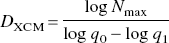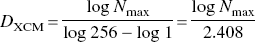Abstract

To the Editor:
A recent article published in the Journal of Cerebral Blood Flow & Metabolism (Hermán et al., 2001) documented the findings of fractal branching pattern on the pial vascular networks in the cat brain. Using the standard box-counting method (Cross, 1997) and an extended counting method (Sandau and Kurz, 1997), the authors reported similar fractal dimensions of approximately 1.3 for both the arterial and venous vasculature of brain cortex from six cats. These fractal dimensions were found to be invariant under geometric transformations, such as translation and rotation of the raw images. After a further assessment of accuracy for the two fractal methods using known fractal objects, the authors concluded that the fractal model seemed to be correct in describing the scale-invariant bifurcation patterns in the pial vascular networks of cats.
Although we agree that the pial vascular networks appear random, such that a quantitative characterization of the arborization of complex vasculature seems difficult using traditional Euclidean geometry, we are specifically concerned about the suitability of the extended counting method used by the authors for fractal analysis. Before discussing the pitfalls of the extending counting method, we first take a brief look at the procedure used by Herman et al. After intensity correction of a digitized video image taken from suprasylvian gyrus, the vasculature was defined by manual tracing of the image's laser printout. The manual tracing was further digitized at a spatial resolution of 600 dots per inch, after which the resulting digital image was skeletonized to yield a one pixel-wide vascular trace. For fractal analysis using the extended counting method, a large square box of width q0 with a square grid of small boxes of width q1 inside was overlaid onto the skeletonized image (Fig. 1). The number of small boxes covering any part of the vasculature was counted as N. Subsequently, the large box was moved to a new place and the counting process repeated until every part of the vessel structure had been visited. The extended counting fractal dimension DXCM, alternatively termed the capacity dimension, was then calculated as
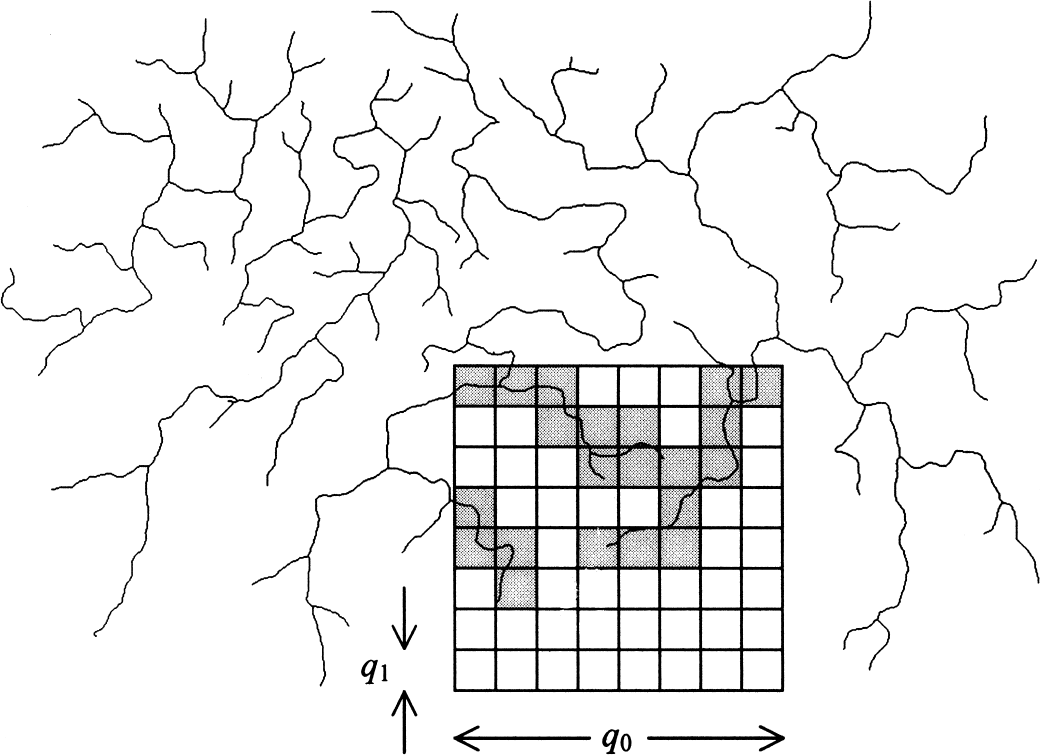
Concept of the extended counting method. A large square box of width q0 and a square grid of small boxes of width q1 inside is overlaid onto the skeletonized vessel image. The number of small boxes covering any part of the vasculature (shown in gray) is counted. Subsequently, the large box is moved to a new place and the counting process repeated until every part of the vessel structure has been visited.
One problem associated with the extended counting method to compute the fractal dimension is that the DXCM as defined by Eq. 1 is unrelated to the branching pattern of the vasculature. To see how this happens, recall that q0 = 256 with q1 − 1 was chosen in the study by Herman et al. Substituting q1 −1 and q0 = 256 into Eq. 1 yields
The second pitfall with the extended counting method is that DXCM is insensitive to loss of vasculature in areas that are not highly vascularized. This happens because Nmax, from which DXCM is derived, represents the maximum local vessel density. As a parameter taking the maximum of all local values, DXCM is not supposed to change with the elimination of other local areas that are less vascularized. In other words, if there were two images showing local spots that are highly vascularized in a similar manner, these two images would have identical DXCM values regardless of how different they are in the other areas. Figure 2, the schematic vessel network shown in Fig. 1, illustrates this situation. The application of the extended counting method on this image showed that Nmax was obtained in the 256 × 256-square region of interest located at the upper left portion of the image (thick line). Thus, reductions in vasculature outside the square have no effect on the computation of DXCM. In this example, both Figs. 2B and 2C, which were obtained by the deliberate elimination of vessels outside the square region of interest, possess exactly the same DXCM of 1.327 as Fig. 2A. Their identical fractal dimensions, however, by no means indicate that these images have similar vascular branching patterns throughout the entire image region.
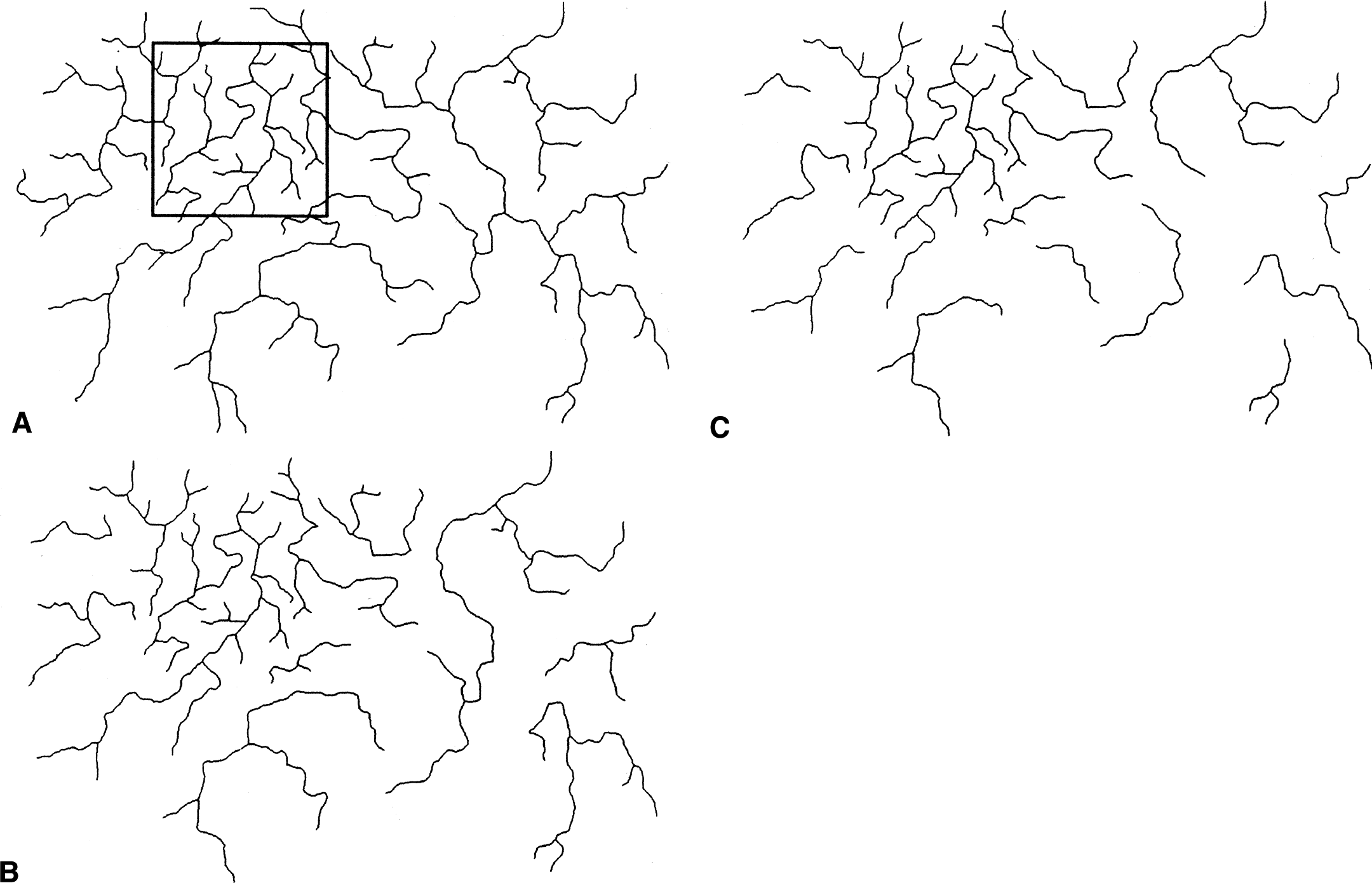
A schematic vessel network
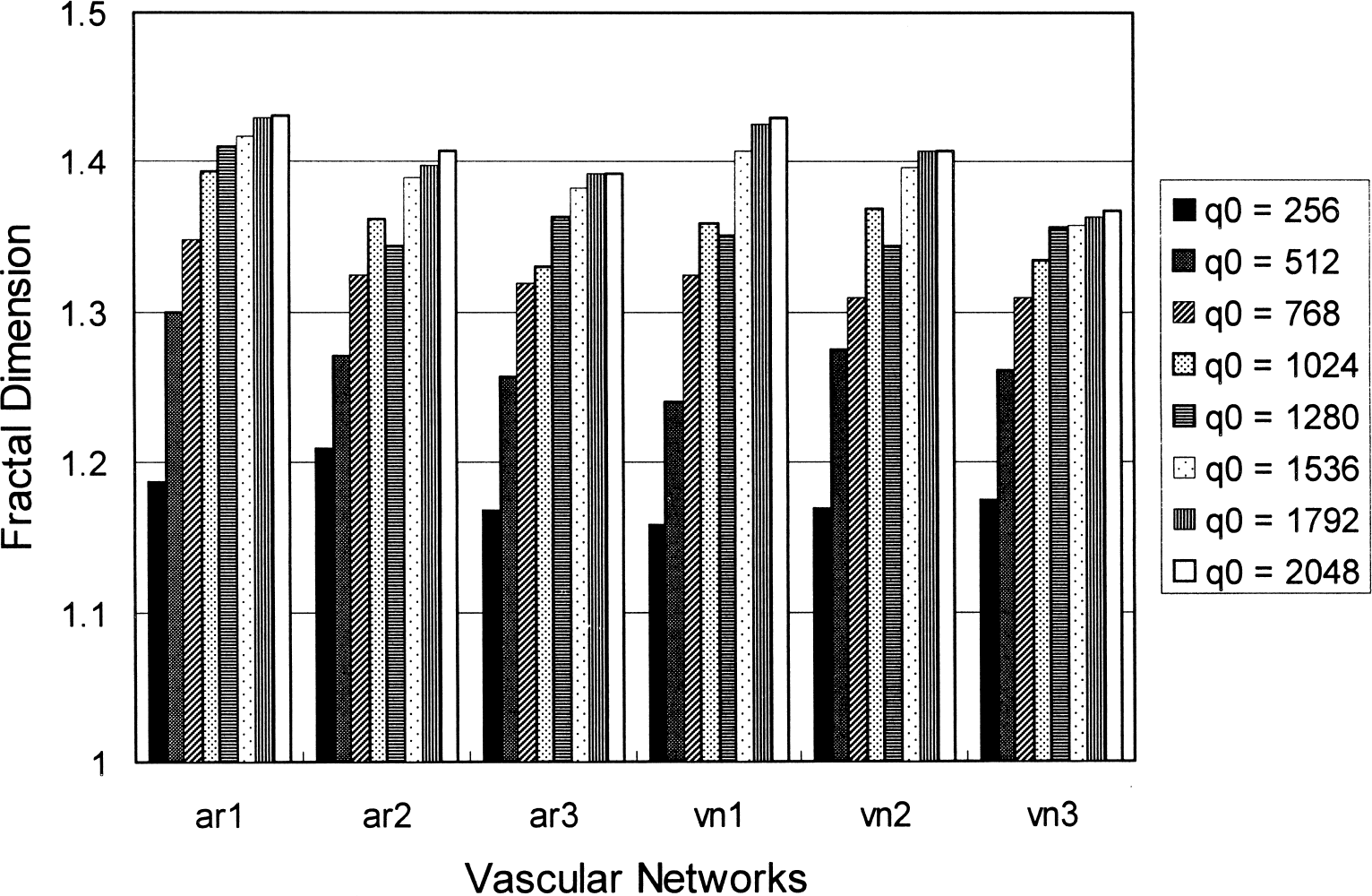
Computation value of the extended counting dimension plotted as a function of q0 for six vascular networks (ar: arterial network; vn: venous network). “Fractal dimensions” vary significantly for small q0, suggesting that the parameter is imprecisely defined.
Another natural consequence resulting from the previously mentioned localization property is that the computation value of DXCM may be prone to variations with different choices of q0, even when q0/q1 is kept constant. Once again, recall that Nmax represents the maximum vessel density within any region of interest that is q0 × q0 pixels in area. Therefore, the choice of a q0 that is small compared with the spatial scale of structural heterogeneity would result in strong dependence of DXCM computation on the selection of q0. To demonstrate this dependency, we digitized three arterial and three venous vascular networks, as shown in Fig. 4 of the article by Herman et al., at 2,400 dot-per-inch resolution. (Note that because the actual size of the laser printouts used by Herman et al. is unknown, their information regarding the 600 dots-per-inch resolution is arbitrary). The extended counting method was performed with q0 = 256, 512, 768, 1,024, 1,280, 1,536, 1,792, and 2,048 at a constant q0/q1 ratio of 256. The resulting changes in DXCM were plotted as a function of q0 for the six vascular networks in Fig. 3. As can be seen from the plot, the computation values of DXCM vary significantly for small q0, suggesting that DXCM is imprecisely defined.
One way to circumvent the above pitfall, as pointed out by Herman et al., is to use a sufficiently large q0 such that the parameter DXCM is not highly localized. Indeed, for all vascular networks shown in Fig. 3, DXCM values became consistent for q0 greater than or equal to 1,536 pixels, corresponding to approximately half the width of the digitized image. At this large q0, however, notice that DXCM would simply represent the mean vessel density within the entire image. In such a case, not only is DXCM unrelated to vascular branching pattern, but the same information can also be obtained through traditional yet more comprehensible image-analysis methods. Therefore, DXCM defined in this manner has no fractal meaning.
We conclude that the extended counting method yields an imprecisely defined “fractal dimension” that is unrelated to the complex branching pattern of pial vasculature. The susceptibility of the computation value of DXCM to the arbitrary choice of q0, and to q0/q1, indicates that DXCM is not a scale-invariant parameter. Whether the branching pattern in the pial vascular networks of cat brain possesses fractal properties remains an open issue awaiting further investigation. The extended counting method, however, is certainly an unsuitable algorithm and may lead to significant confusions in the interpretation of fractal analysis of the vascular arborization process.