Abstract
We studied the expression of prolactin (PRL) mRNA in the mammary gland of resting, pregnant, lactating, and weanling rats using in situ and solution reverse transcriptase-polymerase chain reaction (RT-PCR). In mid- to late pregnancy and throughout lactation, PRL mRNA was detected in both in situ and solution RT-PCR. These PRL mRNA signals were clearly identified in the cytoplasm of alveolar and ductal mammary epithelial cells by the in situ RT-PCR method. In mid- to late pregnancy, such as at the initiating point of PRL mRNA expression, we confirmed in some cases a lack of PRL mRNA by solution RT-PCR. In addition, in the early weaning phase, no signals were detected by solution RT-PCR. However, slight focal signals were detected in some poorly vacuolated cytoplasm of regressing acinar cells by in situ RT-PCR. These findings suggest that PRL mRNA in rat mammary gland begins in mid- to late pregnancy in parallel with the development of the mammary gland, continues throughout lactation, and declines in the early phase of weaning, with regression of mammary epithelial cells.
P
To clarify the changes, we examined a recent PCR technique that combined reverse transcription (RT) to amplify and clearly detect low levels of mRNA in cells and tissues (Chen and Fuggle 1993; Heinford et al. 1993; Jin et al. 1995; Sanno et al. 1997). In situ hybridization is a useful technique to demonstrate gene expression in individual cells, but its application has been limited because of difficulties in detecting low copy numbers of mRNAs (Sanno et al. 1997). The rat mammary gland has a tendency to show nonspecific hybridization during the lactating period because of abundant milk proteins and because of its cellular fragility. The purposes of this study were to detect PRL mRNA in the rat mammary gland, to demonstrate the exact date of initiation of PRL transcription, and to show localization of PRL mRNA in tissue sections using a novel sensitive technique, in situ RT-PCR.
Materials and Methods
Study Groups
Thirty-six (12- to 20-week-old) female Wistar rats (Clea Japan; Tokyo, Japan) that had been mated at 12 weeks of age were used in this study. The rats were divided into nine groups of four rats each comprising nonpregnant (resting), Days 7 and 14 of pregnancy, Days 1, 7, and 14 of lactation, and Days 1, 7, and 14 after weaning. Four rats in each group were anesthetized with ethyl ether and sacrificed by exsanguination via the abdominal aorta. The mammary gland obtained from each animal was used for preparation of mRNA and in situ reverse transcriptase-polymerase chain reaction (RT-PCR). In addition, the pituitary gland of each animal was prepared for mRNA and served as a positive control for RT-PCR. Kidney and heart from resting rats served as controls for solution RT-PCR. All experiments were approved by the Japan Tobacco, Inc. Toxicology Research Laboratory institutional animal care and use committee.
RNA Extraction and Solution RT-PCR
The mammary gland tissues were homogenized in Trizol reagent (Life Technologies; Gaithersburg, MD) and total RNA was extracted according to the manufacturer's protocol. The yield was 0.2-37.8 μg/mammary gland. First-strand cDNA synthesis and amplification of cDNA were performed according to the method of Jin et al. (1995) and Fields et al. (1993). First-strand cDNA was synthesized using a firststrand synthesis kit (Stratagene; La Jolla, CA). The RT reaction was performed in a final volume of 50 μl with 5μg of total RNA, 300 ng oligo (deoxythymidine) primer, 1 × RT buffer, 1.0 mM of each deoxyribonucleotide [deoxy (d)-ATP, dCTP, dTTP, and dGTP], 40 U RNase inhibitor, and 50 U Moloney murine leukemia virus reverse transcriptase (MMLV-RT) at 37C for 60 min, heated at 95C for 5 min, and stored at −20C.
The sequences of the primers used for amplification are shown in Table 1. The primers were of the same sequences confirmed by Jin et al. (1995). PCR amplification was performed in 100 μl final reaction volumes containing 5 μl RT reaction product as a template DNA corresponding to cDNA synthesized from 1μg total RNA, 1 × PCR buffer (Promega; Madison, WI), 1.5 mM MgCl2, 0.2 mM of each deoxynucleotide (Sigma; St Louis, MO), 300 ng each of sense and antisense primer for PRL (Table 1), and 2.5 U Taq DNA polymerase. PCR reaction was carried out in a DNA thermal cycler (Perkin-Elmer/Cetus; Norwark, CT) using the following cycle conditions: 95C for 5 min for denaturing, followed by 30 cycles at 94C for 1 min, 60C for 1 min, and 72C for 2 min for denaturing, annealing, and extension, respectively. After the last cycle, a final extension at 72C for 10 min was performed. The control primer for glyceraldehyde-3-phosphate dehydrogenase (Clontech; Palo Alto, CA) was used to check the integrity of the RNA (Table 1).
The PCR products were analyzed by 2% agarose gel electrophoresis and stained with ethidium bromide. Omission of the MMLV-RT in the RT reaction was used as a negative control for the RT-PCR procedure, and the pituitary gland was used for the positive control.
The PCR reaction of PRL mRNA was carried out under conditions identical to those described above, with β-actin control primer (Clontech) as an internal control. In addition, we compared PRL mRNA in pituitary tissue with that of mammary gland at Day 14 of pregnancy. A total of 5 μg RT product was removed every five cycles at 10-40 PCR cycles.
Oligonucleotide primers and probe used for RT-PCR a
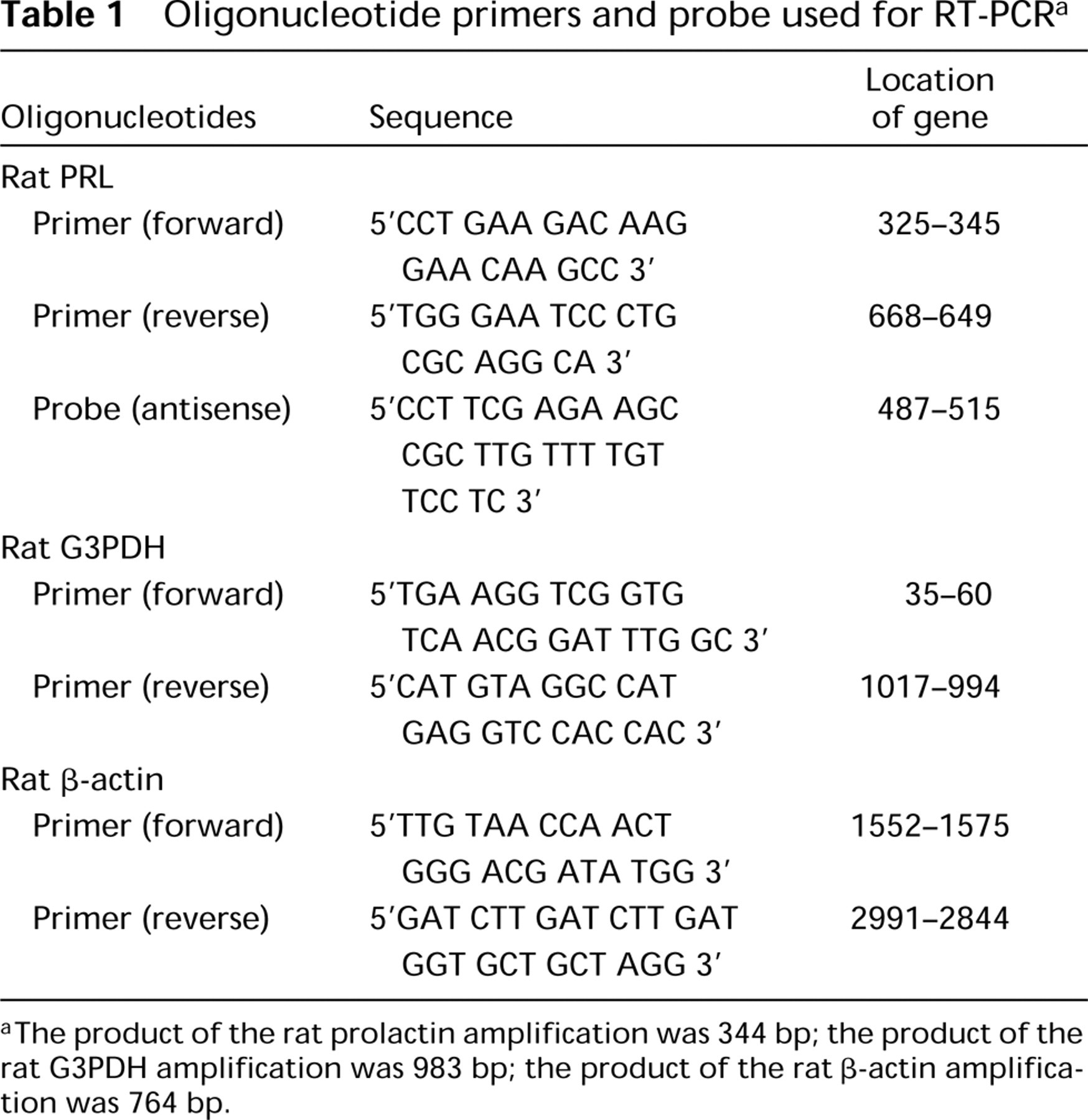
The product of the rat prolactin amplification was 344 bp; the product of the rat G3PDH amplification was 983 bp; the product of the rat β-actin amplification was 764 bp.
In Situ RT-PCR
Paraffin-embedded 4-μm tissue sections were submitted to in situ RT-PCR according to Jin et al. (1995). The oligonucleotide primers and probe used in the study are shown in Table 1.
After deparaffinization, sections were digested with 2μg/μl proteinase K (Promega) in 0.01 M PBS at 37C for 10 min and then rinsed in 0.01 M PBS. A total of 100 U/ml RNase-free DNase (Stratagene) in 0.01 M PBS was used to digest the DNA at 37C for 10 min, and then sections were rinsed in 0.01 M PBS at 80C for 10 min for proteinase K inactivation. After rinsing in 0.01 M PBS at room temperature (RT), the sections were incubated with 1 mg/ml L-α-lecithin (Sigma) in 0.01 M PBS at RT for 30 min and then washed in 0.01 M PBS and diethylpyrocarbonate-distilled water (DEPC-DW). The RT reaction was performed in a final volume of 50 μl/ specimen, 500 ng PRL antisense primer, 5 × first-strand buffer (Boehringer Mannheim; Indianapolis, IN), 0.01 M DTT (Boehringer Mannheim), 0.2 M of each deoxynucleotide (Sigma), 40 U/μl RNase inhibitor, and 8 U/μl Superscript II at RT. The reaction mixture was applied to each slide and covered with Parafilm (American National Can; Greenwich, CT) at 42C for 2 hr. The slides were washed in 2 × SSC, 1 × SSC, 0.5 × SSC, and DEPC-DW.
PCR amplification was performed in 50 μl/slide containing 1 × PCR buffer (Promega), 1.0 mM MgCl2, 0.2 m/M of each deoxynucleotide (Sigma), 0.1 M DTT, 500 ng each of sense and antisense primer for PRL (Table 1), and 5 U Taq DNA polymerase (Promega). Each PCR mixture slide was covered with a TAKARA slide seal for in situ PCR (Takara; Tokyo, Japan) and placed on a thermal cycler (Omnislide; Haybaid, Middlesex, UK) under the following cycle conditions: 95C for 3 min for denaturing, followed by 20 cycles at 94C for 2 min, 60C for 1.5 min, and 72C for 1.5 min for denaturing, annealing, and extension, respectively. After the last cycle, the final extension was maintained at 72C for 12 min. The slide seal was peeled off, fixed in 4% PFA in 0.1 M PB at RT for 5 min, and then washed in 2 × SSC.
Then the amplified products were hybridized with the oligonucleotide probe according to the method of Lloyd et al. (1991). Slides were hybridized with 1 ng/μl biotinlated oligonucleotide antisense probe (Table 1) at 37C overnight. The slides were washed in 2 × SSC, 1 × SSC, and 0.5 × SSC; then hybridization signals were detected with streptavidin-biotin-alkaline phosphatase using nitroblue tetrazolium-bromochloroindolyl phosphate (NBT-BCIP; Dako, Carpinteria, CA).
Omission of the Superscript II RT in the RT reaction and sense probe hybridized after PCR reaction were used as negative controls for the in situ RT-PCR procedure.
To determine the sufficiency of the PCR amplified condition, we carried out 5, 10, and 20 PCR cycles in the in situ RT-PCR procedure, using the mammary gland specimens at Day 7 of lactation.
Results
Expression of PRL mRNA in Rat Mammary Gland
To determine the appropriate PCR cycle number for mammary gland at Day 14 of pregnancy, we amplified the RT mixture from 10 to 40 cycles. In addition, PRL mRNAs were compared with those of the pituitary gland (Figure 1). In the pituitary gland, PRL mRNA was identified at 10 cycles of amplification. The mammary gland, at more than 25 cycles of amplification, showed RT cDNA for PRL. The PCR products for β-actin, a reverse transcribed housekeeping gene, were detected to almost the same degree in both the mammary and pituitary glands. PRL mRNA in the mammary gland was markedly lower than in the pituitary gland. Above 45 cycles, reverse transcribed cDNA for PRL was detected not only in the resting, early pregnancy, and late weaning phase of the mammary gland but also in the kidney and heart (not shown). In subsequent experiments, all mammary gland samples were analyzed at below 30 cycles in PCR amplification.
Typical solution RT-PCR analyses of PRL mRNA are shown in Figure 2. PRL mRNA in the rat mammary gland was detected from mid- to late pregnancy (Day 14 of pregnancy) and throughout lactation. No signals were detected in the resting period, early pregnancy, and mid- to late weaning. The initiating point of PRL mRNA expression in mid- to late pregnancy (Day 14 of pregnancy) showed PRL mRNA expression in three of four cases, and differences varied among individuals (Figure 3).
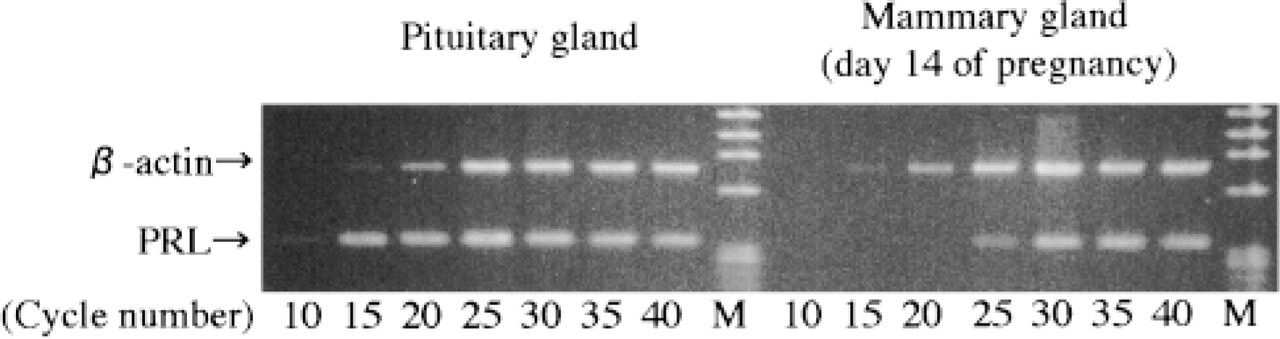
A comparison of PRL mRNA between pituitary gland and mammary gland by solution RT-PCR. Total 5 μg RNA was carried out by PCR amplification every five cycles from 10 to 40 cycles. No significant differences in β-actin mRNA of pituitary gland and mammary gland were found. PRL mRNA of pituitary gland detected fewer cycles than that of the mammary gland. Thirty cycles as a sufficient amplification was chosen for subsequent experiments. M, marker.
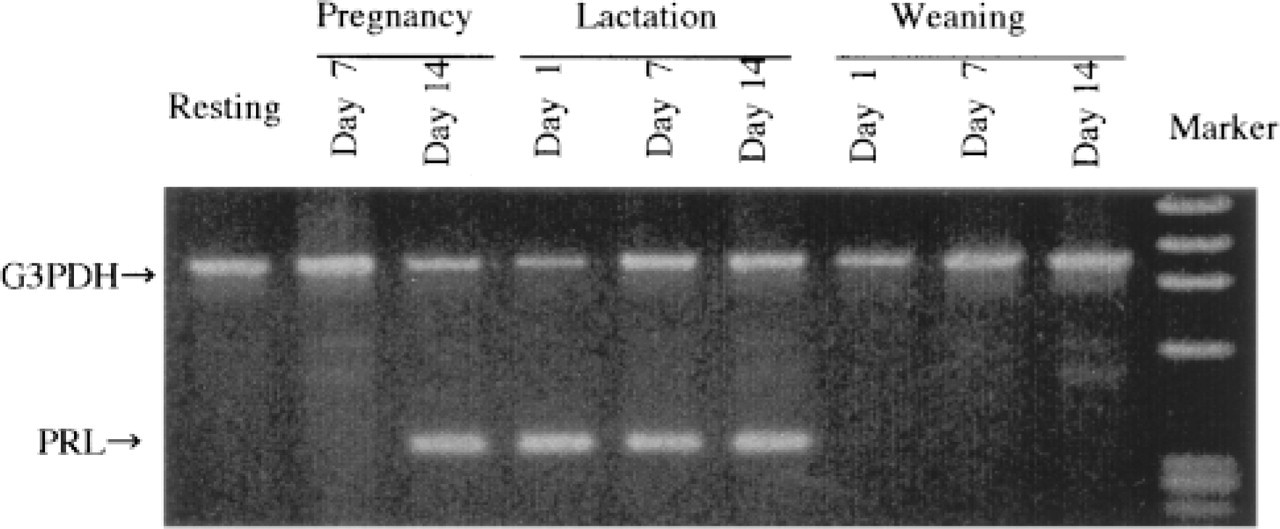
Expression of PRL mRNA of the rat mammary gland in the resting, pregnant, lactating, and weaning periods. Expression of PRL mRNA in the rat mammary gland was indicated in mid- to late pregnancy and throughout lactation.
Localization of PRL mRNA in Rat Mammary Gland
The optimal condition of in situ RT-PCR for the mammary gland was investigated by employing reliable positive signals using various protocols. In particular, to confirm the sufficiency of PCR cycles condition for detection of positive signals for PRL mRNA in the mammary gland at Day 7 of lactation by in situ RT-PCR, we changed the PCR cycles to 5, 10, and 20 cycles. At 5 cycles of PCR amplification, no positive signal was detected (Figure 4A). The signals were weakly detected but not fully exhibited at 10 cycles of PCR amplification (Figure 4B). At 20 cycles of PCR amplification, expression of PRL mRNA in the lactating mammary gland was definitely detected (Figure 4C). Subsequently, all the mammary gland sections were analyzed at under 20 cycles of PCR amplification by in situ RT-PCR.
At 20 cycles of PCR amplification, no signal was detected on the mammary gland in early pregnancy (Figure 5A) and in mid- to late weaning (Figure 5I), nor in the resting state (not shown). Positive signals were detected in mid- to late pregnancy, throughout lactation, and in the early phase of weaning. These positive signals were diffusely observed in the cytoplasm of alveolar and ductal epithelial cells (Figures 5C and 5E), whereas no signals were detected in interstitial cells. There were no local differences in expression of PRL mRNA on the mammary epithelium. In addition, in the early phase of weaning, weak focal signals were detected in some poorly vacuolated cytoplasm of regressing alveolar cells, despite the fact that no PRL mRNA was detected by solution RT-PCR (Figure 5G). The negative controls for adjacent sections without RT enzyme revealed no detectable signal (Figures 5B, 5D, 5F, 5H, and 5J) in the sense probe hybridized after PCR reaction.
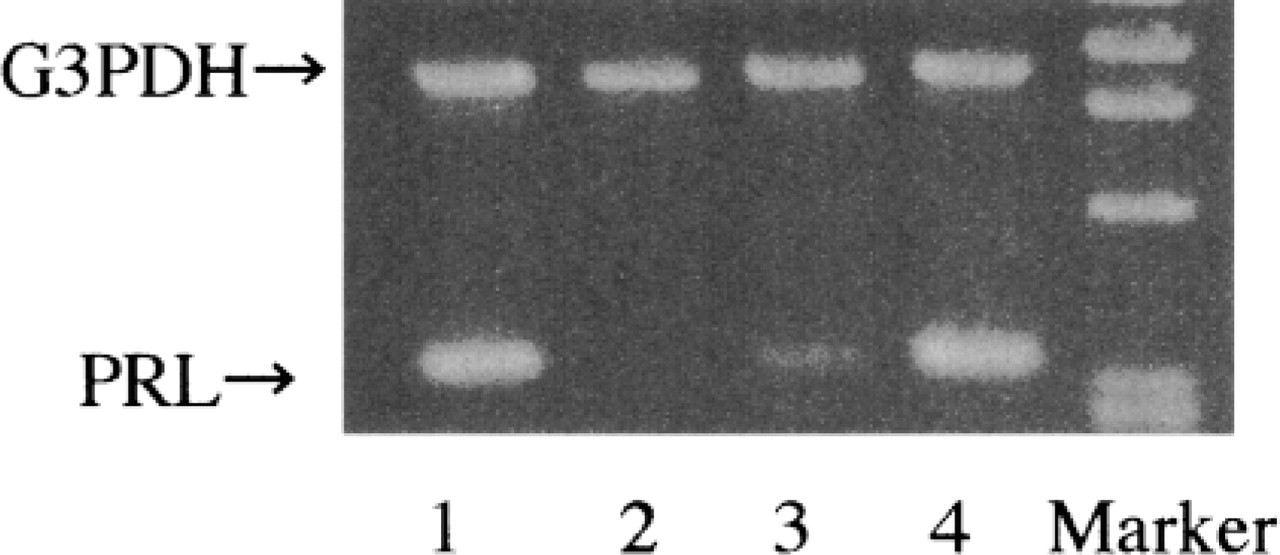
Individual cases of PRL mRNA in the rat mammary gland were observed on Day 14 of pregnancy. Lanes 1-4 show four individual rats. One case showed a lack of PRL mRNA in the rat mammary gland on Day 14 of pregnancy.
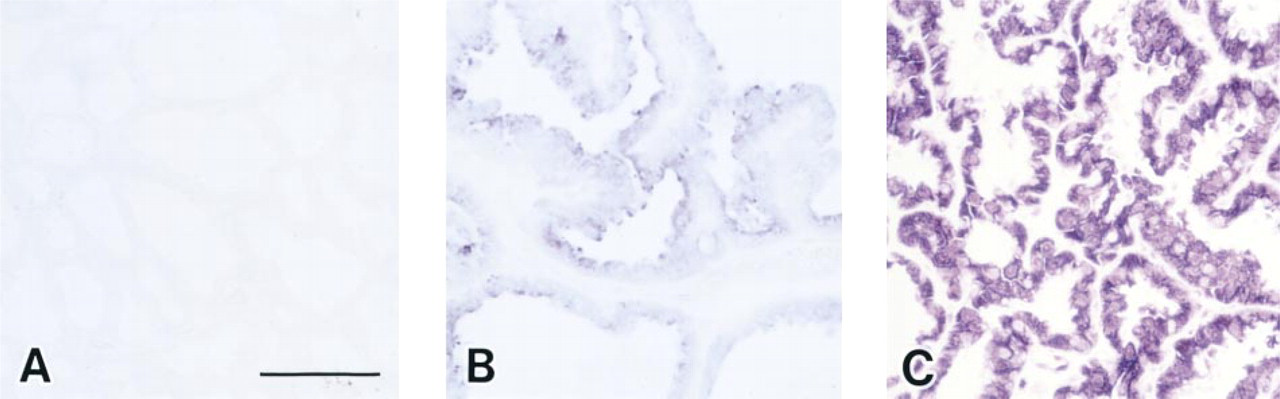
A sufficient amplification condition of PCR cycle number using PRL mRNA of mammary gland on Day 7 of lactation by in situ RT-PCR. At five-cycle amplification of PCR reaction, no signal was detected (
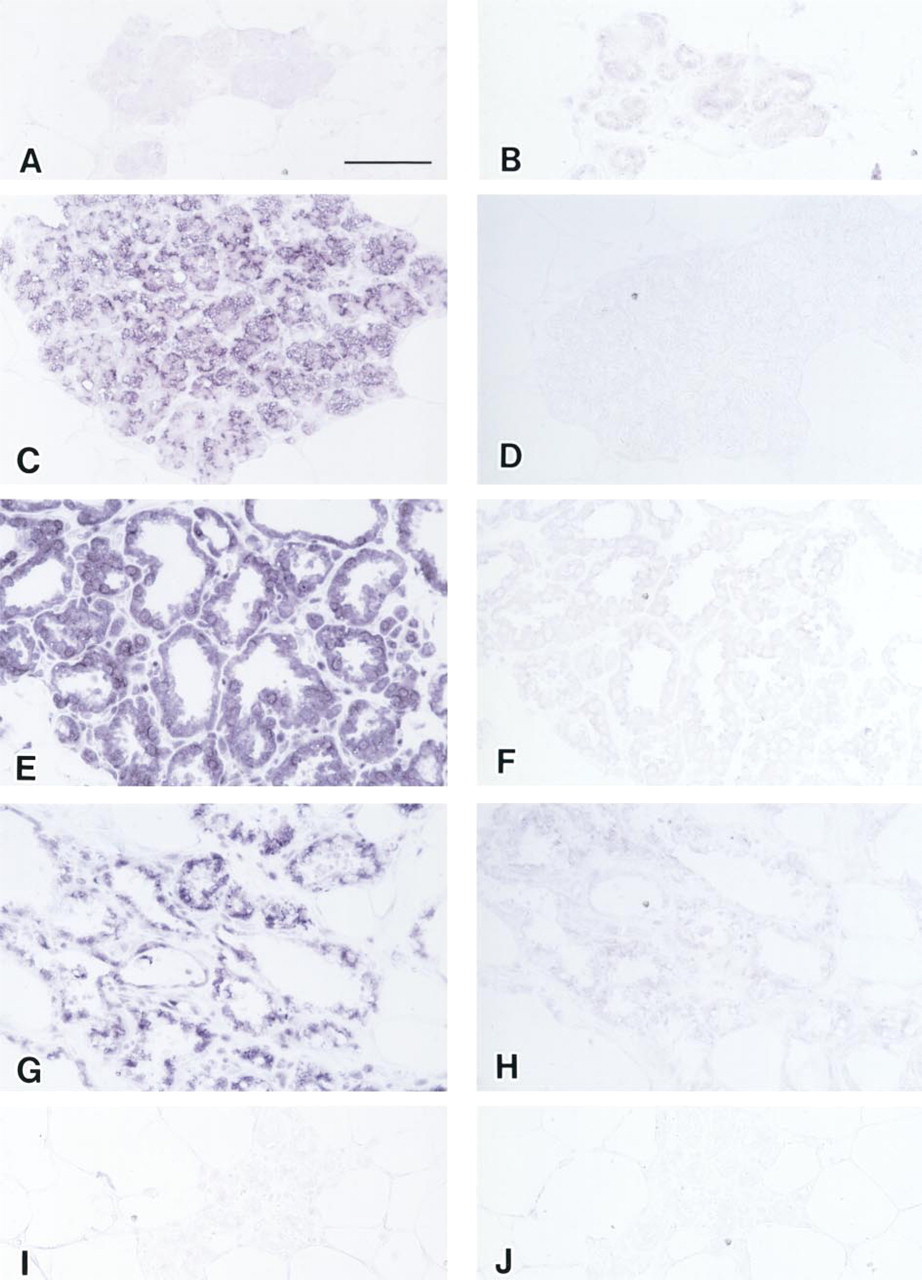
Localization of PRL mRNAs by in situ RT-PCR in the rat mammary gland during pregnancy, lactation, and weaning. No significant positive signals (purple) of PRL mRNA were detected in the early phase of pregnancy (Day 7 of pregnancy;
Discussion
In this study we confirmed that PRL mRNA in the rat mammary gland is expressed during pregnancy and the lactating period, as reported previously (Kurtz et al. 1993; Steinmetz et al. 1993; Mershon et al. 1995; Jonathan et al. 1996; Lkhider et al. 1996). Furthermore, we show that PRL mRNA expression is initiated in mid- to late pregnancy in parallel with the development of the mammary gland and that it declines in the early phase of weaning with the regression of the mammary gland. This is the first report investigating the span of PRL mRNA in the rat mammary gland in the reproductive state.
We demonstrated that PRL mRNA was detected neither in solution nor in in situ RT-PCR at the early pregnancy phase of the mammary gland under appropriate amplification conditions. On the other hand, PRL mRNA was fully exhibited on mammary epithelial cells at Day 14 of pregnancy by in situ RT-PCR.
However, in the same period, expression of PRL mRNA was evident in three of four cases, and differences varied among individuals by solution RT-PCR. This indicates that individual differences in higher expression of PRL mRNA are exhibited by mammary epithelial cells in mid- to late pregnancy. In addition, the development of the mammary gland at this time tended towards wide variations among animals. Therefore, we considered that the lack of PRL mRNA in one case by solution RT-PCR at Day 14 of pregnancy might be due to collection of whole mammary tissue in which a large proportion of fibroadipose tissue was present. It was suggested that PRL mRNA expression was initiated in mid- to late pregnancy in parallel with the development of the mammary gland.
It is well known that pituitary PRL is stimulated by neuropeptides such as TRH and VIP (Abe et al. 1985; Aizawa and Hinkle 1985; Escalada et al. 1997), estrogens (Labrie et al. 1980) and by growth factors such as EGF, IGF-1 (Murdoch et al. 1982; Lara et al. 1994) and is suppressed by a dopamine neurotransmitter (Jonathan 1985). On the other hand, extrapituitary PRL production by decidual cells of the placenta is known to be regulated by different mechanisms, stimulated by progesterone and not suppressed by dopamine (Golander et al. 1979; Huang et al. 1987; Jonathan et al. 1996; Lee and Markoff 1986). However, the mechanisms of induction and/or expression in the mammary gland of PRL have not yet been determined, especially by in vivo studies. In the mid-phase of pregnancy, the plasma PRL was at a low level but progesterone showed a temporary high level during this phase (Garland et al. 1986; Escalada et al. 1997). Plasma PRL levels increase after parturition during lactation (Garland et al. 1986; Lee et al. 1989; O'Malley 1989; Escalada et al. 1997). In addition, Mershon et al. (1995) suggested that estrogen is not required for basal PRL mRNA expression in the rat mammary gland, because PRL and PRL receptor in induced mammary tumor transcripts remained easily detectable after ovariectomy despite a marked decrease in the size of tumors. These results suggest that the initiating mechanism of PRL mRNA might be dependent not only on the plasma PRL and/or estrogen levels as distinct from pituitary PRL regulation. On the other hand, although the hormonal requirements for induction and maintenance of milk production vary, the common factor is that PRL is the hormone primarily responsible for all the major components of milk (Bole-feysot et al. 1998). The initiating mechanism of extrapituitary PRL of the rat mammary gland during the reproductive state is unclear, and further investigations, including that of milk protein genes, are necessary to resolve this issue.
In this study, PRL mRNA in mammary gland was analyzed using solution and in situ RT-PCR instead of the classical PRL binding assay, immunohistochemistry, or Northern analysis. This procedure has several advantages, in that the amplified PCR products showed a greater degree of sensitivity than by Northern analysis or in situ hybridization, allowing identification at very low concentrations of PRL mRNA in the mammary gland (Kurtz et al. 1993; Mershon et al. 1995; Jonathan et al. 1996; Lkhider et al. 1996). In this study, when PRL mRNA levels of the normal pituitary gland were compared with those of the mammary gland in mid- to late pregnancy, we demonstrated that the mRNA levels of PRL in the rat mammary gland were much lower than in the pituitary gland. In addition, PRL mRNA in the mammary gland was detected as the same 344-bp PCR product found in pituitary gland positive control (Jin et al. 1995), which was also confirmed using primers for detection of PRL mRNA in mammary gland.
Interestingly, in the early phase of weaning, PRL mRNA was not detected by solution RT-PCR, but slight signals were detected in some poorly vacuolated cytoplasm of regressing acinar cells by in situ RT-PCR. Jin et al. (1995) reported that genes exhibited more sensitivity in in situ RT-PCR than in solution RT-PCR under conditions using the same cycles for PCR amplification, and that in situ RT-PCR has the advantage of being able to identify localized mRNA. Our results also supported this report, and we confirmed low amounts of PRL mRNA in regressing mammary epithelial cells under sufficient PCR amplification. In addition, this suggests that PRL mRNA in late lactation to early weaning exhibited less variance and more resistance in the cytoplasm of mammary epithelial cells compared to the regressing histological changes in mammary gland features.
The biological role of mammary PRL mRNA expression has been understood to be an autocrine/paracrine factor (O'Malley 1989; Fields et al. 1993; Mershon et al. 1995; Jonathan et al. 1996; Escalada et al. 1997) to play a nutritional role (Kurtz et al. 1993; Jonathan et al. 1996; Lkhider et al. 1996), or to maintain immunological efficacy (Hartmann et al. 1989; Hooghe et al. 1993; Jonathan et al. 1996). The significance of the biological role of PRL mRNA during the reproductive state is not clear from our experiments. However, because our study found that PRL mRNA declined throughout weaning, we have speculated on a role for locally produced PRL that could stimulate milk production in an autocrine/paracrine fashion (Kurtz et al. 1993; Jonathan et al. 1996).
In conclusion, we have demonstrated that PRL mRNA in the rat mammary gland is initiated in mid-to late pregnancy in parallel with the development of the mammary gland, continues throughout lactation, and declines during the early phase of weaning with the regression of mammary epithelial cells.
Footnotes
Acknowledgments
We thank Drs Hisashi Iwatsuka, Kazuhiko Matsumoto, and Yoshifumi Miyakawa (Toxicology Research Laboratories, Central Pharmaceutical Research Institute, Japan Tobacco Inc.) for support and advice. We also thank Johbu Itoh, PhD (Laboratory of Structure and Function Research, Tokai University School of Medicine, Ishehara) for his skillful support in taking photomicrographs.
