Abstract
Background
As TWIST1 methylation is specific to colorectal neoplasia, detection of TWIST1 methylation from faeces samples might be useful for colorectal neoplasia screening. However, because the content of human DNA in faeces is very small, it is very difficult to detect TWIST1 methylation by conventional bisulphite-based methylation assays. Therefore, we developed a new methylation assay without bisulphite treatment, the combined restriction digital PCR assay, and evaluated its sensitivity and specificity in combination with and without the faecal immunochemical test for haemoglobin for colorectal neoplasia detection from faeces samples.
Methods
For the combined restriction digital PCR assay, DNA was treated with three methylation-sensitive restriction enzymes and an exonuclease, followed by measurement of TWIST1 methylation level by droplet digital PCR. Faecal DNA testing and faecal immunochemical test for haemoglobin were performed on 109 patients with colorectal neoplasia and 10 control individuals.
Results
Basic performance testing showed that the combined restriction digital PCR assay enabled detection of 0.14% of the TWIST1 methylation level for the lymphocyte DNA. The combined restriction digital PCR assay from faeces samples had a sensitivity of 22.2% (95% confidence interval, 2.8–60.0%) for non-advanced adenoma, 47.1% (95% confidence interval, 23.0–72.2%) for advanced adenoma, and 33.7% (95% confidence interval, 23.7–45.0%) for colorectal cancer, and a specificity of 100.0%. Combination of faecal immunochemical test for haemoglobin and faecal combined restriction digital PCR assay increased sensitivity to 82.4% (95% confidence interval, 56.6–96.2%) for the detection of advanced adenoma.
Conclusions
We developed the combined restriction digital PCR assay, a possible highly sensitive methylation assay. Combination of faecal combined restriction digital PCR assay with faecal immunochemical test for haemoglobin may provide an alternative screening strategy for colorectal neoplasia, especially for potentially precancerous lesions.
Keywords
Introduction
Colorectal cancer (CRC) is the second most commonly diagnosed cancer in females and the third most in males in the world. 1 It is estimated that 1.4 million new cases and 693,900 deaths occurred worldwide in 2012. 1 Because more than 95% of patients with CRC would benefit from curative surgery if diagnosed at an earlier or precancerous stage, 2 it is important to develop highly sensitive and specific assays to detect precancerous lesions and CRC at the early stage that are non-invasive, inexpensive, and easy to perform.
The main approach to CRC screening throughout the world is the faecal immunochemical test for haemoglobin (FIT), and patients with faecal haemoglobin > 20 µg haemoglobin/g faeces (equivalent to a 100 ng/mL cut-off of haemoglobin in sample buffer) are referred for colonoscopy. 3 Although the sensitivity of the FIT for the diagnosis of colorectal neoplasia is 92.3% for CRC, it falls to 42.4% for the detection of potentially advanced precancerous lesions. 4 Furthermore, the FIT also carries the risk of false-positive results in patients with haemorrhoids, ulcers, and inflammatory bowel disease.5–7 To avoid the disadvantages of the FIT, more sensitive and specific screening methods are required. One solution is faecal DNA tests. A variety of faecal molecular markers including mutations of oncogenes and tumour suppressor genes, microsatellite instability, microRNA, and DNA methylation reportedly could increase the sensitivity of this test in CRC screening.8–14 In 2014, the U. S. Food and Drug Administration approved Cologuard, the first faeces-based colorectal screening test consisting of the combination of FIT and faecal DNA testing of the mutation of KRAS and methylation of BMP3 and NDRG4.4,15
Previously, we reported that TWIST1 hypermethylation is observed frequently in colorectal adenoma and cancer tissues. 16 This finding challenged us to develop faecal DNA testing of TWIST1 methylation for the detection of colorectal neoplasia. However, methylation assays of faeces samples are very difficult to perform because faecal DNA is derived mainly from bacteria, and the content of human DNA in faeces is very small.17,18 In addition, conventional DNA methylation assays require bisulphite treatment of DNA, which causes degradation and loss of DNA.19,20 Therefore, a new assay is required for faecal DNA testing. In the present study, we developed a highly sensitive DNA methylation assay without DNA bisulphite treatment, called the combined restriction digital PCR (CORD) assay of TWIST1, and evaluated its performance in combination with and without FIT for the detection of colorectal neoplasia from faeces samples.
Materials and methods
Clinical materials
We enrolled 234 participants of whom 119 had results that could be fully evaluated (Figure 1). Faeces were collected from 10 healthy subjects without colorectal neoplasia as determined by colonoscopy (control group), nine patients with colorectal non-advanced adenomas (non-advanced adenoma group), 17 patients with advanced colorectal adenoma (advanced adenoma group), and 83 patients with CRC of stages I–IV (CRC group) diagnosed by colonoscopy or surgical resection. All subjects were Japanese. Adenomas of less than 1 cm in size were categorized as non-advanced adenoma.
4
Criteria of advanced adenoma were defined as adenomas of 1 cm or greater in size, or with villous components (tubulovillous or villous), or with high-grade or severe dysplasia.
4
Staging was classified according to the International Union Against Cancer.
21
The median age of the subjects was 55 years (range, 41–81; first and third quartiles, 42.8 and 73.8 years) for the control group, 65 years (range, 46–81; first and third quartiles, 54.0 and 76.0 years) for the non-advanced adenoma group, 65 years (range, 52–79; first and third quartiles, 65.5 and 73.5 years) for the advanced adenoma group, and 68 years (range, 44–92; first and third quartiles, 62.0 and 76.0 years) for the CRC group. The subjects comprised 73 male and 46 female. CRC stages included Stage I in 26 patients, Stage II in 14 patients, Stage III in 34 patients, and Stage IV in six patients (stage information was not available for three patients). The study protocol was approved by the institutional review board of Yamaguchi University Graduate School of Medicine, and informed consent was obtained from each patient and the healthy volunteers.
Enrolment and outcomes. As not enough faeces could be obtained from 16 participants, the faecal DNA was insufficient for the CORD assay.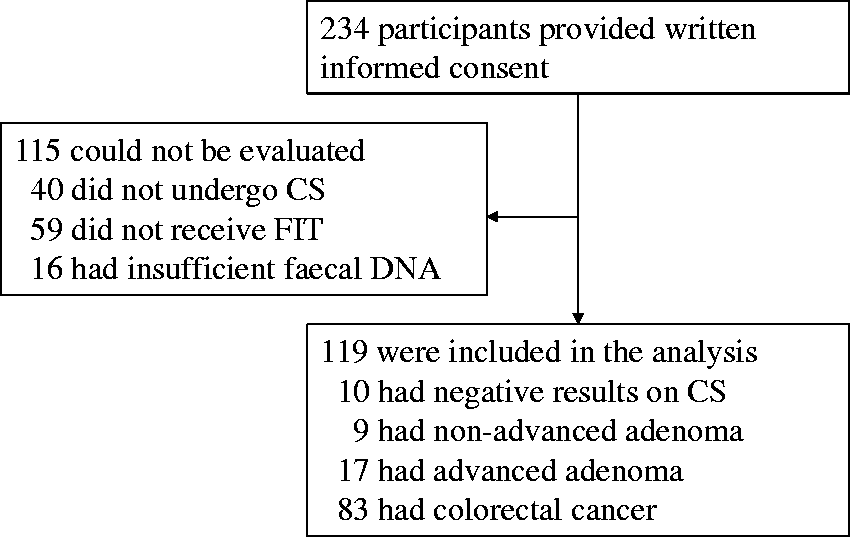
Faecal specimens for DNA testing were collected prior to bowel preparation for colonoscopy or surgical treatment in Yamaguchi University Hospital and were stored at −20℃ until DNA extraction.
FIT
Participants received illustrated, Japanese-language instructions on sampling faeces from one bowel movement by briefly sweeping the tip of a probe several times though the faeces. 22 Fresh faecal specimens were collected into the sampling containers filled with 2 mL of a haemoglobin-stabilizing buffer solution (Eiken Kagaku, Tokyo, Japan) prior to bowel preparation for colonoscopy procedure. 23 FIT was performed using OC-HEMODIA (Eiken Kagaku, Tokyo, Japan), a latex agglutination FIT with analytical characteristics. The OC-Sensor IO instrument processed and quantified the FIT results at the manufacturer-recommended concentration cut-off value of 20 μg haemoglobin/g faeces units (100 ng haemoglobin/1 mL of buffer) for a positive test result. 23
Sample preparation and DNA extraction
DNA from peripheral blood lymphocytes was used as a control for unmethylated TWIST1, and DNA from CRC cell line HCT116 was used as a control for hypermethylated TWIST1. 16 To evaluate linearity between the concentration of methylated TWIST1 in the template DNA and the methylation level of TWIST1 measured by the CORD assay, we prepared DNA mixtures of HCT 116 and lymphocyte DNAs.
Faecal samples were thawed from −20℃, and about 200 mg of each sample was used for DNA extraction using the QIAamp DNA Stool Mini kit (QIAGEN, Tokyo, Japan) according to the manufacturer’s instructions. DNA quantification was performed using NanoDrop 2000 spectrophotometers (Thermo Scientific, Yokohama, Japan).
Methylation assay by CORD
The CORD assay consists of two procedures: One is the enzyme treatment of DNA, and the other is droplet digital PCR, as described below.
Enzyme treatment
Enzyme treatment consisted of two steps. In the first step, 10 µL of eluted DNA was digested for 16 h at 37℃ by the addition of 1 µL of GeneAmp 10× PCR Buffer II, 1 µL of 25 mmol/L MgCl2, 10 units of HhaI, 10 units of HpaII, and 20 units of exonuclease I (Exo I) (all from Thermo Fisher Scientific, Yokohama, Kanagawa, Japan). Exo I was added to eliminate single-stranded DNA that would escape digestion by the restriction enzymes and to avoid PCR amplification of the undigested fraction. 24 In the second step, additional digestion of DNA was performed for 16 h at 60℃ using 10 units of BstUI (New England Biolabs Ltd, Hitchin, UK). After the restriction was complete, the mixture was heated for 10 min at 98℃.
Figure 2 shows target DNA sequences of TWIST1 and hTERT. The TWIST1 had six recognition sites of methylation-sensitive enzymes HhaI, HpaII, and BstUI. When all six of the sites were methylated, the target DNA would escape digestion by these enzymes and would be amplified by PCR. Regarding hTERT, as no recognition sites of theses enzymes existed in the target sequences, hTERT was always amplified by PCR when human DNA existed in both the faecal DNA and the template DNA.
DNA sequences of TWIST1 and hTERT for CORD assay.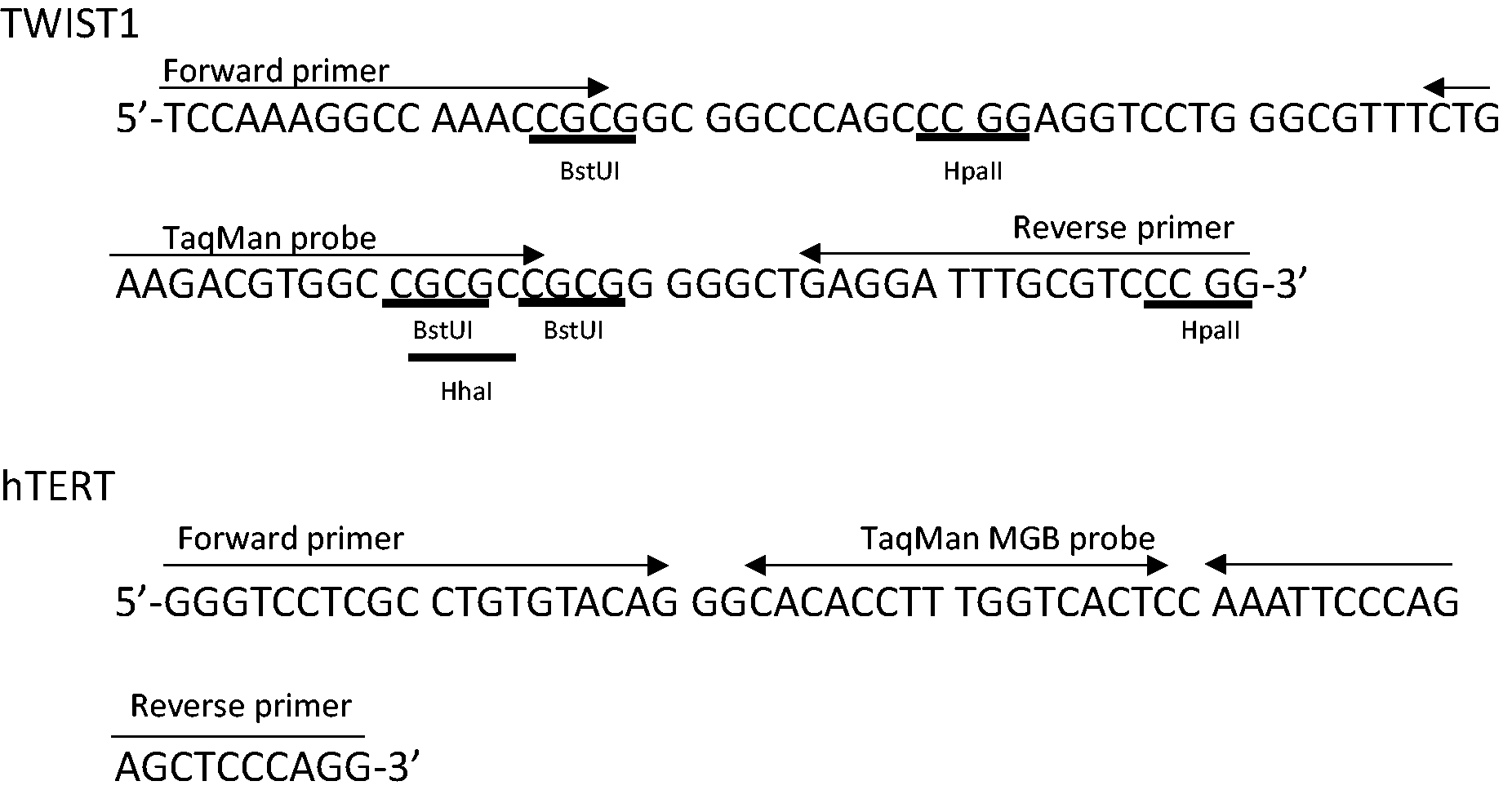
Droplet digital PCR
We performed multiplex droplet digital PCR to count the absolute copy numbers of hTERT and methylated TWIST1. The PCR reaction solution consisted of 2 µL of enzyme-treated DNA, 1× ddPCR Supermix for Probes (BioRad, Tokyo, Japan), 0.25 µmol/L of each primer, and 0.125 µmol/L of each probe in a total volume of 20 µL. For the TWIST1 methylation assay, two sets of a PCR primer pair and a TaqMan probe were designed by Primer Express v3.0 (Thermo Fisher) or Primer3 (Whitehead Institute for Biomedical Research, Massachusetts Institute of Technology) software. We used BLAST (Basic Logical Alignment Search Tool) hosted at the National Center for Biotechnology Information (NCBI) to check that the PCR primers matched only to the intended target (http://www.ncbi.nlm.nih.gov/BLAST). We selected the best primer pair set and the TaqMan probe after comparing the results of PCR using the stool DNAs. The sequences of the primer and probe set of TWIST1 were as follows: forward primer, 5′-TCCAAAGGCCAAACCGC-3′; reverse primer, 5′-CCGGGACGCAAATCCTC-3′; probe, 5′-FAM-CTGAAGACGTGGCCGCGCC-TMARA-3′. The PCR amplicon length is 92 bp from 19,157,854 to 19,157,945 of chromosome 7 (human assembly GRCh37/hg19). Those for hTERT were forward primer, 5′-GGGTCCTCGCCTGTGTACAG-3′; reverse primer, 5′-CCTGGGAGCTCTGGGAATTT-3′; probe, 5′-VIC-CACACCTTTGGTCACTC-MGB-3′. 25 The PCR amplicon length is 60 bp from 1,253,375 to 1,253,434 of chromosome 5 (human assembly GRCh37/hg19). Droplet generation was performed by an automated droplet generator (BioRad) and was followed by PCR. PCR cycling conditions included preheating at 95℃ for 10 min followed by 40 cycles of denaturation at 94℃ for 30 s, annealing at 56℃ for 60 s, and final heating at 98℃ for 10 min. After amplification, the PCR plate was transferred to a QX100 droplet reader (BioRad), and fluorescence amplitude data were obtained by QuantaSoft software (BioRad).
Statistical analyses
To compare variables, the Mann–Whitney U test and Fisher’s test were used, and a P value of less than 0.05 was considered statistically significant. Statistical analyses were performed with StatFlex Ver. 6 (Artec, Osaka, Japan), GraphPad InStat Ver. 3, and GraphPad Prism Ver. 6 statistical software (GraphPad Software, La Jolla, CA, USA).
Results
FIT
Sensitivity and specificity of CORD assay and FIT for the findings of colonoscopy.
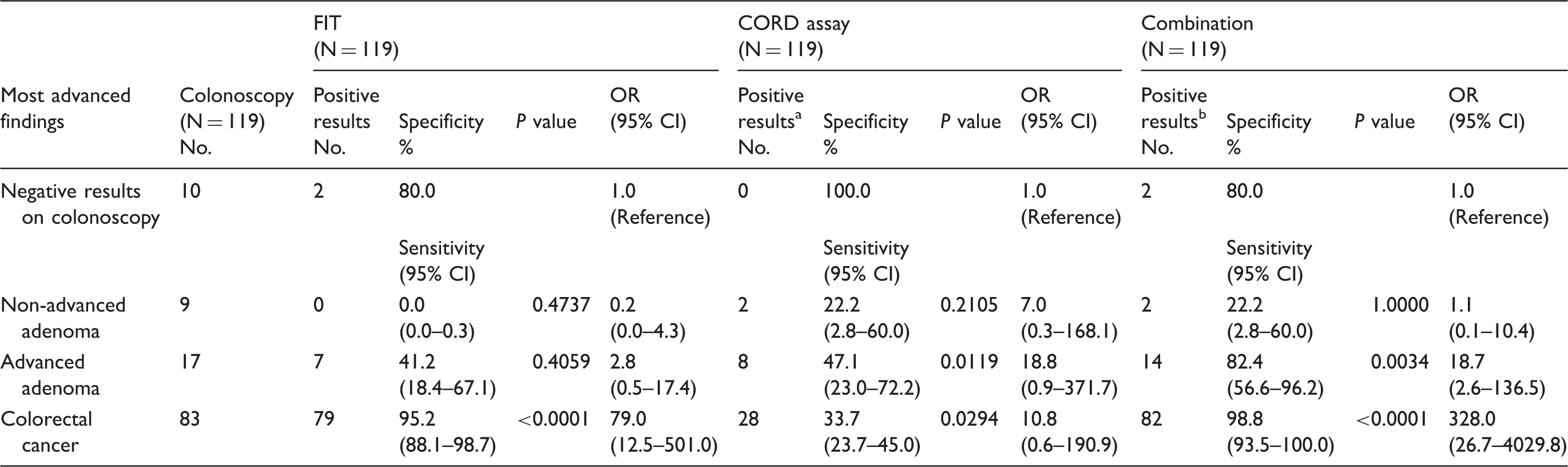
CORD: combined restriction digital PCR; FIT: faecal immunochemical test for haemoglobin; 95% CI: 95% confidence interval; OR: odds ratio.
Criterion for a positive result of the CORD assay is the presence of five or more copy numbers of methylated TWIST1.
Criterion for a positive result with the combination of FIT and CORD assay is either a positive FIT or CORD assay or both are positive.
Quantification of methylated TWIST1 by CORD assay
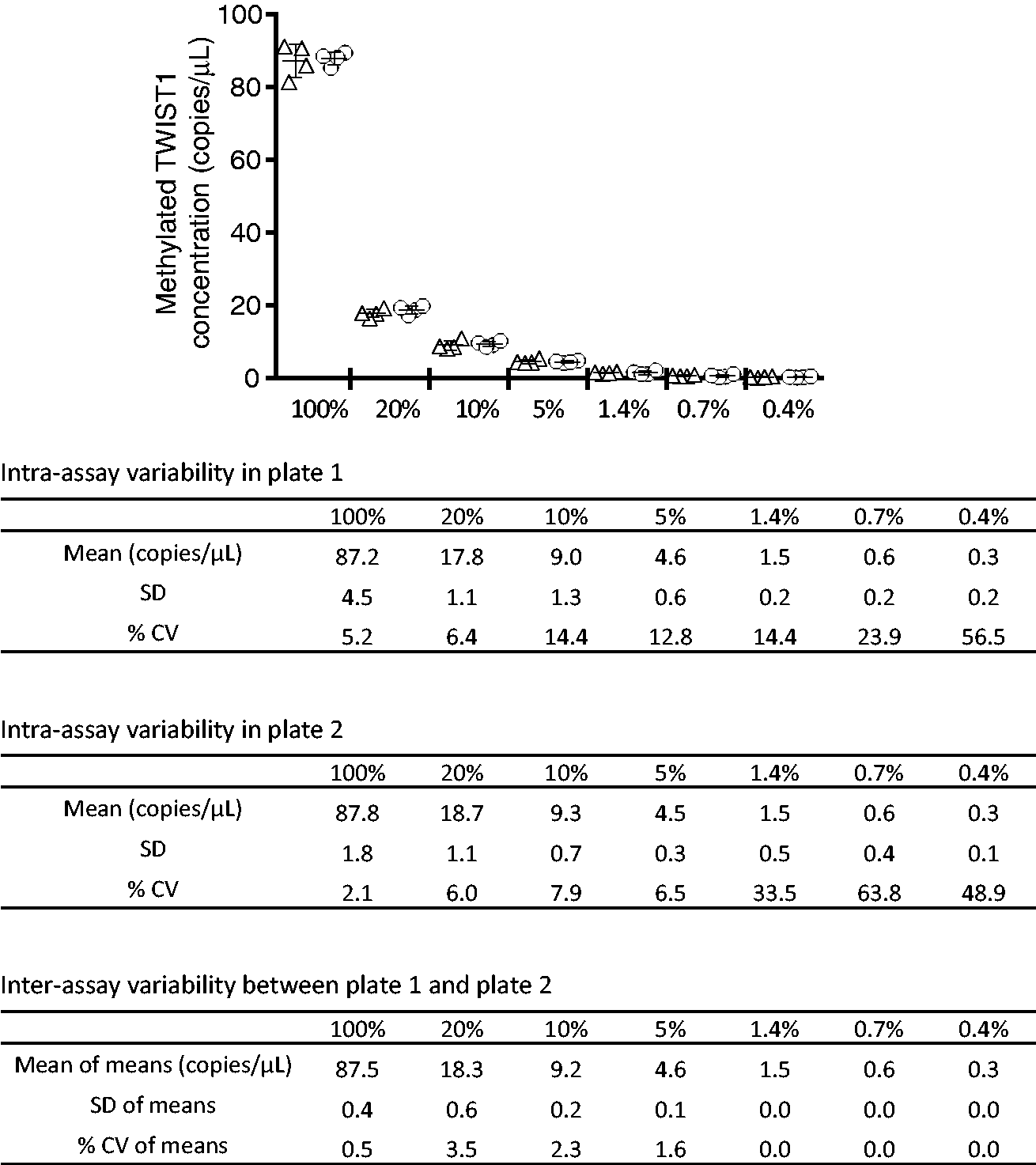
Figure 3(a) and (b) shows the linear relationship between the TWIST1 methylation level evaluated by CORD assay and the concentration of methylated TWIST1 in the template DNA. The CORD assay showed TWIST1 methylation levels of 0.14% for the lymphocyte DNA and 81.1% for the HCT116 DNA (Figure 3(b)). Figure 4 shows the intra- and inter-assay variability of the faecal CORD assay and the mean copy numbers of methylated TWSIT1 that were reproducible.
Basic performance test of the CORD assay. Copy numbers of hTERT and methylated TWIST1 (a) and the ratio of methylated TWIST1 copy numbers to hTERT copy numbers (b) are shown. The error bars represent Poisson 95% confidence intervals. Percentages indicate the ratios of HCT116 DNA to lymphocyte DNA in the template DNA: sample 0% consisted of 0% HCT116 DNA and 100% lymphocyte DNA, whereas sample 100% consisted of 100% HCT116 DNA and 0% lymphocyte DNA. Intra- and inter-assay variability of the CORD assay. Copy numbers of methylated TWIST1 are shown. Percentages indicate the ratios of the stool DNA in the template DNA. The stool DNAs in these experiments were run in quadruplicate on two different plates. The triangles indicate copy numbers of methylated TWIST1 measured on plate 1, the circles indicate copy numbers of methylated TWIST1 measured on plate 2, and the bars indicate the mean with standard deviation.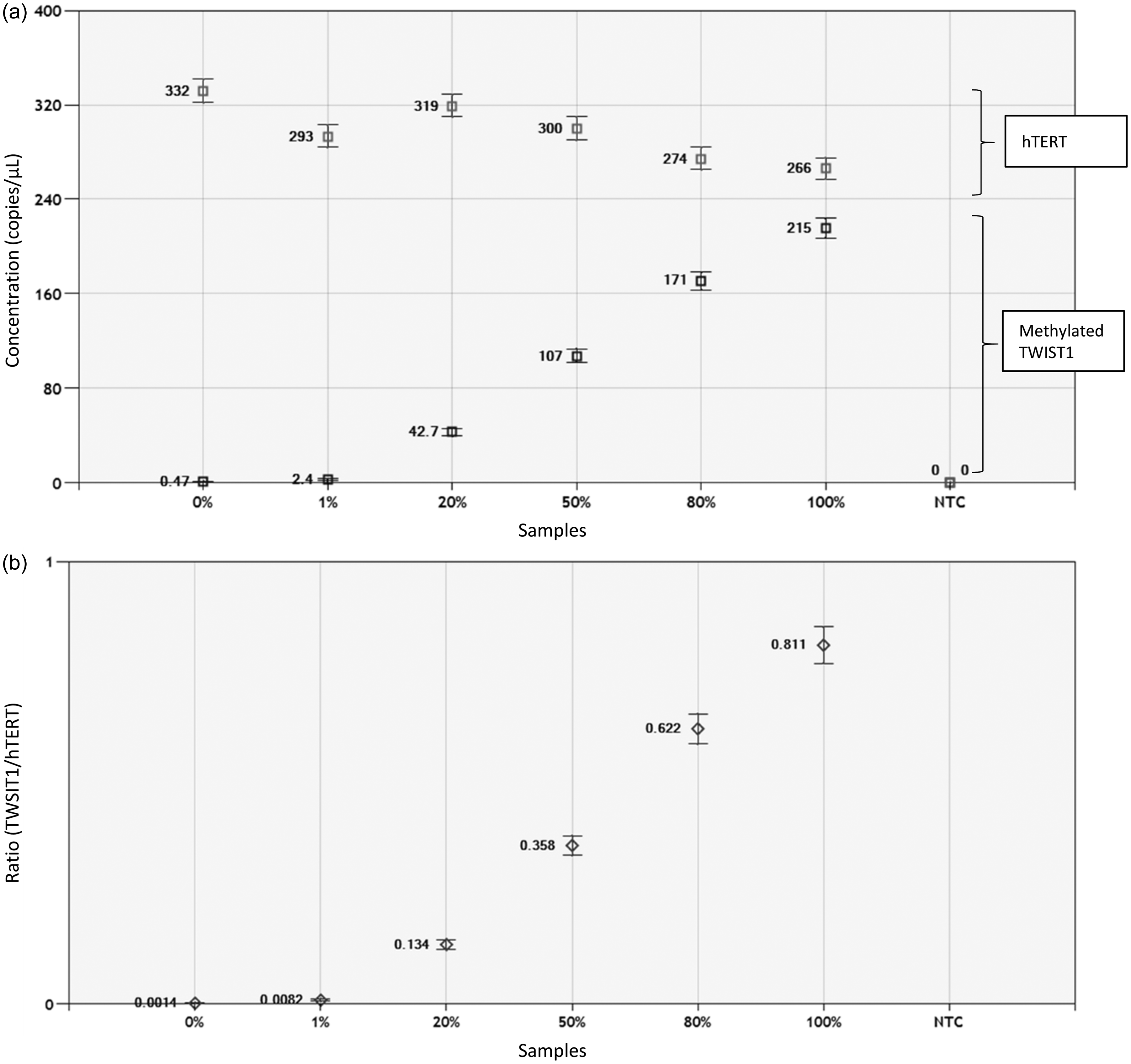
Faecal DNA testing
Faecal DNA testing with the CORD assay (faecal CORD assay) was performed to evaluate the clinical usefulness of the CORD assay for colorectal neoplasia screening. The median copy numbers of methylated TWIST1 were 0.0 (range, 0–2.8) in the control group, 4.2 (range, 0–11.6) in the non-advanced adenoma group, 1.6 (range, 0–240) in the advanced adenoma group, and 1.4 (range, 0–2820) in the CRC group. The area under the receiver-operating characteristic curve was 0.72, and we set five copies of methylated TWIST1 retrospectively as a tentative cut-off point to discriminate between control and CRC. Five or more copies of methylated TWIST1 were found in none (0/10) of the individuals in the control group, in 22.2% (2/9; 95% CI, 2.8–60.0%) of the non-advanced adenoma group, in 47.1% (8/17; 95% CI, 23.0–72.2%) of the advanced adenoma group, and in 33.7% (28/83; 95% CI, 23.7–45.0%) of the CRC group (Table 1 and Figure 5).
Distribution of methylated TWIST1 levels by faecal CORD assay in each group. Each sample is indicated by a closed circle. The box plots show the median with interquartile range (25th percentile and 75th percentile).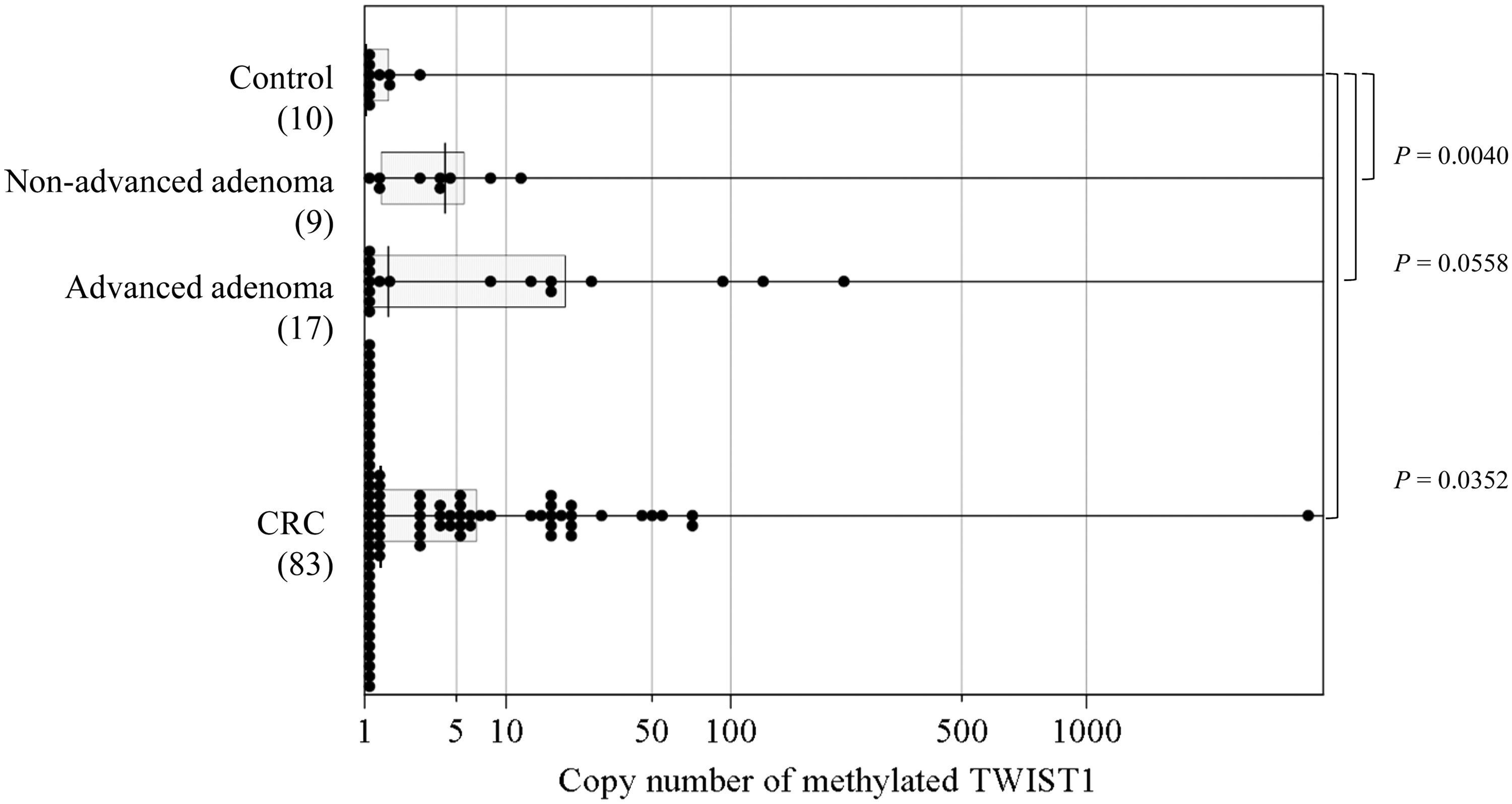
Combination of FIT and faecal CORD assay
The criterion for a positive result with the combination of FIT and CORD assay is either a positive FIT or CORD assay or both are positive. The combination of FIT and faecal CORD assay resulted in a sensitivity of 22.2% (2/9; 95% CI, 2.8–60.0%) for non-advanced adenoma, 82.4% (14/17; 95% CI, 56.6–96.2%) for advanced adenoma, and 98.8% (82/83; 95% CI, 93.5–100.0%) for CRC screening, showing better sensitivity than those by FIT alone (Table 1). The combination of FIT and faecal CORD assay did not worsen specificity as compared to that by FIT alone.
Discussion
Our study resulted in three major findings. First, we established a simple, inexpensive, and highly sensitive methylation assay called the CORD assay that does not require bisulphite treatment of DNA. Second, faecal CORD assay enabled prediction of the presence of colorectal neoplasia with no false positives among the study’s sample population. Finally, as compared to that by FIT alone, the combination of FIT and faecal CORD assay improved the sensitivity for the detection of colorectal neoplasia, especially for the detection of advanced adenoma, without decreasing the specificity.
CORD assay
The CORD assay enabled the detection of 0.14% of the TWIST1 methylation level for the lymphocyte DNA. This is the first report to date to introduce a new, highly sensitive methylation assay. In addition, the CORD assay was applicable for colorectal neoplasia screening from faeces samples. An advantage of the CORD assay is its improvement of the success rates of faecal DNA testing due to the elimination of the bisulphite treatment of DNA. Although we had performed TWIST1 methylation assays using bisulphite-based methods, including methylation specific-PCR, combined bisulphite restriction analysis, and pyrosequencing using the same faeces samples as those of the present study, informative data were obtained in less than 20% of the samples, and PCR amplification failed in the remaining samples (data not shown). Faecal DNA is derived mainly from bacteria, and the content of human DNA is very small. DNA from cells sloughed from the colonic mucosa represents as little as 0.1–0.01% of the total DNA recoverable from faeces.17,18 In addition, although bisulphite treatment of DNA is commonly performed prior to conventional methylation assays, this reaction introduces various DNA strand breaks and results in highly fragmented single-stranded DNA 19 and the loss of ∼90% of the DNA. 20 The bisulphite-based methylation assays require at least 10 copies of the target gene in the template DNA after bisulphite treatment of the DNA. 26 Thus, considering that ∼90% of the DNA will be lost during bisulphite treatment, 19 each template DNA must have at least 100 copies of the target gene prior to bisulphite treatment. In contrast, the CORD assay enables the highly sensitive detection of methylated genes because the CORD assay does not require bisulphite treatment of DNA, and the methylation level is evaluated by droplet digital PCR, which allows the counting of even one copy of the target gene. 27
Faecal CORD assay
The faecal CORD assay resulted in a sensitivity of 47.1% (95% CI, 23.0–72.2%) for the detection of advanced adenoma and a sensitivity of 33.7% (95% CI, 23.7–45.0%) for the detection of CRC, showing moderate sensitivity as compared to those by conventional faecal DNA tests (sensitivities range from 8 to 86% for advanced adenoma and 25–96% for CRC).4,28 In contrast, the specificity of 100.0% with the CORD assay was superior to that of 79–96% with other faecal DNA testing methods. 28 Although high sensitivity is the most important attribute of cancer screening tests, specificity is also important because it affects the number of individuals with false-positive results. Furthermore, two individuals in the control group had positive results by FIT (false-positive results) but negative results by CORD assay (true-negative results), suggesting that the CORD assay might have higher specificity than FIT. However, as the sample size in this study is small, further studies with larger sample sizes are required to determine the best cut-off point of TWIST1 methylation and to confirm the diagnostic accuracy of the CORD assay for colorectal neoplasia screening.
Combination of FIT and faecal CORD assay
We found that the results of FIT and faecal CORD assay were mutually exclusive in the advanced adenoma group. Of the 14 patients with positive result(s) of FIT and/or faecal CORD assay, six (42.9%) patients had a positive result of FIT alone, seven (50.0%) patients had a positive result of faecal CORD assay alone, and only one (7.1%) patient had positive results of both FIT and faecal CORD assay. In the CRC group, the majority (83/87 [95.2%]) of patients had positive results of FIT. Of the remaining four (4.8%) patients with a negative result of FIT, three patients had positive results of faecal CORD assay. Thus, the combination of faecal CORD assay and FIT can improve the sensitivity as compared to that by FIT alone. Indeed, the combination of faecal CORD assay and FIT resulted in a sensitivity of 82.4% (95% CI, 56.6–96.2%) for advanced adenoma detection and 98.8% (93.5–100.0%) for CRC detection, showing higher sensitivities than those by FIT alone. In contrast, the specificity of 80.0% was the same between the combination tests and FIT alone. Although faecal DNA testing of multiple markers in combination with FIT leads to higher sensitivity for colorectal neoplasia screening than that by FIT alone, the disadvantage of the combination test is the decrease in specificity. 4 To avoid a decrease in specificity in the present study, only a single marker was used for faecal DNA testing, and the combination of faecal DNA testing and FIT resulted in higher sensitivities without a decrease in the specificity for colorectal neoplasia detection as compared to those by FIT alone. However, the sample size in the present study is small, and thus, further studies with larger sample sizes are required to confirm the usefulness of the combination of faecal CORD assay and FIT for colorectal neoplasia screening.
In conclusion, we established a highly sensitive methylation assay called the CORD assay, which, to our knowledge, is the first such assay in the world. Because the CORD assay does not require bisulphite treatment of DNA or methylated DNA immunoprecipitation, the cost is lower, and the experimental technique is easier than those of the conventional methylation assays. Faecal testing combining TWIST1 methylation analysis by CORD assay and FIT showed higher sensitivities for the detection of advanced precancerous lesions and CRC without decreasing the specificity than those by FIT alone. Because this study suggests that the combination of faecal DNA testing of TWIST1 methylation and FIT may be useful to detect individuals with colorectal neoplasia, confirmatory studies using independent data-sets are needed to support our findings.
Footnotes
Acknowledgment
We are very grateful to Ms Naoko Okayama and Mr Hidekazu Mizuno of Yamaguchi University Hospital, and Ms Tomomi Hoshida, Dr Toshihiko Matsumoto, and Dr Kiyoshi Ichihara of Yamaguchi University Graduate School of Medicine for their invaluable help in analysing the data.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by JSPS KAKENHI Grant No. 25460687.
Ethical approval
This study was approved by the Ethics Committee of Yamaguchi University (reference number, H19-45).
Guarantor
YS.
Contributorship
Yutaka Suehiro designed this study, performed the experiments and the analysis, wrote the manuscript, contributed to the discussion, and reviewed/edited the manuscript. YZ performed the experiments and the analysis. Shinichi Hashimoto, Shingo Higaki, Yoshitaro Shindo, NS, and Shoichi Hazama were involved in patient recruitment, contributed to the discussion, and reviewed/edited the manuscript. TT, MO, HN, IS, and TY contributed to the discussion and reviewed/edited the manuscript. All authors reviewed and approved the final version of the manuscript. Yutaka Suehiro and YZ contributed equally to this work.
