Abstract
Hepatitis C virus (HCV) detection in the livers of chronically infected patients remains a debatable issue. We used immunohistochemistry, in situ hybridization (ISH) alone or after microwave heating with FITC-labeled probes, RT-PCR with unlabeled primers followed by ISH (RT-PCR-ISH), and in situ RT-PCR with FITC-labeled primers (in situ RT-PCRd) to localize the virus in 38 liver biopsy specimens from 21 chronically infected HCV patients treated with interferon-α (IFN-α). Biopsies were taken at the beginning and end of IFN-α treatment and 1 year later. Results were compared with that of HCV-PCR in serum. RT-PCR-ISH and in situ RT-PCRd showed HCV signal in all liver biopsies even in responders with seronegative HCV PCR. This signal was intranuclear, diffuse, or peripheral, in hepatocytes, bile ductule cells, and lymphocytes. Cytoplasmic signals were occasionally observed. Whereas the percentage of labeled hepatocytes remained constant, the number of labeled lymphoid follicles decreased after INF-α therapy. Immunohistochemistry resulted in the same pattern of positivity but it was weaker and inconstant. This study indicates the persistency of HCV latency in IFN-α responders 1 year after IFN-α treatment cessation, a finding that certainly deserves confirmation.
Keywords
H
The aims of the present study were to localize HCV by in situ molecular biology techniques (ISH, in situ RT-PCRd, and RT-PCR-ISH) in the livers of patients with chronic hepatitis C, to compare results obtained by these techniques with IHC detection of the viral protein (the superoxide dismutase-HCV recombinant polypeptide C100-3), and to compare these data to those of RT-PCR detection of HCV in serum before and after IFN-α therapy.
Materials and Methods
Patient Selection
Twenty-one white HCV-confirmed seropositive patients (11 women and 10 men) were selected for this study, as all had sera collected at the time of each liver biopsy. The median age was 33 years (range 21–60 years). Patients received the same therapeutic treatment (INF-α) for 6 or 12 months. The criterion for INF-α treatment was a persistently elevated serum alanine transaminase (ALT) for at least 6 months. None of the patients was seropositive for HIV or HBV. None had received previous treatment. For each patient, ALT level determination was performed at the same time as biopsy.
Tissue Sample Processing
Liver biopsies were attempted at three time points: just before INF-α treatment (B1) (
Tissue specimens were fixed in 10% formalin and then paraffin-embedded. Liver samples from alcoholic and hepatitis B patients seronegative for HCV antibodies and for HCV RNA, as well as heterologous tissue samples (e.g., salivary glands and gastric tissue) from these same patients, were used as negative controls. Intrahepatic HCV detection was compared with HCV detection in each corresponding serum sample.
Serum Evaluation of HCV
Two methods were used on frozen serum specimens. (a) HCV RNA quantification by the bDNA method, only positive for at least 0.35 × 106 viral copies/ml, was performed with the quantitative bDNA signal amplification assay (Quantiplex TM HCV RNA 1.0; Chiron Diagnostics, Emeryville, CA) at each time point. All samples were run in duplicate. (b) If the bDNA quantification remained negative, the Nested RT-PCR was tested (Amplicor TM HCV; Roche, Paris, France).
In situ hybridization was performed in a moist chamber for 18 hr at 37C with all the FITC-labeled probes. About 20 ng of probe was applied per section. To detect the hybridization complex, slides were incubated first with mouse anti-FITC antiserum, second with rabbit anti-mouse immunoglobulins, and then with a mouse anti-rabbit monoclonal antibody linked to APAAP complex (Dakopatts, Glostrup, Denmark). Alkaline phosphatase activity was revealed using NBT-BCIP chromogenic substrate. In some cases, citrate antigen retrieval by microwave heating before ISH was performed.
As controls for in situ PCR, liver tissues removed from HCV-positive patients who had a high expression of viral nucleic acids served as initial positive controls. Negative controls included (a) omission of the reverse transcription step and/or DNase pretreatment, (b) incubation of slides in an RNase solution immediately before reverse transcription, (c) for the amplification, omission of the DNA polymerase or primers and incubation of the liver tissue with DNase before PCR amplification and ISH, and (d) the use on HCV infected liver of heterologous human papillomavirus-related primers for amplification followed by ISH with HCV probes or of heterologous HBV probe after amplification with HCV sets of primers. HCV primers and probes were also applied to liver and heterologous tissues (gastric mucosa, salivary glands) from HCV-negative patients and to rat liver. Positive controls of PCR amplification included performance of RT-PCR-ISH with β-actin primers and an FITC-labeled actin probe. The presence of HCV was also verified by a standard RT-PCR assay performed on homogenates obtained from frozen liver tissues of several patients.
Immunohistochemistry
Two mouse monoclonal antibodies (MAbs) corresponding to clone Tordji 32 and to a mixture of clones Tordji 22 and 32 were used (Clonatec; Paris, France). They recognize the superoxide dismutase-HCV recombinant polypeptide (Ortho Diagnostic C100-3 Elisa kit). Sections were treated with primary MAb at 1:50 dilution for 1 hr at RT, then with secondary biotinylated horse anti-mouse IgG (Vector Labs, Burlingame, CA) and with the avidin-biotin-peroxidase complex (Vectastain ABC kit; Vector Labs). Specificity of immunore-action was tested by omitting primary antibodies. Citrate antigen retrieval by microwave heating before IHC was performed in all cases.
Quantification of In Situ RT-PCR Results and Statistical Analysis
For each biopsy, counting of HCV labeled vs unlabeled hepatocytes was done in 10 microscopic fields at X 400 magnification. Counts were expressed as percentage of labeled hepatocytes. The surface of the specimens was roughly estimated by measuring width and length. The number of lymphoid follicles in each biopsy was counted and was expressed per 10 mm2. Counts were made by two independent observers. All values were expressed as mean ± 1 SEM. Differences in results between responders vs non-responders and different times (B1, B2, B3) were statistically analyzed by the nonparametric Mann-Whitney
Results
Histology
Histological features proved the chronic hepatitis with portal nodular lymphoid infiltration and patchy necrosis (Figure 1). Patients were classified as responders (
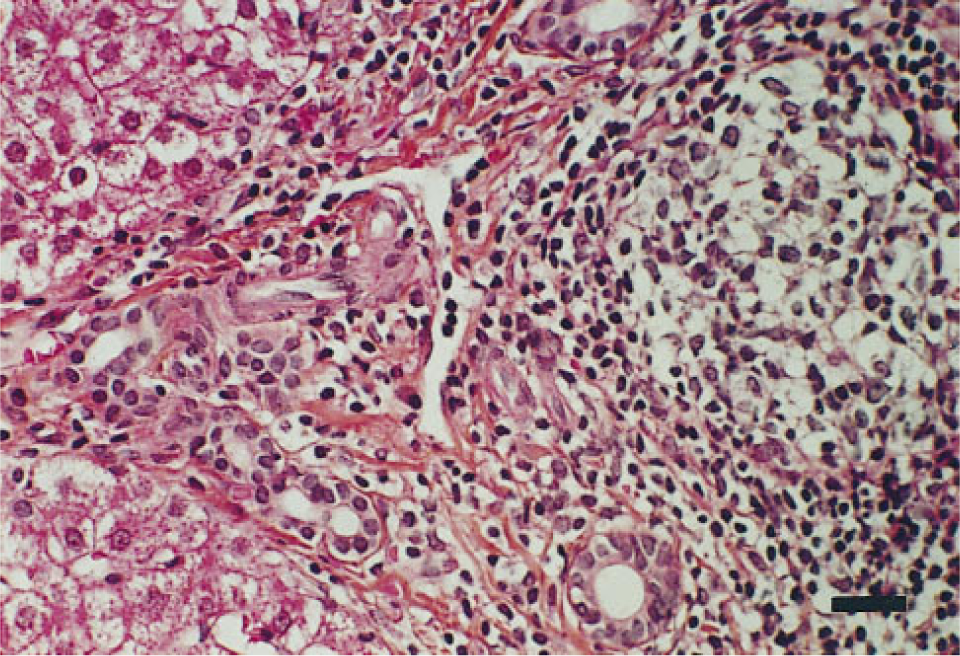
Histological alterations in liver biopsy specimens of HCV chronic hepatitis at B1 time. The portal area contains a nodular lymphocytic infiltrate with patchy necrosis (Mayer's hemalum-eosin-saffron). Bar = 30 μm.
Serum RT-PCR for HCV
RT-PCR results were clear-cut (Figure 3). At B1, all patients were seropositive. At B2, no responders had measurable HCV copies per ml by bDNA assay, but one patient was seropositive by nested RT-PCR. In the non-responders, one was transiently seronegative. At B3, all responders were seronegative and all non-responders were seropositive with relative increased copy numbers compared to B2.
In Situ HCV Detection
Evolution of HCV in situ RT-PCR signal in hepatocytes and lymphoid follicles in responders and non-responders in the course of IFN-α treatment a
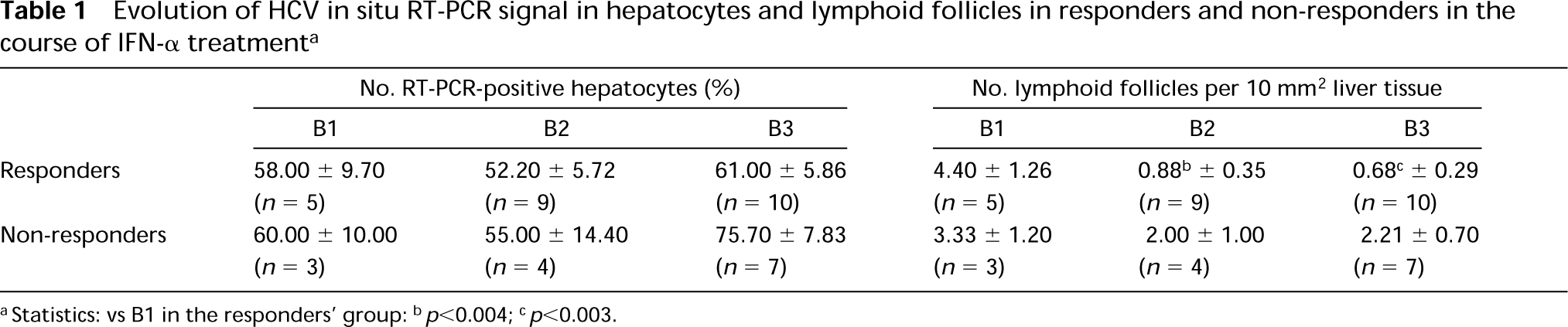

aStatistics: vs B1 in the responders' group
bp<0.004
cp<0.003.
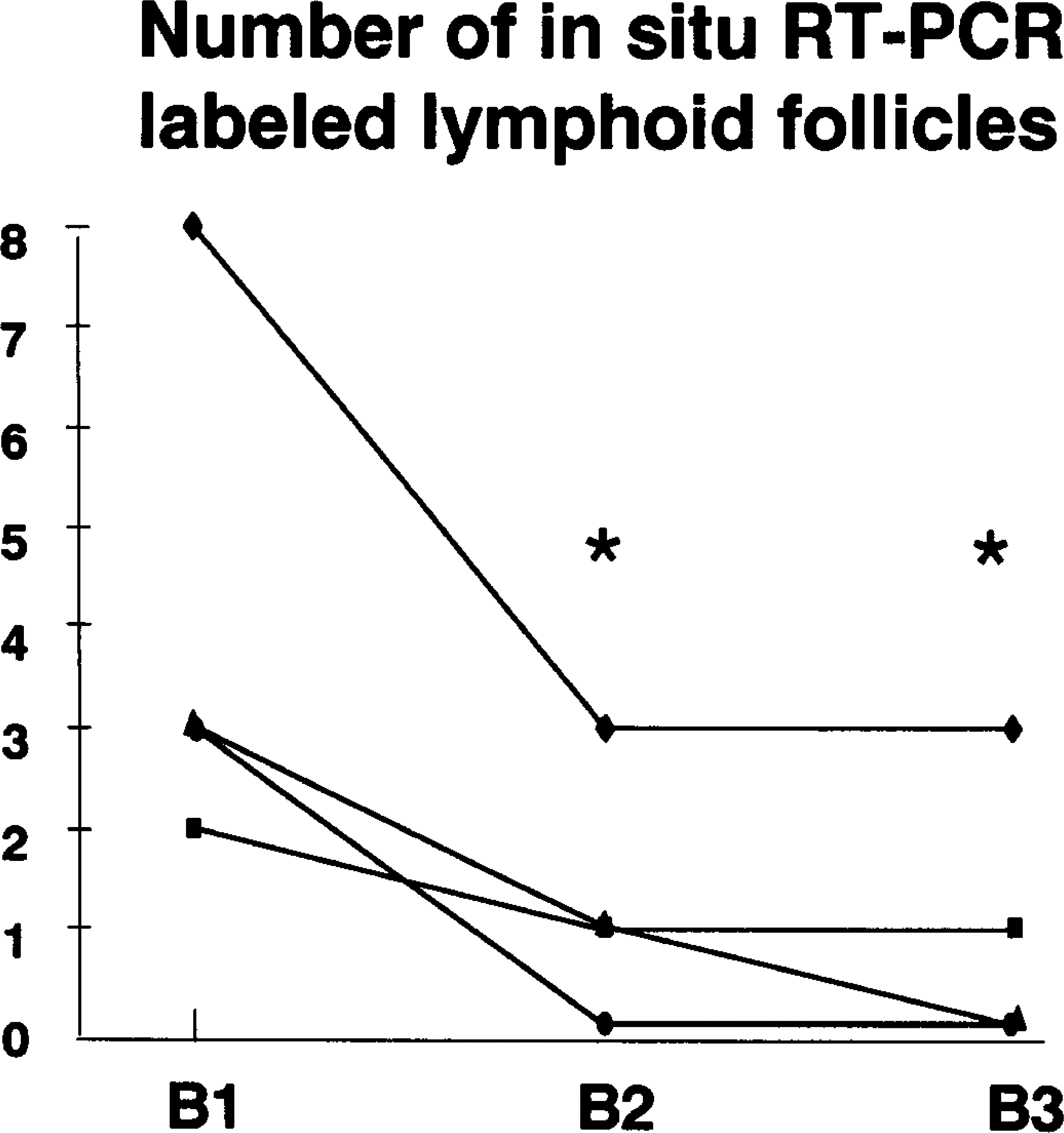
Numbers of lymphoid follicles in liver biopsies at B1 (before IFN-α treatment), B2 (at treatment cessation), and B3 (1 year later) in the same four patients responding to IFN-α therapy. ∗
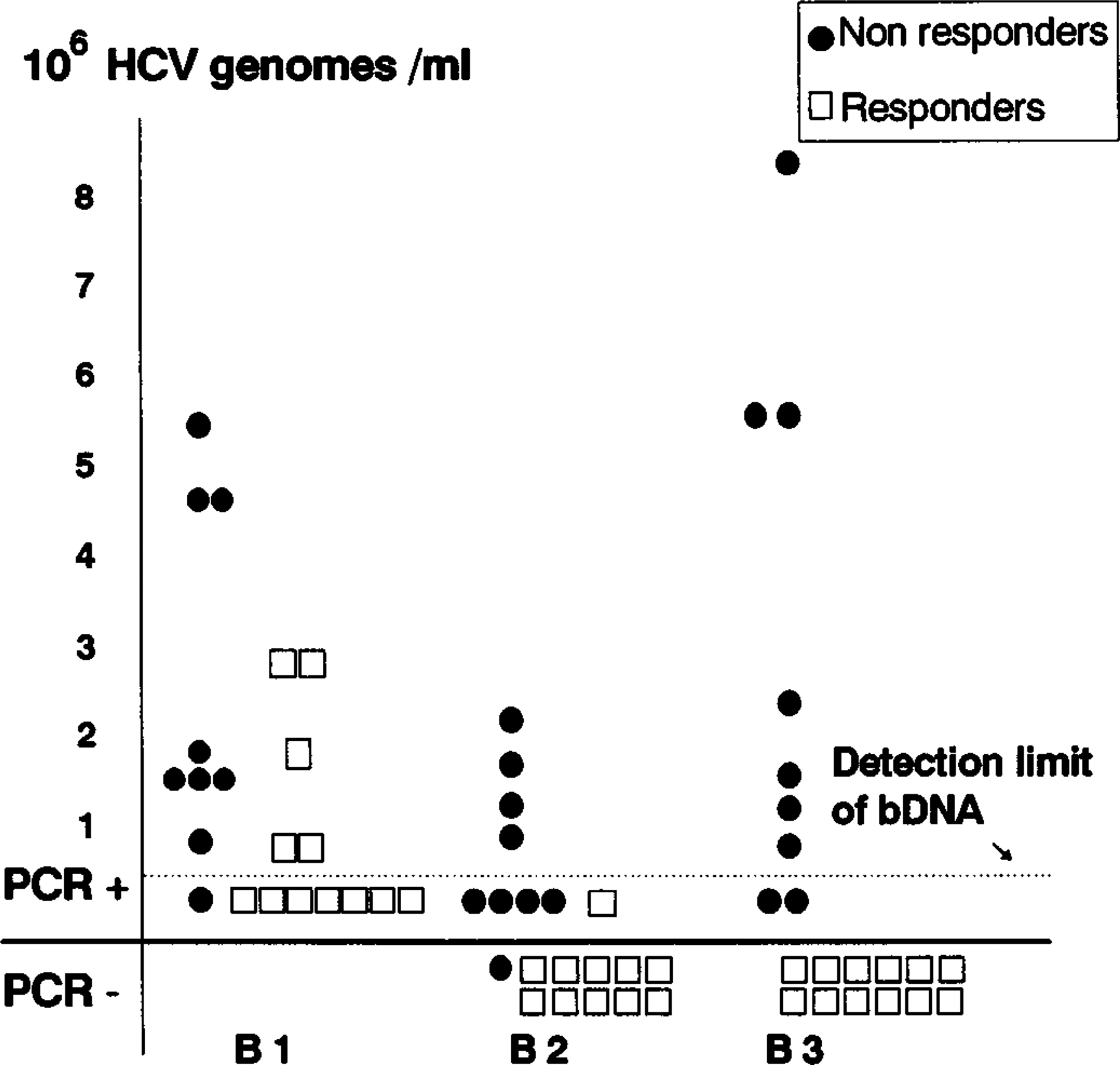
Serum HCV RNA levels in 21 patients according to response to interferon therapy. The bDNA quantification was possible for at least 0.35 × 106 viral copies/ml. Below this value, nested RT-PCR was performed, indicating only the presence or absence of viral RNA in serum.
Quantification of RT-PCR Signal
On the 38 hepatic biopsies studied, there was diffuse panlobular distribution of positive hepatocytes with variable staining intensity. Importantly, responders continued to have HCV nuclear signal at B2 and B3 times (Figure 5). The mean percentage of labeled hepatocytes in those patients remained constant (50–60%) and did not differ from non-responder values (Table 1). This was also checked in the four responders who underwent B1, B2, and B3 biopsies. Nevertheless, in all responders, not only did the number of lymphoid follicles decrease (Table 1) but the nuclear intensity labeling and the number of labeled nuclei in these lymphoid follicles were diminished, although no quantification was performed for this last variable. HCV RT-PCR assays were also performed on RNA extracted from frozen liver biopsies of four patients at B3. All these patients, including one responder to IFN-α therapy, appeared positive for HCV.
Immunohistochemical Results
The results were similar regardless of the antibody used. Treatment of slides by microwave heating did not improve signal specificity. The localization of signal, as with in situ amplification RT-PCR, was intranuclear in hepatocytes, bile duct cells, and portal lymphocytes (Figures 6A-6C) and occasionally cytoplasmic (Figure 6C). However, the immunoreactivity in hepatocyte nuclei was sometimes very weak (Figure 6B), and the number of labeled hepatocytes was always less than that seen with in situ RT-PCR. In addition, IHC was relatively insensitive because half of the samples from HCV livers at B1 were negative (Figure 5), despite the presence of HCV signal by in situ RT-PCR. It is noteworthy that IHC positivity also persisted at B2 and B3 in most responders. No signal was observed in heterologous tissue or in liver tissue from HCV-negative patients used as controls.
Discussion
Our results are in agreement with previous findings describing viral localization at the perinuclear membrane (Lidonnici et al. 1995) or within the nuclei of hepatocytes (Haruna et al. 1993; Nouri-Aria et al. 1993). We noted with RT-PCR-ISH, but rarely with in situ RT-PCRd, as already observed by Nouri-Aria et al. (1993) and occasionally by Nuovo (1994), a diffuse and inconstant staining within the cytoplasm, always associated with the nuclear signal. However, in our opinion, the specificity of the cytoplasmic signal is rather doubtful and could be due to some background. Nevertheless, it was not possible to totally eliminate it in spite of many high-stringency washings. Some authors (Komminoth et al. 1994) stated that the in situ RT-PCRd is not reliable and the nuclear signal not specific because in their control experiments, omitting primers yielded positive nuclear signals similar to those they observed with primers. In our experiment, omitting primers indeed yielded negative results and all controls confirmed the specificity of the signal reaction. It must be pointed out that Nuovo (1994) obtained the same perinuclear signal with a direct incorporation of digoxigenin-labeled nucleotides during the in situ amplification and with primers other than ours.
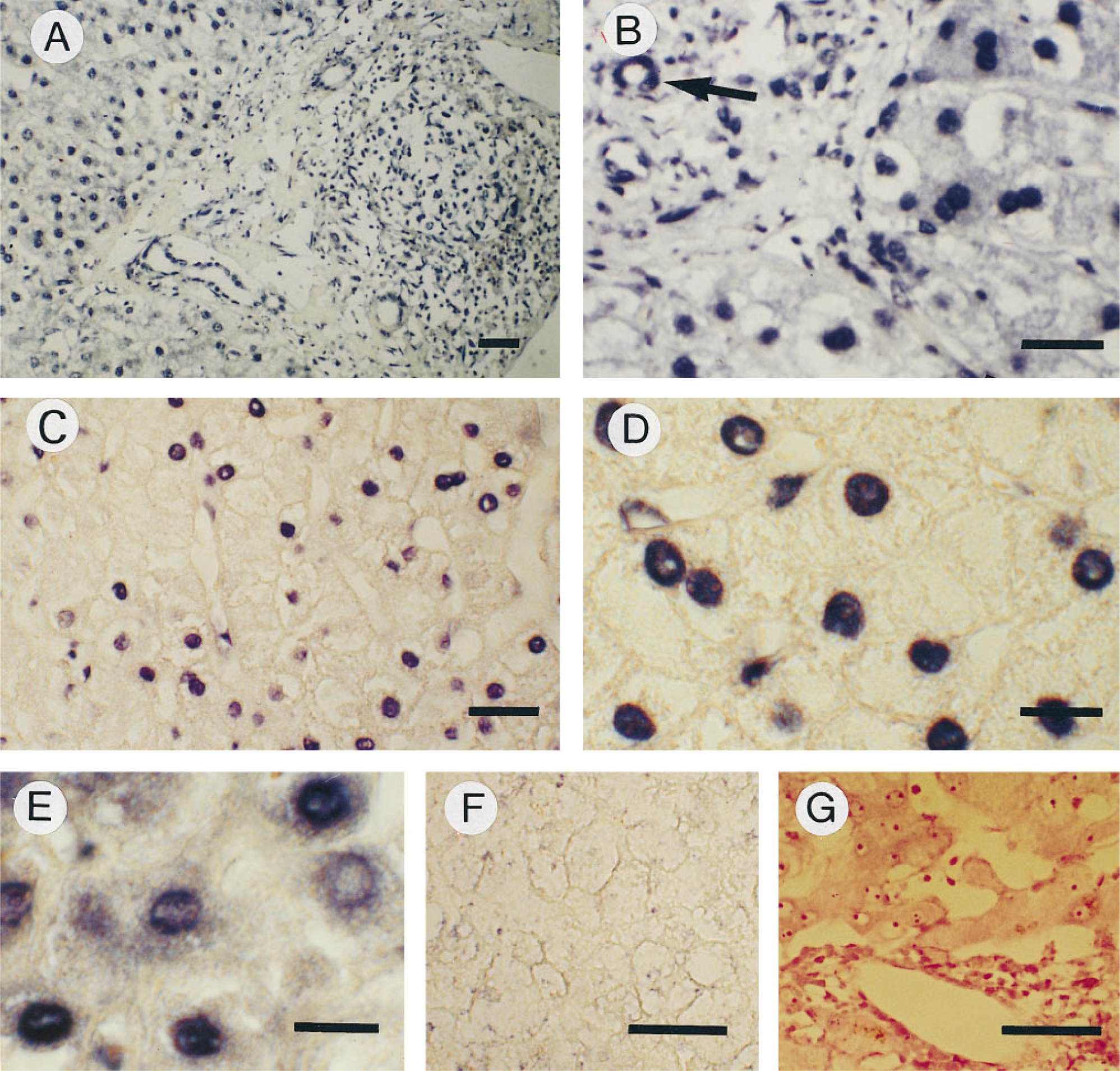
Liver biopsy of a non-responder patient at the end of IFN-α treatment (B2) after RT-PCR-ISH with FITC-labeled probe, at low magnification. In this strongly positive area, intranuclear signal is seen in many hepatocytes, lymphocytes, and bile duct cells (no counter-staining). Bar = 30 μm. (
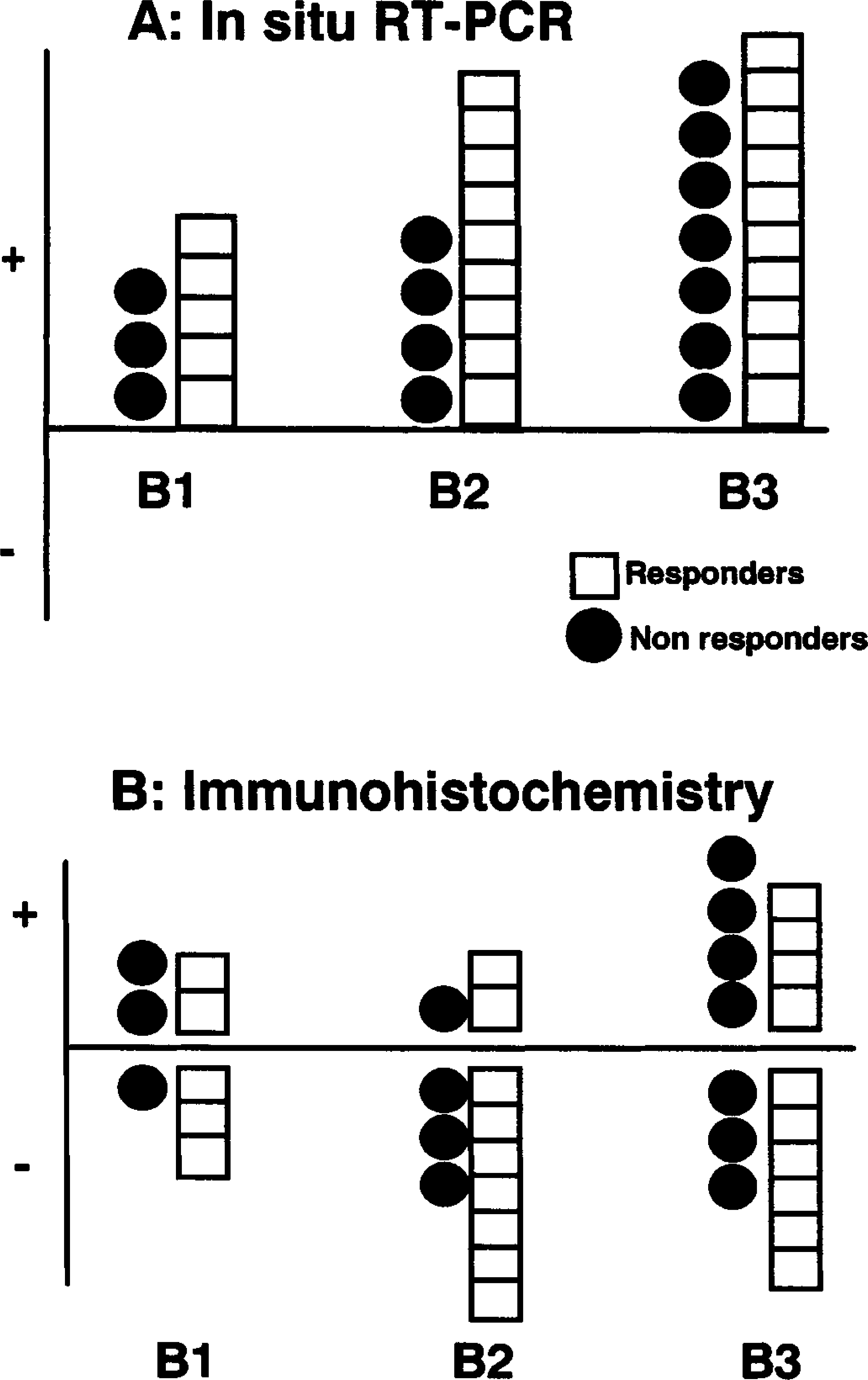
Diagram showing the number of positive patients for HCV detection by in situ RT-PCR (
In the literature, HCV was found mainly in hepatocytes (Negro et al. 1992; Haruna et al. 1993; Nouri-Aria et al. 1993; Lau et al. 1994; Gastaldi et al. 1995) but also in Kupffer cells (Nuovo 1994), mononuclear cells (Nouri-Aria et al. 1993; Lau et al. 1994), sinusoidal cells (Nouri-Aria et al. 1993) and bile duct cells (Nouri-Aria et al. 1993), although rarely (Nuovo 1994). Our own results confirm most of these localizations. Unlike Park et al. (1996), who obtained inconstant results by performing RT-PCR assay after HCV RNA extraction from formol-fixed, paraffin-embedded liver biopsies, we had no problem of sensitivity with the in situ RT-PCR directly realized in such liver tissues. These authors postulated that formalin-induced crosslinks may inhibit extension by reverse transcriptase. We also suggest that RNA extraction after dewaxing at high temperature could induce a large loss of HCV RNA copies, leading to false-negative results.
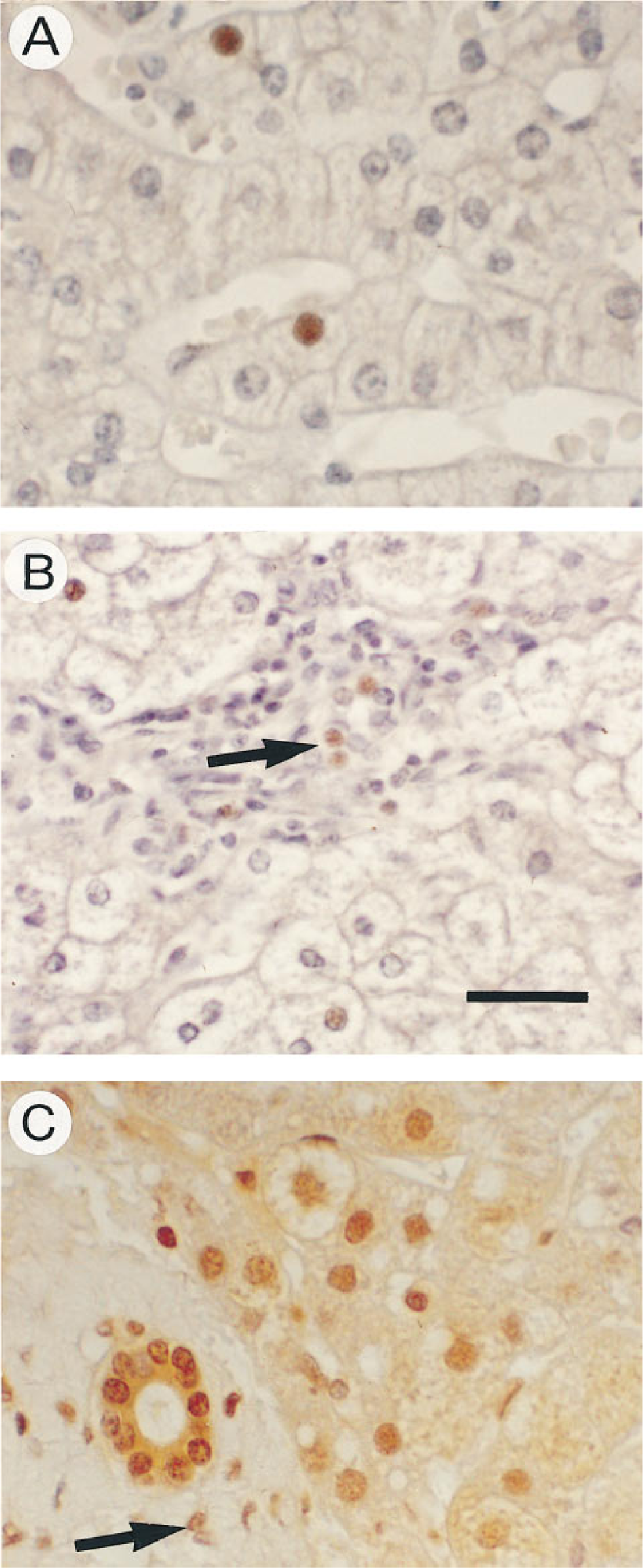
IHC with monoclonal antibodies against the superoxide dismutase-HCV recombinant polypeptide (C100-3 protein) clones Tordji 22 and 32 (counterstaining with Mayer's hemalum). (
As already reported by Chamlian et al. (1993), the first results obtained by IHC with antisera raised against either HCV structural and nonstructural proteins, core antigen, E1, E2, envelope or NS2, NS3, NS4 and NS5 antigens showed a cytoplasmic signal (Hiramatsu et al. 1992; Tsutsumi et al. 1994; Uchida et al. 1994; Nouri-Aria et al. 1995; Barba et al. 1997). When using the two MAb clones Tordji 22 and 32, which are currently the HCV antisera commercially available (Clonatec), Chamlian et al. (1993) obtained exclusively a nuclear signal in hepatocytes. We confirmed this signal but also noted some diffuse cytoplasmic staining in liver samples displaying higher numbers of immunoreactive hepatocytes. All control specimens were negative. However, IHC was less reliable than our in situ RT-PCR techniques.
In this work, HCV RNA was studied for the first time by in situ RT-PCR in infected patients' livers before and after IFN-α therapy. If our findings reflect the true HCV localization, they indicate the persistence of HCV latency within responder livers, even 1 year after IFN-α therapy cessation. Although the number of in situ RT-PCR-positive hepatocytes stayed quite constant in responders (about 50–60%), the number of lymphoid follicles decreased, as well as the HCV nuclear positivity in the lymphocytes. The two latter facts suggest that, as in other forms of chronic hepatitis, the lymphoid population in the liver is one of the factors that mediates disease activity (Navas et al. 1994) or relapse of the disease (Moldvay et al. 1994). Several hypotheses must be raised to explain the discrepancy between serum RT-PCR and in situ RT-PCR results: (a) a lack of sensitivity of serum RT-PCR (level detection 102 HCV genomes/ml) while in situ RT-PCR could evidence a very low viral copy number per cell, such as one or two HPV copies in Siha cells (Nuovo 1994), or (b) a level of viral replication in liver too low to be detected in serum. This viral form, probably in a latent state, may explain the relatively high chronic hepatitis C relapse. Some authors (Saleh et al. 1994) report negative RT-PCR assay in frozen liver tissues from responders to IFN-α therapy. We believe that the conditions of the cryoprotection, the very low level of viral copy numbers, and the presence in tissue of enzymes inhibiting the reaction may lead to false-negative results. In our experiment, RT-PCR assay performed on quickly extracted HCV RNA after liver biopsy was positive in one responder at B3. Certainly, our findings need to be confirmed by further investigations. In addition, we cannot rule out the possibility that the viral persistence is transient and that the virus does disappear in responder livers after a longer time following the IFN-α therapy cessation.
Footnotes
Acknowledgements
Supported by the ARC, grant number 2003. We thank Pr C. Degott and Pr P. Marcellin for having given us the opportunity to study HCV-infected liver biopsies and M. Martinot-Peignoux for performing HCV RNA RT-PCR in serum.
