Abstract
Background:
Although faecal DNA testing of Fusobacterium nucleatum (Fn) is expected to be useful for colorectal neoplasia detection, there is no standardized quantification method of Fn. We performed this study to establish a possible standardized method.
Methods:
In this study, 322 participants including 71 subjects without colorectal neoplasia (control group), 31 patients with non-advanced colorectal adenoma, 93 patients with advanced colorectal adenoma, and 127 patients with colorectal cancer were enrolled. Faecal Fn were quantified by droplet digital PCR (ddPCR) using two PCR primer-probe sets reported previously that are tentatively named Fn1 and Fn2. Fn1 has been used in ddPCR by us and Fn2 has been widely used in quantitative real-time PCR.
Results:
The Fn copy number using Fn1 was five times higher than that using Fn2, with a linear relationship shown between them. Receiver operating characteristic curve analysis showed the area under the curve (AUC) to be almost the same between Fn1 and Fn2 in discriminating between the control group and the colorectal cancer group (AUC = 0.81 and 0.81, respectively), and between the control/non-advanced colorectal adenoma group and the advanced colorectal adenoma/colorectal cancer group (AUC = 0.74 and 0.74, respectively).
Conclusions:
As the diagnostic performance was quite similar between Fn1 and Fn2, ddPCR-based Fn testing using Fn1 and Fn2 could be a possible standardized method for a colorectal neoplasia screening test, considering that Fn levels quantified by Fn1 are about five times higher than those by Fn2.
Introduction
As an overabundance of faecal Fusobacterium nucleatum (Fn) in patients with colorectal neoplasia has been reported by many research groups,1–4 faecal DNA testing of Fn is expected to be a screening test for colorectal neoplasia. Quantitative real-time polymerase chain reaction (qRT-PCR) testing is widely used to evaluate relative copy numbers of Fn. However, the abundance of Fn indicated by the qRT-PCR is influenced by the internal control levels including Bacteroides and Eubacteria because the Fn levels are adjusted by these internal control levels. 1 As the relative quantification makes it difficult to compare the diagnostic accuracy of faecal Fn in colorectal neoplasia between different research groups, establishment of a standardized method, other than relative quantity, is necessary. To solve this issue, droplet digital PCR (ddPCR) may be useful because ddPCR is 1000 times more sensitive than qRT-PCR 2 and enables quantification of the absolute copy number of target genes without the need for calibration samples, and the data can be directly compared between different research groups. Thus, we developed a ddPCR-based test of faecal Fn and reported its usefulness for colorectal neoplasia screening. 2 In the present study, using two PCR primer-probe sets of Fn reported previously, one a set used by us in the above-mentioned ddPCR 2 and the other a set widely used in qRT-PCR,1,5–7 we quantified faecal Fn by ddPCR and compared the diagnostic performance of the two PCR primer-probe sets in colorectal neoplasia.
Materials and methods
Materials
This study was prospectively conducted in Yamaguchi University Hospital, Ajisu Kyoritsu Hospital, St. Hill Hospital, and IMSUT Hospital. The institutional review board of each hospital approved the study protocol (protocol number: H29-228). Our protocol was designed according to the Declaration of Helsinki and ethical guidelines. We included consecutive participants who provided written informed consent for this study from October 2007 to December 2019. Participants with a history of inflammatory bowel disease, colorectal cancer, or colorectal surgery were excluded. After a faecal specimen was collected, all participants underwent colonoscopy. Although the location and size of all lesions were reported by colonoscopists, we used only the most advanced colorectal epithelial lesion and its location to categorize the participants for the analysis. The proximal colon includes the cecum, ascending colon, hepatic flexure, and transverse colon. The distal colon includes all other segments. In this study, 322 participants including 71 subjects without colorectal neoplasia (control group), 31 patients with non-advanced colorectal adenoma (NAA group), 93 patients with advanced colorectal adenoma (AA group), and 127 patients with colorectal cancer (CRC group) were enrolled. The participants in the previous report 2 were included in the present study. From the 322 participants, we selected 49 participants from the Control/NAA groups and 49 participants from the AA/CRC groups for matching by age and sex. Advanced adenoma was defined as adenomas of 1 cm or greater in size, or with villous components (tubulovillous or villous), or with high-grade or severe dysplasia. Staging was classified according to the Union for International Cancer Control. 8
Collected faeces were stored at −20°C until DNA extraction, and the QIAamp DNA Stool Mini Kit (QIAGEN, Tokyo, Japan) was used to extract DNA from about 200 mg of each faecal sample. Control DNA of Fn was extracted from a reference strain of Fusobacterium nucleatum ATCC 25586 (American Type Culture Collection, Manassas, VA, USA) using the QIAamp DNA Stool Mini Kit (QIAGEN). Quantification of double-stranded DNA was performed using a Qubit 2 fluorometer (Thermo Scientific, Tokyo, Japan).
Droplet digital PCR
The copy number of Fn was measured by ddPCR as described previously.2,9 The PCR reaction mixture consisted of 10 ng of DNA, 10 μL of 2×ddPCR Master Mix for Probes (Bio-Rad, Hercules, CA, USA), 0.25 μM of each primer, and 0.125 μM of probe in a total volume of 22 μL. An automated droplet generator (Bio-Rad) was used for droplet generation. We used two Fn primer-probe sets reported previously: one is a primer-probe set we used in previous ddPCR studies,2,9 and the other is a primer-probe set widely used by other investigators in qRT-PCR.1,5–7 We named the primer-probe sets Fn1 and Fn2, respectively. The sequences of the Fn1 primer-probe set are as follows: forward primer, 5′-AAGCGCGTCTAGGTGGTTATGT-3′; reverse primer, 5′-TGTAGTTCCGCTTACCTCTCCAG-3′, and probe, 5′-FAM-CAACGCAATACAGAGTTGAGCCCTGCATT-TAMRA-3′.2,9,10 The sequences of the Fn2 primer-probe set are as follows: forward primer, 5′-CAACCATTACTTTAACTCTACCATGTTCA-3′; reverse primer: 5′-GTTGACTTTACAGAAGGAGATTATGTAAAAATC-3′, and probe, 5′-FAM-TCAGCAACTTGTCCTTCTTGATCTTTAAATGAACC-TAMRA-3′. 1 Fn1 amplifies five fragments of the 16S rRNA gene of Fn, and Fn2 can amplify a single fragment of the nusG gene of Fn (Supplemental Table 1). Cycling conditions included preheating at 95°C for 10 min, followed by 40 cycles of denaturation at 94°C for 30 s and annealing at 60°C for 60 s, and final heating at 98°C for 10 min. The fluorescence amplitude data were acquired by a QX200 droplet reader (Bio-Rad) and QuantaSoft software (Bio-Rad).
Statistical analysis
Data analyses were performed using GraphPad Prism Ver. 9 (GraphPad Software, La Jolla, CA, USA), BellCurve for Excel (Social Survey Research Information, Tokyo, Japan), and EZR (Saitama Medical Centre, Jichi Medical University, Saitama, Japan). 11 Dunn’s test was used for comparison between multiple groups. A linear regression analysis though the origin was performed to compare the amount of the reference strain of Fn ATCC 25586 with Fn copy numbers quantified by Fn1 and Fn2. Deming regression analysis was performed to compare Fn copy numbers by Fn1 with those by Fn2. Receiver operating characteristic (ROC) curve analysis was used to quantify how accurately the Fn test by Fn1 and Fn2 discriminates two groups. A P value < 0.05 was considered to be statistically significant.
Results
Copy numbers of reference strain of Fn
Copy numbers of the reference strain of Fn ATCC 25586 quantified using Fn1 (Figure 1(a)) and Fn2 (Figure 1(b)) are shown. The copy numbers using Fn1 were about five times higher than those using Fn2 (Figure 1(c)). Among the faecal samples, the copy numbers of Fn quantified using Fn1 were also 5 times higher than those using Fn2, with a positive correlation shown between them (r
2
= 0.97) (Figure 1(d)). Copy numbers of Fn. The ddPCR amplitude plots with the horizontal threshold lines of the reference strain of Fn ATCC 25586 using Fn1 (a) and Fn2 (b), and linear regression analysis of Fn1 and Fn2 (c) are shown. The correlation of faecal Fn copies between Fn1 and Fn2 using Deming regression analysis are shown (d). Fn, Fusobacterium nucleatum.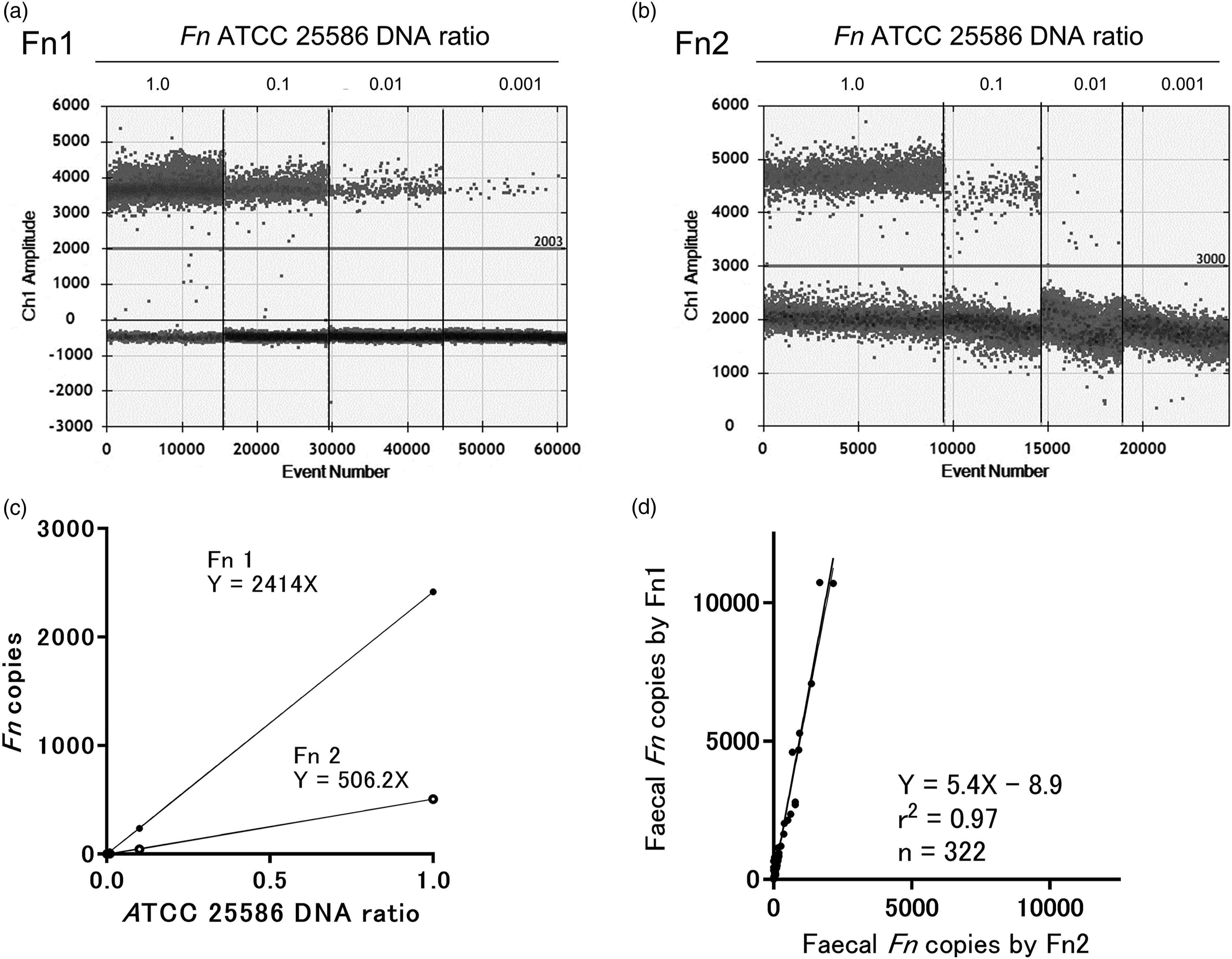
Faecal Fn copy numbers
Clinicopathologic characteristics of the subjects.

*pStage: I: 37, II: 47, III: 39, IV: 4.
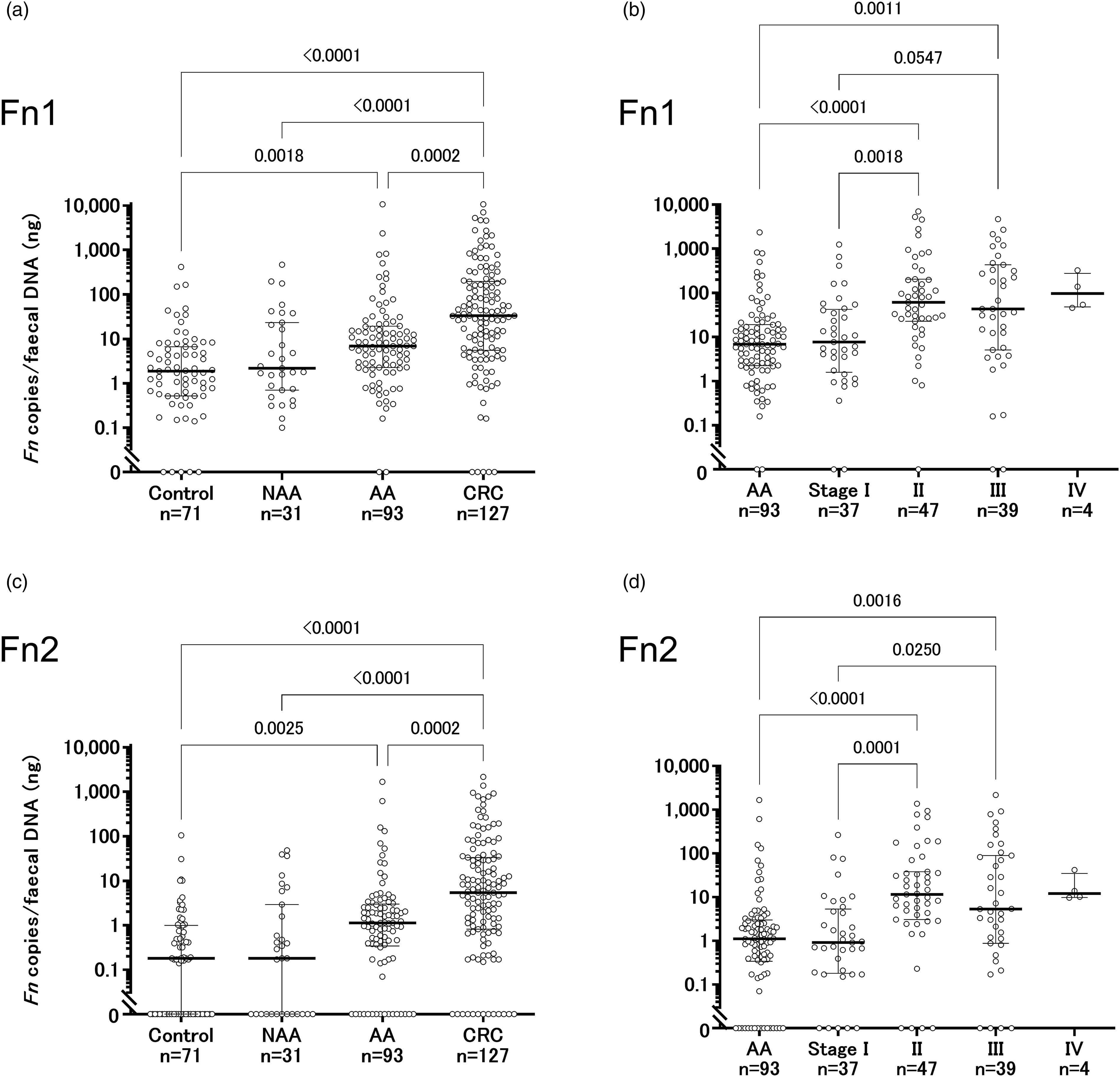
Distribution of faecal Fn. Faecal Fn copies per template faecal DNA (ng) using Fn1 (a, b) and Fn2 (c, d) are shown. Numbers indicate P values using Dunn’s test. AA, advanced colorectal adenoma. CRC, colorectal cancer. Fn, Fusobacterium nucleatum. NAA, non-advanced colorectal adenoma.
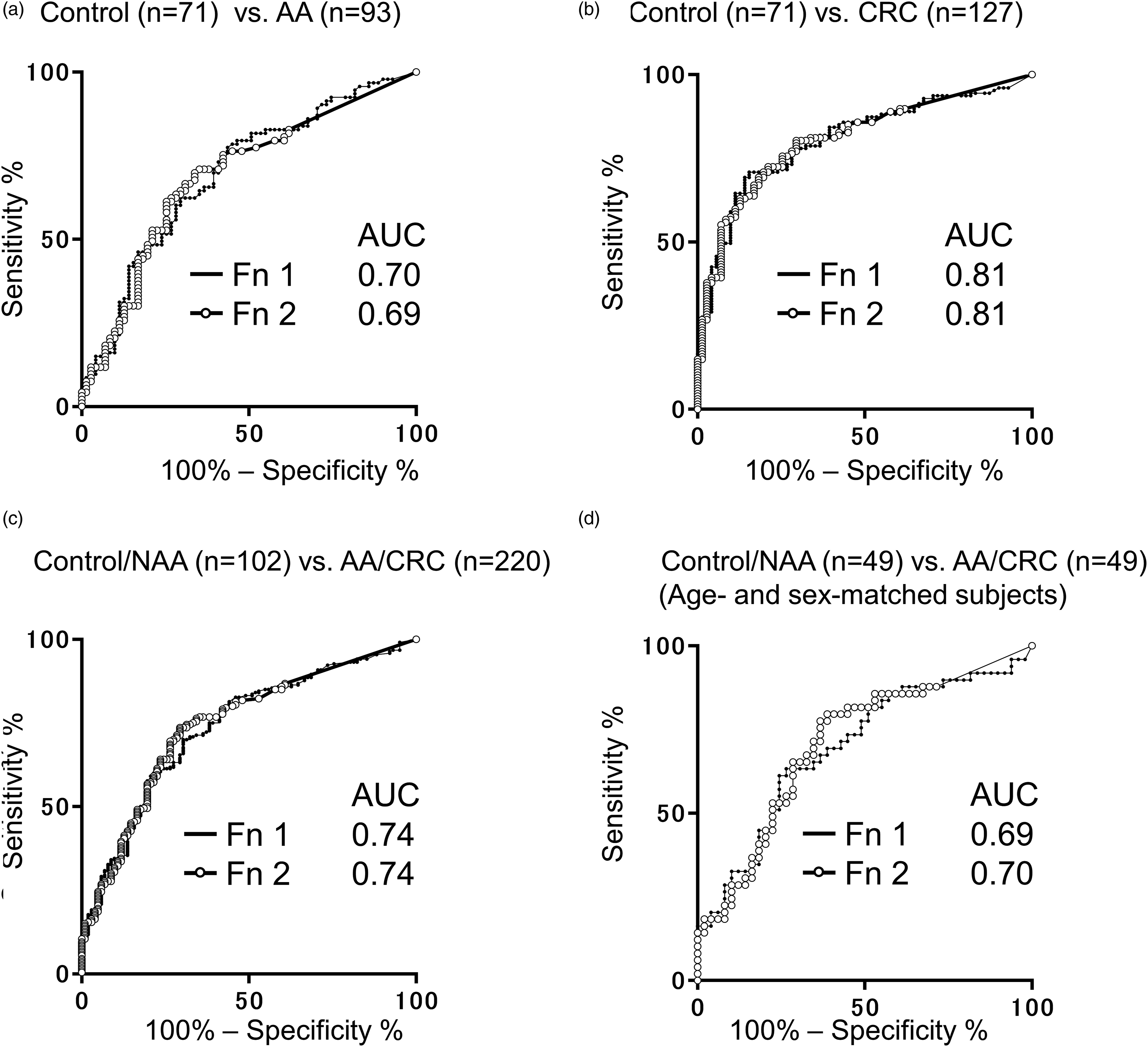
Receiver operating characteristic curve analyses. Comparisons between the control group and the AA group (a), between the control group and the CRC group (b), between the control/NAA groups and the AA/CRC groups (c), and between the control/NAA groups and the AA/CRC groups for age- and sex-matched subjects (d) are shown. AUC, area under the curve, AA, advanced colorectal adenoma. CRC, colorectal cancer. Fn, Fusobacterium nucleatum. NAA, non-advanced colorectal adenoma.
Diagnostic performance.
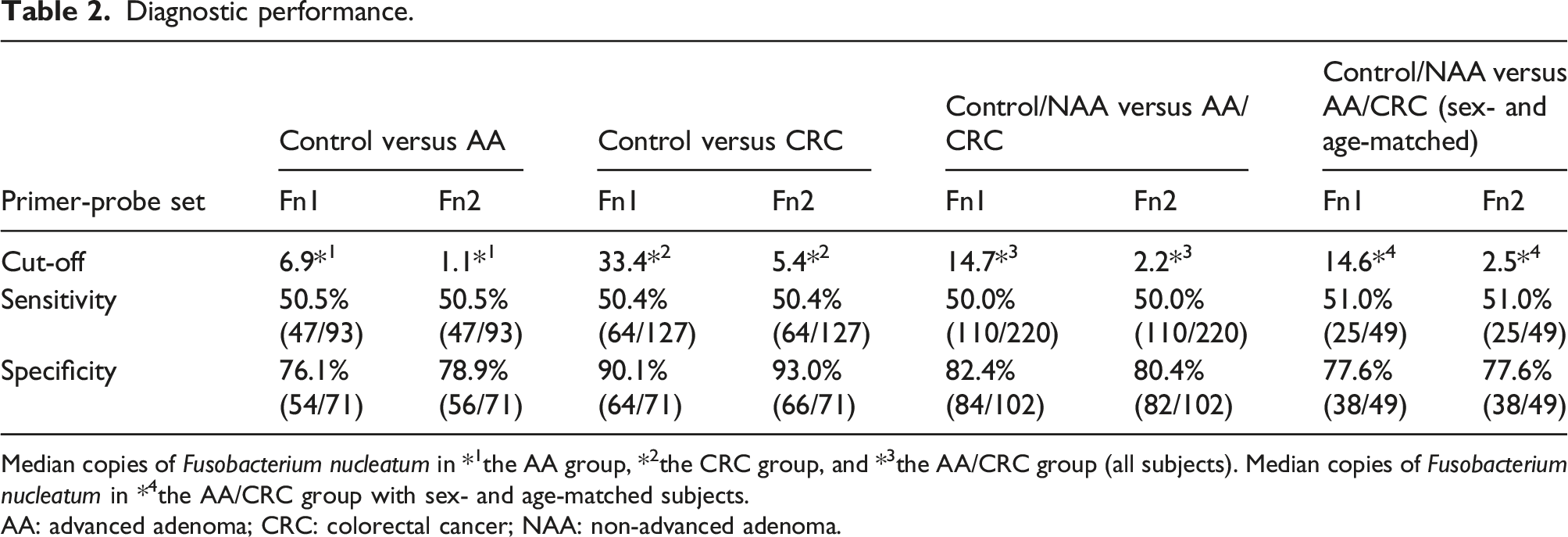
Median copies of Fusobacterium nucleatum in *1the AA group, *2the CRC group, and *3the AA/CRC group (all subjects). Median copies of Fusobacterium nucleatum in *4the AA/CRC group with sex- and age-matched subjects.
AA: advanced adenoma; CRC: colorectal cancer; NAA: non-advanced adenoma.
Discussion
The present study resulted in two important findings. First, the absolute copy numbers of Fn quantified by ddPCR using Fn1 were about five times higher than those using Fn2, and a positive linear correlation was shown between them. Thus, as the Fn1 set amplifies five fragments of the 16S rRNA gene and the Fn2 set can amplify a single fragment of the nusG gene, the measured values coincided with the theoretical values. Faecal Fn was detected in 310 subjects (96.3%) using Fn1 and 253 (78.6%) subjects using Fn2. The difference in the Fn detection rate between Fn1 and Fn2 could reflect the higher detection rate with Fn1 compared to Fn2 primarily in the control/NAA groups because the detection rate of Fn detectable only with Fn1 (Fn1 positive and Fn2 negative) was only 24 of 213 (11.3%) in the AA/CRC groups, whereas it was 35 of 97 (36.1%) in the control/NAA groups. As Fn copy numbers in the control/NAA groups are small and Fn copy numbers quantified by Fn2 are one fifth of those by Fn1, the small number of Fn is detectable only with Fn1 but not with Fn2 primarily in the control/NAA groups. Second, faecal ddPCR-based DNA tests using the Fn1 and Fn2 primer-probe sets have similar diagnostic performance for colorectal neoplasia detection, showing almost the same AUC of the ROC curve analysis, and almost the same sensitivity and specificity, between the two primer-probe sets (Figure 3, Table 2), although the detection rates of faecal Fn were slightly different between Fn1 and Fn2 (96.3% and 78.6%, respectively). To our knowledge, this is the first report to show a comparison of the diagnostic accuracy in colorectal neoplasia between the Fn1 and Fn2 primer-probe sets by ddPCR. Other investigators have shown that Fn is detected in the tissues by qRT-PCR but not in the stool. 12 Furthermore, although Fn2 has been widely used in qRT-PCR for the detection of faecal Fn,1,5–7 excess PCR cycles (up to 60 cycles) of denaturing and annealing steps are required in the qRT-PCR. 1 In contrast, in the present study, ddPCR enabled detection of faecal Fn even using Fn2 with 40 PCR cycles. The highly sensitive detection by ddPCR may be due to its 1000 times higher sensitivity compared with that of qRT-PCR. 13
The limitation of the present study is that the age of the control group subjects was younger than that of the subjects in the other groups Because the microbiota composition of the human gut changes with age, 14 the amount of Fn in faeces might be related to age and not to the presence of colorectal tumor. However, other investigators reported that Fn promotes small and large intestinal tumorigenesis in the APC Min/+ mouse 15 and that FadA, which is adhesin expressed on the surface of Fn, promotes E-cadherin-mediated colorectal tumor growth and induction of proinflammatory cytokines in xenograft mice. 16 Furthermore, in human, a high abundance of faecal Fn was found to be an independent risk factor for metachronous adenomas after endoscopic polypectomy in both retrospective and prospective cohorts. 17 Thus, faecal Fn can be a possible biomarker of the predisposition to colorectal neoplasia. We found an overabundance of faecal Fn in at most 23.9% of the individuals in the control group as shown in Table 2 (100% minus the specificity of 76.1% (54/71), which is the specificity of the Fn1 primer-probe set between the control group and AA group). Careful surveillance is required because we cannot exclude for them the possibility of developing colorectal neoplasia in the future. In this study, we showed the clinical usefulness of Fn in the sex- and age-matched population in the Control/NAA versus AA/CRC groups. However, as the number of patients in the matched population was small, further studies will be required to investigate the relationship between faecal Fn abundance and colorectal neoplasia in age-matched groups with a large number of patients. A prospective study is also needed to investigate whether Fn levels in faeces actually increase the risk of colorectal neoplasia.
In conclusion, as ddPCR gives us the absolute copy number of Fn without the need for calibration samples, the ddPCR-based faecal Fn test enables researchers to directly compare the data of faecal Fn levels for the detection of colorectal neoplasias between different research groups. Because the diagnostic performance was quite similar between the Fn1 and Fn2 primer-probe sets by ddPCR, we propose the ddPCR-based faecal Fn test using the Fn1 and Fn2 primer-probe sets as a possible standardized method for a colorectal neoplasia screening test, considering that the Fn levels quantified using Fn1 are about 5 times higher than those using Fn2.
Supplemental Material
Supplemental Material - Comparison of two primer-probe sets of Fusobacterium nucleatum using droplet digital polymerase chain reaction for the detection of colorectal neoplasia from faecal samples
Supplemental Material for Comparison of two primer-probe sets of Fusobacterium nucleatum using droplet digital polymerase chain reaction for the detection of colorectal neoplasia from faecal samples by Yuko Yamaoka, Mai Sasai, Yutaka Suehiro, Shinichi Hashimoto, Atsushi Goto, Naoki Yamamoto, Nobuaki Suzuki, Shingo Higaki, Ikuei Fujii, Chieko Suzuki, Toshihiko Matsumoto, Tomomi Hoshida, Michiko Koga, Takeya Tsutsumi, Lay A Lim, Yasuo Matsubara, Shinobu Tomochika, Shin Yoshida, Shoichi Hazama, Hiroshi Yotsuyanagi, Hiroaki Nagano, Isao Sakaida, Taro Takami and Takahiro Yamasaki in Annals of Clinical Biochemistry: International Journal of Laboratory Medicine
Footnotes
Acknowledgements
We are very grateful to Ms. Tomoe Senkoji of the Institute of Medical Science, the University of Tokyo for her invaluable help in collecting the clinical data and samples.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by JSPS KAKENHI Grant No. 25460687.
Ethical approval
This study was approved by the Ethics Committee of Yamaguchi University (reference number, H29-228).
Guarantor
YS.
Contributorship
YS designed this study, performed the experiments and the analysis, and wrote the article. YY, MS, and TH performed the experiments and the analysis. YY, SH, AG, NY, NS, SH, IF, CS, TM, MK, TT, LAL, YM, ST, SY, SH, HY, HN and IS were involved in patient recruitment and contributed to the discussion. TT and TY contributed to the discussion and reviewed/edited the article. All authors reviewed and approved the final version of the article.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
