Abstract
Cellular MR imaging is a young field that aims to visualize targeted cells in living organisms. In order to provide a different signal intensity of the targeted cell, they are either labeled with MR contrast agents in vivo or prelabeled in vitro. Either (ultrasmall) superparamagnetic iron oxide [(U)SPIO] particles or (polymeric) paramagnetic chelates can be used for this purpose. For in vivo cellular labeling, Gd3+- and Mn2+- chelates have mainly been used for targeted hepatobiliary imaging, and (U)SPIO-based cellular imaging has been focused on imaging of macrophage activity. Several of these magneto-pharmaceuticals have been FDA-approved or are in late-phase clinical trials. As for prelabeling of cells in vitro, a challenge has been to induce a sufficient uptake of contrast agents into nonphagocytic cells, without affecting normal cellular function. It appears that this issue has now largely been resolved, leading to an active research on monitoring the cellular biodistribution in vivo following transplantation or transfusion of these cells, including cell migration and trafficking. New applications of cellular MR imaging will be directed, for instance, towards our understanding of hematopoietic (immune) cell trafficking and of novel guided (stem) cell-based therapies aimed to be translated to the clinic in the future.
Keywords
Introduction
Magnetic resonance imaging (MRI) is widely used in everyday clinical practice due to its excellent soft tissue contrast and high anatomical detail. Tissue contrast in MR images (MRI) reflects the differential distribution of hydrogen (1H) atoms in particular tissues. As hydrogen atoms (protons) are mainly found within water molecules in organisms, MRI scans essentially visualize the distribution of water molecules in different types of tissues. This relative distribution varies among tissues, and there are other (macro)molecules (i.e., proteins) that can contain bound water in various amounts. In addition, protons can be found in chemically distinct substances, such as lipids (myelin, adipose tissue), and paramagnetic endogenous tissue iron can also be found in certain tissues. All these factors determine contrast between tissues, so that, for instance, the gray matter can be distinguished from the white matter. MRI contrast agents alter the MR signal by changing the relaxivity of 1H atoms in its vicinity. Generally, changes in relaxivity are considered to affect either the spin–lattice or the spin–spin relaxation characteristics of 1H atoms. The spin–lattice relaxation is commonly referred to as T1 relaxation, whereas spin–spin relaxation is known as T2 relaxation. The inverse of the relaxation times are the relaxation rates R1 and R2, with the effectiveness of contrast agents expressed as relaxivity (R1 or R2 per mM of metal).
The ability to preferentially shorten the T1 rather than T2 relaxation time is commonly used to classify contrast agents as T1 agents. In contrast, a decrease of the relaxation time on T2-weighted scans can be induced by the presence of T2 contrast agents. The metal composition contained within MR contrast agents defines their relaxation properties. Paramagnetic metals such as gadolinium, iron, and manganese mainly affect T1 relaxation, whereas superparamagnetic iron oxide (SPIO) particles predominantly reduce the T2 and T2*. (Figure 1). SPIOs are especially effective in reducing the T2* relaxation time due to the induction of strong field inhomogeneities. A net result is the creation of hypointense signal. Both the extensive preclinical and clinical use of MR contrast agents have proven their reliability and safety [1,2].
Cellular imaging can be broadly defined as the noninvasive and repetitive imaging of targeted cells and cellular processes in living organisms. In contrast to molecular imaging that aims to visualize targeted molecules, cellular imaging is much less specific and involves the cell in its entirety. For cellular MRI, cells need to be labeled with MR contrast agents in order to make them stand out from the surrounding tissues. Incorporation of MR contrast agents into cells renders them distinct from the surrounding tissue and therefore allows the in vivo identification and tracking of targeted cells by MRI. Until recently, the in vivo study of particular populations of cells has been precluded by a lack of appropriate technology. Snapshot images from ex vivo histological studies provided the only source of information to further our understanding of cellular dynamics. The loss of a specific population of cells in pathological conditions could only be assessed by ex vivo assessment or indirectly in vivo by atrophy or other morphological changes of the organ of interest. The development of in vivo cellular imaging by MRI, however, allows the repeated assessment of specific cell loss, migration, and regeneration. Although most types of cells do not readily take up MR contrast agents for cellular imaging, the phagocytic activity of certain types of cells, such as macrophages, allows the incorporation of MR contrast agents to visualize their trafficking in vivo. Nonetheless, recently strategies have been devised to chaperone MR-detectable paramagnetic macromolecules and iron oxide particles into cells in vitro and in vivo, providing an efficacious labeling of a wide variety of cells. Ideally, it will be possible to target a particular population of cells in living organisms for cellular MRI to study their tissue distribution, differentiation/phenotype, proliferation, regeneration, survival, and death.

Paramagnetic and superparamagnetic contrast agents. Schematic representation of the most common concepts currently used in contrast agent design. Gd-based chelates are composed of either a monomeric complex that is very small (~600 Da), which is most commonly used in clinical practice today, or of a polymer containing multipe Gd-chelates (> 1000 Da, typically ~10 kDa) that increase the total relaxivity per molecule. Iron oxide-based contrast agents are larger in size, but also have a greater contrast-to-noise ratio. Different formulations of these agents are geared towards improving their blood half-life for macrophage imaging (USPIO), fast uptake by Kupffer cells for liver imaging (SPIO), or for very sensitive detection of single particles or single cells (MPIOs). With increasing size of the iron oxides, T2* effects become more dominant, whereas due to the lack of inner sphere relaxation T1 effects are negligible. Due to their large magnetic moment, SPIO and MPIO particles are preferred for magnetic labeling of cells.
Cellular MRI Based on in vivo Cellular Targeting
Hepatobiliary Imaging
The clearance of MR contrast agents through either the kidney or liver provides the opportunity for cell specific imaging of the hepatobiliary system. Depending on the biodistribution of MR contrast agents in the liver, these can be classified as: (1) nonspecific extracellular lipophobic agents (cleared through the renal system), (2) reticuloendothelial agents, and (3) hepatocyte-directed agents [3]. Extracellular agents of small molecular weight (~600 Da), such as Gd-DTPA, provide information regarding the vascular anatomy and blood flow, but do not enter cells and therefore do not provide any specific information regarding the status of the cells of the hepatobiliary system. Macro-molecular contrast agents (30–5000 nm in diameter) that are cleared from the blood stream by phagocytosis are taken up by the reticuloendothelial system (RES) that involves the Kupffer cells which are resident in the liver. Application of superparamagnetic agents, such as small particles of iron oxide (SPIOs), cleared through the RES from the blood, therefore afford specific imaging of the Kupffer cell fraction of the liver [4]. Incorporation of the contrast agents into Kupffer cells generally leads to a signal loss on T2-weighted images, but in case of a sparse local distribution, it is also possible to see signal enhancement on T1-weighted images. T1-signal enhancement can, for instance, allow a reliable differential diagnosis and hence improves the diagnosis of hemangiomas [5]. Contrast enhancement can last up to 7 days following injection before the agent is broken down by the lysosomes and further metabolized in the body. During this time, a lack of signal attenuation on T2-weighted MRI scans reflects a dysfunctional RES due to a loss of phagocytic Kupffer cells. The lack of contrast agent uptake into Kupffer cells can be observed in pathological conditions, such as malignant (but not benign) lesions or liver cirrhosis [6]. Increased uptake of commercially available ferumoxides, such as Resovist, Feridex, or Endorem [7–9], generally indicate Kupffer cell hyperactivity or increased vascularity. However, the strong signal change due to the increased uptake of these agents can lead to occlusion of other pathologies, such as hepatic lesions [8], and therefore necessitates a noncontrast agent enhanced baseline scan.
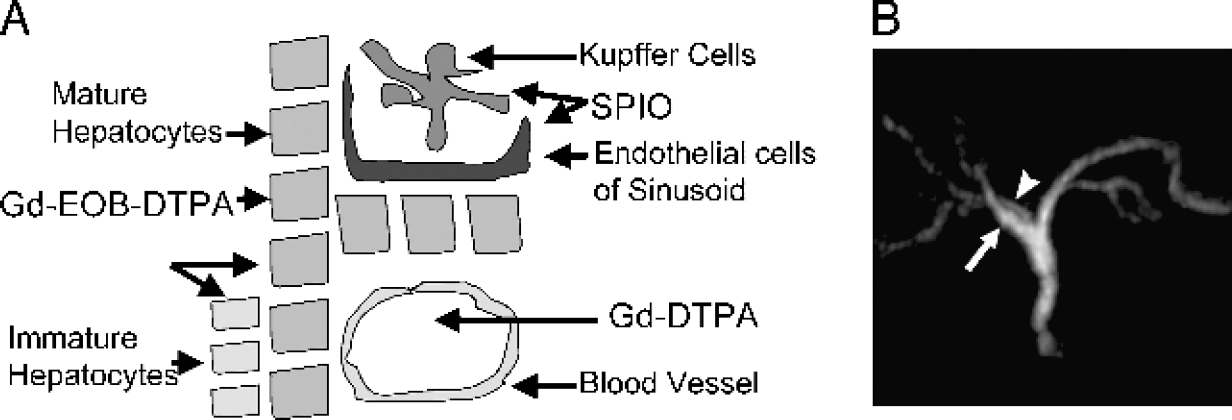
(A) Schematic representation of the different cellular fractions present in the liver and the selective uptake of different contrast agents into the various cell populations. (B) Biliary anatomy in 30-year-old male liver transplant donor candidate following injection of Mn-DPDP. The MR cholangiogram appears normal, whereas conventional T2-weighted MR images were interpreted to be abnormal (not shown). As a normal biliary anatomy was confirmed by surgery, Mn-DPDP enhanced imaging is well suitable for presurgical planning. Adapted from Ref. [17].
The hepatocyte fraction of the liver can also specifically be imaged by cellular MRI. Lipophilic contrast agents are specifically taken up into hepatocytes and excreted through the bile. Superparamagnetic agents are not readily taken up by hepatocytes, but coating with asialofetuin or arabinogalactan to induce specific receptor-mediated endocytosis (RME) has shown some success to promote cellular incorporation [10]. Gadolinium chelates can be rendered hepatocyte-specific by the addition of a lipophilic aromatic chain that leads to hepatocyte incorporation through the membrane-bound organic anion transporter (OATP). Gadoxetate (Gd-EOB-DTPA), a clinically approved hepatocyte imaging agent, for instance, is specifically dependent on the polypeptide Oatp1 activity and is not taken up through Oatp2 [11]. Uptake of gadoxetate through the OATP can be blocked pharmacologically by administration of bromosulfophtalein [12], further supporting the specific uptake mechanism into hepatocytes allowing a very specific in vivo cellular imaging by MRI. From the cytoplasm of the hepatocyte, an enzymatic process excretes these agents through the canalicular aspect of the cell into the bile. A variety of other Gd-based chelates, such as Gd-DTPA-DeA or Gd(Bz-TTDA) [2–14], follow the same pathway of hepatocyte incorporation, but have specifically been designed to further improve the extent and duration of contrast enhancement. Apart from gadoxetate and gadobenate (Gd-BOPTA), Teslascan® (mangafodipir trisodium Mn-DPDP) is also clinically approved as a hepatocyte-specific contrast agent (Figure 2), although its mechanism of action is different from lipophilic Gd chelates. Mn-DPDP is a weakly bound complex that quickly dissociates in vivo. Hepatocytes take up the free manganese and excrete it through the bile. The noncontrast-enhancing ligand complexed with the manganese ions is excreted through the renal system. Mn-DPDP is preferable to manganese chloride (MnCl2) as its ligand slows down the release of the manganese ions and results in an increased hepatocyte uptake. In contrast to Kupffer cell imaging by means of superparamagnetic agents, small lipophilic agents for hepatocyte imaging result in a signal enhancement on T1-weighted MRI scans.
The diagnostic value of hepatocyte-specific imaging lies in the ability to detect lesions of the liver that are characterized by a local decrease in hepatocytes which could be a reflection of nonhepatocyte metastases [6], but can also help to stage hepatocellular carcinomas based on an increase in signal reflecting the large increase of hepatocytes taking-up the cell-specific contrast agent [15]. Mn-DPDP is also incorporated by undifferentiated liver tumors and a combination of Gd-based and Mn-DPDP contrast enhancement therefore potentially allows a differential diagnosis of some liver tumors [16]. The incorporation of manganese into undifferentiated hepatocarcinoma cells is mainly due to the nonspecific nature of cellular uptake.
These different approaches to image the hepatobiliary system also find its application in determining various contributions to the success of liver transplants. A fully functioning transplanted liver would appear to have normal perfusion characteristics and uptake of both RES and hepatocyte-specific contrast agents. Normal biliary excretion and anatomy can be accurately assessed with Mn-DPDP enhanced imaging, whereas conventional T2-weighted imaging cannot [17] (Figure 2). In light of cell transplantation studies, it is not only possible to define the vascular properties of the damaged liver, but it is important to determine to what degree the two major fractions, Kupffer and hepatocytes, of the liver are affected as a specific cell replacement strategy can then be envisaged. Sequential administration of contrast agents can highlight these different aspects of liver damage and regeneration in addition to the potential immunological rejection response to transplanted liver cells [18].
Macrophage Imaging
The development of cellular MRI of Kupffer cells, the resident phagocytes of the liver, using iron oxides [19,20], provided the rationale to further develop this approach to visualize macrophages elsewhere in the body. Following intravenous injection of USPIOs, developed to have a longer blood half-life, the particles are taken up by macrophages including those in lymph nodes [21–23] and bone marrow [24]. Uptake of USPIO into macrophages has been suggested to be dependent on endogenous and exogenous factors such as cytokines [25,26], but is also influenced by particle size [27]. The in vivo cellular targeting of macrophages by intravenous injection of USPIO results in a darkening of the lymph nodes on the MR images [28] and allows the detection of tumor metastases which will remain isointense as these cells do not readily incorporate iron oxide particles.
Cellular imaging of monocyte/macrophage activity has found further applications in several disease models. In the brain of rats and mice with experimental autoimmune encephalomyelitis (EAE), macrophages are present in the multiple inflammatory foci that are found around blood vessels with a disrupted blood–brain barrier. These foci can be visualized by MRI following intravenous injection of MION-46L [29] or USPIO [30–32]. The magnetic labels may either be phagocytized by circulating monocytes or leak through the disrupted blood-brain barrier followed by uptake in resident brain microglia, but can also be used to study the contribution of T cells to lesion development [33]. Detection of macrophage trafficking in vivo can be used to predict the efficacy on anti-inflammatory drugs [34] potentially providing a surrogate imaging marker of treatment efficacy and is currently in the process to find its translation into clinical applications.
In cerebral ischemia, the outer margin of the lesion is first infiltrated by macrophages that can be visualized following USPIO uptake before they spread to also cover the inner core of the lesion [35–37], which in the late chronic phase can also be detected without USPIO incorporation on heavily T2*-weighted scans [38]. Conventional Prussian blue staining often fails to detect the presumably minimal amounts of iron delivered to the brain after peripheral USPIO administration. Therefore, using the same animal model, Schroeter et al [39] developed an improved method of highly sensitive detection of lesion-induced iron uptake following the in vivo USPIO administration with the diaminobenzidine (DAB) and silver/gold enhancement of Prussian blue staining. In the photothrombotic cortical stroke model, Saleh et al. [40] validated the MRI presentation of macrophage accumulation in the ischemic borderzone 6 days after stroke by correlating the in vivo macrophage infiltration with histological detection of USPIOs in macrophages. Detailed analysis of DAB and silver/gold enhanced Prussian blue staining for iron content demonstrated perfect colocalization of the T2*-weighted hypointensity with macrophage accumulation, as depicted on ED1-positive immunohistochemical staining (Figure 3). This study further supported the applicability of in vivo labeling of macrophages at high-field (7T) MRI indicating that sufficient uptake of USPIO occurs. Translation of this approach to human patients with stroke [40] has demonstrate two phases of USPIO imaging. During the first phase, signal loss on T2/T2*-weighted images decreased rapidly after USPIO injection, representing the USPIO blood pool effect. In the second phase, signal enhancement on T1-weighted images increased gradually over time, representing the USPIO uptake by macrophages.
Macrophage activity has furthermore been visualized in the kidney. In the uninjured healthy kidney, no macrophages are present. Injection of the nephrotoxic antibiotic puromycin aminonucleoside induces an influx of macrophages to the kidneys. Following in vivo USPIO injection, both the medulla and cortex appear hypointense with the strongest signal changes at the cortex–medullar junction [41]. Administration of an anti-glomerular basement membrane monoclonal antibody leads to a selective immune-mediated toxicity of the glomerulus. This activation of residing mesangial cells causes an infiltration of macrophages that produces a hypointensity in the cortex [42]. A hypointense band can also be observed on MR images following the infiltration of macrophages in the outer medulla of the kidney after bilateral ischemia and reperfusion [43]. In other areas of the body (e.g., in the knees of animals induced with arthritis), macrophages have also been detected in inflammatory processes [44].

Coronal sections of rat brain, 6 days after photothrombosis, 24 hr after intravenous application of USPIOs. Matching of the T2*-weighted image (A), positioned at the lesion periphery, with in situ iron staining (DAB and silver/gold impregnation enhanced Prussian blue) (B), and ED1 immunohistochemistry (C) identifying macrophages, reveals precise colocalization of the respective signals. Accuracy of spatial agreement between T2*-weighted image (D) and iron staining (E) becomes even more pronounced for sections through the lesion center, where the accumulation of macrophages is depicted as a layer around the lesion.
In atherosclerotic plaques of hyperlipidemic rabbits, monocytes from the blood infiltrate into the aortic wall and differentiate into macrophages detectable in vivo after USPIO administration [45,46]. Iron oxides have also been used to monitor monocyte recruitment into aortic atherosclerotic plaques in apo E knockout mice [25]. Iron-laden macrophages can be detected in the aortic subendothelium and the effect of cytokine injection upon cell infiltration can be studied. Following a retrospective clinical study [47], a recent prospective trial [48] found that uptake occurred mainly in ruptured and rupture-prone plaques and not in stable lesions indicating that the two can be differentiated in order to assess the relative risk for stroke and embolic complications.
Another application of in vivo cellular targeting of macrophages exists in visualizing organ graft rejection. When renal allograft transplants were studied in rats following USPIO injection, a good correlation was found with the hypointense signal intensity, representing infiltrating macrophages, and the graft rejection [49,50]. No significant signal changes were observed when animals were immunosuppressed. Similarly, heterotopic lung transplants in rats without cyclosporine A treatment showed transplant rejection histologically along with a hypointense MR signal following USPIO injection [51]. Syngeneic transplants lacked hypointensity post-contrast, and immunosuppressed animals showed intermediate contrast. Similar results have also been obtained for heterotopic heart transplants [52] (Figure 4).
Cellular MRI Based on in vitro Prelabeling of Cells
Use of Paramagnetic Chelates
Paramagnetic contrast agents, such as Gd-DTPA, are among the most commonly and longest used in medical research. Although a variety of molecules have paramagnetic properties, it is the versatility of gadolinium (Gd3+) [53] that has generated a wide variety of different contrast agents for clinical use (e.g., Gd-DTPA, Gd-DOTA, Gd-BOPTA, Gd-HP-DOTA, Gd-DTPA-BMA, Gd-EOB-DTPA) [1]. It is therefore not surprising that the wide availability of these compounds has spurred some interesting approaches to the labeling of cells for in vivo visualization by MRI.

Allotransplanted rat heart following intravenous injection of USPIO (3 mg Fe/kg body weight). Shown are the long-axis (A,B) and short-axis (C,D) MR images obtained at 4.7 T, 6 days before USPIO infusion (A,C) and 24 hr after USPIO injection (B,D). White and black arrows indicate heart and lung, respectively. A decrease in MR image signal intensity is detected in images of the allogeneic transplanted heart 24 hr after infusion of dextran-coated USPIO particles compared with the preinfusion signal, corresponding to USPIO uptake by macrophages present in the allograft [52].
Prelabeling of cells or cell clusters in situ can provide an exquisite approach to follow the fate of the cells once they are reintroduced into a living organism. Jacobs and Fraser [54] have studied organogenesis. Labeling of embryonic stem cells in vitro allowed a serial assessment of the participation of these cells in various organs in the developing Xenopus [55]. The creation of developmental atlases based on the three-dimensional rendering of cell tracking during development will provide a clearer understanding of the dynamic processes involved in the creation of an organism [56] and define novel avenues to potentially diagnose early developmental malformations in utero.
In order to corroborate the in vivo visualization of cells, Hüber et al [57] developed bimodal paramagnetic contrast agents, such as Gadolinium Rhodamlne Dextran (GRID), that can be detected by both MRI and fluorescent histology (Figure 5). It is therefore possible to track the migration of cells in the damaged brain and to validate their position and phenotype histologically [58,59]. Moreover, in vitro studies demonstrated that GRID was retained within the cells and no evidence of cell leakage and reuptake in a co-culture system were observed, suggesting that this contrast agent allows a reliable distinction between labeled and nonlabeled cells by MRI. After implantation of GRID-labeled neural stem cells into ischemia-damaged hippocampi, neuronal and glial cell differentiation were observed, demonstrating that cells labeled with this contrast agent maintain their potential to respond to the local environment and adopt appropriate phenotypes [58]. The combined use of the paramagnetic Gd-HPDO3A and the fluorescent Eu-HPDO3A (both having very similar cell uptake characteristics) also allows the reliable visualization of labeled endothelial progenitor cells (EPCs) after implantation into the kidney of mice. MRI contrast is produced by means of a hyperintense signal on T1-weighted images due to the small particle size, whereas histological localization of the contrast agents by fluorescent microscopy demonstrated incorporation in membrane-bound cytoplasmic perinuclear endosomes [60]. The use of multimodal agents has the further advantage of the potential in vivo corroboration of the MRI observation by optical imaging technology [61], in addition to more conventional fluorescence histology. Daldrup-Link et al. [61] suggested that Gd-based compounds are preferable to iron-based contrast agents for the design of multimodal agents due to a considerable fluorescent quenching during conjugation to iron oxide particles as a result of nonradiative energy transfer [61,62]. Nonetheless, novel conjugation strategies might overcome these issues. Combining these multimodal properties with paramagnetic chemical exchange saturation transfer (PARACEST) agents might further enhance the potential not only to visualize cells by different imaging modalities, but provide a means by which different type of cells can be tracked separately in vivo. Aime et al. [63] recently reported that two different contrast agents could be distinguished based on the transfer of saturated magnetization to the water molecules at specific irradiation frequencies. Unlike currently used Gd- and Fe-based agents, which commonly affect both T1- and T2-relaxation, the PARACEST agents are far more specific in their contrast generation without contaminating contrast signals at other radio-frequencies. High molecular weight polymers such as poly-

In vitro example of GRID-labeled neural stem cell. The uptake of the bimodal contrast agent GRID can be easily visualized in vitro by means of fluorescent microscopy (in red) and a confocal image can provide a three-dimensional profile of the distribution of the contrast agent within the cell. Here a single (as indicated by the Hoechst counterstain in blue) murine neural stem cell from the MHP36 cell line underwent 16 hr of incubation of GRID inside the culturing media (without any addition of transfection agent) to also allow identification by MRI.
Still, not all paramagnetic chelates are readily taken up into mammalian cells and contrast agents need to be engineered to facilitate cellular incorporation. Improved cellular internalization of paramagnetic contrast agents into cells has been shown for gadolinium particles incorporated in chitosan nanoparticles [66], linked to membrane translocation peptides [67–71], and combined with transfection agents [72]. Additional engineering of these functional properties of contrast agents has resulted in multifunctional compounds that can cross the cell membrane via a transmembrane transport unit (TPU) and guide the paramagnetic chelate by means of a cellular localizing unit (CLU) to particular cellular compartments, such as the cell nucleus [73]. Further strategies that enable an efficient (>95%) and fast (<30 min) labeling approach while preserving cell functioning are desirable to develop this approach into a viable clinical tool to visualize, for instance, transplanted cells. Although all of these approaches resulted in an improved cellular uptake compared with conventional contrast agents, such as Gd-DTPA, these studies only investigated in vitro uptake behavior and it is therefore at present still unclear if these approaches will indeed result in a better in vivo detection of labeled cells by MRI. Still, other studies using paramagnetic chelates suggest that Gd-based contrast agent can provide sufficient signal changes in vivo to allow cellular imaging and therefore strategies aimed at improving or regulating cellular uptake could greatly help future studies [74].
The uptake of manganese chloride (MnCl2) into activated cells via Ca2+ channels in vivo has recently been exploited to probe the functional connectivity of cells [75–77] and the temporal dynamics of regional cellular plasticity involved in learning [78]. The ability to label active cells in vivo opens new vistas to the study of cellular connectivity in three dimensions over time that will greatly benefit our understanding of how different cell assemblies in the brain interact to produce behavior. The ease of labeling cells through their Ca2+ channel also offers a straightforward approach to label cells in vitro prior to transplantation to track these cells in vivo [79]. The potential advantage of using manganese for cellular tracking is the visualization of the functional connections that these cells form in vivo as manganese will be transported across active synapses. This approach therefore allows the validation of how these cells integrate into a functional network in vivo. However, the disadvantage of this approach is that manganese leaking out of these labeled cells would not allow a reliable tracking of the transplanted cells per se.
To achieve cellular labeling in the CNS in vivo, contrast agents will not only have to penetrate the cell membrane, but will also have to cross the intact blood–brain barrier (BBB). The temporal opening of the BBB with, for instance, mannitol allows the penetration of larger molecules into the intact brain parenchyma [80]. Similar to achieving better cellular uptake, molecular coating strategies or the addition of actively transported peptides have been used to allow infiltration of paramagnetic agents across the intact BBB [81]. These novel approaches promise a wide applicability of contrast agents to visualize particular cellular targets by overcoming the inherent limits of large metal ions to spontaneously integrate into mammalian cells.
It is also possible to use reporter genes engineered into cells to serve as markers for cellular imaging. For instance, Louie et al. [74] described a smart contrast agent, EgadMe, that will only produce an MR signal change in the presence of β-galactosidase, a molecular reporter often used in neurodevelopmental studies to identify the progeny of a parent cell. As this contrast agent does not produce a change in the MR signal if it is incorporated into cells without this gene, false positives created by leakage and reuptake of contrast agents in vivo are no longer issues to be concerned about. The engineering of smart contrast agents, such as EgadMe, in combination with functional surface molecules, could in the future allow researchers to repeatedly and selectively visualize particular populations of cells by administering contrast agents in vivo. In a recent study, Cohen et al. [82] constructed C6 tumor cells expressing both EGFP and the H-chain of the iron storage molecule ferritin. This expression was regulated by the presence of tetracycline. Changes in relaxivity (due to iron load and accumulation of ferritin) in response to tetracycline-regulated suppression of the transgene were detected both in vitro and in vivo. The use of such MRI reporters which can be switched ON or OFF depending on the presence of exogenous substrates is a very attractive perspective for noninvasive analysis of the regulation of gene expression in various (transgenic) disease models.
However, despite the versatility of paramagnetic contrast agents, a large number of gadolinium molecules are needed in each cell to allow a robust in vivo visualization, although some strategies, such as metallofullerines [83], have been described to increase relaxivity and therefore could reduce the number of paramagnetic molecules needed for cellular imaging. At present, paramagnetic contrast agents still produce less signal change than superparamagnetic agents, which allow a more robust and efficient detection of a small number of cells. Dalrup-Link et al. [84] demonstrated that twice as many cells labeled with gadopentate dimeguline liposomes were needed for detection by MRI compared to ferumoxides, P7728 liposomes, or polysaccharide nanoparticles. USPIO particles have an inherently stronger effect on MR relaxivity, and therefore, to date, dominate the field of cellular imaging by MRI.
Use of (U)SPIO Particles
Unmodified (U)SPIOs can be used at high concentrations to label mononuclear (T) cells [85,86], monocytes [87], glioma cells and macrophages [88,89], and oligodendrocyte progenitors [90] in vitro, but more efficient uptake methods are desirable. Thus, for an improved uptake of magnetic nanoparticles, the labeling strategies need to be optimized or modified to have an appropriate outer surface layer that not only binds to cellular membranes, but also induces internalization of the particles into the cytoplasm. Trypan blue exclusion-based (i.e., labeling of dead cells) toxicity studies using high concentrations of USPIOs show good tolerance with minimal cell death [91]. Nonetheless, reports of free radicals generation, decreased cell proliferation, and even cell death [92] emphasize the need to evaluate more thoroughly toxicity of the individually chosen labeling strategies. In contrast, particles that do not internalize and stay attached to the outer cell membrane are likely to interfere with cell surface interactions (including cell homing into tissues), may detach easily from the membrane, or can be transferred to other cells. Several modifications of iron oxide particles have now been described that induce an efficient internalization of the contrast agent while trying to minimize any deleterious effects on cellular functions [93].
Early attempts have used magnetoliposomes [94] or lectins [95] to increase the uptake in (nonphagocytic) cells. A significant improvement of labeling could be achieved by linking the particles to the HIV tat peptide [96,97]. This peptide contains a membrane translocating signal that efficiently transports the iron oxides into cells with a nuclear accumulation. The reported nuclear localization, however, appears in contradiction with a recent study that primarily observed a cytoplasmic distribution of the particles [98]. No differences between labeled and unlabeled T cells were observed on cellular functions [99]. Conjugating of a higher number of tat peptides per iron oxide particle results in an increased cellular uptake [100]. Another approach for intracellular magnetic labeling is the use of internalizing monoclonal antibodies (mabs). The mouse anti-transferrin receptor (Tfr) mab OX-26 [101] induces internalization of the Tfr upon binding. Conjugation to OX-26 improves cellular uptake of drugs by RME. MION-46L-OX-26 complexes have been used to magnetically label oligodendrocyte progenitors [102] and neural precursor cells [103] by RME. The particles accumulate in intracellular endosomes in high numbers. The Tfr represents a convenient system as the receptor is expressed in high numbers on many cell types. The natural ligand of Tfr, Tf, has also been used for internalization of iron oxide nanoparticles following conjugation to human Tf [84,104]. Nonetheless, there have been reports that Tf-coated iron oxides localize in the cell membrane without further internalization [105]. Another example of cell labeling using internalizing mabs is the use of biotinylated anti CD-11 mab in conjunction with streptavidinylated dextran-coated iron oxides. These commercially available mabs significantly improved magnetic labeling of dendritic cells [106]. A disadvantage of the use of internalizing mabs, however, is that they are species-specific. Newly synthesized antibodies are therefore required for different species. Additionally, the use of a xenogeneic (i.e., mouse) protein will face regulatory concerns regarding eventual clinical use.
Several other magnetic labels have recently been developed with the important property of having nonspecific uptake by a variety of mammalian cells, regardless of cell origin or animal species. One of these labels is magnetodendrimers or MD-100 [107,108] consisting of iron oxide nanoparticles coated with a generation 4.5 carboxylated dendrimer. Dendrimers are currently used as agents to transfect oligonucleotides into cells [109–111]. The highly charged polymers bind on multiple sites on the cell membrane, induce membrane bending followed by endocytosis [112]. Even at low concentrations of iron (10–25 μg Fe/mL), cells originating from mouse, rat, and human can be easily labeled by simple addition of MD-100 to the cell culture for 24–48 hr [103,108,113–118]. A nonexhaustive list of different cell types labeled with MD-100 include mouse ES cells, mouse fibroblasts, mouse muscle progenitors, rat neural stem cells, rat oligodendroglial progenitors, rat olfactory ensheathing cells, human small cell lung carcinoma cells, human cervix carcinoma cells, human mesenchymal stem cells (MSCs), human neural stem cells, and human embryoid body derived pluripotent stem cells. All cells show a comparable degree of particle uptake into endosomes highlighting the nonspecific, but localizing, nature of the MD-100 uptake. There is no difference between MD-100 labeled and unlabeled cells in terms of proliferation and viability, the ability of neural stem cells to differentiate into neurons or to myelinate axons in vivo [108], and differentiation into myotubes [118].
Anionic magnetic nanoparticles have been developed and used for intracellular labeling of macrophages [119–121]. The negative surface charge of the particles induces an uptake three orders of magnitudes higher as compared to conventional dextran coated SPIOs. The particles also end up in endosomes, the shape of which can be deformed by applying an external magnet [122]. A disadvantage of these particles, as well as of the magnetodendrimers, is that they have not yet been commercially developed and are thus not widely available. Recently, however, a convenient magnetic labeling method that is based on the use of a commercially available (U)SPIO formulation, Feridex® [123,124] or Sinerem [91], has been developed. The method is based on the use of transfection agents, which are also commercially available. By mixing the (U)SPIO formulation and the transfection agent, complexes of the two are formed through electrostatic interactions [125] (Figure 6). When the complexes are added to the cell culture, the transfection agent effectively shuttles the (U)SPIO into the cell through formation of endosomes. Transfection agents that are suitable for this purpose include dendrimers (Superfect), poly-

Mechanism of endosomal magnetic labeling using PLL–Feridex complexes. The dextran coating of the SPLO has a negative surface charge (Zeta potential of −41 mV) [125], whereas the polycation transfection agent PLL has a positive charge. When SPIO and PLL are mixed before incubation with cells, the PLL will coat the SPIO through electrostatic interactions, and the SPIO will obtain a net positive charge. Upon adding the complexes to cells in culture, the SPIO–PLL complexes will bind to the anionic cell membrane at multiple places (due to the large size of the SPIO–PLL complexes, in the MDa range), causing membrane bending and membrane disruption. The cell membrane then forms small invaginations, encapsulating the complexes, following which the outer ends of the cell membrane fuse resulting in the formation of endosomes.

Endosomal magnetic labeling of cells using PLL–Feridex. Shown are Prussian blue stains of (A) mouse D3–ES (embryonic stem)-derived neural progenitors and (B) human EG (embryonic germ)-EBD (embryoid body derived) SDEC stem cells [187]. The iron is present in intracellular endosomes and absent in the nucleus. The concept of using PLL to enhance the intracellular uptake of macromolecules has been around for almost three decades [131]. For comparison, PLL-enhanced uptake of the protein HRP is shown in C–F [131], with diaminobenzidine as peroxidase substrate. PLL was covalently conjugated to HRP, and incubated with mouse fibroblasts for 60 min at 37°C. (C) HRP–PLL at 15 μg/mL, (D) HRP–PLL at 1.5 μg/mL, (E) HRP without PLL at 150 μg/mL, and (F) trypsinized HRP–PLL at 1.5 μg/mL. Only cells incubated with HRP-conjugated PLL show uptake, in a pattern very similar (i.e., endosomal perinuclear staining) as that with Feridex-conjugated PLL.
Apart from the inhibition of chondrogenesis by Feridex, little is known about the effects that contrast agents exert on cellular function. Most commonly, viability and proliferation (mitochondrial metabolic rate, cell division) are the main parameters used to determine if the contrast agent affects cellular function. No significant toxic effects of USPIOs have been reported [108,123,124,126,132,134] and, in some cases, a viability even superior to that of unlabeled cells has been observed [91]. Nonetheless, the long-term stability of iron oxide compounds, including the potential degeneration of the dextran-based outer sphere, and the metabolic fate of iron leeching out of the core have not been adequately addressed in current studies. However, it is presumed that the released iron atoms are incorporated into the normal iron turnover inside the cell. Iron oxide particles covered in styrene/divinyl benzene [135] are thought to be inert inside cells as the outer sphere cannot be degraded within the cell. Still, the large size of these particles could have a significant impact on cellular processes and needs further study. Gd-based contrast agent are generally well tolerated [2], although there are reports of toxic effects after intracerebroventricular injection [136]. Degradation of the chelate of Gd-based compounds would expose the toxic metal particle to the cell and it is likely that this will lead to abnormal cell function or even cell death. The incorporation of MnCl2 through Ca2+ channels also raises concern about its potential neurotoxicity [137]. However, neurotoxicity appears to be a dose-dependent effect and MnCl2 administration in animal models for cellular tracking is well-tolerated [138]. As Mn2+ can be processed within the cell, instability of the compound and exposure of the free particle to the cell is less of a concern. Clearly, the prolonged exposure of cells to MR contrast agents raises different concerns compared to clinically used agents that are only transiently in the brain. Considerably more focused studies on the long-term cellular effects of contrast agents need to be conducted to address these issues appropriately before widespread translation into clinical applications can be considered.
Novel approaches for magnetic cell labeling that have recently been developed aim to overcome some of the aforementioned issues. These strategies include the use of a biolistic gene gun [139,140] and the use of large, styrene/divinyl benzene-coated magnetic microspheres [141–143] or micron-size particles of iron oxide (MPIOs) [135]. It remains to be seen if the latter iron oxide formulation is clinically applicable, and can be used for labeling of nonphagocytic cells. These large iron oxides have the advantage, however, that a single cell can be detected by MRI [144]. Even with poor labeling efficiency, cells incorporate still enough iron (because of the sheer particle size) for a strong contrast on T2*-sensitive sequences [135], which may even enable the detection of single particles [145]. Further developments that can enhance the in vivo effects of contrast agent-labeled cells or provide a better distinction between USPIO-labeled cells and blood vessels on T2*-weighted scans [146] will contribute to an ever increasing ability to track minute quantities of cells in vivo.
Cellular MRI of Cells following Transplantation or Transfusion
Dysmyelinating Disease and Spinal Cord Injury
Investigation of the autoimmune nature of dysmyelinating disease could greatly benefit from the in vivo tracking of lymphocytes as they induce the disease process. In vitro labeling of mouse lymphocytes, such as T-cells, and their tracking in vivo after injection into mice by MRI [33] enables the early visualization of pathogenesis and evolution whilst potentially allowing a direct study of how interventions interfere with these processes (Figure 8). In contrast, in vivo labeling of lymphocytes will be complicated by the uptake of contrast agents into phagocytic cells that are not thought to be involved in the early pathogenetic events, but rather in the response to inflicted cell loss. A specific uptake mechanism for T-cells [147,148] and inhibitory signal to macrophages would be desirable to allow a distinct and specific in vivo labeling of T-cell and macrophage activity in dysmyelinating disease.
The specific loss of cells, such as oligodendrocytes, which myelinate axons leads to severe motor impairments in patients that are not easily addressed by pharmacological means. Transplantation of cells differentiating, for instance, into oligodendrocytes can reverse the pathology and may provide persistent relief of these impairments. The diffuse nature and widespread migration of cells after transplantation, however, make it difficult to relate cell infiltration/migration and functional recovery. Prelabeling of (stem cell-derived) oligodendrocyte progenitors with MR contrast agents affords the visualization and tracking of the movement of the cells in the spinal cord and the brain [90,102,103,108]. Other cells able to myelinate damaged myelin sheets, such as Schwann cells or olfactory ensheathing cells (OECs), have also been shown to incorporate SPIO particles by using dendrimers [117] or fluid phase pinocytosis while preserving their functional ability to remyelinate damaged fibers in vivo [149].
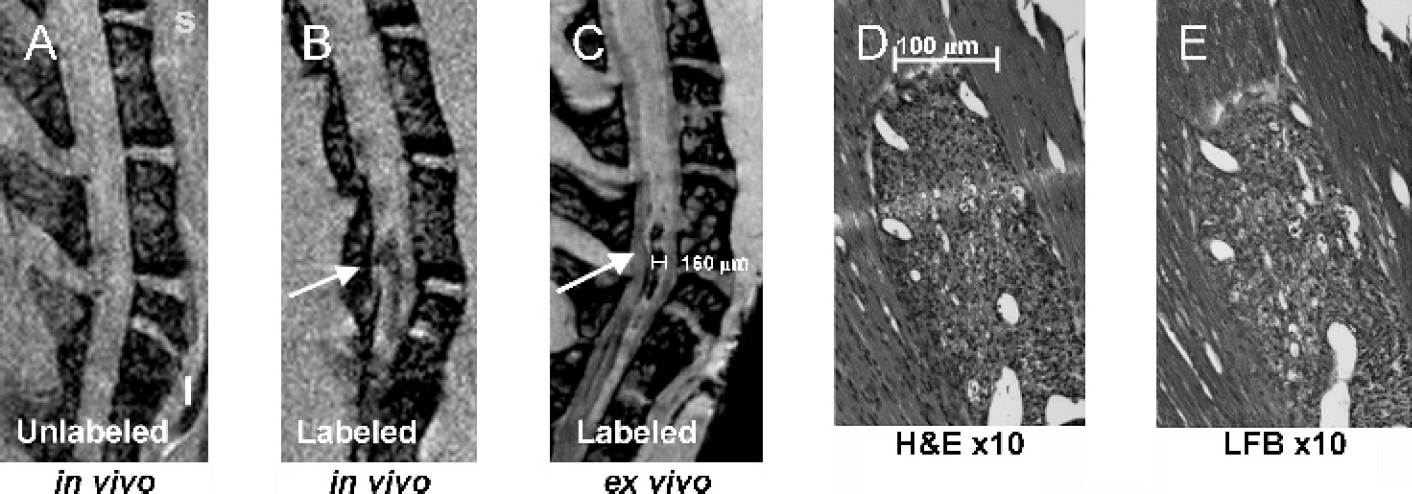
In vivo and ex vivo MRI of spinal cords of mice induced with experimental autoimmune encephalomyelitis (EAE) following adoptive transfer of PLL–Feridex-laheled (B,C) or unlabeled (A) PLP-sensitized T lymphocytes. Hematoxylin/eosin staining (D) and Luxol fast blue staining (E) of the hypointense lesion area in B and C demonstrates co-localization of labeled cells with areas of demyelination. Adapted from Ref. [33].
As with demyelination of the spinal cord, it is also possible to track the migration of transplanted cells in the spinal cord aiming to restore lost function due to transection of the cord. Recently, Lee et al. [117] were able to detect SPIO-labeled OECs for at least 2 months postinjection. OECs migrated in the intact spinal cord but only showed limited migration in the damaged cord. The physical barrier of the transection severely limits the OECs ability to populate both sides of the stump. No behavioral improvement was observed in transplanted animals in comparison to lesioned animals. Transplantation of cells into both sides of the lesion might be needed to increase the functional repair mediated by cell transplantation. It was also observed that in lesioned animals, small hemorrhages resulted in hypointensity on the MR images as well as positive staining for Prussian blue, a histochemical marker of iron often used to indicate the presence of iron oxide particles in transplanted cells. Similar issues arise for transplanted cells in the brain. Further refinement of current methodologies should be adept to overcome these issues.
Neurodegenerative Disorders
Transplantation of fetal cells in neurodegenerative disease is showing promising results in both preclinical and clinical settings. Although in clinical settings cellular therapy for Parkinson's and Huntington's disease is underway, MRI has been mainly used to determine if transplanted cells were responsive to particular pharmacological challenges in preclinical models [150,151] or to ensure the safety of the grafting surgery. However, little research has been devoted to the cellular MRI of transplanted cells in these neurodegenerative disorders. Recently Sandhu et al. [152] reported that uni- or bilateral transplantation of GRID-labeled neural stem cells in a rat model of Huntington's disease allowed visualization of the grafted cells for at least 28 days after injection. Moreover, a quantitative comparison between uni- and bilateral grafts demonstrated that unilateral grafts infiltrated the nontransplanted, but damaged, hemisphere and equally distributed over both hemispheres. The volume of unilateral transplants was maintained at half the volume of the bilateral grafts suggesting that this approach could provide a means to assess graft survival and possibly migration serially and quantitatively in vivo in neurodegenerative disorders.
The potential to assess graft survival and migration in vivo by MRI will help to further refine cell therapy for neurodegenerative disease. In Parkinson's disease, it will be possible to define in preclinical models how many cells need to be transplanted and to survive in relation to dopaminergic cell loss which can be also assessed in vivo by phMRI. Neurodegenerative disease, such as Alzheimer's, which might benefit from a widespread migration and early intervention, will benefit from cellular imaging of transplanted cells by providing a correlation between cell survival, stagnation of neurodegeneration, and cognitive decline.
Cerebral Ischemia
Stroke is the most common cause of adult disability in the Western world. Still to date, therapeutic interventions have met with limited success. Cellular MRI in stroke allows monitoring of the survival and migration of transplanted stem cells aiming to improve behavioral impairments. The prelabeling of cells with iron oxide-based [91,135,139,140] or gadolinium-based [59] contrast agents has allowed different groups to visualize the presence and migration of transplanted stem cells to the site of cerebral ischemia and to brain tumors [153]. Zhang et al. [139,140] calculated that transplanted subventricular zone cells migrate at an average speed of about 65μm/hr to the site of lesion, approximately half the speed of cells migrating along the rostral migratory stream in intact brains. However, to date, it is unclear if or how the speed of migration or infiltration of cells into the site of lesion is related to functional recovery. A common observation in these studies, irrespective of the type of transplanted cells (embryonic stem cell, neural stem cell, or bone marrow stromal cell), is that grafted cells start migration from remote implantation sites within the first two days after transplantation and typically arrive at the site of injury between 10 and 14 days (Figure 9). Some of the above mentioned studies, combined with immunohistochemical staining using cell-specific antibodies, could show a substantial differentiation of the implanted cells into mature neurons and glia in the target zone. The ability to track the pathways along which these cells migrate and to study their dynamic behavior in vivo will in future studies help to address the issue as to how stem cell survival, migration, and/or infiltration promote behavioral recovery and help to guide future cell therapy in patients [154].
Myocardial Infarction
Imaging of transplanted cells after myocardial infarction poses a further technical challenge as the heart is constantly moving and therefore requires a fast image acquisition protocol. Various strategies, such as cardiac gating of image acquisition, have been devised to allow cardiac imaging. However, time constraint and potential movement artifact considerably complicate the image acquisition of implanted cells by MRI.
These technical complications can be overcome and Kraitchman et al. [127], Hill et al. [142], and Garot et al. [155] have demonstrated that implantation of iron-oxide labeled transplanted cells can be detected using clinical MR scanners. Injection of iron-oxide particles alone (without cells) into the area of damage in the heart only results in a transient (<12 hr) signal attenuation on T2-weighted scans; unlabeled control cells do not affect the MR signal in vivo suggesting that the long-term identification of iron oxide-labeled cells is not a mere artifact of the surgical procedure or the incorporation of free iron oxide particles into host cells [156]. The signal attenuation on T2*-weighted images of ischemic myocardium is similar to that of USPIO labeled cells and hence complicates their in vivo detection. However, a comparison of the signal attenuation between the smaller hypointense area on proton density-weighted scans and the larger attenuation on T2*-weighted scans allowed a detection of USPIO-labeled cells [157]. A further strategy to visualize the engraftment and survival of transplanted cells in the damaged myocardium will consist of the infusion of Mn-DPDP which is incorporated into viable cardiocytes via Ca2+ channels [158] and should restore the signal hyperintensity on T1-weighted MR scans.
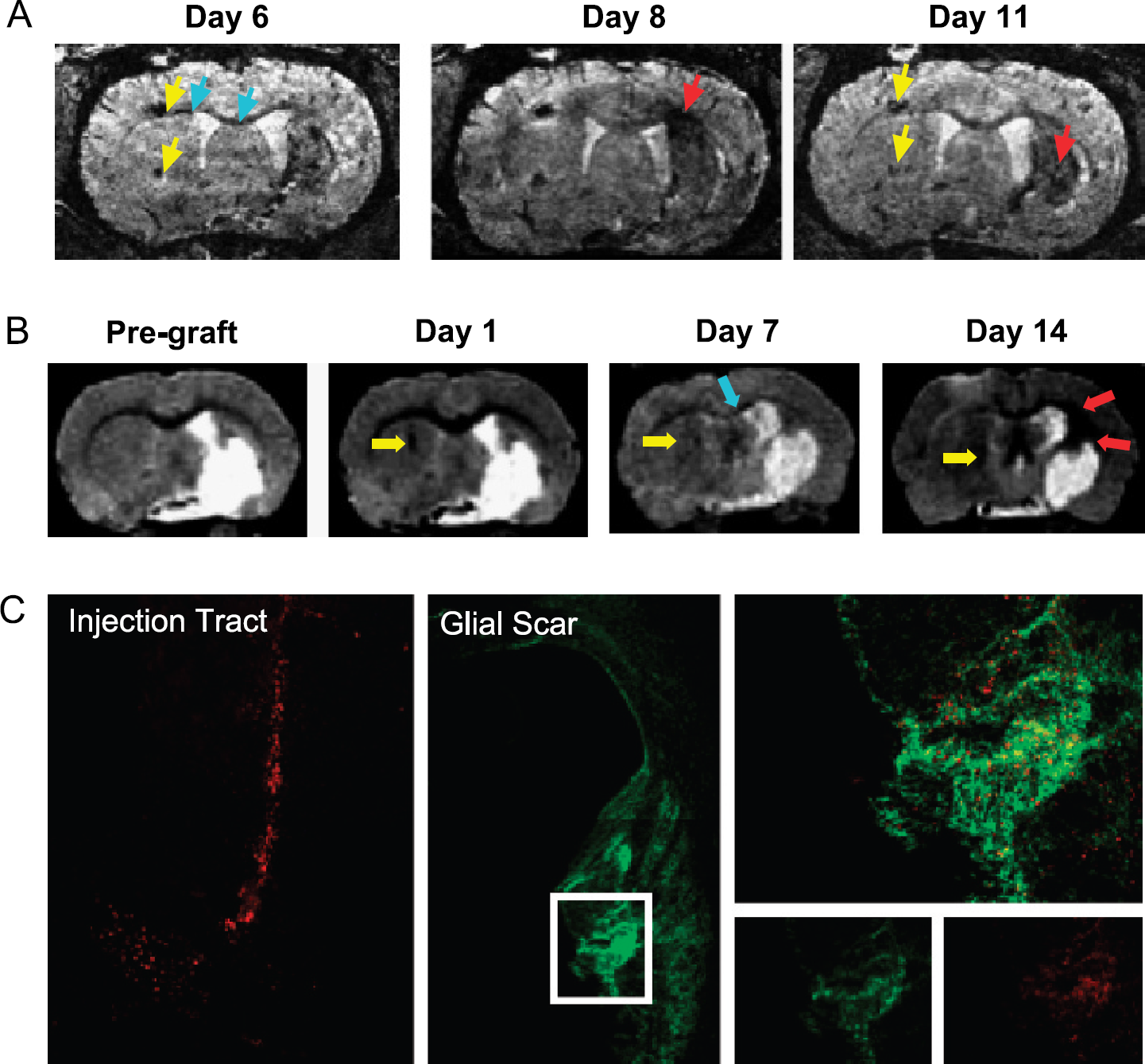
(A) In vivo high-resolution MR images at 78 μm isotropic resolution of a rat brain, using a 3-D gradient-echo imaging sequence, show the migration of implanted stem cells [91]. The yellow arrowheads indicate the primary implantation sites of the embryonic stem cells. As early as 6 days after implantation, cell groups migrate (blue arrows) across the corpus callosum towards the contralateral stroke region. After two more days, the cells can be observed spreading, as a diffuse dark area, into the borderzone of the lesioned area. On Day 11 postimplantation, the embryonic stem cells have accumulated throughout the entire striatum (red arrows). The primary implantation sites are by now clearly depopulated, as can be seen from the diminishing contrast. (B) In a similar fashion, GRID-labeled stem cells' transcallosal migration can be followed serially towards a chronic ischemic lesion. MR1 allows a visualization of the stroke damage prior to the implantation of stem cells providing a baseline image against which subsequent signal changes due to GRID-labeled cells can be evaluated. The potential to serially assess the same animal over time can provide a dynamic assessment of the migration of transplanted stem cells from their injection tract (yellow arrow, Day 1), along the corpus callosum (blue arrow, Day 7), to populating the peri-infarct region (red arrows, Day 14). (C) The bimodal nature of GRID subsequently allows the corroboration of these observations by direct detection of transplanted cells by means of the red fluorescence of the label. It can be observed that at 14 days the injection tract was indeed still filled with GRID-labeled stem cells corresponding to the MR image and that transplanted cells (see insert) indeed infiltrated the peri-infarct area participating in the glial scar (in green GFAP-positive cells) [59]. The serial investigation of stem cell migration in damaged brains and the subsequent corroboration by histology can greatly increase our understanding of how the migration of neural stem cells relates to functional repair after stroke damage.
Prelabeling of endothelial progenitor cells with CD34-coated magnetic beads also allows their tracking in vivo and provides a mean to study the revascularization of areas of damage which ensues repopulation by cardiocytes and could provide surrogate markers of treatment efficacy without being directly related to graft survival [159]. Although transplantation of stem cells for cardiovascular disease is progressing rapidly to clinical applications, at present, the common use of implanted devices, such as pacemakers, in this patient group still remains largely prohibitive for MRI [160]. However, significant advances have been achieved in designing catheters that allow MRI-guided delivery and monitoring of transplanted cells. Clinical translation of these approaches are on the horizon and promise to revolutionize the treatment of patients with heart attacks [160].
Aside from imaging of migration or trafficking, the ability of cellular MRI to verify accurate cell delivery in the targeted organ (region) is likely to become very important in itself. This is the case for MSC delivery in myocardial infarct, as well as MSC delivery to organs (liver, kidney) through their feeding arteries [161,162]. Systemic administration of MSCs through intravenous administration would be highly desirable as it is less invasive. However, the number of MSCs that home to injured areas (e.g., the heart or kidney), may be below the threshold of detection for in vivo MRI, as many cells become initially entrapped in the liver [163].
Hematopoietic and Immune Cell Trafficking
Most interest to date in the use of prelabeled cells for cellular MRI has been in the use of stem cells, but the ability to monitor immune cell trafficking is likely to have a significant future impact on our understanding of the in vivo immune response. In addition to the earlier discussed study by Anderson et al. [33] on trafficking of myelin-sensitized T cells into the spinal cord of animals with EAE, antigen-specific cytotoxic T cells have been found to specifically home to OVA+ [164] and HER2/neu+ [165] tumors. As cellular MRI allows serial monitoring of the same animal/tumor over time, the biodynamics (tissue site of cell infiltration vs. time) can be determined in a robust manner. SPIO-labeled hematopoietic bone marrow cells have a different distribution pattern as compared to free SPIO particles. In contrast to free particles, cells were found to rapidly home to bone marrow following intravenous injection [166].
Comparison of Cellular MRI with Other Noninvasive Techniques
The high-spatial resolution and the potential to repeatedly image the same subject over time provide the appeal of MRI to visualize the survival, migration, and integration of cells in vivo. Nonetheless, other techniques also possess the potential for cellular imaging in the living organism. Especially studies on hematopoietic stem cell transplantation have used fluorescent dyes, such as PKH26, to prelabel cells in vitro to allow in vivo detection of transplanted/infused cells by means of optical imaging [167–169]. However, this approach does not allow deep tissue penetration, and therefore, is limited to easily accessible targets. Alternatively, invasive procedures, such as a craniectomy, are required to allow access of the imaging probe to the anatomical structure of interest [167]. Exposure to light to visualize the fluorescent dyes can also cause phototoxicity [170], killing the labeled cells complicating the serial assessment of these cells by optical imaging methods. The attractiveness of this approach lies in the commercial availability of fluorescent dyes for cellular tacking that are widely used on a variety of cell types and provides a powerful approach to study, for instance, cellular differentiation by combining the fluorescent marker for the identification of transplanted cells with immunohistochemical methods.
A very promising new development in cellular imaging consists of the possibility to detect bioluminescence from deep tissue structures [171]. Typically, a gene encoding for luciferase is engineered into cells [172]. The substrate luciferin (supplied through intravenous injection) is oxidized by luciferase in the presence of ATP and oxygen, producing a photon in the course of this bioluminescent reaction that can be detected by very sensitive cameras, thus allowing repeated in vivo cellular imaging [173]. A drawback of bioluminescent imaging is that the scattering of light by tissue limits the spatial resolution of this technique considerably compared to MRI. In addition, clinical or whole-body large-animal imaging is, at present, not feasible, and pigmented strains are less suitable for bioluminescent imaging. Still, bioluminescent imaging has allowed Kim et al. [174] to track the in vivo migration of neural stem cells from the contralateral hemisphere to the site of lesion in mice with middle cerebral artery occlusion while allowing a corroboration of these results by histology. Migration of neural stem cells into gliomas across hemispheres has also been visualized [175], and the trafficking of immune cells has been monitored as well [176,177]. The potential to track a particular type of cell, such as a neural stem cell or immune cell, by bioluminescence, while using the same techniques to detect light, emitted at a different wavelength from distinct reporter genes to study gene expression in these cells clearly will provide interesting novel avenues to study the basic biology of transplanted cells by combining molecular (gene expression) and cellular imaging (location of cells).
Various other techniques, such as positron emission tomography (PET) and single photon emission tomography (SPECT), also have the potential to visualize cells in vivo. The current spatial resolution of both PET and SPECT is at least 10 times lower than that observed with MRI, but it is the higher specificity of radioligands that conveys the attractiveness of these techniques to study transplanted cells in vivo [178]. For instance, Jacobs et al. [179] have shown that engineering of reporter genes into cells in vitro allows the identification of these cells by means of detection of the transgene expression in vivo. Hence, the reporter molecules could serve as a surrogate marker of the cells to be imaged and no prior loading of the cells with a contrast agent would be required as radioligands are small enough to cross the intact BBB. Differential expression of proteins/receptors, such as the transferrin receptor, on particular cells can also serve as a marker to identify these transplanted cells in vivo [180,181]. Although both PET and SPECT provide excellent approaches to study specific molecular targets and determine cellular differentiation or metabolic consumption, the repeated assessment of animals with radioligands raises issues regarding the safety of this approach for longitudinal studies and its translation into a clinical setting. The prelabeling of cells in vitro with radioligands prior to in vivo imaging (as exemplified by MR cellular imaging) is also less likely to be apposite for the longitudinal tracking of cells due to the short half-life of radioligands. Nonetheless, Chin et al. [183] have demonstrated that it is possible to visualize MSCs by SPECT over a period of 14 days after preincubation of cells with 111In oxine. It is also unclear if the persistent presence of radioligands in these immature cells could have any deleterious effects on their genetic make-up while proliferating. The use of specific reporter markers in transplanted cells detectable by repeated administration of a radioligand could provide a very attractive approach to monitor graft survival and function over extended time periods spanning over years without much concern about a lingering agent within administered immune [183,184] or stem cells [185].
The use of noninvasive imaging technology based on reporter genes, such as bioluminescence or PET imaging, can greatly enhance our understanding of cellular behavior in vivo. These techniques have the advantages that one can follow rapidly dividing cells for prolonged times, as well as the ability to determine cell survival. However, the need for on-site synthesis of radioligands severely limits the wider applicability and practicality of PET and SPECT as cellular imaging techniques, whereas limits on tissue penetration and need for genetic engineering will complicate translation of optical imaging and bioluminescence into clinical applications. With MRI, it is impossible to differentiate live from dead cells, and label dilution by cell division is a significant limitation. Nonetheless, the wider availability of MRI scanners in both clinical and preclinical settings further strengthens the applicability of cellular MRI as a translational technique that can be easily implemented in many centers. Given the high-resolution of MRI at near-cellular levels, it is the unique complementary information that can be derived from the different imaging approaches that highlight their importance in cellular imaging.
Clinical Issues Addressed by Cellular MRI
Many diseases are characterized by specific cellular loss or the infiltration of immune cells. Cellular MRI offers the opportunity to visualize the cellular aspects of pathology noninvasively in patients. The ability to track the presence or lack of these cells can therefore provide additional information that can be factored into clinical decision making as to the diagnosis or treatment of the patient. For instance, the presence of a large fraction of undifferentiated dividing cells within a tumor can indicate the malignancy of the neoplasia and help clinicians to decide on the best course of action. Currently, structural MRI does not provide this finesse and only visualizes gross morphological changes that are mostly downstream consequences of cellular change. Clinical studies using USPIO to target macrophages in vivo already provide evidence of the significance that this approach can yield to patient care. The introduction of anti-inflammatory agents in stroke or multiple sclerosis can be guided by the visualization of macrophage infiltration into the CNS [34], whereas the immunosuppressive regime can be adjusted to counteract graft rejection that can be monitored based on the presence of macrophages within grafted tissue/organs [50]. As contrast agents increase in specificity, further clinical application will emerge that will center cellular imaging as a core analytical technology for diagnosis and follow-up of patients.
For cellular MRI using prelabeled cells following adoptive transfer, verification of correct delivery in the target organ will no doubt hold great value for the practicing radiologist. A first clinical study using magnetically labeled dendritic cells has now started in Holland [186], and one of the major findings was that ultrasound-guided targeted delivery of therapeutic dendritic cells failed in 50% of the patients, despite the procedure being performed by experienced radiologists. Until the development of cellular MRI, there was no other way to assess the efficacy of delivery. Accurate delivery will determine failure or success of cell therapy and may explain the large variation of treatment outcome in patients receiving autologous dendritic cell vaccines. A similar clinical role of cellular MR imaging is likely to become important for monitoring targeted stem cell injection into the infarcted myocardium.
Conclusions and Perspectives
Cellular MRI has now found broad applications in biology and medicine. The approach of prelabeling of cells followed by longitudinal in vivo imaging investigations has now started in patients [186], and the various potential applications are emerging rapidly. For the further clinical development of this young field, both short- and long-term toxicity studies are warranted. In particular, comprehensive studies on the fate of the contrast agents in vivo following their metabolism have to be conducted to determine any potential adverse effects and potential remedies before further clinical trials can be envisaged. The in vivo targeting of cells for cellular (macrophage) imaging has now been well established in experimental paradigms and has rapidly found its translation and relevance in clinical applications. Cellular MRI has an increasing appeal as it allows the serial in vivo study of dynamic processes and verification of accurate cell delivery which previously were largely inaccessible.
Footnotes
Acknowledgments
The authors are supported by grants RO1 NS045062 (JWMB), Multiple Sclerosis Society PP0922 (JWMB), UK Medical Research Council ROPA Grant (MM), UK Department of Trade and Industry (MM), and Competence Network Stroke (BMBF/ projects B1 and B5) (MH).
