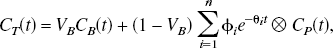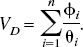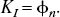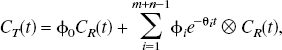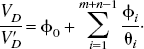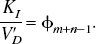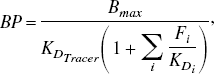Abstract
A kinetic modeling approach for the quantification of in vivo tracer studies with dynamic positron emission tomography (PET) is presented. The approach is based on a general compartmental description of the tracer's fate in vivo and determines a parsimonious model consistent with the measured data. The technique involves the determination of a sparse selection of kinetic basis functions from an overcomplete dictionary using the method of basis pursuit denoising. This enables the characterization of the systems impulse response function from which values of the systems macro parameters can be estimated. These parameter estimates can be obtained from a region of interest analysis or as parametric images from a voxel-based analysis. In addition, model order estimates are returned that correspond to the number of compartments in the estimated compartmental model. Validation studies evaluate the methods performance against two preexisting data led techniques, namely, graphical analysis and spectral analysis. Application of this technique to measured PET data is demonstrated using [11C]diprenorphine (opiate receptor) and [11C]WAY-100635 (5-HT1A receptor). Although the method is presented in the context of PET neuroreceptor binding studies, it has general applicability to the quantification of PET/SPECT radiotracer studies in neurology, oncology, and cardiology.
Keywords
The development of positron emission tomography (PET) over the last 2 decades has provided neuroscientists with a unique tool for investigating the neurochemistry of the human brain in vivo. The ever-increasing library of radiolabeled tracers allows for imaging of a range of biochemical, physiologic and pharmacological processes. Each radiotracer has its own distinct behavior in vivo, and their characterization is an essential component for the development of new imaging techniques and their translation into clinical applications. Estimation of quantitative biologic images from the rich four-dimensional (4D) spatiotemporal data sets requires the application of appropriate tomographic reconstruction and tracer kinetic modeling techniques. The latter are used to estimate biologic parameters by fitting a mathematical model to the time–activity curve (TAC) of a region of interest or voxel. Calculations at the voxel level produce parametric images but are associated with an increase in noise for the TACs. Analysis strategies must therefore be robust to noise, yet fast enough to be practical.
There is a range of quantitative PET tracer kinetic modeling techniques that return biologically based parameter estimates. These techniques may be broadly divided into model-driven methods (Kety, 1951; Sokoloff et al., 1977; Phelps et al., 1979; Mintun et al., 1984; Huang and Phelps, 1986; Gunn et al., 2001) and data-driven methods (Gjedde, 1982; Patlak et al., 1983; Patlak and Blasberg, 1985; Logan et al., 1990; Logan et al., 1996; Cunningham and Jones, 1993). The clear distinction is that the data-driven methods require no a priori decision about the most appropriate model structure. Instead, this information is obtained directly from the kinetic data.
Model-driven methods use a particular compartmental structure to describe the behavior of the tracer and allow for an estimation of either micro or macro system parameters. Well-established compartmental models in PET (Fig. 1) include those used for the quantification of blood flow (Kety, 1951), cerebral metabolic rate for glucose (Sokoloff et al., 1977), and for neuroreceptor ligand binding (Mintun et al., 1984). Further developments have produced a series of reference tissue models that avoid the need for blood sampling (Blomqvist et al., 1989; Cunningham et al., 1991; Hume et al., 1992; Lammertsma et al., 1996; Lammertsma and Hume, 1996; Watabe et al., 2000). Parameter estimates are obtained from a priori specified compartmental structures using one of a variety of least-squares fitting procedures: linear least squares (Carson, 1986), nonlinear least squares (Carson, 1986), generalized linear least squares (Feng et al., 1996), weighted integration (Carson et al., 1986), or basis function techniques (Koeppe et al., 1985; Cunningham and Jones, 1993; Gunn et al., 1997).
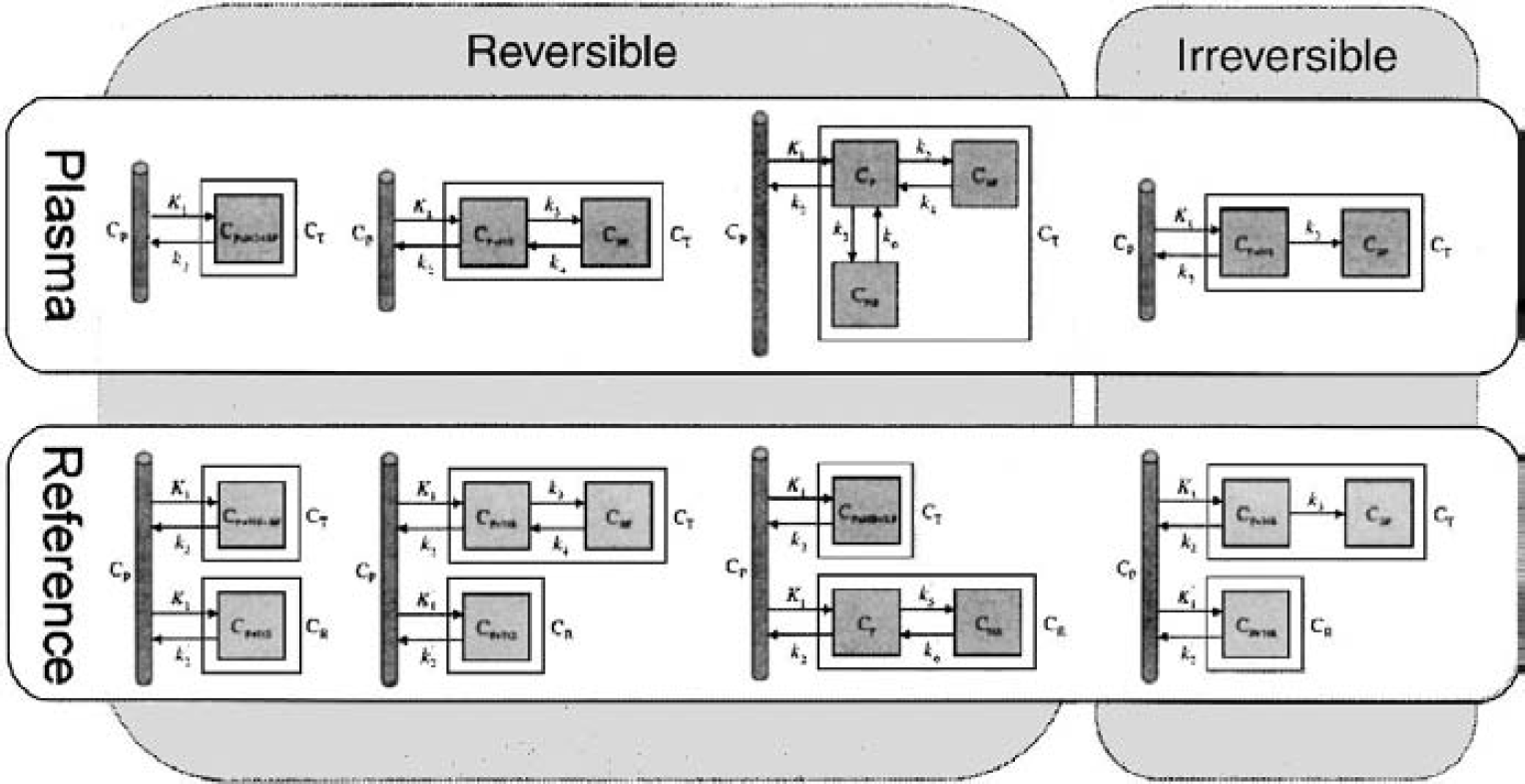
A range of positron emission tomography (PET) compartmental models commonly used to quantify PET radiotracers. These include models for tracers that exhibit reversible and irreversible kinetics and models that use either a plasma or reference tissue input function.
Data-driven methods such as graphical analysis (Gjedde, 1982; Patlak et al., 1983; Patlak and Blasberg, 1985; Logan et al., 1990; Logan et al., 1996) or spectral analysis (Cunningham and Jones, 1993) derive macro system parameters from a less constrained description of the kinetic processes. The graphical methods (Patlak and Logan plots) use a transformation of the data such that a linear regression of the transformed data yields the macro system parameter of interest and are attractive and elegant owing to their simplicity. However, they require the determination of when the plot becomes linear, they may be biased by statistical noise (Slifstein and Laruelle, 2000), and they fail to return any information about the underlying compartmental structure. Appendix A gives a formal derivation of the Logan plot and shows that it is valid for an arbitrary number of compartments for both plasma and reference tissue input models when the data are free from noise. Spectral analysis (Cunningham and Jones, 1993) characterizes the systems impulse response function (IRF) as a positive sum of exponentials and uses nonnegative least squares to fit a set of these basis functions to the data. The macro system parameters of interest are then calculated as functions of the IRF (Cunningham and Jones, 1993; Gunn et al., 2001). Spectral analysis also returns information on the number of tissue compartments evident in the data and is defined as a transparent technique. Schmidt (1999) showed that for the majority of plasma input models, the observation of all compartments led to only positive coefficients, and as such the spectral analysis (Cunningham and Jones, 1993) solution using nonnegative least squares is valid. It is straightforward, however, to deduce that for reference tissue input models, negative coefficients can be encountered and that this approach is strictly not valid [see Appendix C in Gunn et al. (2001)].
Recently, we have published general theory for both plasma input and reference tissue input models (Gunn et al., 2001). This work shows that a general PET tracer compartmental system may be characterized in terms of its IRF, and that this is independent of the administration of the tracer and is therefore applicable to both bolus and bolus-infusion methodologies. The term tracer excludes multiple injection protocols involving low specific activity injections that are classed as nontracer studies and lead to nonlinear compartmental systems. In brief, a general PET tracer compartmental model leads to a set of first-order linear differential equations. This set of equations may be solved to yield an expression for the total tissue radioactivity concentration in terms of either a plasma input function or a suitable reference tissue input function. These results are derived from linear systems theory, which leads to the deduction that the IRF comprises solely exponentials and a delta function (Gunn et al., 2001). This work forms the foundation for the presented method: data-driven estimation of parametric images based on compartmental theory (DEPICT).
DEPICT allows for the estimation of parametric images or regional parameter values from dynamic PET data. DEPICT requires no a priori description of the tracers fate in vivo; it derives the model description from the data and it returns the number of compartments (model order). This transparent modeling technique has application to a wide range of PET radiotracers, but the emphasis here is with respect to the analysis of radioligands that bind to specific neuroreceptor sites.
THEORY
This section introduces the theory behind DEPICT. First, the general forms for plasma input and reference tissue input models are presented; second, key parameters for neuroreceptor binding studies are cast in this framework; and third, the general parameter estimation approach is constructed using basis functions and solved via basis pursuit denoising. The method encompasses the majority of linear compartmental systems, which are applicable to tracer studies with an arbitrary input function. It assumes that there is only one form (the parent compound) in which radioactivity enters the tissue from the arterial plasma. Specifically, those models that are defined by definitions 1 and 2 in Gunn et al. (2001) are considered. These sets of models encompass all noncyclic systems, the subset of cyclic systems in which the product of rate constants is the same regardless of direction for every cycle and requires the eigenvalues of the system to be distinct.
Plasma input models
The general equation for a plasma input compartmental model is given by
Reference tissue input models
The general equation for a reference tissue input compartmental model is given by
Parameters for neuroreceptor binding studies
For reversible neuroreceptor binding studies, the binding potential may be calculated from the macro parameters, using either a plasma or reference tissue input function, under the assumption that the volume of distribution of the free and nonspecific binding is the same in the target and reference tissues. Throughout this article, the term “binding potential” is used interchangeably to refer to BP [the binding potential as originally defined by Mintun et al. (1984)], BP.f2 and BP.f1 where f2 and f1 are the tissue and plasma free fractions respectively. For a reference tissue input analysis, BP.f2 is the only binding potential estimate possible, whereas with a plasma input analysis it is possible to obtain estimates for BP.f1, BP.f2, and if a separate measure of the plasma free fraction exists, BP. The binding potential is a useful measure of receptor specific parameters, which includes both the maximum concentration of receptor sites and the affinity,
Calculation of commonly used binding parameters from the general impulse response function
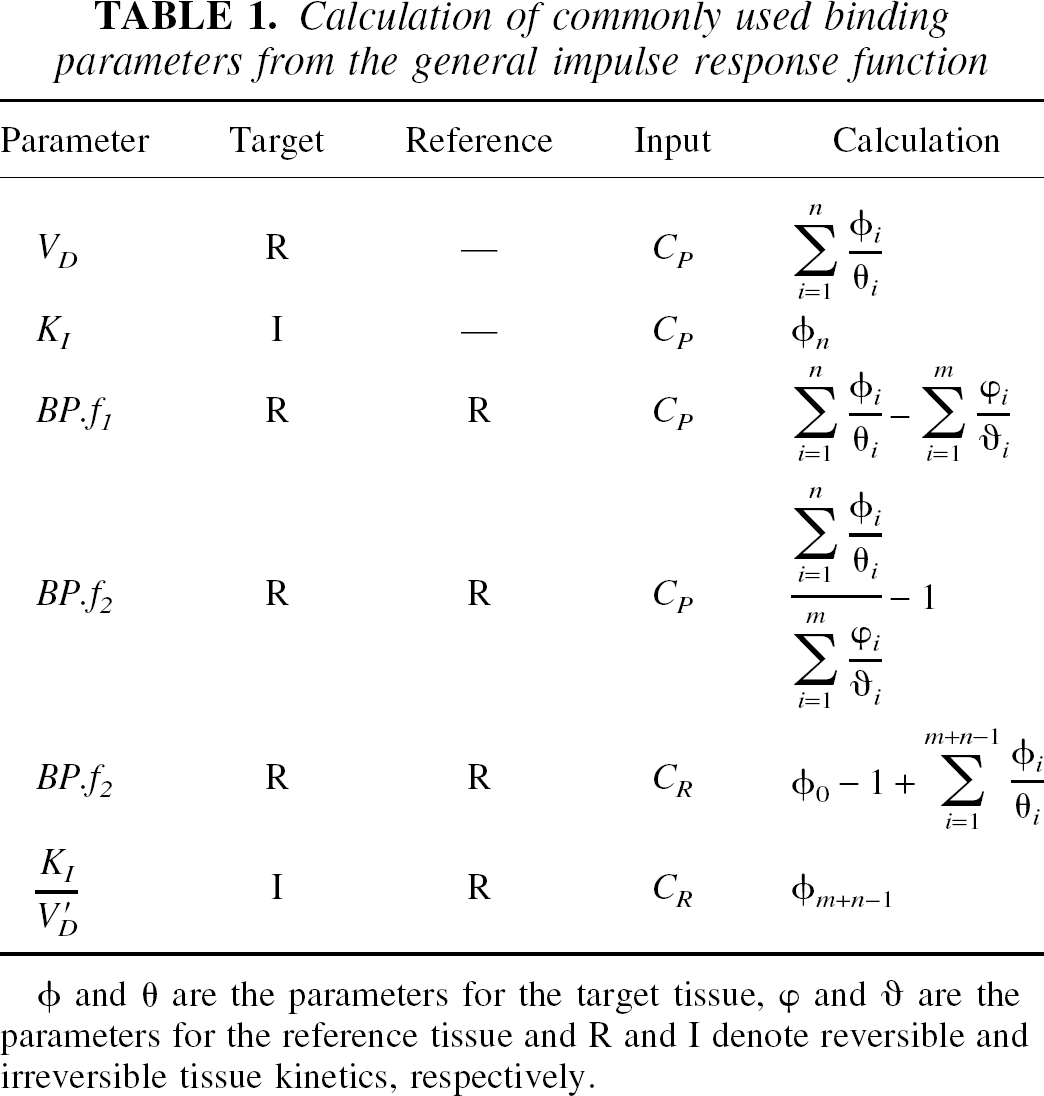

φ and θ are the parameters for the target tissue, φ and υ are the parameters for the reference tissue and R and I denote reversible and irreversible tissue kinetics, respectively.
Construction of the general model in a basis function framework
Plasma input
1
and reference tissue input PET compartmental models are characterized by
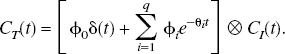
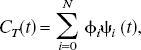
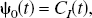
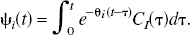
A set of N values for θ
i
may be prechosen from a physiologically plausible range θmin ⩽ θ
i
⩽ θmax. Here, the θ
i
values are spaced in a logarithmic manner to elicit a suitable coverage of the kinetic spectrum. Other possibilities for the spacing of the basis exist such as an equiangular scheme (Cunningham et al., 1998). For data that have not been corrected for the decay of the isotope, θmin may be chosen as (or close to the decay constant (θmin = λ min−1) for the radioisotope and θmax may be chosen as a suitably large value (θmax = 6 min−1). For reversible systems, where the calculation of VD or BP.f2 is the goal, the choice of a value for θmin, which is slightly bigger than λ, can suppress the calculation of infinite VD and BP.f2 values from noisy data. PET measurements are acquired as a sequence of (F) temporal frames. Thus, the continuous functions must be integrated over the individual frames and normalized to the frame length to correspond to the data sampling procedure. The tissue observations, y, already exist in this form and correspond to
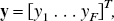
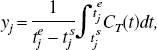
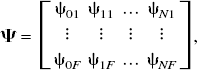
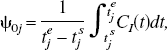
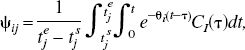
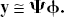
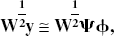
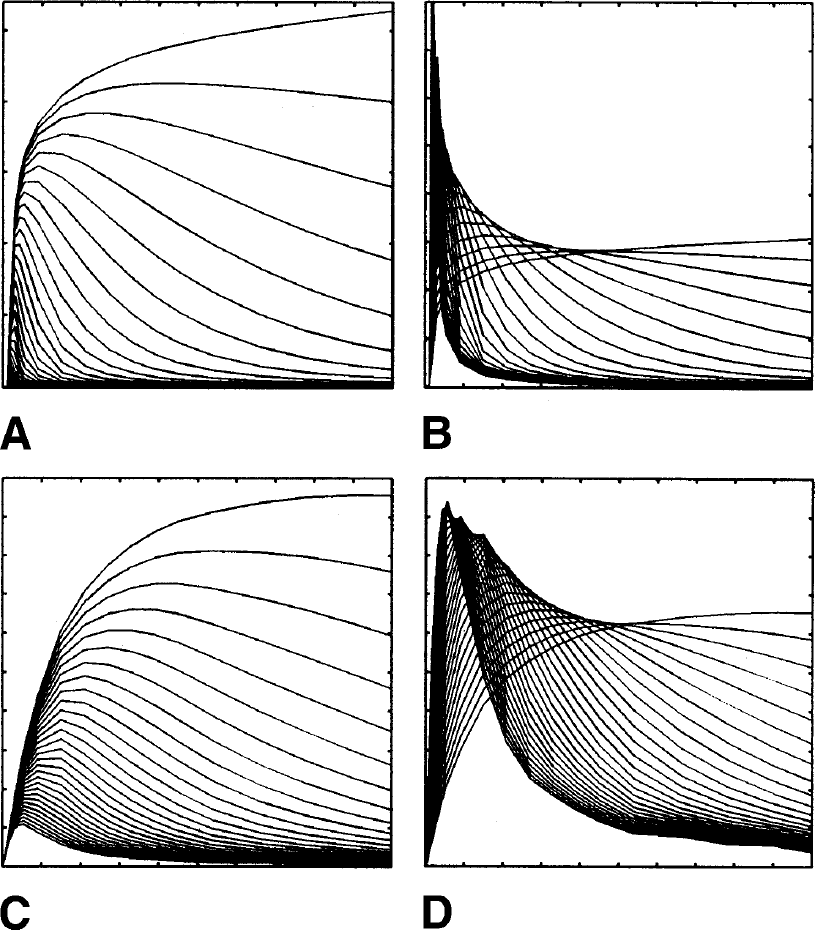
Example dictionaries (Ψ).
Solution of the general model by basis pursuit denoising
Standard least-squares techniques are not applicable because of the overcomplete basis, which leads to an underdetermined set of equations. This ill-posed problem requires an additional constraint to impose a unique solution on the estimation process. This constraint is chosen to be consistent with prior knowledge about the solution, namely, that the solution will be sparse in the basis coefficients. This constraint has also been used in the wavelet community, for the estimation of sparse representations from overcomplete dictionaries (Chen, 1995; Chen et al., 1999). The motivation for sparseness is consistent with the expectation that the data are accurately described by a few compartments (such as the models in Fig. 1). To transform the problem so that a unique solution exists, it is necessary to change the metric from ordinary least squares. The introduction of a regularizer or penalty function to the standard least-squares metric offers a framework for this,
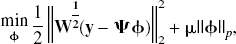
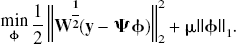
With the introduction of slack variables, basis pursuit denoising can be written as a simple bound constrained quadratic program (Chen, 1995; Chen et al., 1999):
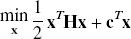
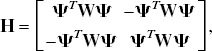
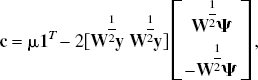
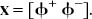
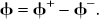
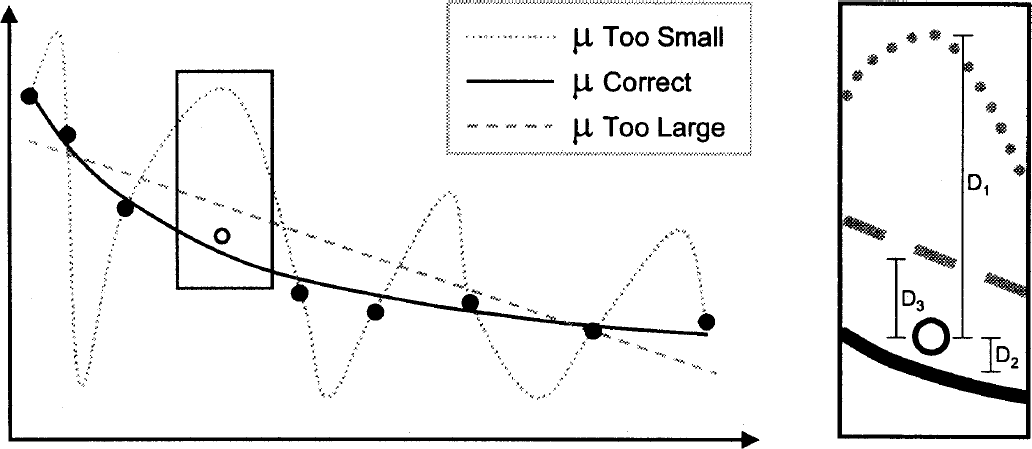
Effect of the regularization parameter (μ) and selection via leave one out cross-validation (LOOCV): When the μ value is too small, the data (•) are overfitted, and when the value is too big the data are underfitted. The “best” μ value is chosen using the method of LOOCV. A data point is omitted (○) and the data are fit for a set of μ values. The error in the model's prediction of the omitted data point is then calculated in a least-squares sense (i.e., Di2). This is repeated for the omission of each data point in turn, and the resultant generalization errors are summated. A μ is selected by choosing the value of μ that minimizes this generalization error. The figure shows how the parsimonious model minimizes the generalization error (i.e., D22 < D32 < D12).
Model order and transparency
The presented method is transparent because it returns information about the underlying compartmental structure. The number of nonzero coefficients returned corresponds to the model order, which is related to the number of tissue compartments. The number of nonzero components is counted as the number of distinct peaks within the spectrum. The method will often include two peaks next to each other in order to approximate an exponential in between them. This occurrence is treated as a single nonzero coefficient. For plasma input models, the model order equals the number of tissue compartments (ignoring the blood volume component) and for reference tissue models, the number of nonzero coefficients corresponds to the total number of tissue compartments in the reference and target tissues. Although the model order dictates the number of compartments, any decision about the true model configuration is limited by the problem of indistinguishability. Figures 4 and 5 depict the sets of models that are equivalent in terms of model order for plasma input and reference tissue input models. If one requires a compartmental description of the tracer, and the set is not singular, then it is necessary to invoke biologic information about the system in order to select among the possible models.
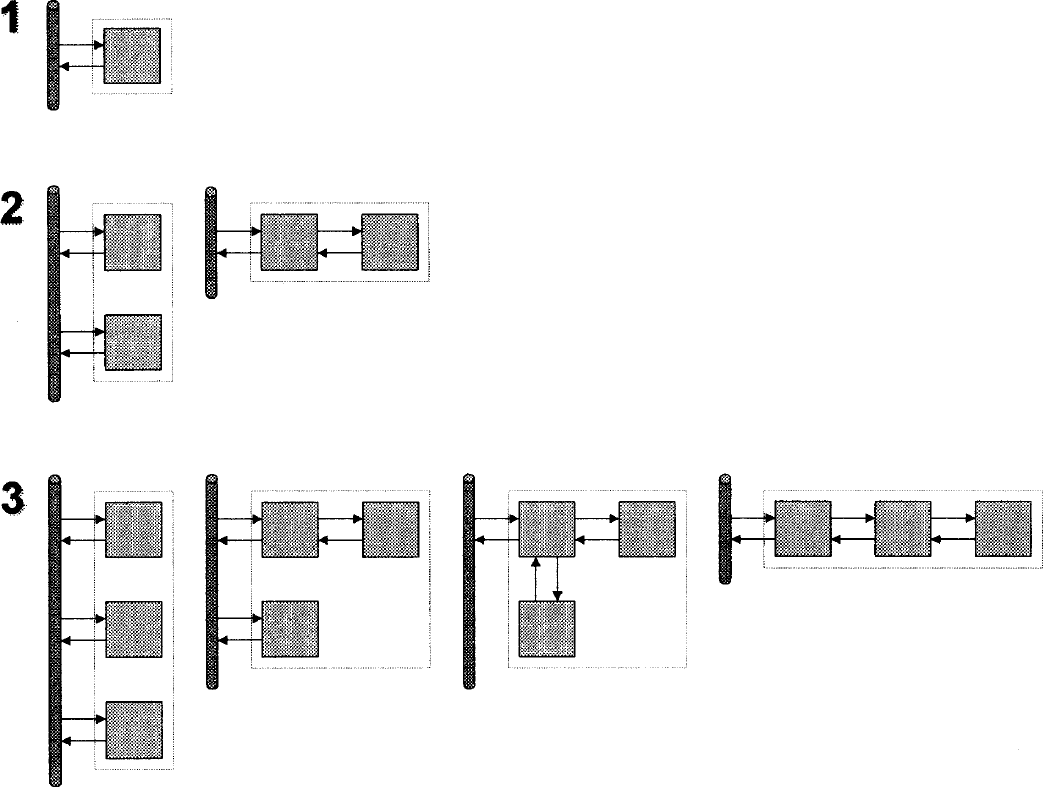
Indistinguishability for plasma input models.
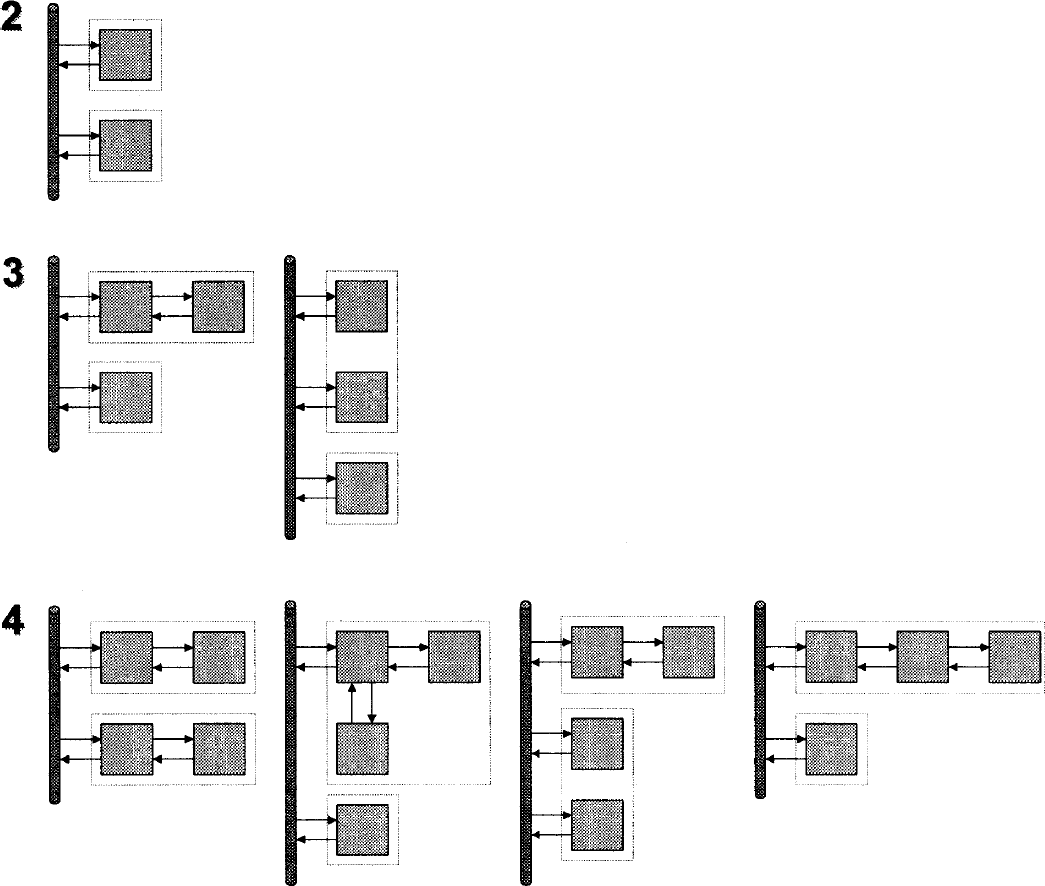
Indistinguishability for reference tissue input models.
MATERIALS AND METHODS
For DEPICT and spectral analysis, the tissue and plasma data were uncorrected for the decay of the isotope. Instead, a decay constant was allowed for in the exponential coefficients (decay constant for [11C]: λ = 0.034 min−1). For the Logan analysis, the tissue and plasma data were precorrected for the decay of the isotope.
DEPICT
The basis pursuit denoising approach was implemented with 30 basis functions (logarithmically spaced between 0.048 and 6 min−1). The number of basis functions was chosen to be 30 based on a balance between precision and computation time (data not shown). The weighting matrix was determined from the true TAC activity and the frame duration as described previously (Gunn et al., 1997). The regularization parameter, μ, was determined by numerically minimizing the LOOCV error across a discrete set of logarithmically spaced μ values (10−3.5 ⩽ μ ⩽ 100.5). For the one-dimensional simulations, the parameter μ was determined as the mean value from the 1,000 realizations. For parametric imaging, the μ value was obtained from a series of LOOCV estimates (μ
i
) obtained from random voxel locations contained within a brain mask. This process examined at least 20 voxels and continued until the relative precision of 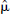 was less than 50%,
was less than 50%,
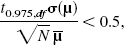
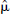 was then estimated as the mean of the vector μ. DEPICT is available at http://www.bic.mni.mcgill/∼rgunn.
was then estimated as the mean of the vector μ. DEPICT is available at http://www.bic.mni.mcgill/∼rgunn.
One-dimensional simulations
Simulations were performed to assess the stability of the three methods to different noise levels. A measured input function from a PET scan was used in conjunction with a two-tissue compartmental model to simulate noise-free data representative of a target tissue [K1 = 0.4 (mL plasma) · min−1 · (mL tissue)−1, k2 = 0.2 min−1, k3 = 0.4 min−1 and k4 = 0.1 min−1) and a reference tissue [K′1 = 0.4 (mL plasma) · min−1 · (mL tissue)−1, k′2 = 0.4 min−1, k′5 = 1 min−1 and k′6 = 1 min−1]. Ninety minutes of data were simulated for a total of 24 temporal frames (3 × 10 seconds, 3 × 20 seconds, 3 × 60 seconds, 5 × 120 seconds, 5 × 300 seconds, 5 × 600 seconds). The noise-free simulated data are shown in Fig. 6. For a range of noise levels, 1,000 realizations of noisy target tissue TACs were generated by adding normally distributed noise proportional to the activity/frame duration at each point. The noise variance was scaled so that a value of 1 corresponded to the largest noise-free TAC value. These data were then analyzed in two different ways: (1) using the plasma input function, the noisy target tissue TACs were fitted with the basis pursuit method, the Logan plot, and spectral analysis to derive VD estimates; and (2) using the reference tissue input function, the noisy target tissue TACs were fitted with the basis pursuit method and the Logan plot to derive BP.f2 estimates.
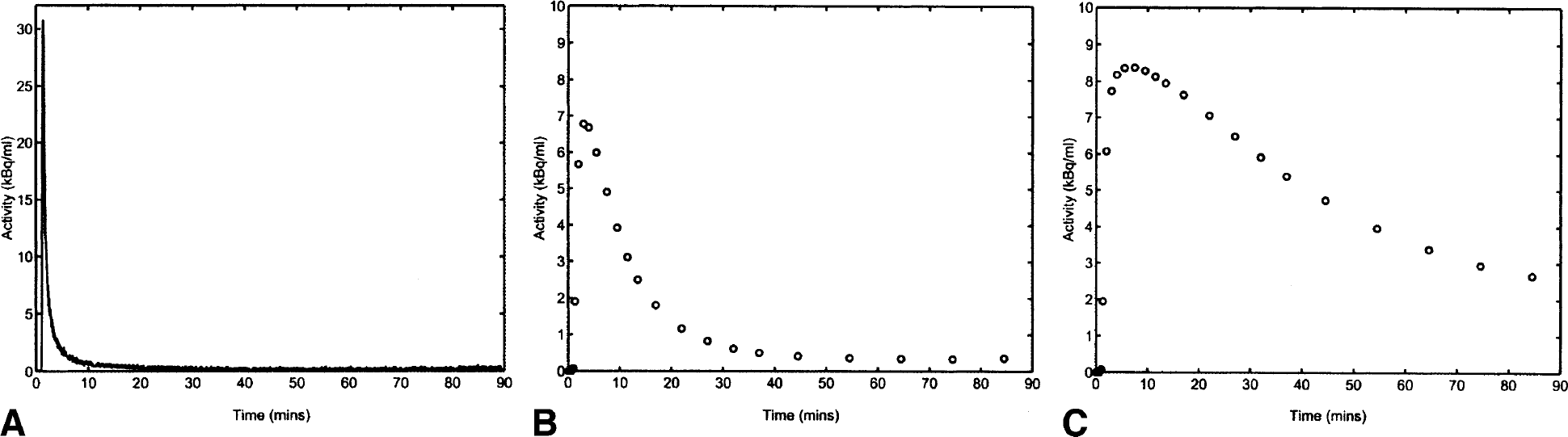
One-dimensional simulation data: noise-free time–activity curves (TAC).
Four-dimensional simulations
Simulated dynamic PET data were generated using PETSim, a dynamic PET simulator (Ma et al., 1993; Ma and Evans, 1997) as described previously (Aston et al., 2000). In short, the simulator takes a magnetic resonance image volume segmented into 28 regions, each of which is assigned an activity value, and generates a simulated PET volume. The resolution and noise characteristics were chosen to correspond to the ECAT HR+ PET camera (CTI, Knoxville, TN, U.S.A.) in 3D mode with a resolution of 4 × 4 × 4.2-mm full width at half maximum (FWHM) at the center of the field of view and images were reconstructed using a 6-mm FWHM Hanning filter. A simulated [11C]SCH 23390 data set was generated using a measured plasma input function and a two-tissue compartmental model to simulate regional tissue kinetics using rate constants for different brain regions taken from human PET data, K1 (0.08 to 0.13 mL plasma · min−1 mL tissue−1), k2 (0.24 to 0.37 min−1), k3 (0 to 0.14 min−1), k4 (0.1 min−1) for 34 time frames (4 × 15 seconds, 4 × 30 seconds, 7 × 60 seconds, 5 × 120 seconds, 14 × 300 seconds). The cerebellum was simulated with a one-tissue compartment model. Two dynamic data sets were generated; a noise-free data set and the other consistent with an injection of 370 MBq of activity. The two data sets were analyzed with DEPICT to estimate parametric images of VD and model order.
Measured data sets
Two measured data sets, which were taken from ongoing clinical studies (MRC Cyclotron Unit, Hammersmith Hospital, London, U.K.), were analyzed with DEPICT. Both data sets were acquired on the ECAT EXACT3D (CTI) (Spinks et al., 2000). The data were reconstructed with model-based scatter correction and measured attenuation correction using the method of filtered backprojection (ramp filter, 0.5 Nyquist). The reconstructed images had a resolution of 4.8 × 4.8 × 5.6 mm at the center of the field of view:
[11C]Diprenorphine (opiate receptor): 130 MBq of the radioligand was injected into a normal male volunteer, and acquisition consisted of 32 temporal frames of data (1 × 50 seconds, 3 × 10 seconds, 7 × 30 seconds, 12 × 120 seconds, 6 × 300 seconds, 3 × 600 seconds). Continuous arterial sampling and metabolite analyses were performed during the scan, which allowed for the generation of a metabolite corrected plasma input function as described previously (Jones et al., 1994). DEPICT was used to estimate parametric images of VD and model order using a plasma input analysis.
[11C]WAY-100635 (5-HT1A receptor): 243 MBq of the radioligand was injected into a normal male volunteer and acquisition consisted of 22 temporal frames of data (1 × 15 seconds, 3 × 5 seconds, 2 × 15 seconds, 4 × 60 seconds, 7 × 300 seconds, 5 × 600 seconds). A region of interest was defined on the cerebellum for the extraction of a reference region TAC. DEPICT was used to estimate parametric images of BP.f2 and model order using a reference tissue input analysis.
RESULTS
One-dimensional simulations
A summary of the results from the 1D noise simulations is presented in Fig. 7 for the plasma input model. DEPICT was able to obtain a good fit to the data. Both DEPICT and spectral analysis performed well in terms of parameter and model order estimation. Table 2 shows that DEPICT produces the lowest mean square error (MSE) of the three methods. The reference tissue input simulations are summarized in Fig. 8 and Table 3. Again DEPICT obtained a good fit to the data and a reasonable model order estimation. Table 3 shows that DEPICT produces the lowest MSE of the two methods.
One-dimensional simulation: summary parameters for VD estimation with a plasma input function
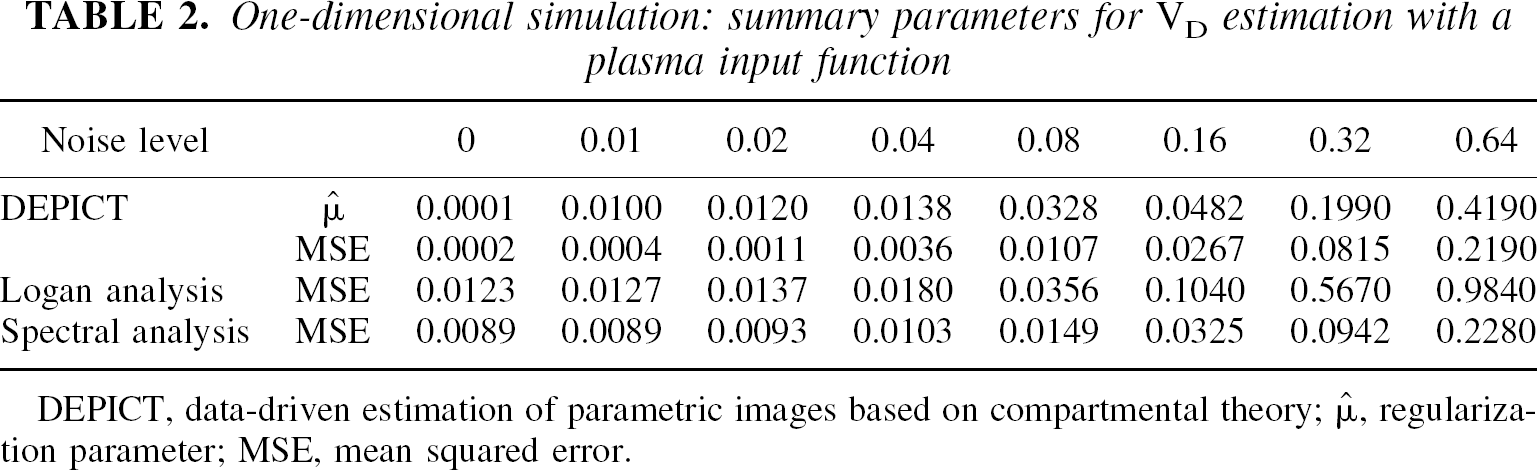
DEPICT, data-driven estimation of parametric images based on compartmental theory; 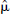 , regularization parameter; MSE, mean squared error.
, regularization parameter; MSE, mean squared error.
One-dimensional simulation: summary parameters for BP.f2 estimation with a reference tissue input function
DEPICT, data-driven estimation of parametric images based on compartmental theory; 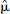 , regularization parameter; MSE, mean squared error.
, regularization parameter; MSE, mean squared error.
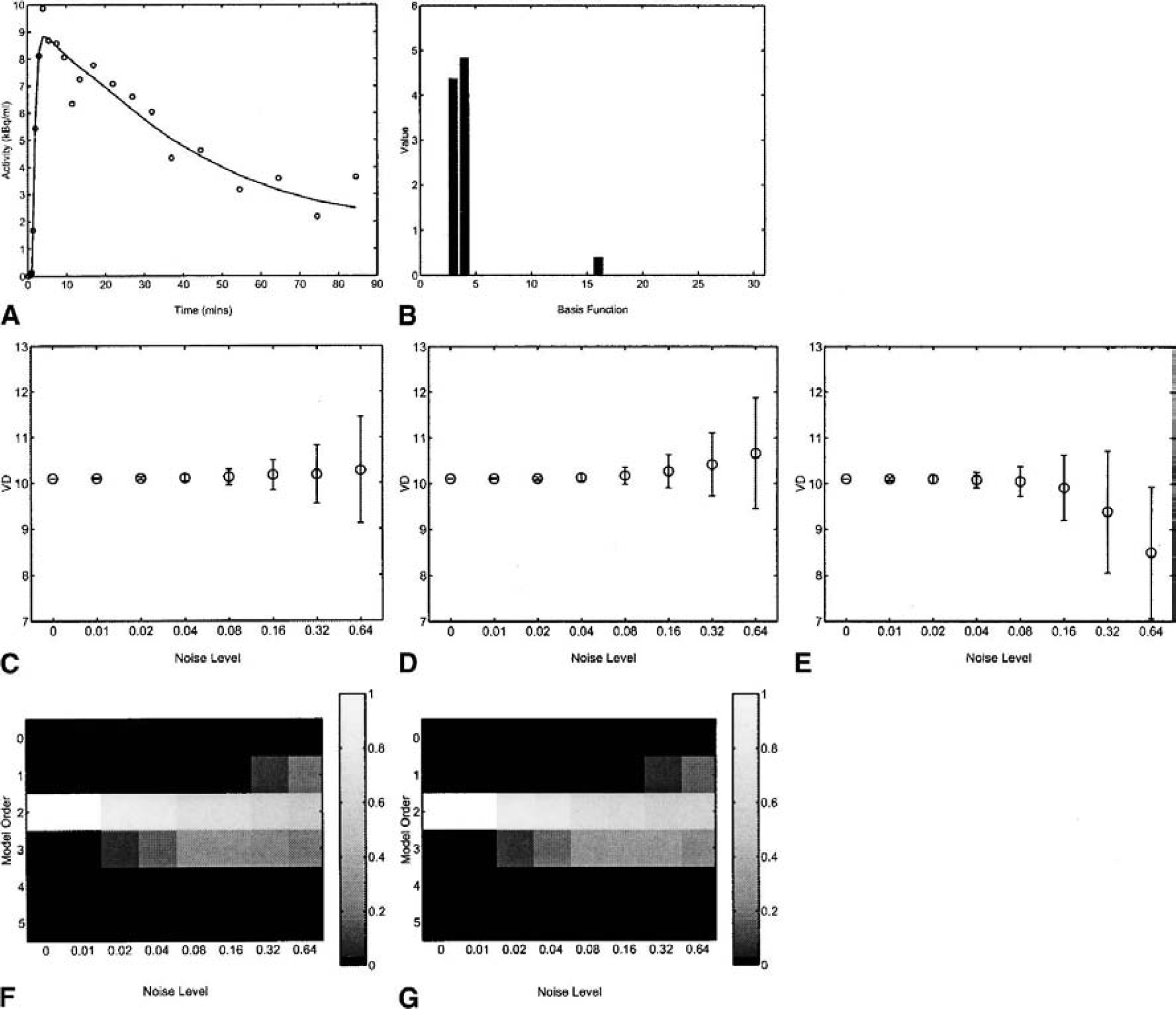
One-dimensional simulation: plasma analysis.
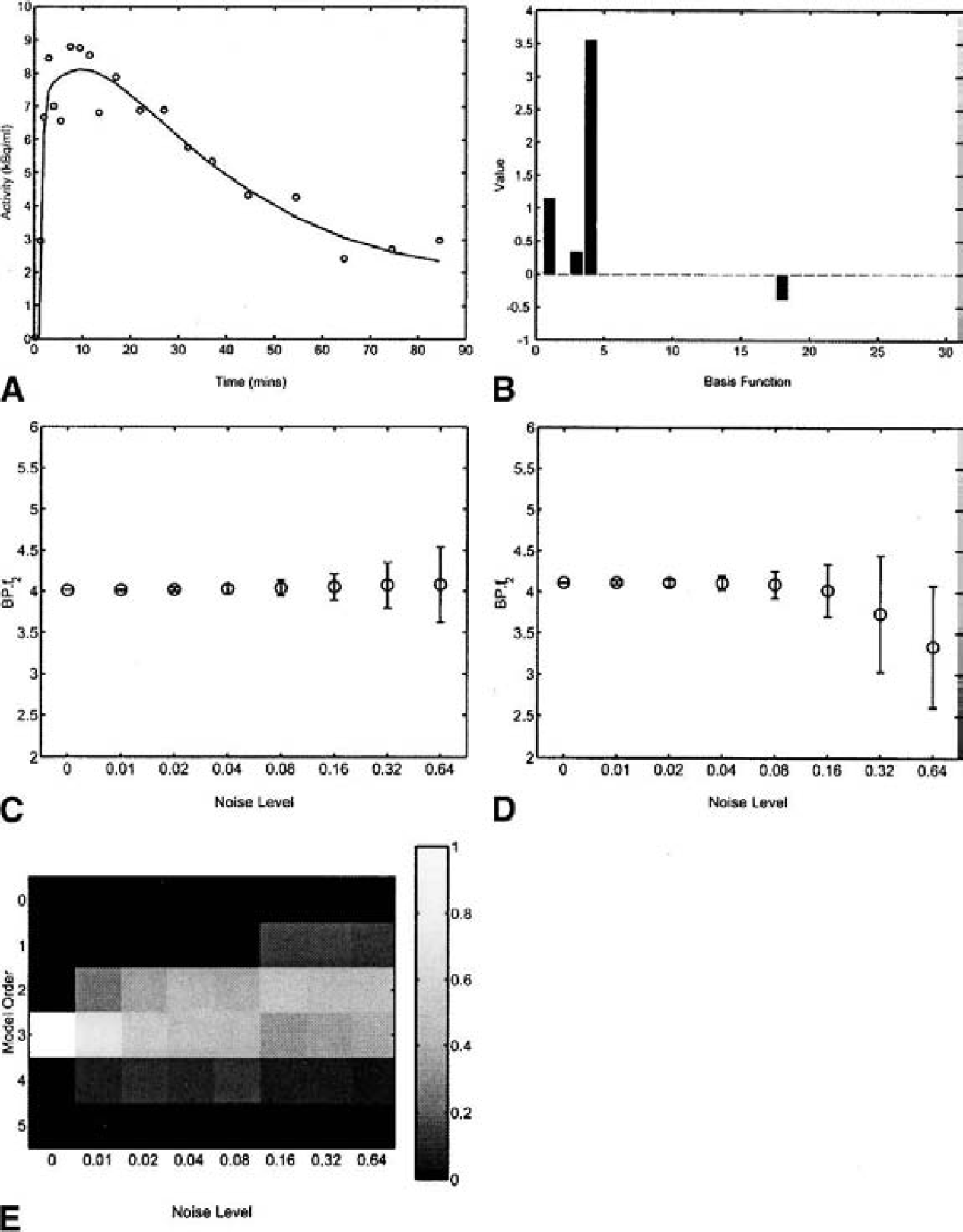
One-dimensional simulation: reference tissue analysis.
Four-dimensional simulations
Parametric images estimated from the noisy 4D realization using DEPICT were of good quality and are presented in Fig. 9. DEPICT accurately estimates the number of tissue compartments used in the simulation (Fig. 9B), with two tissue compartments estimated on average within the cortical regions and one tissue compartment on average within the cerebellum.
The regularization parameter was calculated as  = 0.045 (Fig. 10A). DEPICT was also applied to the noise-free data set, and this allowed for the calculation of the percentage error in the VD estimate. The error distribution is given in Fig. 10B.
= 0.045 (Fig. 10A). DEPICT was also applied to the noise-free data set, and this allowed for the calculation of the percentage error in the VD estimate. The error distribution is given in Fig. 10B.
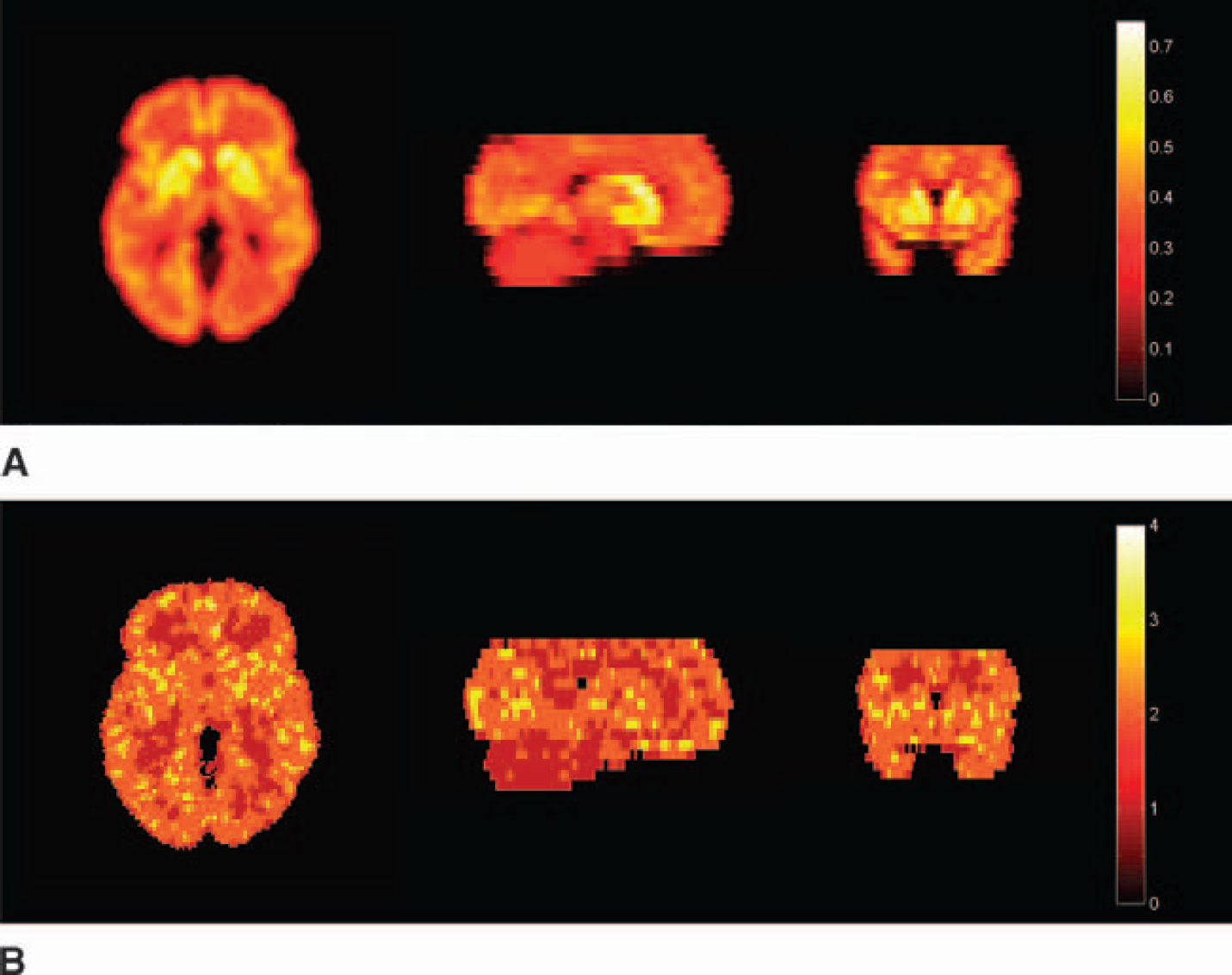
Four-dimensional simulation: parametric images estimated using DEPICT.
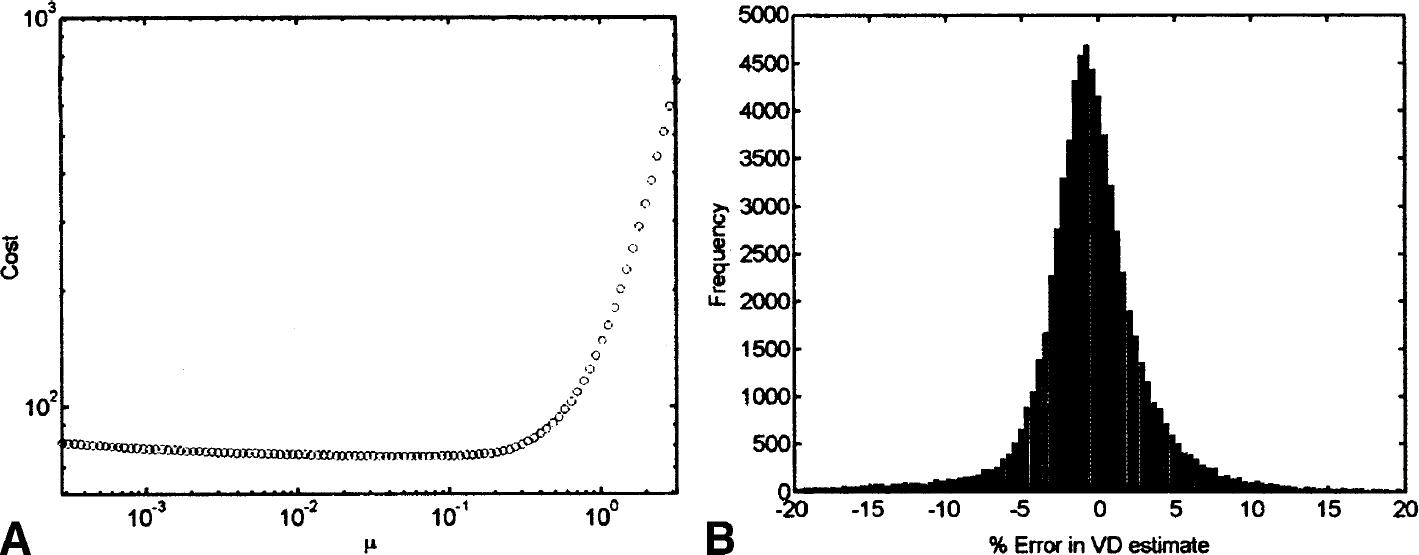
Four-dimensional simulation: cost function and error in VD estimate.
Measured data
Both the parametric images of VD for [11C]diprenorphine and BP.f2 for [11C]WAY-100635 were of good quality and reflected the known distribution of opiate and 5-HT1A receptor sites, respectively. The model order images for both radioligands showed less structure than the 4D simulation, but both reflected the model order expected for [11C]diprenorphine (Jones et al., 1994) and [11C]WAY-100635 (Gunn et al., 1998; Farde et al., 1998; Parsey et al., 2000) (Fig. 11). These parametric images took 1 hour to compute with DEPICT on a desktop workstation (equivalent computation times for parametric images generated by the Logan analysis and Spectral analysis would be 5 minutes and 30 minutes, respectively). The regularization parameters were calculated as  = 0.0095 for [11C]diprenorphine and
= 0.0095 for [11C]diprenorphine and 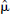 = 0.015 for [11C]WAY-100635 (Fig. 12).
= 0.015 for [11C]WAY-100635 (Fig. 12).
Measured data: parametric images estimated using data-driven estimation of parametric images based on compartmental theory (DEPICT).
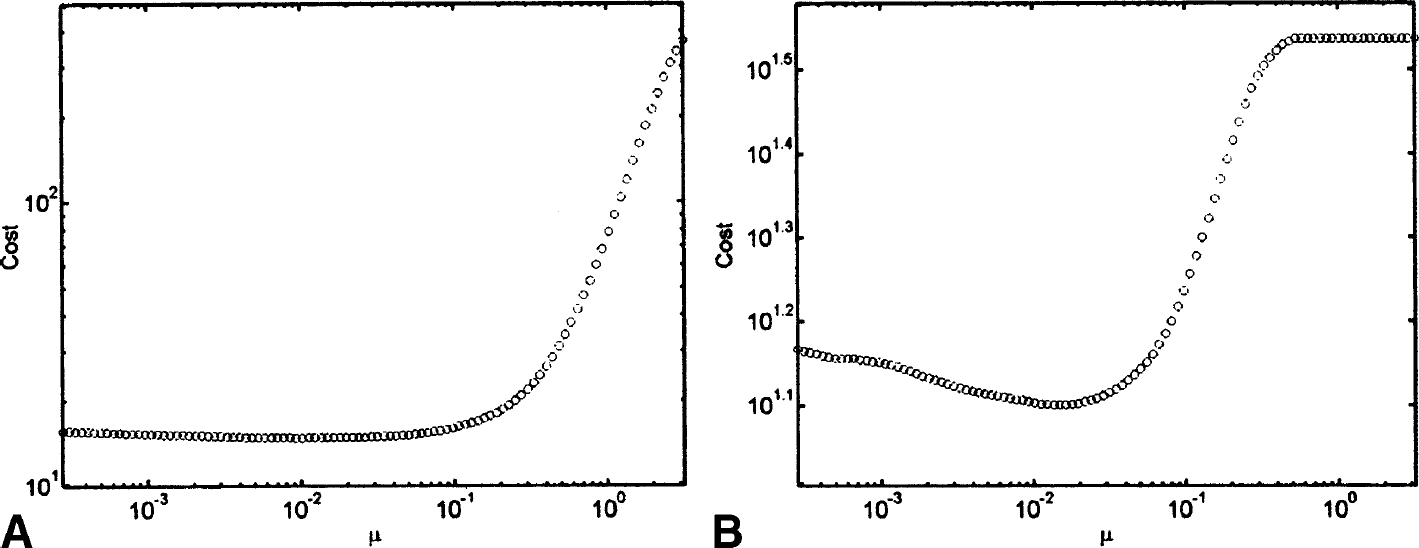
Measured data: cost functions for regularization parameter (μ).
DISCUSSION
The current article has introduced DEPICT, a tracer kinetic modeling technique for the quantitative analysis of dynamic in vivo radiotracer studies that allows for the data-driven estimation of parametric images based on compartmental theory. DEPICT requires no a priori decision about the tracers fate in vivo, instead determining the most appropriate model from the information contained within the data. Although the method is classed as data-driven, it is founded on compartmental theory (Gunn et al., 2001) and this enables parameter estimates to be interpreted within a traditional compartmental framework. The system macro parameters are simply determined from the estimated IRF. The method can be applied to dynamic radiotracer studies involving either a bolus or bolus-infusion tracer administration scheme. DEPICT is general, and although the examples here are concerned with tracers exhibiting reversible kinetics, the method is equally applicable to systems with irreversible kinetics. In addition to parameter estimates, DEPICT returns model order estimates that correspond to the number of numerically identifiable compartments in the system. There may be additional compartments that are not supported by the statistical quality of the data; however, this is not in any way a practical restriction.
DEPICT uses a basis function approach (Koeppe et al., 1985; Cunningham and Jones, 1993; Gunn et al., 1997) to the parameter estimation problem. The constructed problem is ill posed, and its solution requires an appropriate constraint. Spectral analysis approaches this problem using the nonnegative least-squares algorithm (Cunningham and Jones, 1993), whereas DEPICT uses the method of basis pursuit denoising (Chen, 1995), which involves a one-norm penalty function on the coefficients. Both methods lead to a sparse solution, but DEPICT does not constrain the coefficients to be positive, which makes it appropriate for application to the general reference tissue model. Basis pursuit denoising is a technique that extracts a subset of terms from an overcomplete dictionary, and thus it is possible to provide a model that is interpretable, while retaining good approximating capability. The principle behind the approach is to trade off the error in approximation with the sparseness of the representation.
The three data-driven methods investigated all performed well for low noise levels, as determined from the 1D simulations, with DEPICT returning the lowest mean squared error. The Logan analysis demonstrated a bias at higher noise levels, which has been documented recently (Slifstein and Laruelle, 2000). To address the issue of noise-induced bias in the Logan analysis, two approaches have since been developed (Logan et al., 2001; Varga and Szabo, 2002). These modifications would have improved the performance of the Logan analysis at high noise levels but were not considered here, because the aim was to introduce DEPICT and compare it against the data-driven methods in common use. DEPICT and spectral analysis allow for the estimation of the model order. The model order was well characterized by both methods for the 1D plasma input simulations and using DEPICT for the reference tissue input simulations. Spectral analysis is not valid for the general reference tissue model because it does not permit negative coefficients, which can occur in the impulse response function.
In summary, this article has introduced a new method, DEPICT, which delivers parametric images or regional parameter estimates from dynamic radiotracer imaging studies without the need to specify a compartmental structure. DEPICT is applicable to both plasma and reference tissue input analyses. The results presented demonstrate that DEPICT is highly competitive with existing data-driven estimation methods. Furthermore, DEPICT is a transparent data-driven modeling approach because it returns not only macro parameter values, but also information on the underlying model structure.
Footnotes
1
Here, the blood volume term for plasma input models is treated as a plasma volume term (i.e., CB(t) = CP(t)) which enables us to express both the plasma and reference tissue input cases in the same framework. This is simply so that we may be concise in our notation. To use a whole blood volume term Ψ0 = CP is replaced by Ψ0 = CB in ![]() .
.
APPENDIX A: LOGAN PLOT DERIVATION
Here, a formal derivation of the Logan plot is presented for both a plasma and reference tissue input function. The plasma input Logan plot corresponds to the original presentation by Logan et al. (1990). The reference tissue input analysis presented here differs from Logan et al. (1996) in that it proves that the plot is valid for an arbitrary number of compartments in the reference tissue as well as the target tissue. Both derivations presented exclude the presence of vascular contribution to the tissue signal.
APPENDIX B: GLOSSARY

| Symbol | Description | Units |
|---|---|---|
| Cj(t) | Concentration time course of radioactivity in the target tissue | kBq · mL−1 |
| CR(t) | Concentration time course of radioactivity in the reference tissue | kBq · mL−1 |
| Cp(t) | Concentration time course of radioactivity in plasma | kBq · mL−1 |
| CB(t) | Concentration time course of radioactivity in whole blood | kBq · mL−1 |
| C(t) | Concentration time course of radioactivity of the input function | kBq · mL−1 |
| ϕ | Parameter vector for coefficients of the general impulse response function | |
| θ | Parameter vector for exponents of the general impulse response function | |
| tsj | Start time for the jth temporal frame | min |
| tej | End time for the jth temporal frame | min |
| n | Number of tissue compartments in the target tissue | |
| m | Number of tissue compartments in the reference tissue | |
| q | Number of exponential terms in the impulse response function | |
| Ψ | Matrix of basis functions or “Dictionary” | kBq · mL−1 |
| y | Measured positron emission tomography tissue data | kBq · mL−1 |
| μ | Regularization parameter | |
| VD | Volume of distribution of the target tissue | (mL plasma) · (mL tissue)−1 |
| V′D | Volume of distribution of the reference tissue | (mL plasma) · (mL tissue)−1 |
| KI | Irreversible uptake rate constant from plasma for the target tissue | (mL plasma) · min−1 · (mL tissue)−1 |
| BP | Binding potential | (mL plasma) · (mL tissue)−1 |
| VB | Fractional blood volume | |
| f 1 | Plasma free fraction | |
| f 2 | Tissue free fraction | |
| Bmax | Maximum concentration of receptor sites | nmol/L |
| KDTracer | Equilibrium dissociation rate constant | nmol/L |
| Fi | Free concentration of competing ligand | nmol/L |
| KDi | Equilibrium disassociation rate constant of competing ligand | nmol/L |
| ⊗ | Convolution operator |