Abstract
The microvasculature of the human brain plays an important role in the development and maintenance of the central nervous system and in the pathogenesis of brain diseases, and is the site of differential gene expression within the brain. However, human brain microvascular-specific genes may not be detected in whole-brain gene microarray because the volume of the brain microvascular endothelium is relatively small (0.1%) compared with the whole brain. Therefore, the differential gene expression within the human brain microvasculature was evaluated using suppression subtractive hybridization with RNA isolated from human brain microvessels. Gene identification was restricted to the first 71 clones that were differentially expressed at the brain microvasculature. Twenty of these were genes encoding proteins with known function that were involved in angiogenesis, neurogenesis, molecular transport, and maintenance of endothelial tight junctions or the cytoskeleton. Eighteen genes coding for proteins of an unknown function were identified, including five genes containing satellite DNA sequences. The results provide the initial outline of the genomics of the human brain microvasculature, and have implications for the identification of both targets for brain-specific drug transport and changes in microvascular gene expression in brain diseases.
The blood vessels comprising the brain microvasculature form an effective barrier between the blood and the cells of the brain. The permeability properties of the brain endothelial barrier are distinct, and include epitheliallike tight junctions and a diversity of transporters for both small and large molecules. Because the brain microvascular endothelium serves as the interface between blood and brain, the brain capillary endothelium would be expected to selectively express genes that encode for transporter proteins and proteins that respond to multiple cytokine and chemical signals. A comprehensive molecular analysis of the differential gene expression within the brain microvasculature is critical in identifying its roles in neurogenesis, angiogenesis, and drug transport to the brain (Pardridge, 2001).
Tissue-specific gene expression within the whole brain has been analyzed with gene microarray and DNA sequencing. However, the brain microvasculature comprises only 10−3 of total brain volume (Pardridge, 2001), and the sensitivity of gene microarray can be as low as 10−4 (Schena et al., 1995). Therefore, gene microarrays derived from total brain may not effectively identify the majority of genes that are differentially expressed in brain at the microvasculature. Conversely, the genomics of the brain microvasculature can be analyzed after the purification of RNA obtained from freshly isolated brain microvessels (Boado and Pardridge, 1991). Suppression subtractive hybridization (SSH) methodology is well suited for analyzing gene expression in the brain microvasculature and for subtracting the commonly expressed genes widely expressed in many tissues (Diatchenko et al., 1996). In contrast to serial gene-expression analysis, SSH does not necessitate comprehensive sequencing capacity because only the differentially expressed genes are analyzed. In addition, unlike differential display techniques, SSH can examine differential expression of low-abundance transcripts (Diatchenko et al., 1996). In the present study, microvessels were isolated from fresh human brain removed at the time of neurosurgery, and the poly(A)+ RNA was immediately extracted from the human brain microvessels to generate tester complimentary DNA (cDNA). The SSH methodology was used on the human brain microvessels to identify differentially expressed genes compared with liver and kidney, and the first set of differentially expressed genes was analyzed.
MATERIALS AND METHODS
Suppression subtractive hybridization
The SSH procedure was performed using the PCR-Select cDNA Subtraction Kit from Clontech (Palo Alto, CA, U.S.A.). Poly(A)+ RNA was purified from microvessels isolated from fresh human brain (age, 2 months to 8 years) obtained during neurosurgery for intractable epilepsy secondary to cerebral dysplasia (Mathern et al., 1999). The integrity of the microvessel preparation was confirmed by microscopy showing the absence of contaminating brain cells (Fig. 1A), and by showing the presence of intact messenger RNA (mRNA) through actin Northern blot analysis (Fig. 1B). Human kidney, liver, heart, and lung total RNA was obtained from Clontech, and poly(A)+ RNA was purified using the mRNA Separator Kit (Clontech). Two micrograms human brain capillary, liver, or kidney poly(A)+ RNA was reverse transcribed using avian myeloblastosis virus reverse transcriptase and oligodeoxythymidine primers, as described previously (Li et al., 2001). The second strand of cDNA was synthesized and the products subsequently digested with RsaI to create smaller, blunt-ended fragments necessary for efficient subtraction. After confirmation of digested product, adaptors 1 and 2R were ligated independently to two samples of the tester (brain-microvascular derived) cDNA for use in postsubtraction PCR amplification. Equal amounts of kidney and liver driver cDNA were mixed and hybridized with the different adaptor-ligated tester cDNA to equalize the levels of high- and low-expressed genes, and to enrich for differentially expressed genes as a function of hybridization kinetics. The two hybridization samples with different PCR adaptors were mixed and fresh kidney and liver driver cDNA was added for a second hybridization. This process further enriches differentially expressed genes and forms double-stranded cDNA molecules with different adaptors on each strand. These enriched genes are then put through two cycles of polymerase chain reaction (PCR) with primers designed for the adaptor sequences to amplify tester-enriched genes and to reduce the recovery of background fragments.
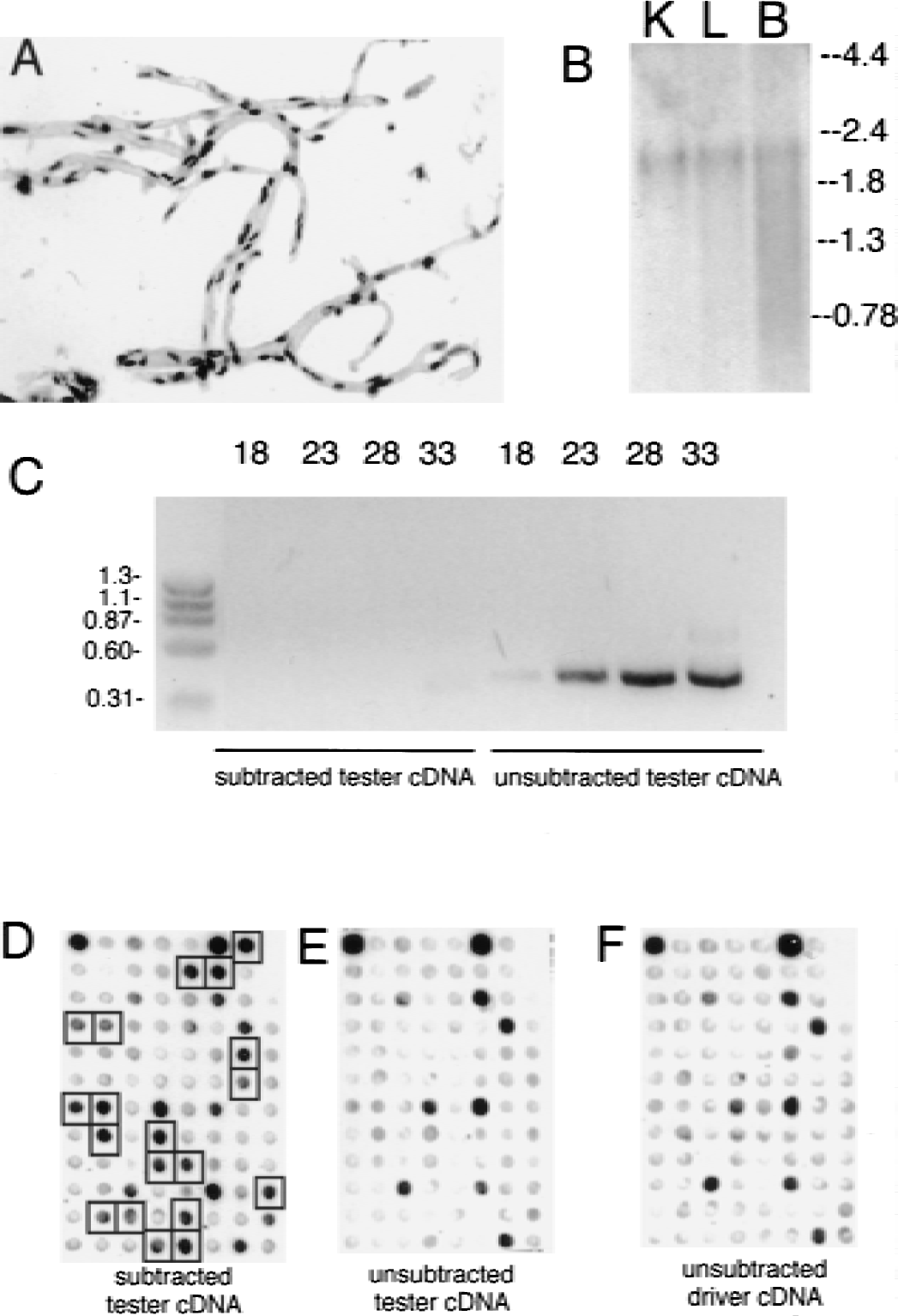
Enrichment of brain microvasculature-specific genes.
Gene microarray screening
A fraction of the brain microvascular-enriched cDNA PCR products was column purified (Qiagen, Valencia, CA, U.S.A.), and subcloned into pCR-2.1 by deoxythymidine/deoxyadenosine ligation (Invitrogen, San Diego, CA, U.S.A.). The ligation products were transformed into XL-1 Blue-competent Escherichia coli (Stratagene, San Diego, CA, U.S.A.) for blue/white selection of clones containing the SSH-derived insert. Based on the results from the ligation and subcloning of this small fraction of subtracted cDNA, the total library of subtracted cDNA products would yield a total 1.2 × 106 clones. Five hundred colonies harboring clones containing the insert were chosen and used to inoculate 96-well plates with ampicillin (100 μg/mL) and Lennox broth medium. After growing for 2 days at 37°C, the cultures were blotted to a nitrocellulose filter using a vacuum. The filters were denatured with 0.5 mol/L sodium hydroxide, neutralized, and baked at 80°C. Radiolabeled probes were produced using the random primer method (Amersham, Chicago, IL, U.S.A.) with column-purified subtracted tester cDNA, unsubtracted tester cDNA, and unsubtracted driver cDNA templates. Standard Southern blotting protocol was used (1.5X sodium chloride/sodium phosphate/ethylenediaminetetraacetic acid, 1% sodium dodecyl sulfate, 0.5% dry milk, stored at 65°C overnight, 3 × 105 cpm probe/mL) to probe the membranes, and an oligonucleotide-blocking solution (Clontech) was used to block the signals from the adaptor molecules present on each cDNA insert.
Tissue distribution analysis
The concentration of the human mRNA samples for the reverse transcription-coupled PCR (RT-PCR) reactions were measured and 15 ng total mRNA was reverse transcribed using an oligodT18 primer and a 32P-dCTP tracer. The size distribution and amount of mRNA reverse transcribed was determined by gel electrophoresis and exposure to x-ray film. The normalized mRNA samples were then treated with DnaseI (DNA-free) to remove genomic DNA contamination. The complete removal of genomic DNA was tested for each tissue mRNA sample using an RT-PCR reaction (Platinum Quantitative RT-PCR system; Invitrogen [formerly Life Technologies], San Diego, CA, U.S.A.) either with both reverse transcriptase and Taq polymerase or with Taq polymerase alone. Gene-specific PCR primers were designed to have melting points of 60°C, and the oligonucleotides were purchased from Biosource International (Camarillo, CA, U.S.A.); 10 ng each mRNA were used in RT-PCR reactions (50°C for 30 minutes, 95°C for 5 minutes, followed by PCR cycling at 95°C for 30 seconds, 55°C for 30 seconds, and 72°C for 2 minutes), and 5-μL aliquots were extracted after various numbers of cycles and analyzed by agarose gel electrophoresis.
RESULTS
The human brain microvessel preparation is free of adjoining brain cells (Fig. 1A) and comprises microvascular endothelial cells, pericytes, or smooth muscle cells, which share the basement membrane with the endothelial cell, and astrocyte foot processes, which remain adhered to the basement membrane surface of the isolated brain capillary. The intact capillary, and not purified endothelial cells, was used to generate RNA for the gene microarray because brain capillary function arises from the paracrine interactions between the endothelium, pericyte, and astrocyte foot process.
The human brain capillary tester cDNA was subtracted with driver cDNA obtained from human liver and kidney RNA. Poly(A)+ RNA was isolated from these brain microvessels and human kidney and liver tissue, and the presence of full-length mRNA was confirmed by actin Northern blotting (Fig. 1B). The genes commonly expressed between the brain microvasculature and liver and kidney were then subtracted using the SSH process to yield a cDNA library that was enriched for genes expressed at the brain microvasculature. The subtraction efficiency of genes expressed both in tester and driver tissues was determined by PCR analysis of the ubiquitous glyceraldehyde 3-phosphate dehydrogenase (G3PDH) gene, both in the unsubtracted microvasculature cDNA library and in the kidney- and liver-subtracted brain microvasculature-derived cDNA library. The G3PDH gene was effectively subtracted during the SSH process because the PCR product was detectable only after 33 cycles in the subtracted, microvasculature-enriched pool, whereas it appeared after just 18 cycles in the unsubtracted starting material (Fig. 1C). After subcloning the microvasculature-enriched library, the clones were microarrayed in a 96-well format and genes that were enriched compared with the starting material were detected by Southern blotting with subtracted tester cDNA (Fig. 1D), unsubtracted tester cDNA (Fig. 1E), or unsubtracted driver cDNA (Fig. 1F). The final subtracted tester cDNA library comprised 1.2 × 106 clones, 500 of which were arbitrarily selected for Southern blot screening. Of these 500 clones, 71 were enriched during the SSH process and subsequently sequenced. The identified genes are catalogued in Table 1. The clones were designated hLK1 to hLK71 to reflect a human (h) brain microvasculature library subtracted by human liver (L) and kidney (K).
SSH of human brain microvasculature reveals previously characterized and novel brain microvasculature-specific genes
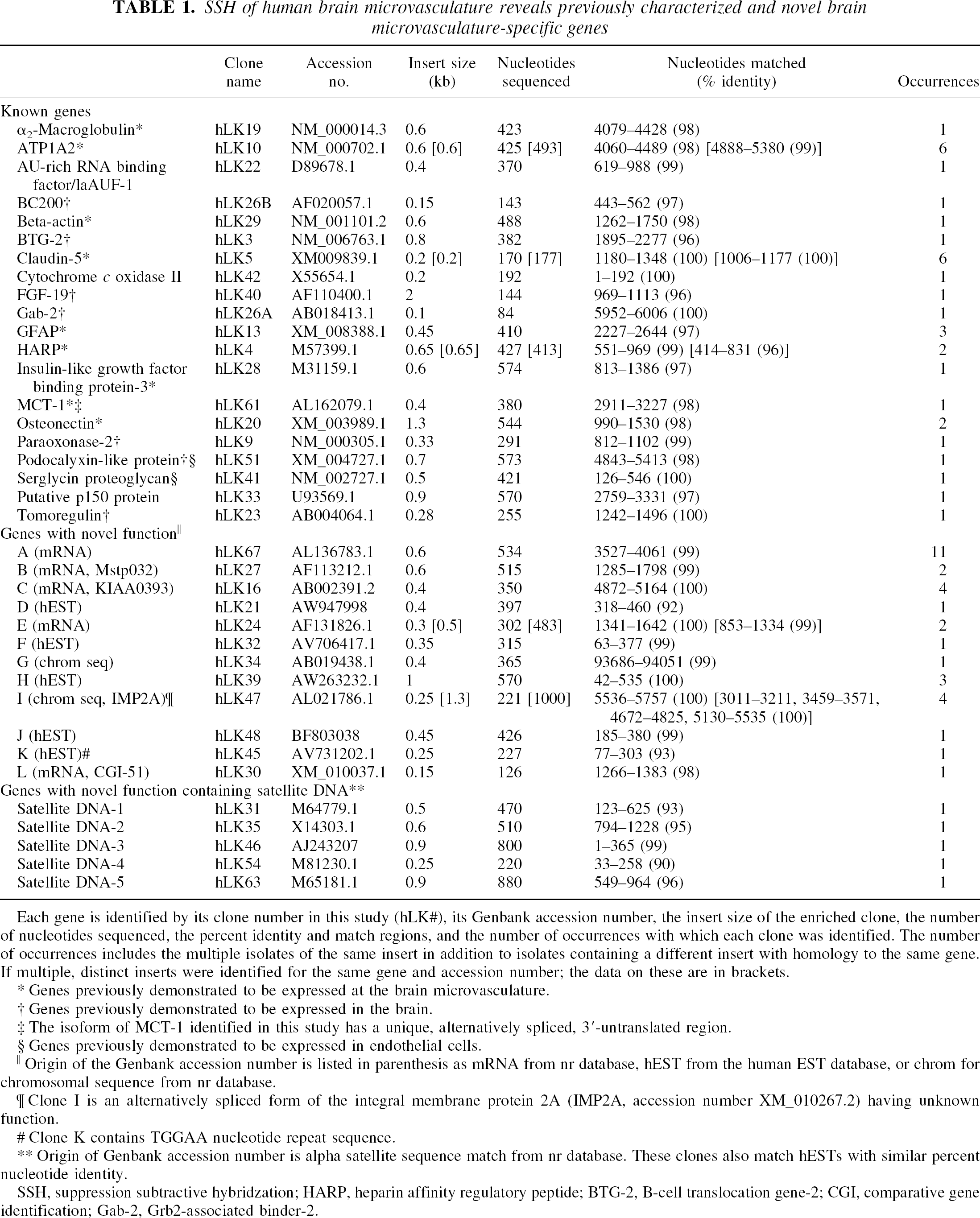
Each gene is identified by its clone number in this study (hLK#), its Genbank accession number, the insert size of the enriched clone, the number of nucleotides sequenced, the percent identity and match regions, and the number of occurrences with which each clone was identified. The number of occurrences includes the multiple isolates of the same insert in addition to isolates containing a different insert with homology to the same gene. If multiple, distinct inserts were identified for the same gene and accession number; the data on these are in brackets.
Genes previously demonstrated to be expressed at the brain microvasculature.
Genes previously demonstrated to be expressed in the brain.
The isoform of MCT-1 identified in this study has a unique, alternatively spliced, 3′-untranslated region.
Genes previously demonstrated to be expressed in endothelial cells.
Origin of the Genbank accession number is listed in parenthesis as mRNA from nr database, hEST from the human EST database, or chrom for chromosomal sequence from nr database.
Clone I is an alternatively spliced form of the integral membrane protein 2A (IMP2A, accession number XM_010267.2) having unknown function.
Clone K contains TGGAA nucleotide repeat sequence.
Origin of Genbank accession number is alpha satellite sequence match from nr database. These clones also match hESTs with similar percent nucleotide identity.
SSH, suppression subtractive hybridzation; HARP, heparin affinity regulatory peptide; BTG-2, B-cell translocation gene-2; CGI, comparative gene identification; Gab-2, Grb2-associated binder-2.
This study identified 20 known genes, 12 genes encoding proteins with an unknown function (designated clones A to L), and 5 genes with significant homology to α satellite DNA-containing elements encoding proteins with an unknown function (Table 1). Many clones were identified multiple times and as different fragments of the same gene coding sequence, and the total number of occurrences of each gene in the pool of 71 is listed in Table 1. The process of identifying brain microvasculature-enriched gene expression was robust because almost all the identified genes with known function have been associated with endothelial tissue or the brain microvasculature (Table 1). Genes such as the adenosine-uridine–rich binding factor, cytochrome c oxidase II, and p150 may be common to many tissues. Although the paraoxonase-2 and podocalyxinlike protein genes are expressed in the brain, they are also expressed in the liver and kidney, respectively. Certain kidney- or liver-expressed genes were enriched through the SSH process, even though the liver and kidney cDNA pools were used as the driver in this subtraction of microvasculature cDNA. This finding is likely a result of a highly focal site of expression of these genes in liver or kidney. The podocalyxinlike protein is expressed in kidney principally at the podocyte processes, which invest the glomerulus, similar to the investment of the brain microvasculature by astrocyte foot processes.
The tissue distributions of the genes encoding proteins with novel function were analyzed by RT-PCR. Messenger RNA from human tissues was used as the template for the RT-PCR reaction, and contaminating PCR signal originating from genomic DNA was removed by treatment with endonuclease. Samples were removed from the RT-PCR reaction after 12, 18, 24, 32, and 40 cycles and analyzed by agarose gel electrophoresis. The selective expression of a PCR product for the human brain microvasculature at 24 PCR cycles and the detection of the same PCR product in peripheral tissues at 32 PCR cycles represents an enrichment of gene expression at the human brain microvasculature of 100- to 200-fold (Fig. 2). Levels of total mRNA were normalized between tissue samples by both absorbance and an oligodT-primed reverse transcription reaction for analysis of the total mRNA product for each tissue. As Fig. 2 illustrates, the levels of mRNA were adequately normalized as determined by the nearly identical RT-PCR profiles of the G3PDH and γ-actin housekeeping genes. The Glut1 glucose transporter gene was amplified as an example of a gene selectively expressed in brain at the microvascular endothelium (Fig. 2). Clone hLK5 was claudin-5, which is a tight junctional protein specific to endothelial cells. Both the Glut1 glucose transporter and claudin-5 exhibit similar patterns of RT-PCR with abundant PCR product found in the brain microvasculature mRNA and lower product found in total brain. The signal in total brain RT-PCR may largely originate from the endothelial contribution in the total brain sample. However, there is little or no claudin-5 or Glut1 RT-PCR product observed in RT-PCR of total peripheral tissue mRNA. Because the Glut1 and claudin-5 vasculature-derived product was seen in total brain RT-PCR, one would also expect that any appreciable expression levels of these genes in peripheral vasculature would also result in RT-PCR product using the total tissue sample. Therefore, based on the tissue distribution revealed by the RT-PCR study, clones A, D, E, H, and the satellite DNA-3 genes were classified as brain microvasculature-specific genes having relatively little expression in peripheral tissues. Novel clones B, C, F, and I were classified as brain microvasculature-enriched genes because these were expressed in some peripheral tissues but were comparatively enriched at the brain microvasculature. Clones G and J did not show a significant increase in expression at the brain microvasculature relative to peripheral tissues. With the exception of clone C, none of the novel genes yielded protein homologies on a protein database search and their function remains unknown. Clone C has a high amino acid homology (95%) with the HERC2 protein, yet appears to be distinct and displays high homology (100%) to another protein of unknown function, designated KIAA0393 (Table 1). This protein has a 3′ untranslated region in the mRNA that is different from HERC2, which functions as a guanine nucleotide-exchange factor involved in protein trafficking. Mutations HERC2 can lead to neuromuscular and spermiogenic abnormalities (Ji et al., 1999).
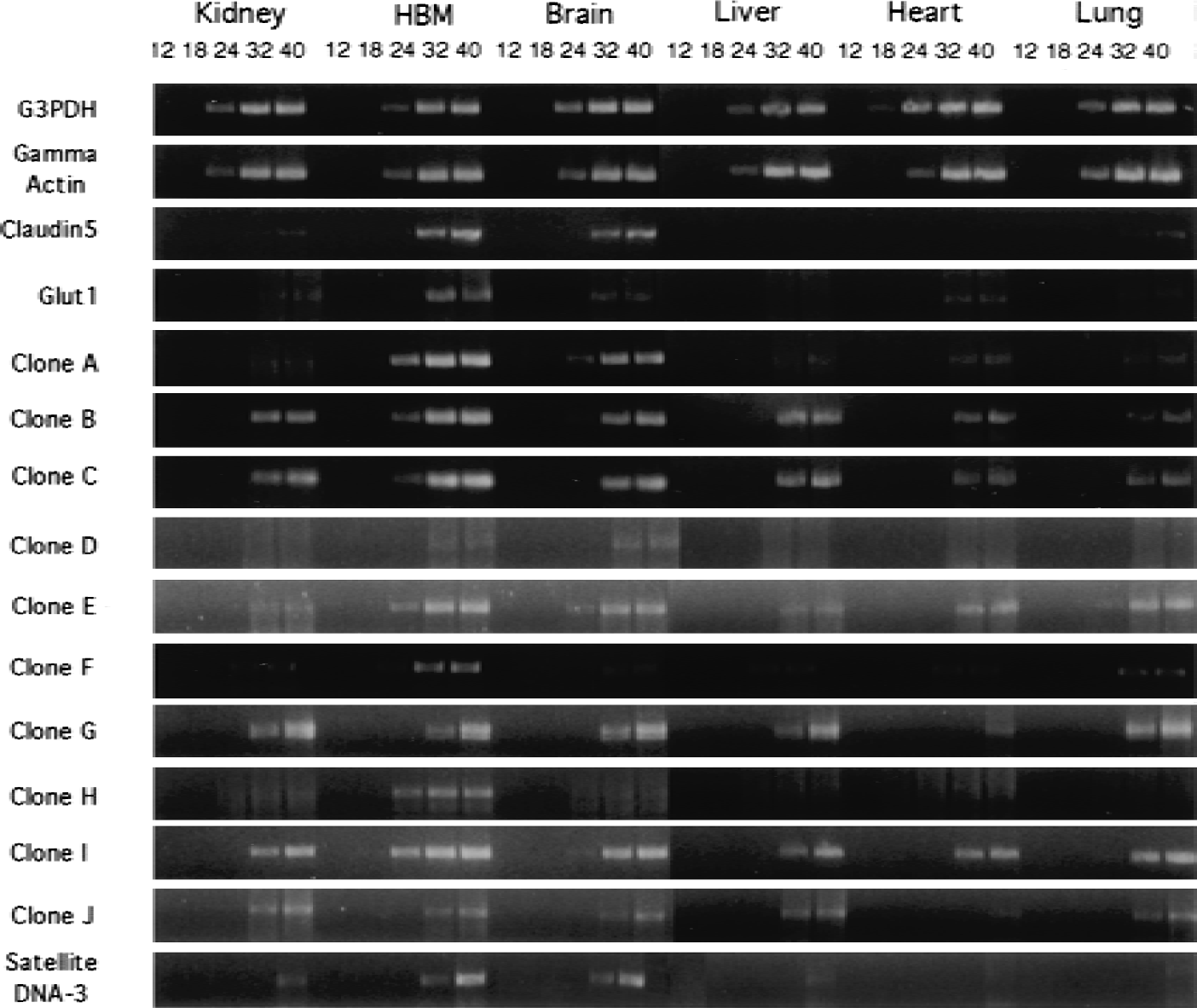

Tissue distribution of gene expression for those genes encoding proteins with unknown function. The RT-PCR reactions were performed with gene-specific primers and a 10-ng human mRNA template. The mRNA concentration was normalized between tissues using absorbance (A260) measurements and by performing an oligodT18-primed reverse transcription reaction using 32P-dCTP to analyze total full-length mRNA content. Normalization was confirmed by similar tissue distribution of G3PDH and γ-actin RT-PCR products. The samples were treated with DNasel to remove any contaminating genomic DNA and the absence of genomic DNA was confirmed with control PCR reactions that did not contain reverse transcriptase, but only Taq polymerase. Aliquots of PCR product were removed after 12, 18, 24, 32, and 40 cycles and analyzed by 2% agarose gel electrophoresis and ethidium bromide staining.
A distinct class of genes of novel function also contained satellite repeat sequences (Table 1). The satellite class of DNA repeats comprises 3% of the human genome and consists of perfect or slightly imperfect tandem repeats of the satellite sequence (Lander et al., 2001). The satellite sequences identified in this study are referred to as α satellite repeats. These sequences have a 171-bp repeat unit and are generally located near centromeric regions. Some of the regions in this DNA class are actively transcribed and code for a human deubiquitinating enzyme (Saitoh et al., 2000), active ribozyme motifs in cave crickets (Rojas et al., 2000), and in the 5′ untranslated region of a human G-protein α subunit (Habecker et al., 1993). An RT-PCR analysis with one of the satellite DNA clones, satellite DNA-3, indicated that the clone was of RNA origin and was brain microvasculature specific (Fig. 2). All of the satellite DNA clones matched hEST database entries and α-satellite stretches of DNA sequence (Table 1). These findings provide additional evidence that some repetitive DNA sequences in the human genome are transcribed in a tissue-specific pattern, and may have functional advantages for the organism.
DISCUSSION
The brain microvascular-enriched genes of known function that were detected in this study can be functionally grouped to illustrate the multiple roles that the brain microvasculature plays in the development and maintenance of the human brain. In the current study, these gene groupings detected for the human brain microvasculature were compared with brain microvascular-enriched genes detected previously from rat brain microvasculature (Li et al., 2001), and the combined results are shown in Fig. 3. Several genes participate in the regulation of the endothelial tight junction and cytoskeleton. Claudin-5 is a transmembrane protein that is a component of the tight junctions of endothelial cells, and is located in the fetal mouse brain exclusively in the microvasculature (Morita et al., 1999). The podocalyxinlike protein is a membrane glycoprotein that has also been identified on the luminal surface of high endothelial venules (Sassetti et al., 1998). Glial fibrillary acidic protein is a neurofilament protein in astrocyte foot processes, and the finding of its mRNA in the brain microvascular preparation suggests that certain astrocyte transcripts are transported from the cell body to the foot process investing the microvasculature. Serglycin is a proteoglycan synthesized by endothelial cells, is located in cytoplasmic vesicles, and is constitutively secreted from the endothelium where it interacts with the extracellular matrix (Kulseth et al., 1999). The extracellular ground substance of brain is altered in neurogenesis and is controlled by protease action in the brain interstitium. Several proteases or cytokines are inhibited by binding to α2-macroglobulin, which is synthesized at the brain microvasculature (Mollgard et al., 1988). Paraoxonase-2 is another lipoprotein-related gene product, is found in the brain, and has been associated with atherosclerosis of the vasculature (Hegele et al., 1999).
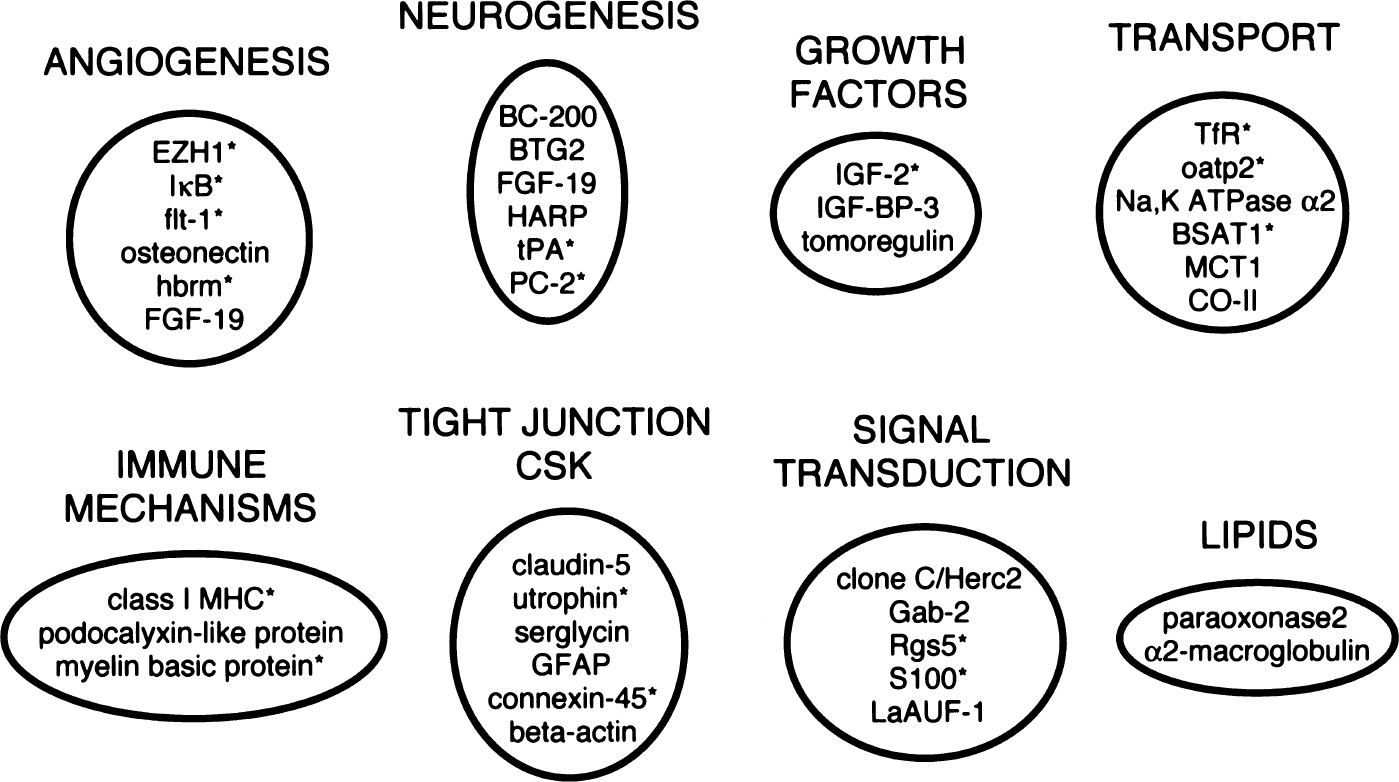
Clustering of brain microvascular gene products by function. Gene grouping is a composite of genes with known function isolated in the present study for the human brain microvasculature, and from a previous study of the rat brain microvasculature (Li et al., 2001). An asterisk (*) notes the brain microvascular-enriched gene products detected in the rat.
Genes encoding for nutrient or peptide transporters are selectively expressed at the brain microvasculature (Fig. 3). An ion transporter, ATPase subunit α2, is a monomer in a family of α and β subunits that form heterodimers to catalyze the active transport of Na+ and K+ across the plasma membrane, and is enriched in the microvasculature compared with brain tissue (Zlokovic et al., 1993). A nutrient transporter, monocarboxylate transporter-1, mediates blood-brain barrier transport of lactic acid and the ketone bodies, which are energy substrates in the developing brain (Leino et al., 1999). Transport at the brain microvasculature is likely regulated by signal transduction phenomena mediated by brain capillary-enriched genes. Gab-2 is a human adaptor molecule with prominent brain expression (Zhao et al., 1999) that undergoes tyrosine phosphorylation by different inducers including the receptor tyrosine kinase f1t3, which plays a role in endothelial proliferation (Zhang and Broxmeyer, 2000).
The brain microvascular endothelium expresses factors implicated in the induction and regulation of both angiogenesis and neurogenesis. HARP, a member of the heparin-binding growth factor family that is distinct from the fibroblast growth factor family, promotes neurite outgrowth and angiogenesis (Souttou et al., 2001), and is expressed by brain endothelial cells grown in tissue culture (Papadimitriou et al., 2000). FGF-19 is a member of the fibroblast growth factor family of cytokines with angiogenic and neurotrophic effects and is selectively increased in the fetal brain (Nishimura et al., 1999). Tomoregulin, a member of the epidermal growth factor/neuregulin family, stimulates the phosphorylation of the erbB-4 receptor, suggesting roles in the normal development and maintenance of the brain (Uchida et al., 1999). Insulin-like growth factor binding protein-3 is a carrier for insulinlike growth factors and releases the growth factor by plasmin-mediated proteolysis at the endothelial cell surface (Booth et al., 1999). In addition to cytokines or growth factors that stimulate growth, antiproliferative genes are also expressed at the human brain microvasculature. Osteonectin, a calcium-binding glycoprotein that inhibits angiogenesis by binding vascular endothelial growth factor, is selectively increased in the endothelium of newborn brain or injured adult brain (Kupprion et al., 1998). The p53 inducible gene, PC-3 (also called BTG-2) acts as an antiproliferative protein by causing the downregulation of cyclin D1 levels and impairment of the G1/S cell-cycle transition (Guardavaccaro et al., 2000). These findings outline a central role for the microvasculature in both the development of the blood vessel network of the brain and in the promotion and regulation of growth and division of neuroglial cells.
Neurogenesis may also be regulated by BC-200, which is clone hLK26B (Table 1). BC-200 is a small cytoplasmic RNA molecule that is specifically expressed in brain of humans and primates. BC-200 RNA lacks a 5′ poly(A) tail, but was cloned using the SSH methodology because of the presence of a long stretch of adenosine residues in the central portion of the RNA transcript. The BC-200 RNA is enriched in neuronal dendrites and forms part of the 11.4S ribonucleoprotein particle (Cheng et al., 1997). A prominent part of this study is the finding that several clones isolated from a subtracted human brain microvascular cDNA library contain satellite DNA sequences (Table 1). The tissue specificity of one of these clones, satellite DNA-3 (Table 1), was examined with RT-PCR. This clone was selectively expressed at the human brain microvasculature (Fig. 2). Unlike BC-200, the satellite DNA-3 clone lacked a poly(A) region, inferring that this sequence was part of a larger open-reading frame. Repeat-containing DNA sequences also include the class of transposon-derived long-interspersed nuclear elements. Clone hLK33 matches the p150 protein (Table 1), which is derived from a long-interspersed nuclear element that contains two open-reading frames. One open-reading frame encodes for a 40-kd nucleic acid-binding protein, and one encodes for the p150 protein, which has both reverse transcriptase and endonuclease activity. Some long-interspersed nuclear elements are transcriptionally active and can retrotranspose into chromosomal DNA (Sassaman et al., 1997). The expression of multiple satellite DNA sequences at the brain microvasculature appears to be a special characteristic of humans, because previous work on the genomics of the rat brain microvasculature led to the detection of no satellite DNA sequences expressed in the rat (Li et al., 2001).
The vascular genomics of the human brain reveals panels of known genes, genes of unknown function, and satellite DNA-containing sequences (Table 1). This work can be extended in at least three directions. First, the function of the proteins with unknown characteristics that are selectively expressed at the human brain microvasculature (clones A, D, E, and H, Table 1) needs to be determined because these gene products could be novel conduits for drug or gene targeting to the human brain (Pardridge, 2001). Second, the subtracted library should be screened for additional known genes that have not previously been shown to play a prominent role in the regulation of the human brain microvasculature, so that the functional template in Fig. 3 can be expanded. Such work will reveal the prominent role played by the brain microvasculature in brain function, including the regulation of transport, neurogenesis, and response to disease. Third, the vascular genomics of human brain disorders, such as brain cancer, can be elucidated subsequent to the initial isolation of microvessels from human brain cancer tissue, and this can provide the platform for new strategies of treatment of neurologic disease.
Footnotes
Acknowledgments:
The authors thank Dr. H. V. Vinters of the UCLA Department of Pathology/Neuropathology for review of tissue specimens.
