Abstract
Effective development of therapeutics that target the molecular basis of disease is dependent on testing new therapeutic moieties and delivery strategies in animal models of human disease. Accelerating the analyses of these models and improving their predictive value through whole animal imaging methods, which provide data in real time and are sensitive to the subtle changes, are crucial for rapid advancement of these approaches. Modalities based on optics are rapid, sensitive, and accessible methods for in vivo analyses with relatively low instrumentation costs. In vivo bioluminescent imaging (BLI) is one of these optically based imaging methods that enable rapid in vivo analyses of a variety of cellular and molecular events with extreme sensitivity. BLI is based on the use of light-emitting enzymes as internal biological light sources that can be detected externally as biological indicators. BLI has been used to test spatio-temporal expression patterns of both target and therapeutic genes in living laboratory animals where the contextual influences of whole biological systems are preserved. BLI has also been used to analyze gene delivery, immune cell therapies, and the in vivo efficacy of inhibitory RNAs. New tools for BLI are being developed that will offer greater flexibility in detection and analyses. BLI can be used to accelerate the evaluation of experimental therapeutic strategies and whole body imaging offers the opportunity of revealing the effects of novel approaches on key steps in disease processes.
In Vivo Bioluminescent Imaging
In vivo bioluminescent imaging (BLI) has been used as a noninvasive means of tracking pathogens or tumor cells in animal subjects early in the disease process [1]–[8], and to develop new animal models that incorporate reporter genes into the rodent genome as markers of transcription that reveal developmental changes or response to various stimuli [9]–[11]. Such transgenic strategies have permitted real-time assessment of a wide variety of biological events noninvasively in living animals, and have enabled relatively high throughput in vivo screens that are likely more predictive of the human response to therapy than traditional analyses of animal models that require excision of target tissues. BLI is an optical imaging modality that is based on the ability of light to penetrate tissues, although limited by absorbance and scattering, and incorporates the use of genes encoding luciferases as internal biological sources of light. BLI is an ideal imaging modality for understanding the parameters that govern gene delivery and effective expression, or inhibition of expression in model systems, and can be used to accelerate the development of therapies that are based on the delivery of biomolecules to target tissues.
The luciferase reaction has been exploited for decades as the basis of in vitro assays that are sensitive and have a broad dynamic range [12]–[16]. Firefly luciferase has been used, for example, to assay the levels of ATP with extreme sensitivity. Compared to other techniques, such as colorimetric or spectrophotometric measurement, bioluminescent indicators offer the advantages of high sensitivity and wide dynamic range. Developments in low light detection technologies have made it possible to measure the low levels of light that escape the scattering and absorbing environment of the living body, and have thus enabled the use of luciferase reactions as in vivo indicators of biological events. The bioluminescent proteins that have been described to date emit light that appears blue to yellow-green in color with emission spectra that peak at wavelengths of 490 to 620 nm [17]. Signal attenuation by the absorbing and scattering properties of live mammalian tissues is significant for wavelengths under 600 nm [18,19]. It would be assumed, therefore, that the light from luciferase reactions would not be detectable through more that a few millimeters of mammalian tissue. However, the emission spectra of most luciferases are broad and contain a significant red component (above 600 nm). For instance, 30% of the emission spectra of luciferases from fireflies (Photinus spp.), the most widely used bioluminescent proteins, is above 600 nm.
Luciferase from the North American firefly (P. pyralis) is the most commonly used bioluminescent reporter in biomedical research, and is also the most extensively studied bioluminescent reaction [12,13,19]–[28]. The fundamental biochemistry of light production by luciferases is the oxidation of a substrate, which produces, as a decay product of the reaction, an almost heatless light. The enzyme substrates and the structures of the luciferase enzymes vary significantly, and in the organisms of origin the mechanisms controlling the speed and intensity of the luminescence are regulated in a variety of ways. Co-factors, such as the nucleotide ATP or other sources of energy, are required for the conversion of the substrate and light production [21,23,24,28]–[30]. Modifications to the enzymes have been made to optimize their use as reporter genes in mammalian cells, and these modifications have eliminated many of the regulatory mechanisms that occur in insects and other species of origin [20,31]. These modifications include optimization of codon usage, removal of peroxisome targeting sites, and elimination of transcription factor binding sites that may interfere with the desired regulation in transcriptional reporter designs.
Imaging modalities in the optical regime have the advantages of being sensitive, and rapid as well as accessible to biomedical laboratories, due to relatively low instrumentation costs. This has led to the development of a number of optical imaging approaches for in vivo analyses [5,8,32]–[47]. Of these, some are particularly well suited for evaluating novel intervention strategies in small laboratory animals as in vivo data can be obtained in nearly real time, and the reporter genes can easily be built into vectors, integrated into mammalian genomes, and transferred into organs and tissues [3,5,10,19]. Visible red and near-infrared (NIR) light can penetrate relatively deeply into tissues and are quantifiable under certain circumstances. Background signals can be minimal, if not completely absent, as there are few, if any, sources of light in mammals especially in the NIR wavelengths. The low background can result in extraordinary signal-to-noise ratios, leading to the extreme sensitivity of these in vivo assays.
Integration of Approaches: Connecting Correlative Cell Culture Assays to Animal Models
Reporter genes that can be detected in correlative cell culture assays and in vivo offer the significant advantage of linking cell culture assays and in vivo imaging. The luciferase reaction can be monitored in cell lysates in living cells, in living organs, and in whole animals. Therefore, bioluminescent reporters allow for an integrated set of assays in which the same reporter can be used in cell culture correlates of biological processes, and then in vivo to test predictions made in cultured cells. Such integration of bioassays results in more predictable animal models where the data are highly correlative, and established cell culture assays can be validated in living animals. Integration of culture and in vivo assays was demonstrated in a study by Scheffold et al. [48] where cytotoxicity assays were modified to use luciferase labeled target cells to determine the sensitivity of a tumor target to the taumoricidal activity of an enriched population of T cells with natural killer cell makers that were redirected to the tumor cells using a bispecific antibody. The cytotoxicity assay, using the same luciferase labeled target cells, antibody and lymphocyte population, was then performed in vivo revealing the cytotoxic effects of this combination therapy.
The complexity of this therapeutic strategy, involving engineered antibodies and cytotoxic T cells, required a fairly involved animal study with a number of treatment arms including cellular and acellular controls. The experimental therapy was compared to Herceptin (a therapeutic antibody that recognizes the same tumor target as the bispecific antibody), and to four other controls including two T-cell controls and two antibody controls. The use of in vivo imaging as a real-time readout enabled repeated analyses of each animal in the various treatment groups. As the signal in each animal at each time point could be compared to previous time points (i.e., internally controlled data), the statistical significance of each treatment group could be improved. The data from this study demonstrated that development and testing of complex treatment regimens can be greatly accelerated using imaging, and that imaging can provide more robust datasets than can more traditional assays of tumor burden [48].
Imaging in experimental animal models will undoubtedly have an increasing role in the development of novel therapies, and will be key to the successful development of complex therapies that involve immune cells, radiation, DNA-based therapies or chemotherapies as combination therapies or as complementary treatments. The optimization of single-agent treatment regimens can be greatly accelerated using imaging assays to validate the therapeutic target and to reveal the success of delivery and efficacy. Given the complexity of multistep and multiagent therapeutic strategies, the use imaging to assess outcome and determine the mechanisms of action will have a considerable impact on the preclinical stages of development.
In the transition from cell culture assays to animal models, conservative use of laboratory animals and optimization of the amount of data that can be obtained from a given animal study will be dramatically affected by the incorporation of imaging methods into animal protocols [49]. Imaging technologies, such as BLI, which are rapid and accessible, can increase the number of quantitative data points that can be collected for a given animal in a study and improve the statistical significance. By imaging the whole intact live animal, all organs can be studied simultaneously and the contextual influences of intact organ systems can be retained. Taken together, these advantages create more predictive animal models since imaging of animal models offers the benefits of longitudinal study designs and internal experimental controls. At the termination of the study, the image data can be used to further improve animal studies by guiding appropriate endpoint tissue sampling for histology or biochemical analyses.
Autoreactive Immune Cells as the Trojan Horse: Local Delivery of Therapies Using Immune Cell Homing
The migration of immune cell populations is key to an organism mounting an effective immune response, and understanding these patterns as they occur in the context of living rodent models has recently been the subject of considerable investigation. Intravital fluorescence microscopy has been used to visualize cellular interactions and migration in real time as they occur in tissues that have been exposed for interrogation [50]–[52]. However, intravital microscopy methods are limited to fairly superficial tissue sites (several millimeters), and are subject to motion artifacts due to respiration and cardiac function [53]. Therefore, the set of microscopic techniques used for intravital assays can reveal the local dynamics of immune cells in exposed lymph nodes, but it is not suitable for investigating a variety of different tissues and cannot be used for whole body analyses The utility of intravital microscopy is in the analyses of cell-cell interactions and cellular movement within a given tissue. In contrast, BLI is one of several macroscopic optical imaging modalities that offer rapid whole body analyses of cell movement, and its noninvasive nature permits relatively long study periods of days or weeks. The microscopic and macroscopic analyses are complementary and studies that incorporate both levels of investigation offer the greatest benefit.
As the migratory pathways of immune cells within the body relate to the temporal changes of the immune response, localization of these cells may also be useful for local delivery of therapies. The dynamic changes in immune cell populations that take place in the context of an intact living mouse have been investigated using BLI, and there are several examples of monitoring immune cell trafficking patterns. Understanding the migration patterns of autoreactive immune cells has been used to optimize the delivery of therapeutic proteins to sites of immune-mediated tissue destruction [4,54,55].
Adoptive immunotherapy, where expression of a proinflammatory antagonist is expressed in an autoreactive immune population, has been monitored in a mouse model for multiple sclerosis in humans, experimental autoimmune encephalomyelitis (EAE), using BLI [4]. The autoantigen-reactive CD4+ T-cell population, specific for myelin basic protein (MBP) was transduced with a retrovirus that encoded both a dual reporter protein, comprised of GFP and luciferase as a fusion, and the 40 kDa monomer of interleukin-12 (IL-12 p40) as a therapeutic protein [4]. The transduced cells could be selected using flow cytometry, and following adoptive transfer, the migratory patterns of these cells were monitored. BLI demonstrated that the effector cells that would typically cause immune-mediated destruction of myelin trafficked to the central nervous system of symptomatic animals, and if the CD4+ T cells expressed the immune modulator, IL-12 p40, there was a reduction in the clinical score for the disease.
This imaging and therapeutic approach was further substantiated in another mouse model of autoimmunity. In an experimental model of rheumatoid arthritis type II, collagen-specific CD4+ T hybridomas and primary CD4+ T cells were used as vehicles for local delivery of IL-12 p40 [55]. The CII-reactive T-cell hybridomas were transduced with the reporter constructs and IL-12 p40 and BLI was used to demonstrate that the transferred cells accumulated in inflamed joints of immunized mice. Transfer of CII-specific T cells or hybridomas expressing IL-12 p40 significantly reduced joint stiffness, while transfer of cells transduced with the vector only as a control had no effect in this model. Moreover, transfer of the T cells specific for MBP that were effective in the EAE model and produced equivalent amounts of IL-12 p40 also had no effect on joint inflammation. Therefore, the migration to, and retention within the target tissue, mediated by the specificity of the T-cell receptor, appeared to be an essential requirement for effective tissue-specific delivery.
Modifying antigen-specific T cells, the effectors of disease, for local expression of immunoregulatory proteins and systemic delivery of these cells, is a promising strategy for treating autoimmune disease and has the potential for treating other diseases. The inherent problems of nonspecific toxicity following systemic administration of inflammatory agonists, for example, cytokines, can be obviated by local delivery to the site of tissue destruction using the migration of specific immune cells to these sites. Revealing the trafficking patterns of immune cells and understanding the mechanisms that control these patterns are key for the effective development of cell-based therapies in general, as well as the enhanced cellular therapies described here. In these two examples, BLI enabled the trafficking studies and accelerated the optimization of protocols for local delivery of immunoregulatory proteins.
Lymphocytes, dendritic cells, and monocytes are highly mobile cells that can travel throughout the body in response to a variety of tissue- and disease-specific signals. The techniques that have traditionally been used to monitor immune cell distribution in tissues, such as flow cytometry and immunofluorescence and microscopy, cannot readily assess the dynamic changes that take place on a whole body scale. Therefore, imaging methods that reveal these global patterns are necessary for a systems approach to development of immune cell therapies. BLI has utility for revealing the migratory pathways of immune cells within rodent models of disease, and can be used to support clinical trials. However, advances in clinical imaging techniques will be necessary to reveal the temporal patterns of immune cell trafficking in human therapeutic studies.
Nucleic Acid-based Therapies
Nucleic acid-based therapies offer the potential of gene replacement, potentiating conventional therapies, or reducing expression of viral gene products, oncogenes, or dominant mutant proteins. Development of therapeutics that are based on gene transfer has been constrained by our inability to rapidly and noninvasively analyze delivery, expression, and efficacy in animal models. A variety of imaging strategies based on several different modalities have been developed for the purpose of advancing novel therapeutic strategies that utilize biomolecular therapies, and each offers significant advantages to the field. The utility of BLI for this purpose has been demonstrated in several different therapeutic areas including oncology.
Effective evaluation of new treatment strategies for malignant disease will require that animal models be utilized that closely resemble the human diseases they are designed to model. Orthotropic and spontaneous tumor models more closely resemble human disease than do subcutaneous xenograft models [56]–[58]. However, these types of approaches are frequently more difficult to study since the lesions arise in more unpredictable patterns, and can be located deep within the body at sites that are not accessible for caliper measurements. Disease in these models can involve multiple organ systems, and occur in the face of an intact immune system. Consequently, these models will contribute to our understanding of complex disease mechanisms, and imaging is essential for assessment of tumor burden at deep tissue sites and access to information pertaining to expression of molecular targets. Moreover, as nucleic acid-based therapies are developed, accessing expression data in vivo will become key to optimizing these therapies.
An excellent example of how imaging can be used to optimize a DNA-based cancer therapy was published by Rehemtulla et al. [8], where the gene encoding yeast cytosine deaminase (CD), an enzyme that converts the nontoxic compound 5-fluorocytosine (5FC) into the drug 5-fluorouracil (5FU), was used as a therapeutic gene in an orthotropic rat glioma model (9L cell line). In this study, BLI was used to determine the levels of gene expression following adenoviral-mediated delivery of the therapeutic gene and diffusion-weighted MRI was used as a surrogate marker of tumor cell killing. Evaluation of this multicomponent anticancer therapy without imaging would have required serial sacrifice of many animals and the use of labor-intensive molecular assays on a more limited set of samples.
The use of two imaging modalities, one to assess delivery and the other to assess efficacy, demonstrates the strength of multimodality imaging in animal models. Such approaches that fully utilize the strengths of imaging will result in more robust analyses of these model systems, as demonstrated in this study, and will significantly accelerate the development of new therapeutic strategies for cancer. In the case of cancer gene therapy, it is useful to know the location and magnitude of expression to evaluate the therapeutic effect. For treatment regimes, such as the one in this study, that require optimal dosing of a DNA-based therapy and a prodrug, noninvasive measurements of delivery and efficacy are extremely helpful in the development of these approaches.
In the current paradigm of target-directed therapies, events that occur early in the disease process represent important targets for intervention, and therefore, development of both models that incorporate key genetic elements of oncogenesis and assays that provide access to information about these targets are essential for evaluating new intervention strategies. Tumor models where malignancies spontaneously develop as a result of introduction of mutations and transgenes into mice have allowed the study of the initial or early events in the initiation of disease [58,59]. For instance, a conditional transgenic mouse model that develops retinoblastoma-dependent sporadic cancer has been studied using firefly luciferase. The reporter gene was incorporated into this model along with the transforming gene so progress of the full disease course could be followed using BLI [11]. The onset of disease, subsequent disease progression, and response to therapy in this tumor model were more readily evaluated using a noninvasive approach than could have been accomplished using assays that require sampling of tissues. The early molecular events represent key targets for the effective treatment of oncogenesis, and coupling spontaneous tumor models with sensitive imaging modalities will facilitate the analysis of the key processes in the initiation and progression of malignancy.
In many inherited human diseases, lifelong expression of the wild-type gene is the therapeutic objective. Many gene delivery strategies result in transient expression and cannot meet this requirement. Successful lifelong expression of a reporter gene was demonstrated in an in utero gene transfer model where the target cells were those lining the peritoneal wall [60]. In this model, a recombinant adeno-associated viral (rAAV) vector carrying a modified luciferase gene was injected into the peritoneal cavity of fetal mice. BLI was used to monitor the reporter gene expression patterns throughout the lifetime of the mice beginning at gestational age 18 days (Figure 1). Effective biodistribution of luciferin is essential for using luciferase as an in vivo reporter and evidence that the substrate, luciferin, can cross the placental barrier was presented in this study. The luminescent signals from the transduced fetuses were apparent while the animals were in utero. After the treated animals were born the signals were apparent from the abdominal region and the signals did not diminish in intensity over the 24-month study period—this is essentially lifelong expression. In utero gene transfer offers the possibility of gene replacement at a time where immune tolerance, to the expressed therapeutic protein, may be induced.
Prior to this study, it had been assumed that intraperitoneal administration of rAAV would result in transduction of cells primarily in the liver. When a secreted serum protein was used as a marker of transduction, high levels of the protein were detectable in the blood suggesting efficient transduction. However, analyses of selected organs did not provide evidence of efficient transduction in these tissues [60]. Since whole body imaging of reporter gene expression using BLI can rapidly reveal expression patterns, the signal from cells lining the peritoneal wall was immediately apparent in the first experiment in this study. The information obtained in this experiment indicated the primary target tissue of this gene transfer protocol, and the spatio-temporal expression patterns of the reporter gene could be used to direct the subsequent study designs and accelerate development of this gene transfer method.
Lifelong persistent expression of the reporter gene following in utero gene transfer via AAV. The luciferase reporter gene was delivered to fetal mice via intraperitoneal injection of rAAV at gestational age 15. At gestational age 18 the fetal signal was detectable through the maternal abdominal wall (A). Four days after birth the luciferase signal is apparent over the abdomen of the newborn pups (B). At 1 month of age the signal is still apparent over the abdominal area (C) and this signal persisted for the lifetime of the mice [60].
Advances in gene targeting will further enhance these approaches in that it may be possible to direct the integration of the therapeutic DNA to regions of the genome where the likelihood of adverse events is diminished [61]–[64]. This will increase the safety of these potentially life-saving therapeutic approaches. As the use of fetal diagnostic tests for genetic diseases increases, the development of strategies for in utero gene replacement will become increasingly important. In the study by Lipshutz et al. [60], a tissue was targeted that would have utility for gene replacement where the defect is in a secreted protein. In this example, local expression of a secreted protein from the therapeutic gene transferred to cells lining the peritoneal wall could have a systemic effect.
Small Interfering RNAs
Many nucleic acid-based therapies have been directed at gene replacement (e.g., for treating genetic loss of function diseases), or gain of function (e.g., delivery of genes encoding prodrug converting enzymes for cancer therapy). However, the treatment of many diseases could benefit from transient control of endogenous gene expression. For this purpose, RNA interference (RNAi) has been investigated in mice using in vivo imaging strategies. RNAi is a conserved surveillance mechanism that was first described in Caenorhabditis elegans and is now being applied to controlling expression in mammalian cells and in rodent models of human disease [65]–[67]. Long double-stranded RNAs are sensed, by cells as a pathologic condition (i.e., infection by RNA viruses where double-stranded RNA is present). These long double-stranded RNAs serve as triggers of cellular defense and are processed into ˜21 nucleotide duplexes that act as guides for the destruction of RNAs that share homology to the trigger sequence (for a review, see Denli and Hannon [68]). Small interfering RNAs (siRNAs) mimic the 21 nucleotide intermediates in the RNAi pathway and can silence genes in somatic cells without activating nonspecific suppression by double-stranded RNA-dependent protein kinase [69], and is therefore a pathway separate from the one mediated by interferons. RNAi presents a new tool for controlling gene expression and has recently been demonstrated, in a rodent model, to inhibit replication of hepatitis B virus in vivo [70].
BLI has been used to demonstrate that reporter transgene expression could be suppressed significantly by synthetic siRNAs and by small-hairpin RNAs (shRNAs) that had been transcribed in vivo from DNA templates [71]. In this study, luciferase mRNA or a sequence from hepatitis C virus (HCV) was the target of RNAi, and a modified hydrodynamic transfection method [72] was employed to deliver naked siRNAs to cells in the liver of adult mice. An siRNA with homology to the firefly luciferase gene, or an unrelated siRNA, was co-injected with a luciferase-expression plasmid, and luciferase expression was assayed using BLI. In each experiment, serum measurements of a co-injected human [alpha]-1 antitrypsin (hAAT) plasmid [73] served to normalize transfection efficiency and to monitor nonspecific translational inhibition. Specific, siRNA-mediated inhibition of luciferase expression was demonstrated in adult mice (p < .0115) compared to no change in expression following delivery of unrelated siRNAs (Figure 2). On average, treatment with luciferase-specific siRNAs resulted in an 81% (±2.2%) reduction in bioluminescent signal.
Subsequently, RNAi directed against a region of the HCV genome was tested. Targeting of the virus itself in mice is not practical because this human pathogen does not replicate in mouse cells. Therefore, the NS5B region (nonstructural protein 5B, viral-polymerase-encoding region) of this virus was fused with the luciferase coding sequence and the fused mRNA served as the target sequence. siRNAs were again co-transfected with this construct in vivo, and luciferase expression was monitored via BLI. An siRNA targeting the NS5B region reduced luciferase expression from the chimeric HCV NS5B protein-luciferase fusion by 75%, suggesting that RNAi may serve as a basis of therapies for HCV and other viral pathogens. However, siRNA must be synthesized in vitro, as they cannot be expressed in vivo; this may severely limit their utility as therapeutics.
Unlike siRNAs, functional shRNAs can be expressed in vivo from DNA templates using RNA polymerase III promoters (discussed by Tuschl [74]). These small RNAs were at least as effective as siRNAs in suppressing gene expression in vivo [71,72]. Expression of a cognate shRNA (pShh1-Ff1) inhibited luciferase expression by up to 98% in three independent experiments and the empty shRNA-expression vector had no detectable effect. These data indicated that plasmid-encoded shRNAs could induce a potent and specific RNAi response in adult mice. RNAi may eventually complement gain-of-function approaches by enabling silencing of disease-related genes with DNA constructs that direct the expression of shRNAs. The full potential of this strategy remains to be determined and imaging will undoubtedly play a role in resolving key issues in the development of RNAi as a therapeutic strategy. Parameters of this potential therapy that can be evaluated by imaging include the evaluation of delivery strategies, construct design, and persistence of gene silencing.

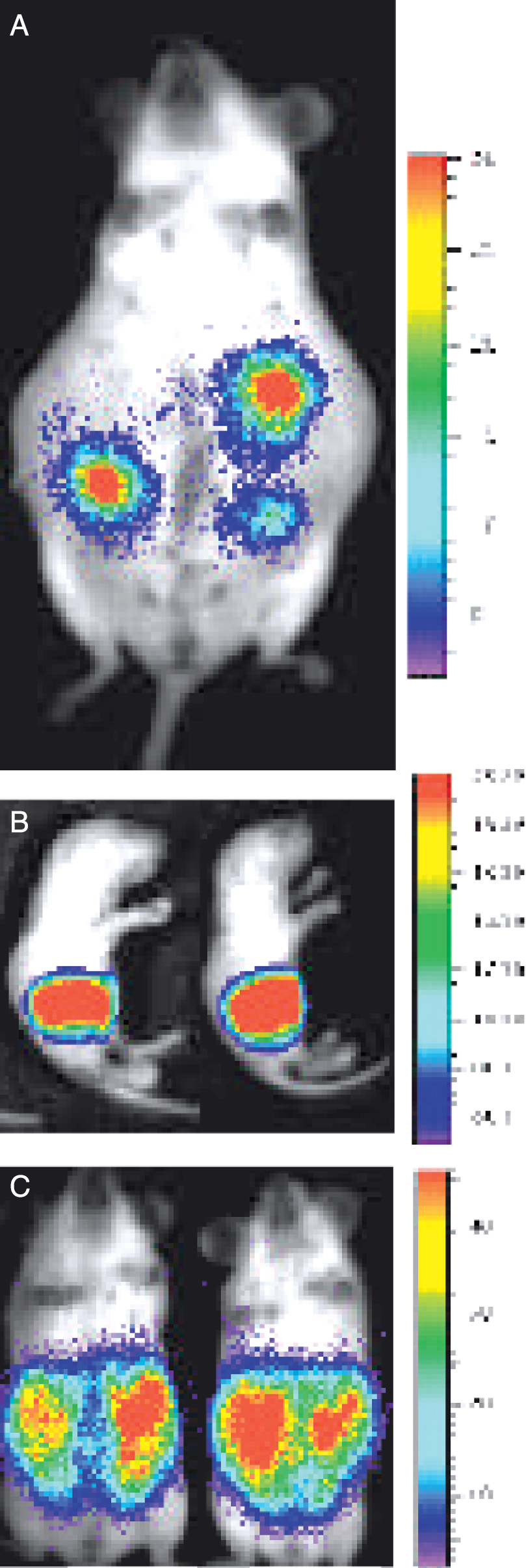
Reduction of luciferase signal in after treatment of mice with siRNAs that target luciferase message. Significant signal was detected from mice that were hydrodynamically transfected with 2 μg of luciferase plasmid pGL3-control (Promega) and 20 μl of RNasin (Promega) in a final volume of 1.8 mL of phosphate-buffered saline (A). Reduction in signal intensity was apparent in mice that received 40 μg of luciferase siRNA (Dharmacon, Lafayette, CO) together with the reporter gene construct and RNasin (B). These images were obtained at 72 hr after transfer of nucleic acid, For imaging mice were anesthetized and given 3 mg of luciferin intraperitoneally 15 min prior to imaging. These results demonstrated that RNAi was able to reduce expression in a living mammal [70].
Emerging Reporter Gene Strategies for BLI
BLI has largely been used as a marker of transcriptional regulation or, when expressed constitutively, as a marker of tumor burden or cell migration. Recently, there have been several developments, in the use of luciferase as a reporter protein, that may increase the number of functional assays that can be performed in vivo using BLI. These include use of dual-function reporters that are fluorescent and bioluminescent, modifications to the luciferase gene that lead to the expression of sensor proteins, and modification that lead spectral changes that can be used for improved detection or added function. To date, many of these modifications that show promise have not been used to directly test biomolecular therapies. Nonetheless, review of these modifications is warranted given their potential for improving in vivo assays.
Dual Reporter Strategies
Several groups have used gene fusions and bicistronic mRNA strategies to create dual-function reporter genes that have enhanced utility over singe reporter approaches. Gene fusions were first used in a study by Day et al. [75] to evaluate gene expression patterns in fruit flies. In this study, a reporter gene fusion comprised of coding sequences from firefly luciferase and GFP was used to assess expression levels by fluorescence microscopy and photometric assays. This approach was used in the BLI studies of immune cell trafficking (above) and has been used to link imaging modalities by using reporter genes that can be detected by BLI and by PET imaging [76,77], or by BLI and fluorescent microscopy and flow cytometry [4,55,78]–[81].
Luciferases other than that from the North American firefly have been evaluated as reporter genes with potential in vivo utility. The luciferase from the sea pansy (Renilla reniformis) is a blue emitting luciferase that uses the substrate coelenterazine [82]. This enzyme derives the energy for the reaction from the substrate and does not require ATP [82]. Development of bioluminescent proteins that use alternate substrates as in vivo reporters may enable multiplexing of the in vivo assays as the reporters can be distinguished biochemically, and the ability to monitor two, or more, biological processes in a single animal would greatly increase the utility of BLI. Renilla luciferase is one of many coelenterazine-utilizing luciferases that may offer the opportunity for multiparameter analyses in BLI. Distinguishing bioluminescent reporters spectrally is, at present, not possible since the current bioluminescent reporters emit light that is largely absorbed by mammalian tissues, and transmitted light from any of these reporters is due to the red components of the emission spectra.
The luciferases from Renilla and firefly utilize two different substrates and thus this pair of enzymes is, in one sense, ideal for dual detection. In cell culture assays, this reporter pair has become a powerful tool for measuring transcriptional fluctuations relative to stable gene expression [83] and kits for such assays are commercially available (Promega, Madison, WI). Successful transition from the cell culture assays to in vivo applications will depend on the biodistribution of coelenterazine. Biodistribution of luciferin has been analyzed and shown to be appropriate for in vivo use of firefly enzymes [9], however, limited biodistribution of coelenterazine may significantly limit the detectability of Renilla luciferase expression in vivo. Renilla luciferase has been used in vivo and expression has been detected using several different low light imaging systems [84]. The patterns of expression were variable and less predicable than those from the firefly enzyme. Moreover, tissues largely absorb the blue emission of this enzyme and the sensitivity of detection will be less than that of the firefly enzyme, assuming similar expression levels and specific enzymatic activities.
Renilla luciferase has been used as a fusion protein with glial-derived neurotropic factor (GDNF), and has been studied in a glial cell line transplanted into the central nervous system of rats [85]. In this study, the levels of the GDNF reporter fusion could be measured optically in the cerebrospinal fluid, but was not evaluated by whole body imaging. Although expression levels were not assayed in vivo in this study, the report demonstrated that Renilla luciferase could be functionally fused to other open reading frames and used to assay expression levels. The ability to image emission from the Renilla luciferase in vivo has been demonstrated in vivo [84,86], and noninvasive detection of both the firefly and Renilla luciferase in the same living mouse has been reported [84]. Addressing the biodistribution issues for coelenterazine, by testing various analogs or by blocking p-glycoprotein activity and by generating coelenterazine-utilizing enzymes that emit at longer wavelengths, may result in effective development of this luciferase as reporter with utility in in vivo assays. Fusions between luciferases and fluorescent proteins have also been evaluated with the intent of using the resonance energy transfer from luciferase to GFP—converting blue emission to green—as a tool to assess the proximity of the two proteins [87]–[92].
Towards In Vivo Assays for Protein-Protein Interaction
Bioluminescence resonance energy transfer (BRET), also known as chemiluminescent resonance energy transfer (CRET) and luminescence resonance energy transfer (LRET), is the process of transferring energy from a light-emitting enzyme to a fluorophore with the net result of a color shift to a longer wavelength [88]–[92]. Energy transfer from Renilla luciferase to GFP has been used to study protein-protein interactions in cultured cells using the known association of two proteins [88]. In this study, insulin-like growth factor II (IGF-II) and insulin-like growth factor binding protein 6 (IGFBP-6) were the fusion partners for GFP, and Renilla luciferase, respectively. Here, the association between the two fusion proteins was assessed spectrophotometrically. When the proteins were not associated the blue emission from Renilla luciferase was detected and when the binding regions of the fusion partners brought the reporter genes into close proximity, the green light was emitted. The wavelengths of emission in this assay (in the blue-green region of the spectrum) are in the range that is attenuated in vivo due to absorption by the primary absorbing pigment in the body, hemoglobin. Although, this demonstration of BRET suggests that it may be possible to monitor protein-protein interaction in vivo, at the present wavelengths it will have limited in vivo utility due to attenuation of signal.
Indirect methods of assessing protein-protein interaction have also been described. Components of the well-studied yeast two-hybrid system have been adapted for use in mammalian cells. The fusions, VP16-MyoD and GAL4-ID, have been used to activate transcription of reporter genes in cell culture [93]. These gene fusions have, more recently, been used to activate expression of firefly luciferase with the intent of establishing a BLI assay for protein-protein interaction. In this study, the cell culture assay was reproduced in living mice using firefly luciferase transcription as a reporter for the interaction of VP16-MyoD and GAL4-ID. The yeast two-hybrid system has traditionally been used to screen libraries of fusion partners to identify novel protein-protein interactions in yeast cells. In vivo screening strategies based on this approach in living rodent models may enable identification of protein-protein interactions that only occur in the context of intact organ systems and may lead to development of therapies whose mechanism of action is the disruption of protein-protein associations.
Another approach for the study of protein-protein interaction has been described using the phenomenon of protein-protein splicing that was first described in bacteria [94]–[96]. The initial description of this approach adopted the nomenclature for RNA splicing where the internal regions of proteins that are “spliced out” have been called inteins and the external, or spliced proteins, are referred to as exteins—similar to the introns and exons of RNA splicing. Ozawa et al. [97] described a split luciferase that could be reassembled to form an active protein after directed splicing to create a reporter gene strategy that would interrogate protein-protein interactions in cell culture assays. The approach was to fuse two complementary inteins to two parts of the luciferase such that upon protein-protein interaction, the inteins were excised out of the inactive precursor fusion proteins, and the luciferase exteins ligated into a contiguous luciferase polypeptide with enzymatic activity.
The intein that was used was a split DnaE from Synechocystis sp.; the N- and C-termini of DnaE were fused to the N- and C-terminal fragments of a split luciferase. The crystal structure of luciferase had previously revealed two globular domains joined by a flexible hinge region [23]. The splicing regions were placed in the flexible hinge [97]. The intein components were connected to two well-characterized proteins that was known to interact in mammalian cells, the phosphorylated insulin receptor substrate 1 (IRS-1) and its target N-terminal SH2 domain of PI3-kinase (SH-2). Luciferase in this configuration served as a marker of protein-protein interaction since the folding of the DnaE intein could be activated by insulin through the interaction of the IRS-1 and SH2 domains, in a dose-dependent manner, leading to protein splicing at the luciferase hinge and reconstitution of activity.
The luciferase activity in the reconstituted protein was a small fraction of that observed in the wild-type luciferase, likely a consequence of the amino acid residues left in the hinge region by the splicing event. The flexible polypeptide comprising the hinge region and holding the two globular regions of the enzyme together forms the critical substrate-binding region [21,23,30]. The amino acid remnants of the protein splicing event may significantly compromise the ability of the resulting enzyme to produce light by changing the size of the substrate-binding cleft. There is no particular reason that the splicing sequences could not be placed elsewhere in the protein where amino acid additions would not affect the enzymatic activity. This may improve the activity of the resulting enzyme. The split luciferase strategy was reproduced by Paulmurugan et al. [98] in a cell line that could be used in living animals, and the interaction of the engineered proteins could be measured in living subjects using BLI. This group then extended this approach to Renilla luciferase using selected sites for splicing that would allow reconstitution of the enzyme activity. The interacting proteins, MyoD and ID, were used in this study to draw together the split luciferase [93]. Modification of two different luciferases into sensors of protein-protein interaction suggests that several steps in a given biological process could be monitored noninvasively in living animals.
Signal Amplification Strategies
Many promoters that are of interest as therapeutic targets may not have high levels of expression and would therefore be difficult to study using conventional reporter gene strategies in vivo. The sensitivity of the current detectors requires that the reporter gene be expressed at high levels; for this reason, strong constitutive promoters are usually used. To circumvent this limitation and enable the study of weak promoters, a transcriptional amplification step has been investigated [99]. The relatively weak form of the prostate-specific antigen (PSA) gene was used to drive expression of a transcriptional activator, GAL4 fused to VP16 activation domains, in the LNCap human prostate cell line. A synthetic promoter consisting of a basal promoter and repeats of the GAL4 binding domains was used to drive expression of luciferase. In this configuration, expression of the transcriptional activator from the PSA promoter led to nearly a 50-fold amplification while maintaining the tissue specificity of the PSA promoter. Zhang et al. [100] have used this amplification method in an androgen-responsive DNA-based therapy that was directed against prostate cancer. The advantage of linked cell culture and in vivo assays through the luciferase reporter enabled the rapid optimization and testing of this system.
Summary and Conclusions
The tools of the molecular biologists have previously been limited to studying gene regulation in cells and in cell lysates. The use of reporter genes, with in vivo detectable signatures, has extended these tools to study the spatio-temporal expression patterns of genes that have been transferred, via viral and nonviral means, to tissues and organs. This use of reporter genes in vivo offers the opportunity to evaluate expression in the living body and interrogate the biology of intact cells, noninvasively in living animals where the contextual influences of the intact functional organ systems are retained. The advances in cellular and molecular imaging will allow for more comprehensive analyses of responses to novel biomolecular therapies within intact animals.
The use of multimodality approaches that offer exquisite structural information linked to functional data will improve the localization studies of molecular therapies. Moreover, multiple reporter systems can be used to expose related pathways and enable us to approach questions within the complexity of intact physiology in whole animal models. BLI has been used to generate high-quality spatio-temporal information about expression and gene delivery, and has proven to be amenable to highly specific and even multifactorial genetic experiments. The utility, cost, flexibility, and especially the high-quality predictive data generated from whole animal cellular and molecular imaging define a key advantage for the development of novel biomolecular therapies, and BLI is rapidly becoming a fundamental tool in this paradigm shift.
BLI holds promise for accelerating analyses of biomolecular therapies and offering a rapid means of conducting in vivo analyses that improve the predictive quality of assessment of these approaches. All imaging strategies provide datasets from relevant, intact animal systems, and as such, increase both the efficiency and the effectiveness of designing and testing new approaches to intervention. Imaging requires fewer animals per study and reduces the time and cost associated with the development of novel therapies. Animal-to-animal variation can be eliminated by using the zero time point as an internal control for each animal in a study group, and thus improving the biostatistics in animal modeling. Given the accessibility and speed of BLI, in vivo efficacy, pharmacokinetics, toxicology, and target validation studies can be conducted on a larger number of delivery strategies and constructs than has been possible in the past. BLI offers the potential of applying, in vivo, many of the tools that molecular biologists have used in cells and in tissue lysates over the past several decades. This opens new opportunities for understanding the effects of gene transfer as a therapeutic strategy.
Footnotes
Acknowledgments
This work was supported, in part, by unrestricted gifts from the Mary L. Johnson and Hess Research Funds (CHC) and by the following grants from the NIH: R33CA88303, R24CA92862, and P20CA86312 (CHC), and the Juvenile Diabetes Foundation.
