blotting Analysis
Proteins were separated by SDS-PAGE on a 10% acrylamide Tris-glycine gel (Novex; San Diego CA). Proteins were transferred to a PVDF nylon membrane by electrophoresis. The membrane was blocked for 1 hr with 5% milk in Tris-buffered (pH 7.4) saline containing 0.1% Triton X-100. Primary antibody incubations were performed using a 1:500 dilution of crude serum for 1 hr at room temperature (RT). A goat anti-rabbit horseradish peroxidase conjugate (Amersham; Piscataway NJ) was used as the second antibody. Bands were detected using ECL reagents (Amersham) and exposing the blot to Hyperfilm MP (Amersham).
Immunohistochemistry
Cryostat sections of 8-week-old Swiss-Webster mouse pancreases and NOD mouse were thaw-mounted on Superfrost Plus glass slides (Fisher Scientific; Pittsburgh, PA) and fixed for 15 min in 4% paraformaldehyde. Sections were then blocked with 10% normal donkey serum in PBS and incubated with anti-LRP5 AB #884 for 2 hr at RT. Bound antibody was detected with Texas Red-conjugated donkey anti-rabbit (Jackson Immunoresearch; Avondale, PA). Sections were counter-stained with 4′-6-diamidino-2-phenylindole (DAPI; Molecular Probes, Eugene, OR) to visualize nuclei. LRP5-positive sections were then rinsed, blocked again with 10% normal donkey serum, and then incubated with either guinea pig anti-glucagon antisera (Linco Research; St. Charles, MO), or guinea pig anti-insulin (Dako; Carpinteria, CA). Signal was visualized with FITC-conjugated anti-guinea pig IgG (Jackson Immunoresearch). Fluorescent images were captured using a CCD Spot Cam (Phase 3 Imaging Systems; Milford, MA), recompiled in Adobe Photoshop (San Jose, CA), and co-localization was indicated by yellow fluorescent signal. Sections of mouse spleen and thymus were similarly prepared, stained, and analyzed using the same double immunofluorescence staining technique with anti-LRP5 AB #884 and the anti-pan-macrophage marker BM8 (BMA Biomedicals; Augst, Switzerland).
Fresh frozen human liver specimens (NDRI) were cryo-sectioned at 8 μm and fixed in 4% paraformaldehyde in PBS for 1 hr. Antigen retrieval was performed on the sections by immersion in AR-10 antigen retrieval solution (Biogenex; San Ramon, CA) and pulse-microwaved to 100C for 10 min. Sections were rinsed, blocked in 10% normal donkey serum in PBS, and stained with a mixture of anti-LRP5 antibody #171 and either macrophage or dendritic marker clones RFD7 (Serotec; Oxford, UK), RFD1 (Serotec), or 19.2 mannose receptor (Pharmingen; San Diego, CA), for 18 hr at 4C in PBS with 1% normal donkey serum. Secondary antibody was FITC- or Texas Red-conjugated donkey anti-mouse or donkey anti-rabbit (Jackson Immunoresearch).
In Situ Hybridization
The plasmid Bluescript SK+ containing a 181-bp cDNA insert coding for LRP5 was linearized with either HindIII or BamHI (Life Technologies; Lexington, KY) to create template DNA. T7 and T3 polymerases were used with Biotin RNA Labeling Mix (Boehringer Mannheim; Mannheim, Germany) to in vitro-transcribe antisense and sense riboprobes, respectively. In situ probes specific for mouse LRP5 were prepared in the same manner, linearizing the Bluescript plasmid containing a 178-bp specific fragment for mouse LRP5. All chromogenic in situ hybridization was carried out as previously described (Lynch et al. 1999) on 8-μm paraffin tissue sections of 8-week-old Swiss-Webster mice perfused with 4% paraformaldehyde for optimal RNA preservation. mRNA in situ signal was detected histochemically using the Tyramide Signal Amplification—Indirect Kit (NEN Life Sciences; Boston, MA) in combination with the ABC Elite Kit (Vector Labs; Burlingame, CA). Signal was visualized with 3,3′-diaminobenzidine tetrahydrochloride (DAB; Sigma, St. Louis, MO) in nickel enhancement buffer (Digene; Gaithersburg, MD). Fluorescent in situ hybridization was similarly performed on cryostat sections (6 μm) processed for hybridization in a similar manner. Briefly, sections were fixed in 4% paraformaldehyde, digested with 10 μg/ml of proteinase K for 10 min, re-fixed in 4% paraformaldehyde, and blocked for endogenous peroxidases in 3% H2O2/100% methanol for 20 min. In situ hybridization was carried out as above using the TSA Direct Red FISH kit (NEN Life Sciences) according to the manufacturer's instructions. Sections were counterstained with DAPI (Molecular Probes). Rhesus monkey eyes were enucleated and fixed by immersion in 4% paraformaldehyde for 8 hr. The lenses were removed, the eyes were dissected radially and cryoprotected in graded sucrose solutions, and a sucrose/Tissue-Tek OCT Compound (Miles; Naperville, IL) mixture and frozen on dry ice (Barthel and Raymond 1990). The TSA Direct Blue FISH kit (NEN Life Sciences) was used for signal detection on the monkey retina sections to avoid obscuration by pigmented epithelium. Sections stained with TSA blue coumarin reagent were mounted with Vectashield mounting medium containing propidium iodide (Vector Laboratories).
Combined In Situ Hybridization/Immunohistochemistry
Immunohistochemistry was performed sequentially on sections that showed optimal fluorescent signal for LRP5 mRNA. After in situ hybridization, sections were rinsed in PBS, blocked in 10% normal donkey serum, and incubated with appropriate profiling antibody (see Table 1) according to the manufacturer's recommended conditions. Antibody binding was detected with either the appropriate FITC-conjugated antibody (Jackson Immunoresearch) or FITC-conjugated streptavidin (Vector Labs). Sections were coverslipped with Prolong anti-fade mounting medium (Molecular Probes). Images were digitally acquired and reassembled using a Nikon E1000 epifluorescence microscope and a Spot Cam CCD camera (Phase 3 Imaging Systems).
Results
Characterization of Anti-LRP5 Antibodies
Antibodies directed against either the C-terminal 17 amino acids of human LRP5 (AS171), or a 253-amino-acid portion of the extracellular domain of mouse LRP5 (AS884), were characterized by Western blotting analysis (Figure 1). Cell membranes were prepared from HEK-293 cells infected with an adenovirus containing either the human LRP5 cDNA, the mouse Lrp5 cDNA, or a control insert. The mature LRP5 protein has a predicted molecular weight of 176,666. A band of approximately MW 175,000 is detected by the AS884 antibody in membranes prepared from cells infected with the mouse LRP5 cDNA but not from cells infected with either the human LRP5 cDNA or a control adenovirus. Therefore, it appears that the AS884 antibody is specific for mouse LRP5. In contrast, AS171 antibody recognizes a band of approximately 175 kD in both human and mouse and therefore is not selective for the species of the receptor. This result is not unanticipated because there is only a single amino acid difference between mouse and human in the 17-amino-acid peptide used for immunization.
Materials used in this study
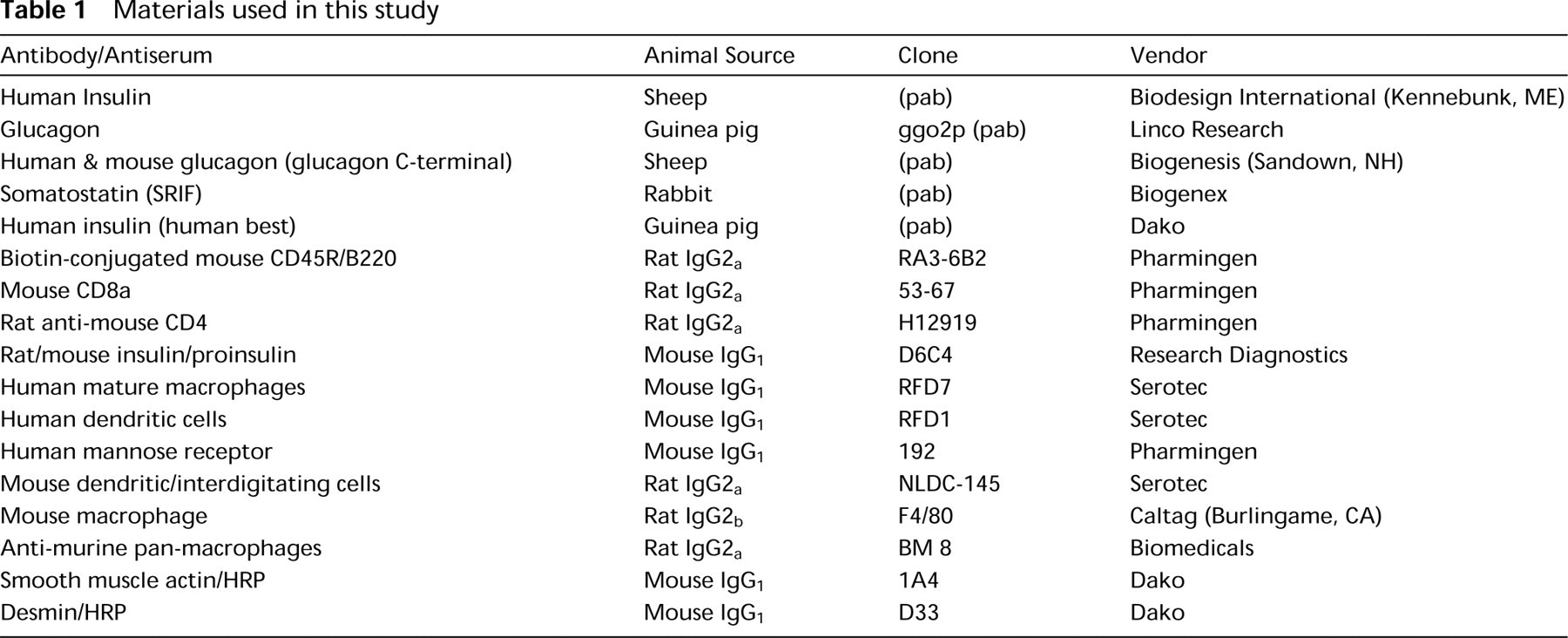
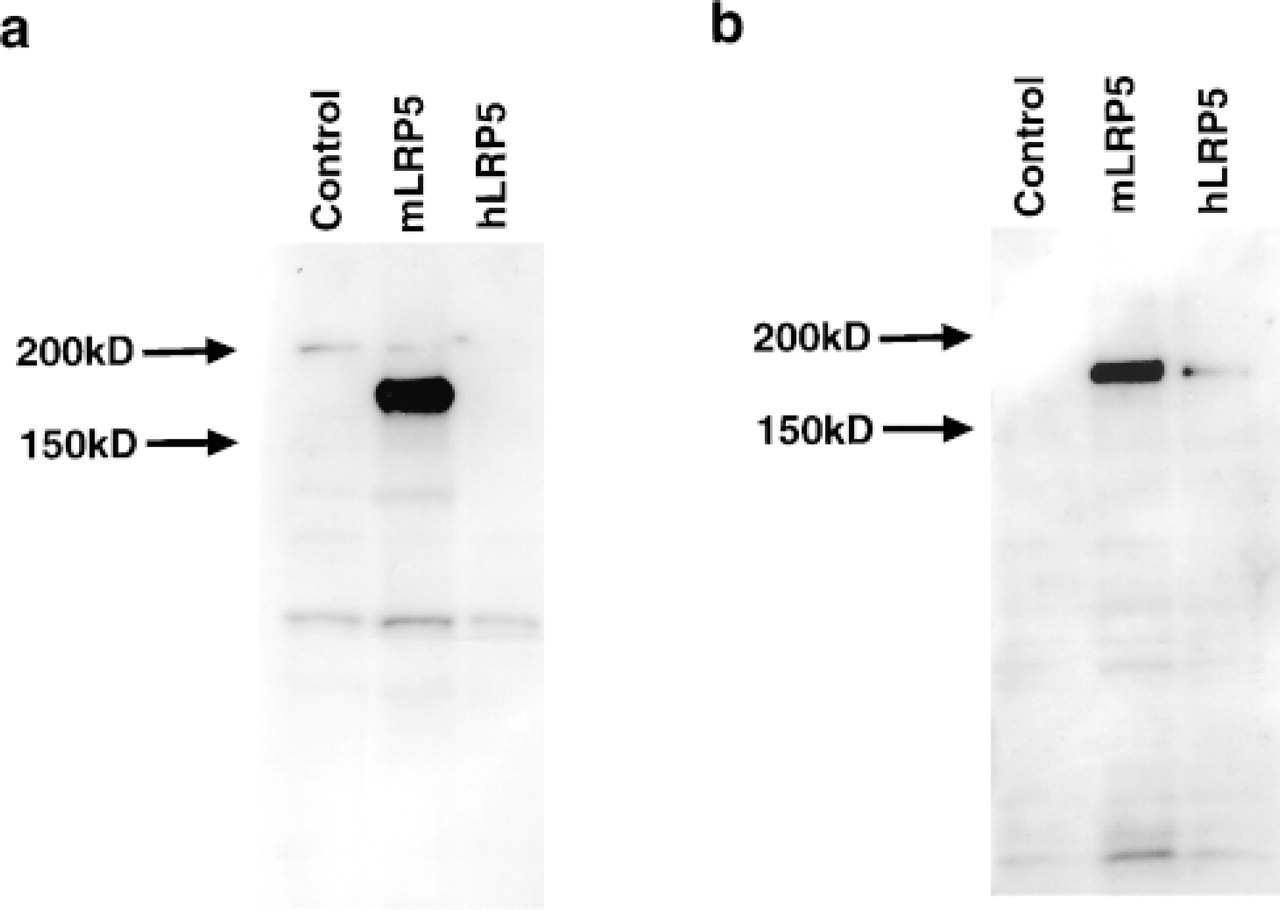
Western blotting characterization of anti-LRP5 antibodies. (
LRP5 Is Expressed in Pancreatic Islets
Because LRP5 was cloned as a susceptibility gene for Type I diabetes (Hey et al. 1998), expression was examined by in situ hybridization and immunohistochemistry in normal human pancreas as well as in Swiss-Webster and NOD mice. In the human and monkey pancreas, strong expression of LRP5 was seen in islets. Co-localization studies with cell type-specific antibodies identified the LRP5-expressing cells as insulin-producing β-cells (Figures 2j-2l). In addition to the β-cells, there were isolated LRP5-expressing cells in the parenchyma of the pancreas which, by appearance and immunohistochemical characterization, were identified as macrophages (Figures 2n-2o).
In Swiss–Webster and C57BL/10SnJ mice, pancreatic LRP5 expression was also found mainly in the islets but, in contrast to the human and monkey pancreas, immunolocalization studies demonstrated the LRP5-expressing cells in the mouse to be glucagon-expressing α-cells (Figures 2a-2c). In the 1-month-old NOD mouse, expression of LRP5 was indistinguishable from that in non-diabetic Swiss-Webster mice, showing localization to the α-cells. Figure 2h shows the typical location of the α-cells in the periphery of the islet. In 3-month-old NOD mice, there was severe leukocytic infiltration of the islets characteristic of NOD insulitis. No insulin immunoreactivity was seen in these islets, indicating that the expected β-cell destruction had indeed occurred. In these islets, LRP5-expressing cells were still present and still co-expressed glucagon (Figure 2i). No LRP5 expression was seen in the cells infiltrating these NOD islets, although isolated LRP5-positive macrophage-like cells similar to those seen in the human and monkey pancreas were also seen in the mouse pancreatic parenchyma (Figure 2m). Immunohistochemical analysis revealed that these cells were immunoreactive for the pan-macrophage marker BM-8 (Table 1; and data not shown).
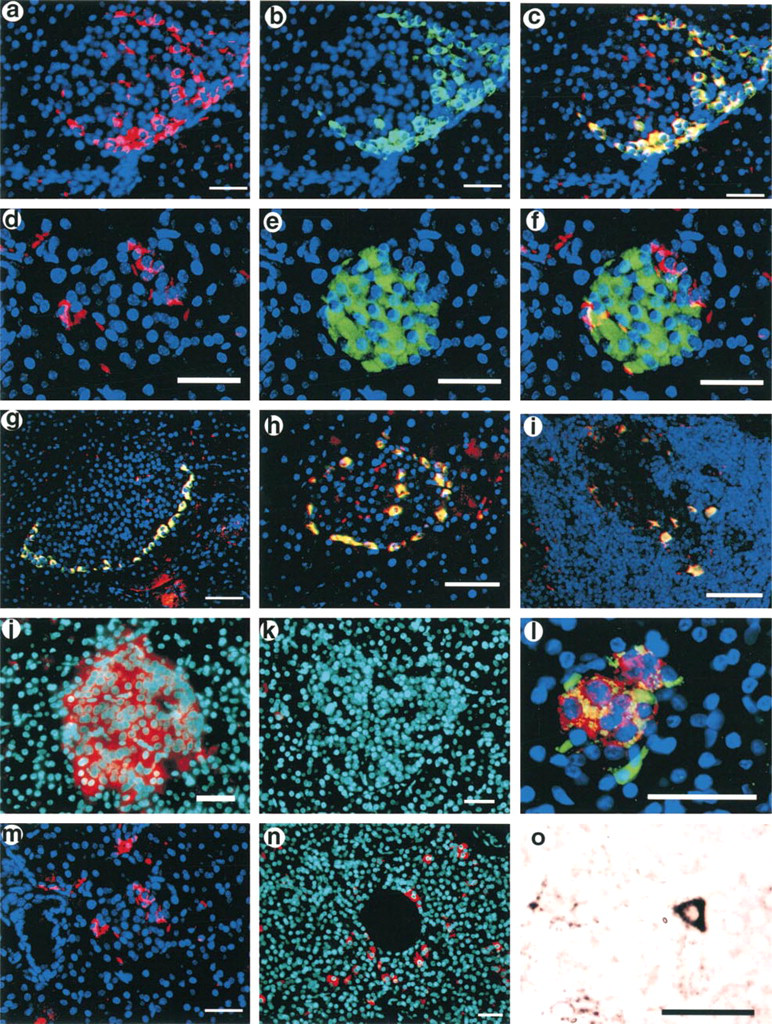
Expression of LRP5 in the islets of Langerhans. (
Macrophage expression. (
LRP5 Is Expressed in Macrophages of Human and Mouse Spleen and Thymus, Human Liver, and in a Macrophage Cell Line
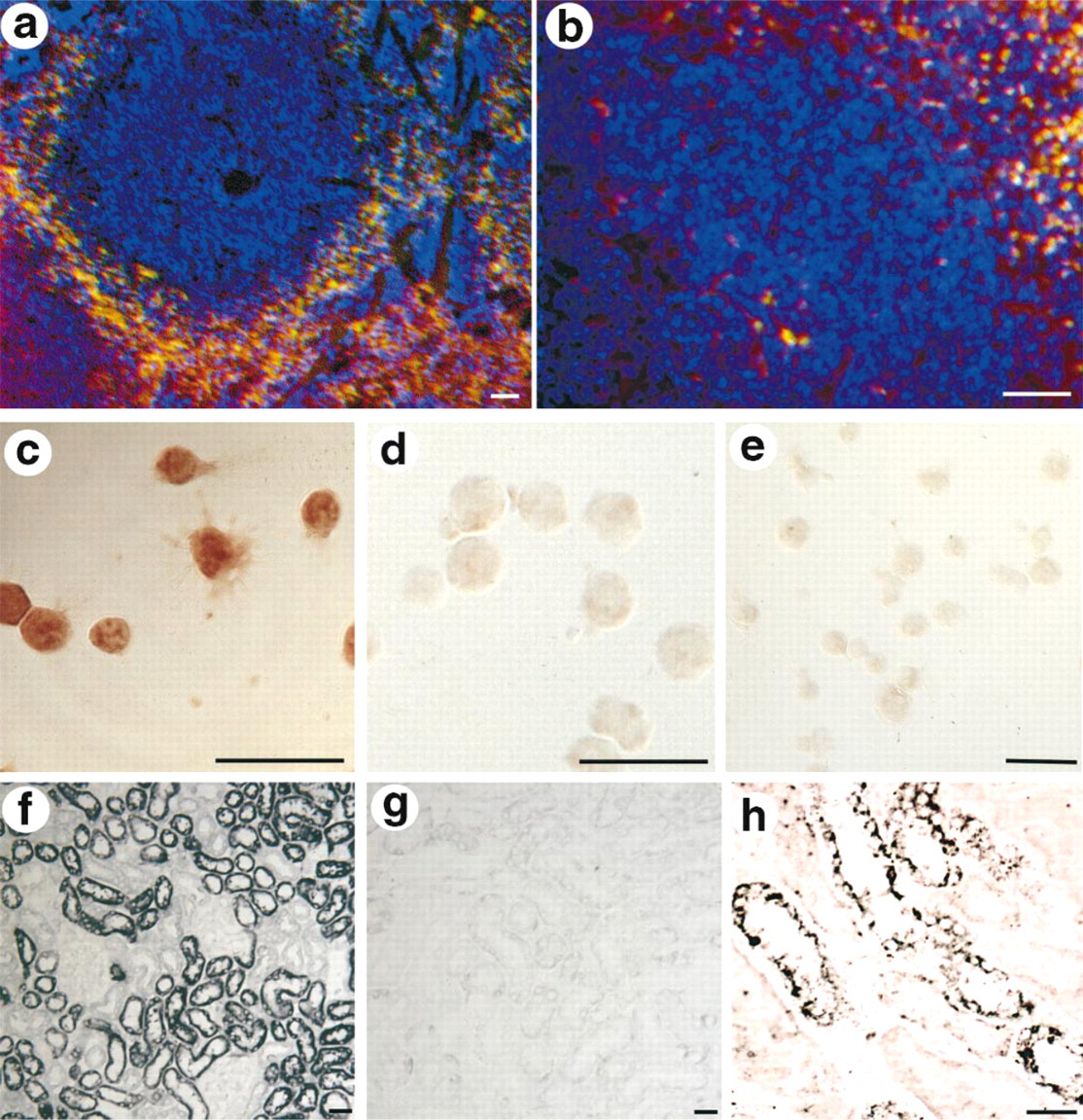
Because LRP5 has been implicated in an autoimmune disease (Type 1 diabetes) and expression was seen in macrophages of the pancreas, LRP5 expression was examined in other tissues with larger populations of macrophages. In human and mouse spleen, LRP5-expressing cells were seen in the non-filtering areas of the red pulp, the location characteristic of macrophages (Van Krieken and te Velde, 1997), and also were positive for the macrophage markers F4/80 and BM-8 (Table 1; Figures 3a and 3b). In mouse thymus and human thymoma, LRP5 expression was seen in both the cortex and medulla, in cells that were immunoreactive for macrophage markers, again consistent with macrophage identity (data not shown). Further suggesting that LRP5 is expressed in macrophages, strong LRP5 immunoreactivity was also seen in the THP.1 macrophage cell line but not in the Jurkat T-cell line (Figures 3c–3e). In the THP.1 cells, immunoreactivity was primarily membrane-localized, consistent with the expected transmembrane topology of LRP5. In the human liver, LRP5 was seen in Kupffer cells, particularly in those liver specimens with pathological evidence of hepatic congestion. These cells were identified by their bean-shaped morphology lining the liver sinusoids, and by co-expression with the dendritic cell markers RFD7, RFD1, and the mannose receptor (data not shown).
Renal Tubules Express LRP5
Northern blotting analysis had shown that LRP5 is most highly expressed in the liver and kidney (Hey et al. 1998). To define the kidney expression at a cellular level, in situ hybridization was carried out. In both mouse (Figures 3f and 3g) and human (Figure 3h) kidney, the convoluted tubules strongly expressed LRP5, whereas the glomeruli and collecting ducts showed no expression.
Vitamin A-storing Stellate Cells of the Liver Express LRP5
Given the prominent LRP5 expression in the liver by Northern blotting (Hey et al. 1998) and the high-level expression of other LDL receptor family members in the liver, the expression of LRP5 in the liver was determined in detail. As noted above, LRP5 expression was seen in the Kupffer cells of the liver. However, the most prominent LRP5 expression in the liver of both human and mouse was seen in cells of the space of Disse, interposed between the sinusoidal lining cells and hepatocytes (Figures 4a–4h). This location is prototypical of hepatic stellate cells, also referred to as vitamin A-storing, perisinusoidal, or Ito cells (Kawada 1997; Senoo et al. 1998). Stellate cells also characteristically express the cytokeratin desmin and smooth muscle actin. Therefore, to confirm stellate cell identity, LRP5 in situ hybridization was combined with desmin or smooth muscle actin immunochemistry on the same sections of human liver. Both desmin and smooth muscle actin co-localized with LRP5 (Figures 4d–4h), confirming stellate cell identity.
LRP5 Is Expressed in the Retinal Pigmented Epithelium
Macrophages of the pancreas, spleen, and thymus, the THP.1 macrophage cell line, Kupffer cells of the liver, and hepatic stellate cells all express LRP5, and all perform phagocytic functions. Furthermore, the stellate cells of the liver are the main site of uptake and storage of vitamin A. We therefore asked if the other major phagocytic cell type that metabolizes and stores vitamin A, the cells of the retinal pigmented epithelium (RPE), also express LRP5. In situ hybridization on Rhesus monkey (Figure 4i) and Swiss–Webster mouse retina (not shown) demonstrated strong LRP5 expression in the RPE of both species. In addition, there was weaker expression in the inner segments of the photo-receptor layer (Figure 4i), which is the location of the retinal rods and cones, which utilize vitamin A-derived retinoids as their visual pigment.
LRP5 Is Expressed in CNS Neurons
In addition to its expression in the RPE and photoreceptors, LRP5 was seen to be expressed in the ganglion cells of the retina (Figure 4i). These are the projection neurons of the retina, which share many properties with other CNS neurons (Dowling 1987). We then asked whether LRP5 was expressed more broadly in the CNS. Northern blotting analysis demonstrated that LRP5 is expressed widely in the human brain (data not shown). To define this expression more precisely, in situ hybridization analysis of LRP5 in mouse, monkey, and human brain was carried out. It showed that LRP5 is expressed selectively in the cerebral cortex, hippocampus, and hypothalamus (Figure 5). The LRP5-expressing cells were neuronal in morphology (Figures 5c and 5e) and co-labeled with neuronal markers (data not shown). In the hippocampus, expression was seen most strongly in the CA2–4 subregions (Figure 5f).
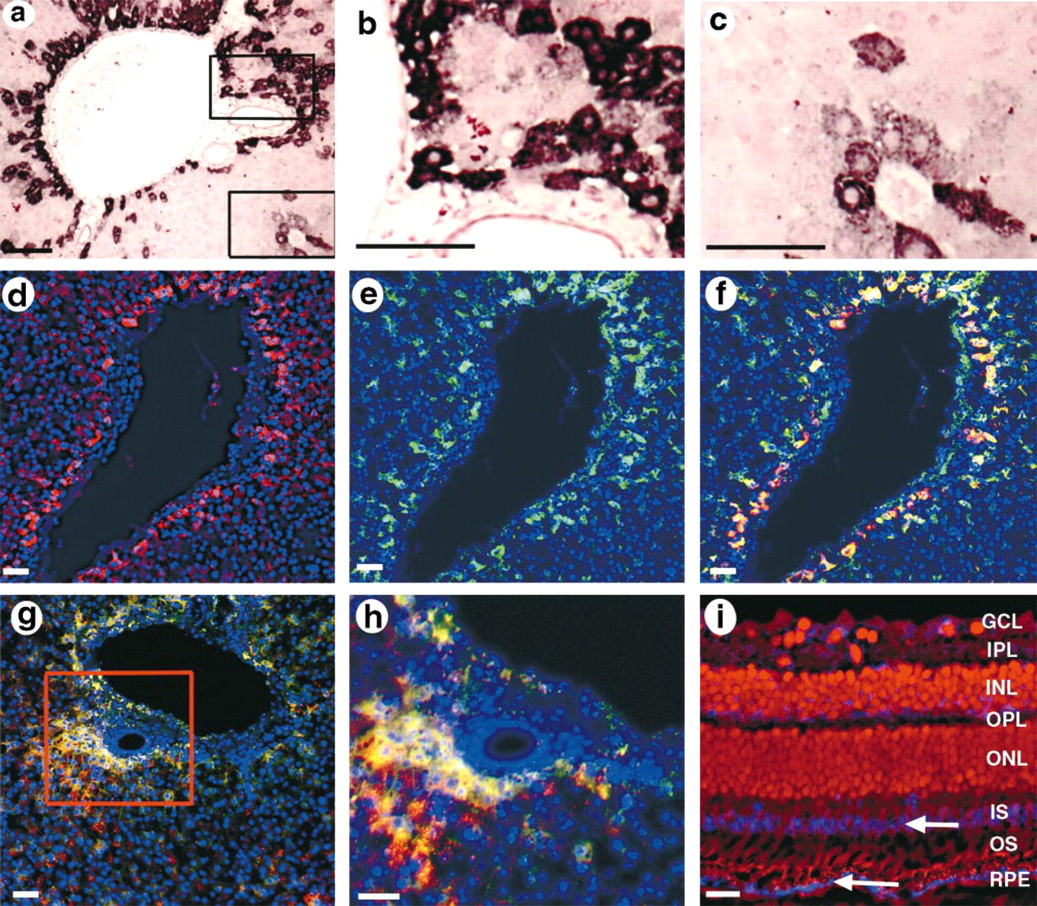
LRP5 expression in vitamin A-metabolizing cells. (
Discussion
LRP5, a novel member of the LDL receptor superfamily, was originally cloned on the basis of its genetic association with Type I diabetes in humans (Hey et al. 1998). Genetic studies have provided statistical evidence for the association of markers that are in linkage disequilibrium with LRP5 (Nakagawa et al. 1998). However, additional analysis is required to confirm the hypothesis that polymorphic variation in LRP5 confers risk for developing diabetes. The two associated markers are in the 5′ untranslated region of LRP5, and the association does not appear to be due to protein coding polymorphism (unpublished data). This raises the possibility that variation in transcriptional regulation, and therefore in gene expression level or specificity, may account for the association of LRP5 with Type I diabetes. As a start to determining the normal function of LRP5 and its potential role in type I diabetes, we undertook a detailed study of LRP5 expression in normal human, mouse, and NOD mouse tissues, with the goal of obtaining functional information that could be tested experimentally. Cellular resolution of LRP5 expression by in situ hybridization combined with cell type-specific immunohistochemistry demonstrated LRP5 expression in four functionally important cell types: the distributed mononuclear phagocyte system, the islets of Langerhans, retinoid-storing and -metabolizing cells, and CNS neurons. Each of these findings suggests potential functions for LRP5 and potential etiological connections to disease.
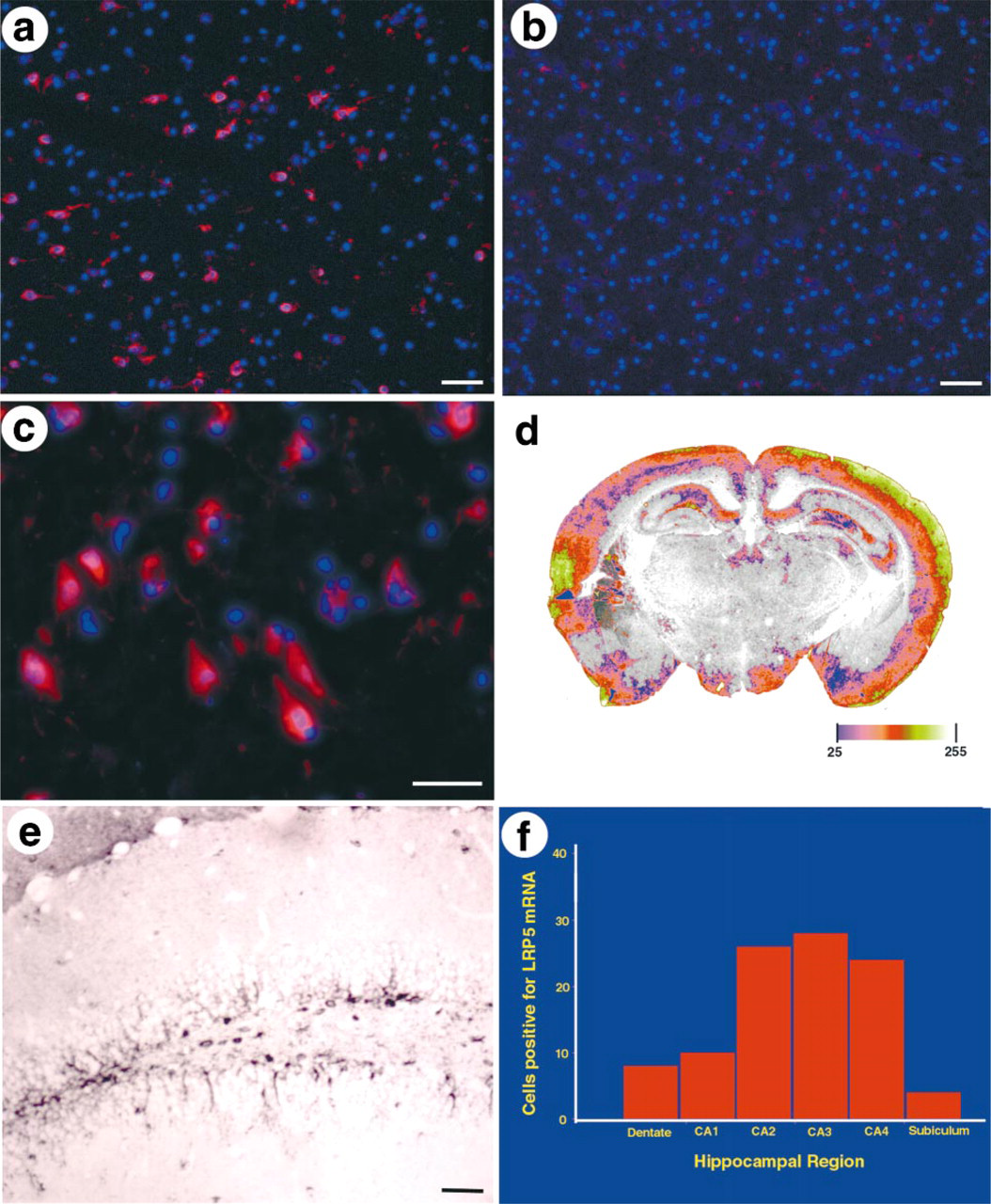
LRP5 expression in the brain. (
In the distributed phagocytic system, LRP5 expression was observed in tissue macrophages of the pancreas, spleen, and thymus, the macrophage cell line THP.1, the Kupffer cells of the liver, and the pigmented epithelium of the retina. Macrophages and dendritic cells are the first cells to infiltrate the islets of NOD mice (Delovitch and Singh 1997) and are believed to play a critical role in the destruction of β-cells in Type I diabetes via production of inflammatory cytokines that attract cytotoxic T cells (Arnush et al. 1998; Bradley et al. 1999). This finding raises the possibility that altered LRP5 may increase the risk of type I diabetes by altering the function of effector immune cells.
High-level LRP5 expression was seen in pancreatic islets, the target of immune destruction in Type I diabetes. Islet expression was seen in all species examined but, intriguingly, the islet cell type that expresses LRP5 was seen to differ between primates and rodents. In human and African green monkey pancreas, LRP5 was expressed in insulin-expressing β-cells, the cells destroyed in Type I diabetes. In contrast, in wild-type Swiss-Webster mice and in NOD mice at various stages of diabetes development, LRP5 was clearly expressed in the glucagon-expressing α-cells. Consistent with this finding, the number of LRP5-expressing cells was maintained at a normal level even in fully diabetic NOD mice. The reason for this species difference is not clear, but it suggests a difference in the role of LRP5 in the pathogenesis of Type I diabetes in the two species. The fact that β-cells are normally antigen-presenting, whereas α-cells are not, supports this contention (Papaccio et al. 1998). To date, the region of the mouse genome that contains the mouse LRP5 orthologue has not been linked to NOD diabetes.
Several cell types that transport, store, and metabolize retinoids expressed LRP5, including the stellate (Ito) cells of the liver, the tubule epithelium of the kidney, and the pigmented epithelium and photoreceptors of the retina. Vitamin A has been implicated in the development and/or progression of autoimmune diabetes in the BB rat (Driscoll et al. 1996), as well as in humans (Basu and Basualdo 1997). Therefore, the expression of LRP5 in these specialized cells is of potential pathophysiological significance. Vitamin A metabolism is complex and involves several poorly understood receptor-mediated events in which LRP5 can be speculated to play a role. For example, the vitamin A-containing chylomicron remnants are taken up by hepatocytes and other cell types via the LDL receptor and LRPs, which potentially include LRP5. In hepatocytes, retinol is transferred (bound to retinol-binding protein, RBP) to the hepatic stellate cells via a receptor that has not yet been identified (Blomhoff 1994); LRP5 is also a candidate for this receptor. Under physiological conditions, stellate cells regulate retinoid homeostasis and hepatic sinusoidal blood flow and, in response to liver injury and inflammation, produce multiple proinflammatory cytokines (Kawada 1997; Senoo et al. 1998). Similar vitamin A-storing cells are found in the human pancreas and display similar responses to inflammation (Ikejiri 1990; Nagy et al. 1997). It can be speculated that some of the isolated LRP5-expressing cells we observed in the human and monkey pancreas are stellate cells; this possibility is currently under study. The expression of LRP5 in the retinal pigmented epithelium and retinal photoreceptors further suggests that LRP5 may play a role in vitamin A physiology or cholesterol transport processes, both of which are essential for normal RPE function and are implicated in disease (Holz et al. 1994). To assess the potential role of LRP5 in any of these processes, it will be important to determine the ligand(s) for LRP5, and particularly whether cholesterol, or vitamin A complexed to RBP or in chylomicron remnants, can be taken up via LRP5.
LRP5 expression was detected in CNS neurons, and several data suggest the possibility of a role for LRP5 in Alzheimer's disease pathogenesis. In situ hybridization studies showed that in human, monkey, and mouse, LRP5 is highly expressed in the cerebral cortex, hippocampus, hypothalamus, and retina, all of which are preferentially affected in Alzheimer's disease. Alzheimer's disease has been genetically associated with the LRP1 ligands α2-macroglobulin and APOE. APOE has been reported to also bind LRP5 (Kim et al. 1998). Although there is no reported genetic linkage of Alzheimer's disease to the 11q13 LRP5 region, the related gene LRP6 (Brown et al. 1998) is also expressed in the human hippocampus and maps to the region of chromosome 12 linked to Alzheimer's disease (Scott et al. 1999). It will be important to assess the possible genetic association of LRP5 and LRP6 with Alzheimer's disease and other disorders.
Comparison of LRP5 expression with that of other members of the LRP gene family reveals several similarities that suggest further clues to the function of LRP5 and other members of this gene family. Cellular localization data are available for only four of the eight currently known LRPs. LRP1, like LRP5, is expressed selectively in macrophages, CNS neurons, hepatocytes, and smooth muscle cells (Gliemann 1998). LRP2, also termed GP 330 or megalin, is expressed in renal tubule epithelium (Gliemann 1998; Korenberg et al. 1994), as is LRP5 (Figures 3f–3h). On a functional level, although many of the LRPs can mediate lipoprotein endocytosis, this may not be their primary function (Willnow et al. 1999). LRP1 binds and endocytoses a variety of proteinases and inhibitor complexes, including α2-macroglobulin, tissue and urinary plasminogen activator, lipoproteins, including APOE-enriched lipoproteins and lipoprotein lipase, and extracellular matrix proteins, such as thrombospondin (Gliemann 1998; Strickland et al. 1995). LRP5 may share some of these ligands, given its expression in macrophages and CNS neurons. In addition to binding lipoproteins, LRP2 (megalin) specifically mediates the uptake and storage of vitamin B12 by the kidney (Moestrup et al. 1996). The expression pattern of LRP5 suggests a possibly analogous role for LRP5 in vitamin A homeostasis. The finding of LRP5 expression in the pancreatic islets and macrophages is consistent with the possibility, based on genetic data, that LRP5 is a Type I diabetes gene. Given that the genetic association of LRP5 with Type I diabetes is found in the 5′ region of the gene, we speculate that diabetes risk may be conferred by changes in LRP5 expression level or tissue expression specificity. Testing of the effect of diabetes-associated LRP5 polymorphisms on the expression of luciferase or other reporters will begin to address this question, and should be done in islet or macrophage cell lines, given the normal LRP5 expression pattern demonstrated here.
Our knowledge of the biology of the LRPs is certainly incomplete, but the data currently suggest that this gene family is important in multiple regulatory networks, including lipoprotein metabolism, immune regulation, and vitamin homeostasis. It is likely that the LRPs, already genetically associated with Type 1 diabetes and with Alzheimer's disease (Kamboh et al. 1998), will be implicated in other conditions affecting human health and therefore may provide novel therapeutic targets for these diseases.
