Abstract
To evaluate the postulated role of extrastriatal D1 receptors in human cognition and psychopathology requires an accurate and reliable method for quantification of these receptors in the living human brain. [11C]NNC 112 is a promising novel radiotracer for positron emission tomography imaging of the D1 receptor. The goal of this study was to develop and evaluate methods to derive D1 receptor parameters in striatal and extrastriatal regions of the human brain with [11C]NNC 112. Six healthy volunteers were studied twice. Two methods of analysis (kinetic and graphical) were applied to 12 regions (neocortical, limbic, and subcortical regions) to derive four outcome measures: total distribution volume, distribution volume ratio, binding potential (BP), and specific-to-nonspecific equilibrium partition coefficient (
The D1 receptors are the most abundant dopaminergic receptor subtype in neocortical areas including the prefrontal cortex (PFC) as well as in the hippocampus and amygdala (for reviews, see Seeman, 1992; Meador-Woodruff et al., 1996). Studies in nonhuman primates have shown that activation of D1 receptors in the PFC is involved in working memory (Arnsten et al., 1994; Sawaguchi and Goldman-Rakic, 1994; Williams and Goldman-Rakic, 1995) and that D1 receptors in the hippocampus are implicated in short- and long-term memory (Huang and Kandel, 1995; Otmakhova and Lisman, 1996; Bernabeu et al., 1997; Seamans et al., 1998). An accurate and reliable method for measuring these receptors in extrastriatal regions of the living human brain will permit the study of their role in mediating cognition in health and disease states.
The radiolabeled benzazepine [11C]SCH 23390 (
Recently, two new benzazepines developed by Novo Nordisk (Malov, Denmark) (Andersen et al., 1992) have been evaluated as PET radiotracers in baboons: [11C]NNC 756 (
The purpose of the present study was to further evaluate [11C]NNC 112 as a radiotracer with which to quantify D1 receptors in the human brain. Specifically, the aim of this study was to define the optimal scanning protocol and analytic method to derive accurate and reliable D1 receptor parameters in striatal and extrastriatal regions. Six healthy volunteers were studied twice, after injection of [11C]NNC 112. Twelve brain regions were studied, ranging from high (caudate and putamen) to low (neocortex, limbic regions, and thalamus) receptor density regions. Two approaches were compared for measurement of D1 receptor parameters: compartmental kinetic and graphical analysis. Four outcome measures were compared: total distribution volume (
MATERIALS AND METHODS
Human subjects
We report here the results of six consecutive weekly experiments. Six healthy volunteers participated in this study (age 33 ± 9 years, range 23 to 45 years; these and subsequent values given as means ± SD; five men and one woman). All subjects were scanned twice on the same day to evaluate the reproducibility of the outcome measures. The study was approved by the Columbia Presbyterian Medical Center Institutional Review Board, and subjects provided written informed consent after receiving an explanation of the study. The absence of pregnancy and medical, neurological, and psychiatric history (including alcohol and drug abuse) was assessed by history, review of systems, physical examination, routine blood tests including pregnancy test, urine toxicology, and electrocardiography.
Radiochemistry
[11C]NNC 112 was prepared by
Positron emission tomography protocol
Subject preparation included placement of arterial and venous catheters, fiducial markers, and polyurethane head immobilizer. An arterial catheter was inserted in the radial artery after completion of the Allen test and infiltration of the skin with 1% lidocaine. A venous catheter was inserted in a forearm vein on the opposite side. Four fiducial markers filled with 11C (∼3 μCi/marker at the time of injection) were glued on the subject's head. Head movement minimization was achieved with a polyurethane head immobilizer system (Soule Medical, FL, U.S.A.) molded around the head of the subject. This system provides better restraint than a thermoplastic mask (unpublished data). The PET imaging was performed with the ECAT EXACT 47 (Siemens/CTI, Knoxville, TN, U.S.A.; 47 slices covering an axial field of view of 16.2 cm, axial sampling of 3.375 mm, three-dimensional mode in-plane and axial resolution of 6.0 and 4.6 mm full width at half-maximum at the center of the field of view, respectively) (Wienhard et al., 1992). A 10-minute transmission scan was obtained before radiotracer injection. [11C]NNC 112 was injected intravenously over 45 seconds. Emission data were collected in the three-dimensional mode for 120 minutes as 21 successive frames of increasing duration (3 × 20 s, 3 × 1 min, 3 × 2 min, 2 × 5 min, 10 × 10 min). Images were reconstructed to a 128 × 128 matrix (pixel size of 2.5 × 2.5 mm2). Reconstruction was performed with attenuation correction using the transmission data and a Sheppe 0.5 filter (cutoff 0.5 cycle/projection rays). Subjects were allowed to rest outside of the camera for 15 to 30 minutes between the two injections.
Input function measurement
After radiotracer injection, arterial samples were collected every 5 seconds with an automated sampling system for the first 2 minutes (Graham and Lewellen, 1993) and manually thereafter at longer intervals. A total of 31 samples were obtained per experiment. After centrifugation (10 minutes at 1,800
Six selected samples (collected at 2, 8, 16, 30, 50, and 70 minutes) were further processed by protein precipitation using acetonitrile followed by high-pressure liquid chromatography to measure the fraction of plasma activity representing unmetabolized parent compound. Plasma samples (0.5 mL) were added to 0.7 mL of acetonitrile in a centrifuge tube. After mixing, the samples were centrifuged at 14,000 rpm for 3.5 minutes. The acetonitrile solution was separated and analyzed by high-pressure liquid chromatography. The system consisted of a Waters 510 isocratic pump, a Rheodyne injector with a 2-mL loop, a Phenomenex C18 ODS column (10-μm particle size, 250 × 4.6 mm, stainless steel), and a gamma detection system (Bioscan Flow Count unit). The column was eluted with a solvent mixture of acetonitrile/0.1 mol/L aqueous ammonium formate (30:70) at a flow rate of 2.5 mL/min. Five fractions collected over 12 minutes were counted. A standard [11C]NNC 112 solution was processed with each experiment. Parent eluted with fractions 3 and 4. For each sample, the fraction parent was estimated by the ratio of decay-corrected activity in collections 3 and 4 to that in the total collection.
A biexponential function was fitted to the six measured fractions parent and used to interpolate values between and after the measurements. The smallest exponential of the fraction parent curve, λpar, was constrained to the difference between λcer, the terminal rate of washout of cerebellar activity, and λtot, the smallest elimination rate constant of the total plasma (Abi-Dargham et al., 1999). The input function was calculated by the product of total counts and interpolated fraction parent at each time. The measured input function values [
For the determination of the plasma free fraction (
Magnetic resonance imaging acquisition and segmentation procedures
Magnetic resonance imaging (MRI) was performed on a GE 1.5 T Signa Advantage system (Milwaukee, WI, U.S.A.). After a sagittal scout (localizer) image performed to identify the anterior-posterior commissural (AC-PC) plane (1 minute), a transaxial T1-weighted sequence with 1.5-mm slice thickness was acquired in a coronal plane orthogonal to the AC-PC plane over the whole brain with the following parameters: three-dimensional SPGR (spoiled gradient-recalled acquisition in the steady state); repetition time 34 milliseconds; echo time 5 milliseconds; flip angle 45°; slice thickness 1.5 mm and zero gap; 124 slices; field of view 22 × 16 cm; 256 × 192 matrix, reformatted to 256 × 256, yielding a voxel size of 1.5 × 0.9 × 0.9 mm; and time of acquisition 11 minutes.
The MRI segmentation was performed within MEDx (Sensor Systems, Sterling, VA, U.S.A.), with original subroutines implemented in MATLAB (Math Works, Natick, MA, U.S.A.). Steps for MRI segmentation included correction for field inhomogeneities, fitting of the voxel distribution to a combination of three Gaussians functions, voxel classification, and postfiltering. To correct for field inhomogeneity, histograms of one anterior coronal slice and one posterior coronal slice were computed, as described below.
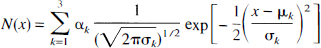
where α
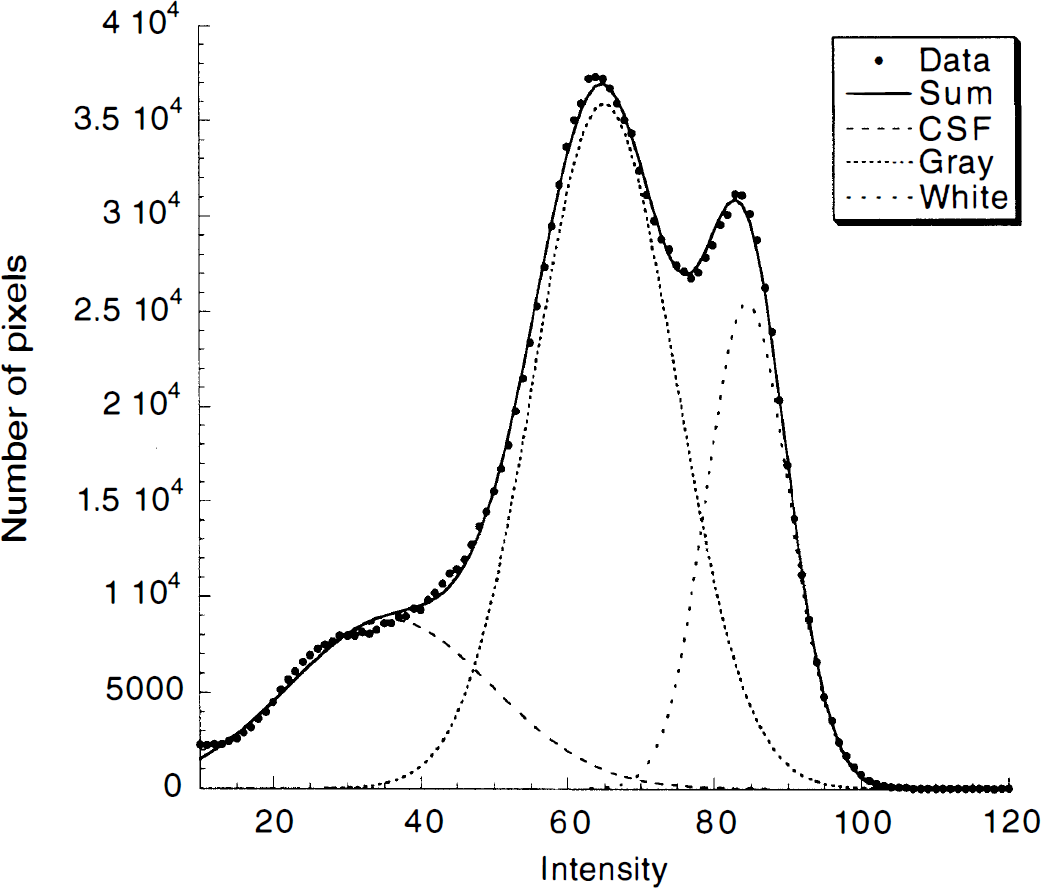
Distribution of voxel intensity from a T1-weighted SPGR (spoiled gradient-recalled acquisition in the steady state) magnetic resonance imaging file and fitted sum of three Gaussians estimating voxel intensity of cerebrospinal fluid and gray and white matter. The solid line represents the sum of the fitted Gaussians. The intersection between the Gaussians was used as threshold for pixel classification.
Image analysis
Image analysis was performed with MEDx according to the following steps: frame realignment, PET-MRI registration, and time-activity curve measurement.
1. Frame realignment. To correct for head movement during the acquisition, all frames were coregistered to the first frame of the study, using a least-squares algorithm for within-modality coregistration (automated image registration) (Woods et al., 1992). Images were thresholded to the level of the 11C fiducial markers, and only the signals from the markers were used for automated image frame-to-frame registration. The
2. PET-MRI registration. After frame-to-frame registration, the 21 frames were summed, and the summed PET image was coregistered and resampled to the MR image, using automated image registration (Woods et al., 1992). The summed PET image was used for the coregistration because it contains counts from the initial flow-dependent activity distributions that enhance detection of boundaries of regions with low receptor density, such as the cerebellum. The parameters of the spatial transformation matrix of the summed PET data set were then applied to each individual frame. Thus, each PET frame was resampled in the coronal plane to a voxel volume of 1.5 × 0.9 × 0.9 mm3.
3. Region-of-interest (ROI) tracing and activity sampling. The ROI boundaries were drawn on the MR image according to criteria based on brain atlases (Talairach and Tournoux, 1988; Duvernoy, 1991) and on published reports (Pani et al., 1990; Kates et al., 1997; Killiany et al., 1997). Neocortical regions included dorsolateral prefrontal cortex (DLPFC), medial prefrontal cortex, orbitofrontal cortex (OFC), anterior cingulate cortex, parietal cortex, temporal cortex, and occipital cortex. Subcortical regions included caudate, putamen, thalamus, amygdala, hippocampus, and cerebellum. Two methods were used for final ROI definition. A segmentation-based method was used for neocortical regions, and a direct identification method was used for subcortical regions. For neocortical regions, “large” regions were first drawn to delineate the boundaries of the ROIs. Within these regions, only the voxels classified as GM were used to measure activity distribution. This process is illustrated in Fig. 2 for DLPFC and OFC. Because of the mixture of GM and WM in central gray structures (especially thalamus), the segmentation-based approach was not used to define subcortical ROIs, and the boundaries of these regions were identified by anatomical criteria.
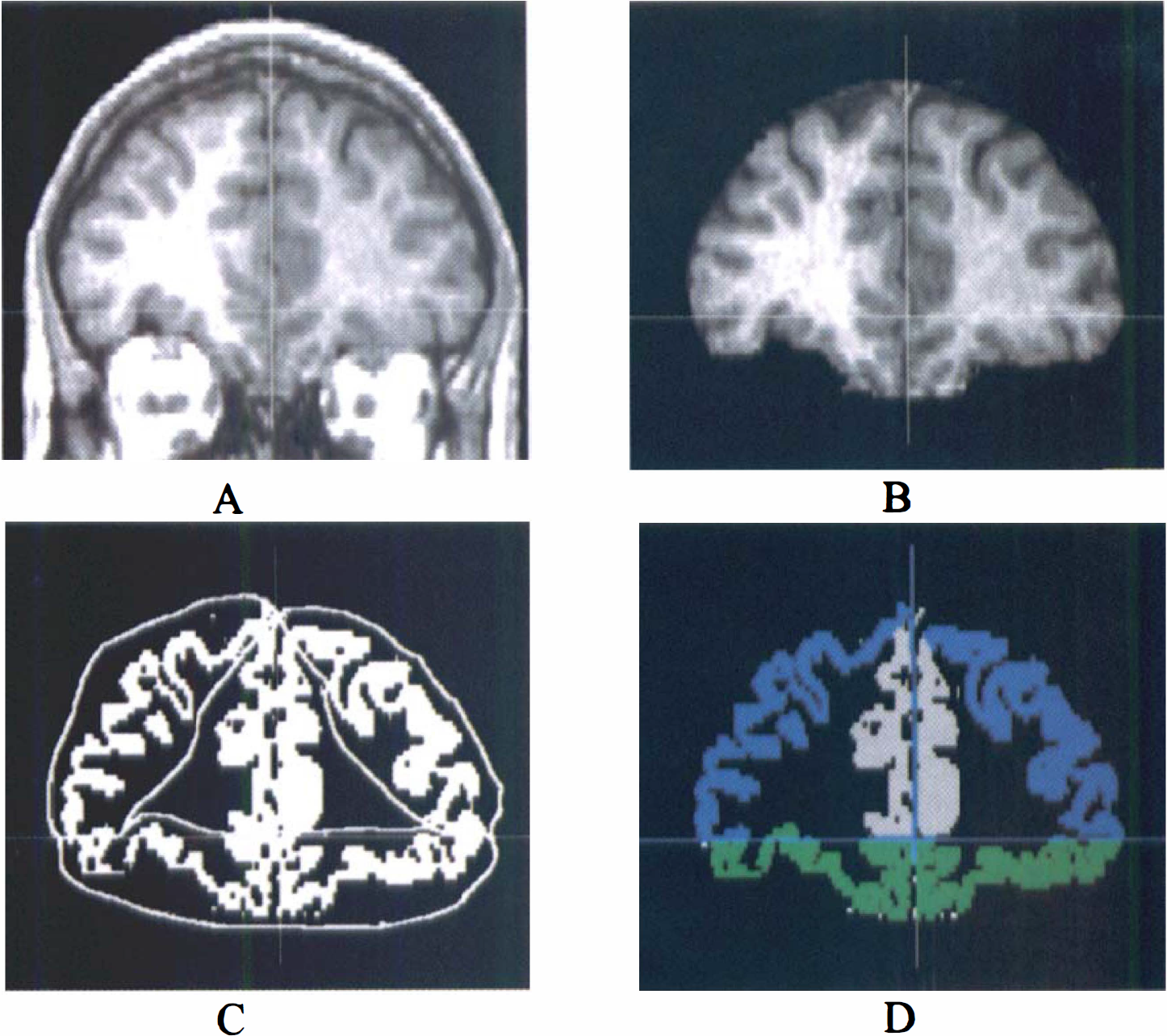
The process of region-of-interest (ROI) definition for dorsolateral prefrontal cortex (DLPFC) and orbitofrontal cortex (OFC) in one coronal plane.
Quantitative analysis
Derivation of [11C]NNC 112 regional distribution volumes was performed using kinetic and graphical analyses. A three-compartment model (i.e., two-tissue compartment model) provided the general framework for each method. The model included the arterial plasma compartment (
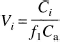
Kinetic analysis
Kinetic analysis was performed according to the differential equations:
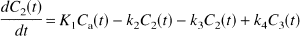
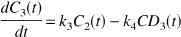
with the kinetic parameters defined as
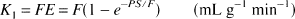
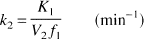
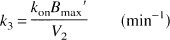

where
In the ROIs, volumes of distribution were then calculated according to
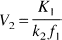
and
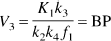
Substituting in Eq. 10 the terms
The BP, as defined above, is the only expression of BP that is dependent on receptor parameters (i.e.,
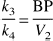
The ratio of total to nonspecific distribution volume is termed the DVR:
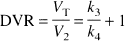
DVR is reported to facilitate comparison with results of studies reporting reproducibility of [11C]raclopride DVR (Volkow et al., 1993), [18F]altanserin DVR (Smith et al., 1998), and [11C]SCH 23390 DVR (Chan et al., 1998).
Results from kinetic analysis were evaluated with four criteria: identifiability, validity, stability, and reliability.
Identifiability. The term “identifiability” refers to the error in the parameter estimation due to the inherent uncertainty of this estimation. The identifiability was assessed by the standard error of the parameter at convergence. The standard error of the parameters was given by the diagonal of the covariance matrix (Carson, 1986) and expressed as percentage of the parameters (coefficient of variation; %CV). This standard error should not be confused with the variance of the parameter in the investigated population (between-subject SD or %CV) or with the lack of reproducibility (within-subject SD or %CV). It should also be noted that uniform weighting of the least-squares estimation might be associated with an overestimation of the standard error of the parameters if the measurement variances range over orders of magnitude (Carson, 1986). Validity. Physiological plausibility was assessed by comparing regional BP values with distribution of D1 receptors in the human brain Stability. Experimental data were collected for 120 minutes. The relationship between BP derivation and the duration of the scan was evaluated by fitting shorter-duration data sets (110, 100, 90, 80, 70, 60, 50, 40, and 30 minutes) and comparing the results with the reference value obtained with the 120-minute data set. For each region and each duration, the averages ± SD (n = 12) of the results expressed as percentage of the reference value were calculated to provide an estimate of the bias (average) or dispersion (SD) induced in the outcome measure by analyzing shorter data sets. The solution was considered stable after time Reliability. The test/retest variability was calculated as the absolute value of the difference between test and retest values, expressed as percentage of the mean value of both measurements. To evaluate the within-subject variability relative to the between-subject variability, both within-subject SD and between-subject SD were calculated and expressed as percentage of mean value (within-subject %CV and between-subject %CV). The reliability of the measurements was assessed by the intraclass correlation coefficient (ICC) calculated as follows (Kirk, 1982):
where BSMSS is the mean sum of squares between subjects, WSMSS is the mean sum of squares within subjects, and n is the number of repeated observations (n = 2 in this study). This coefficient estimates the reliability of the measurement and assumes values from −1 (no reliability; i.e., BSMSS = 0) to 1 (maximum reliability, achieved in case of identity between test and retest; i.e., WSMSS = 0).
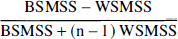
Graphical analysis
Regional time-activity curves were graphically analyzed according to the equation
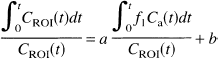
where the values of the slope
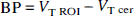
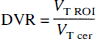
and
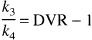
Reliability of the graphical analysis was evaluated as described for the kinetic analysis.
Noise analysis
Simulations were performed to evaluate the sensitivity of the kinetic and graphical analysis to experimental noise. This analysis was carried out for the cerebellum and one ROI (caudate). First, noise-free data [
Statistical analysis
For each outcome measure and each subject (n = 6), the average of test and retest values was calculated, and results are given as means ± SD of these six average measurements. Thus, unless otherwise specified, SD refers to between-subject SD, that is, to the estimated variability in the investigated population (each subject counts only once, n = 6). When the SD refers to variability between the experiments rather than between subjects (such as for the injected dose), the SD is followed by n = 12. Statistical analysis was performed with repeated-measures analysis of variance. Relationships between continuous variables were analyzed with the Pearson product-moment correlation coefficient. A two-tailed probability value of 0.05 was selected as the significance level.
RESULTS
Plasma analysis
[11C]NNC 112 underwent rapid metabolism measured in human plasma (Fig. 3). At 30 minutes, only 18 ± 6% (n = 12) of the total activity corresponded to the parent compound. The six measures (2, 8, 16, 30, 50, and 70 minutes) of fraction parent were fit to a sum of two exponentials. In all cases, the unconstrained fit of the fraction parent curve yielded a value of 0 for the smallest exponent (λpar). This solution was consistent with the low rate of metabolism after 30 minutes and the absence of data after 70 minutes. However, this solution was not consistent with the observation of a faster washout rate in the cerebellum compared with the total plasma activity after 30 minutes. This problem was addressed by constraining λpar to the difference between λcer, the terminal rate of washout of cerebellar activity, and λtot, the smallest elimination rate constant of the total plasma. For each experiment, the terminal cerebellar washout rates were calculated by fitting the cerebellum data from 30 to 120 minutes to a one-exponential function (λcer = −0.00764 ± 0.00287 min−1, n = 12). Similarly, the smallest exponential of the total plasma activity was derived by fitting the total plasma activity from 30 to 120 minutes to one exponential (λtot = −0.00322 ± 0.00083 min−1, n = 12). The smallest exponential of the fraction parent curve was constrained to the difference between the two functions λcer and λtot (−0.00450 ± 0.00296 min−1, n = 12), and this fit was used to correct total activity by the fraction parent.
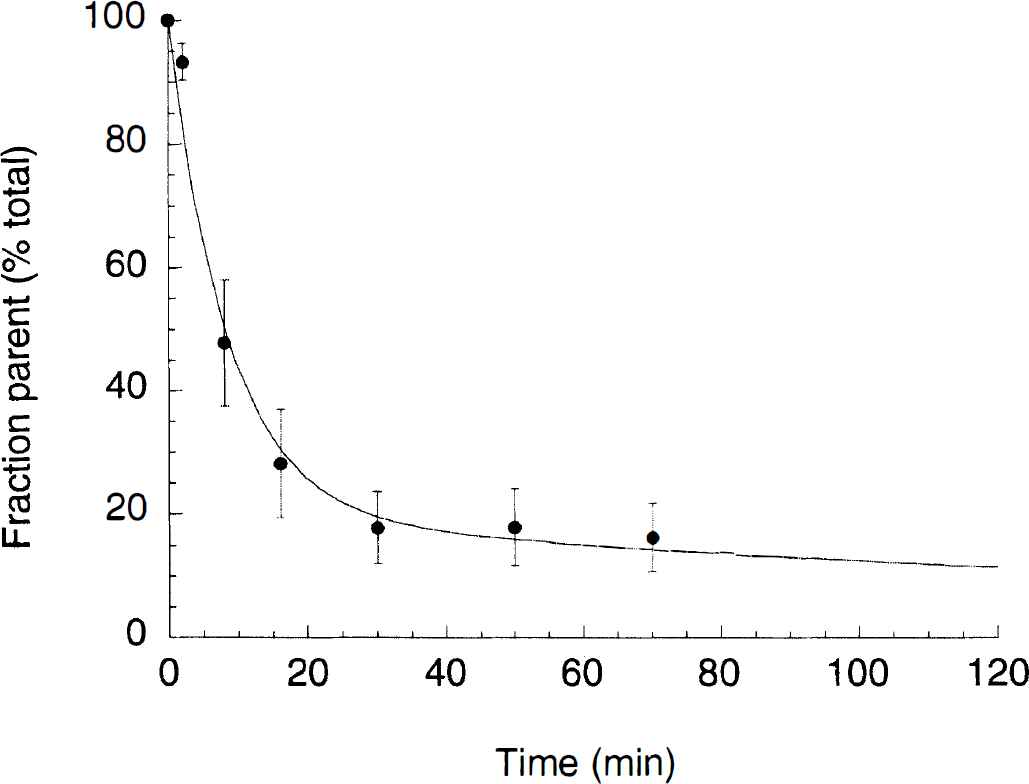
Fraction of plasma activity associated with parent compound (% total) following [11C]NNC 112 injection in humans. Each point is the average ± SD of 12 measurements (6 subjects studied twice). The solid line represents the sum of two exponentials, with second half-life constrained to −0.00440 min−1. This value corresponded to the average value of the difference between cerebellum and total plasma terminal half-lives.
The input function was fit to a sum of three exponentials (Fig. 4), and fitted values were used as input function for kinetic and graphical analysis. The average [11C]NNC 112 plasma clearance was 96 ± 31 L/h. Significant between-subject differences in plasma clearance were detected (
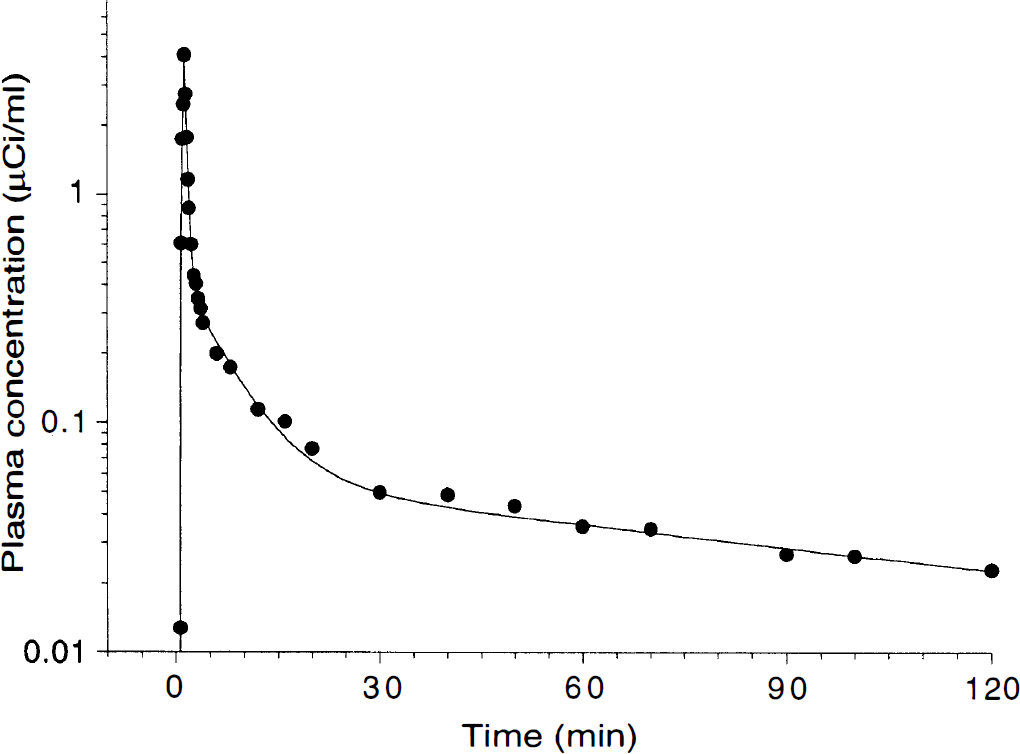
Plasma concentration of [11C]NNC 112 following injection of 18 mCi in one healthy volunteer. Points represent measured values. The solid line represents the result of a three-exponential fit to the measured values. Clearance was 100 L/h.
The mean plasma
Brain uptake
Highest brain regional uptake was observed in caudate and putamen, and lowest uptake was observed in the cerebellum and thalamus (Fig. 5). In other regions (neocortex and limbic system), activity levels were intermediate between striatal and cerebellar regions. Activity peaked early in the cerebellum, neocortical, and limbic regions (<15 minutes) and later in striatum (20-40 minutes). Appreciable washout was observed in all regions, suggesting that the association and dissociation rates were fast enough to allow derivation of D1 receptor BP within the time frame of the scan. At 60 minutes, the ROI/cerebellar activity ratios were as follows: putamen, 3.72 ± 0.58; caudate, 3.26 ± 0.58; amygdala, 1.77 ± 0.32; hippocampus, 1.48 ± 0.23; anterior cingulate, 1.78 ± 0.29; DLPFC, 1.53 ± 0.20; temporal cortex, 1.69 ± 0.33; occipital cortex, 1.30 ± 0.17; and thalamus, 1.19 ± 0.21.
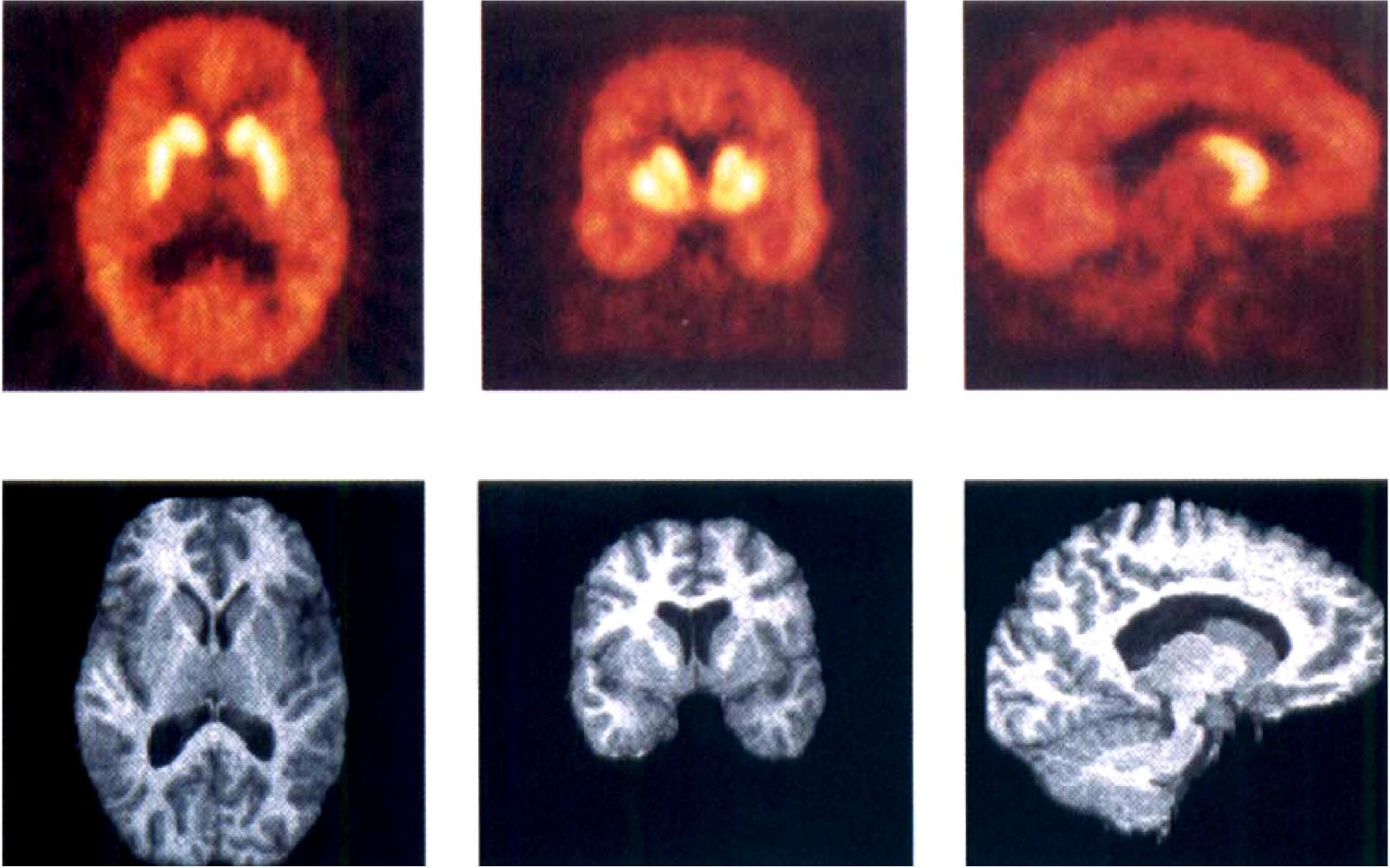
[11C]NNC 112 in a human volunteer. (
Kinetic analysis
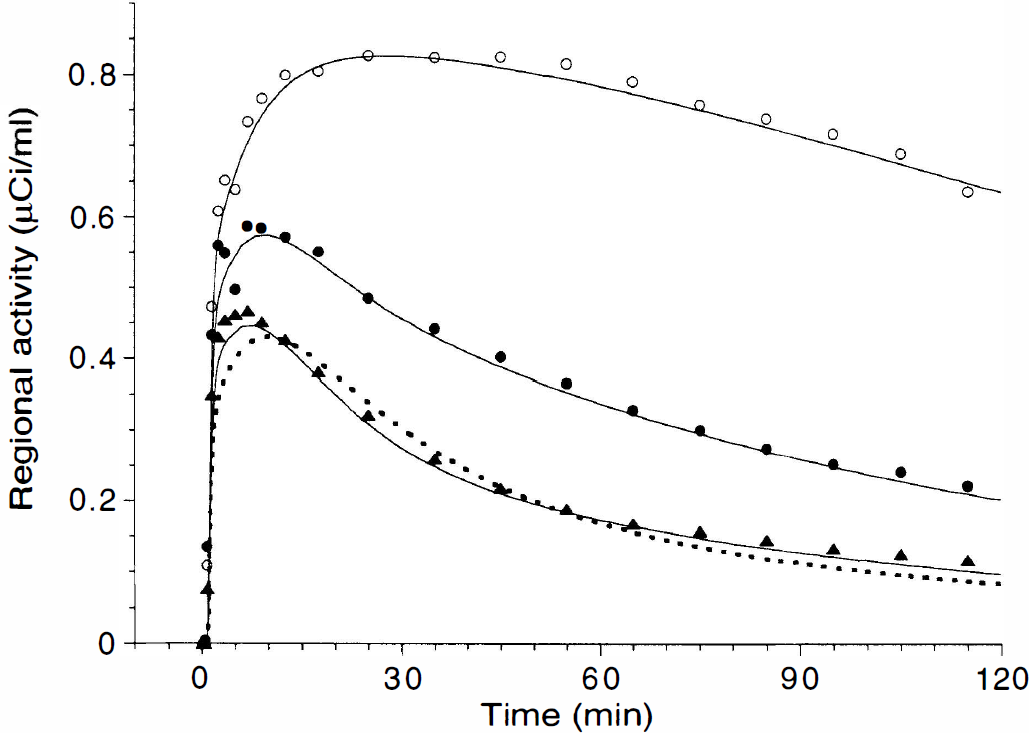
Brain time-activity curves measured in putamen (open circles), dorsolateral prefrontal cortex (filled circles), and cerebellum (triangles) after injection of 18 mCi of [11C]NNC 112 (same subject as in Fig. 4). The solid line represents the results of a three-compartment model fit, and the dashed line illustrates a two-compartment model fit in the cerebellum. In the cerebellum, the three-compartment model provided a slightly but significantly better fit.
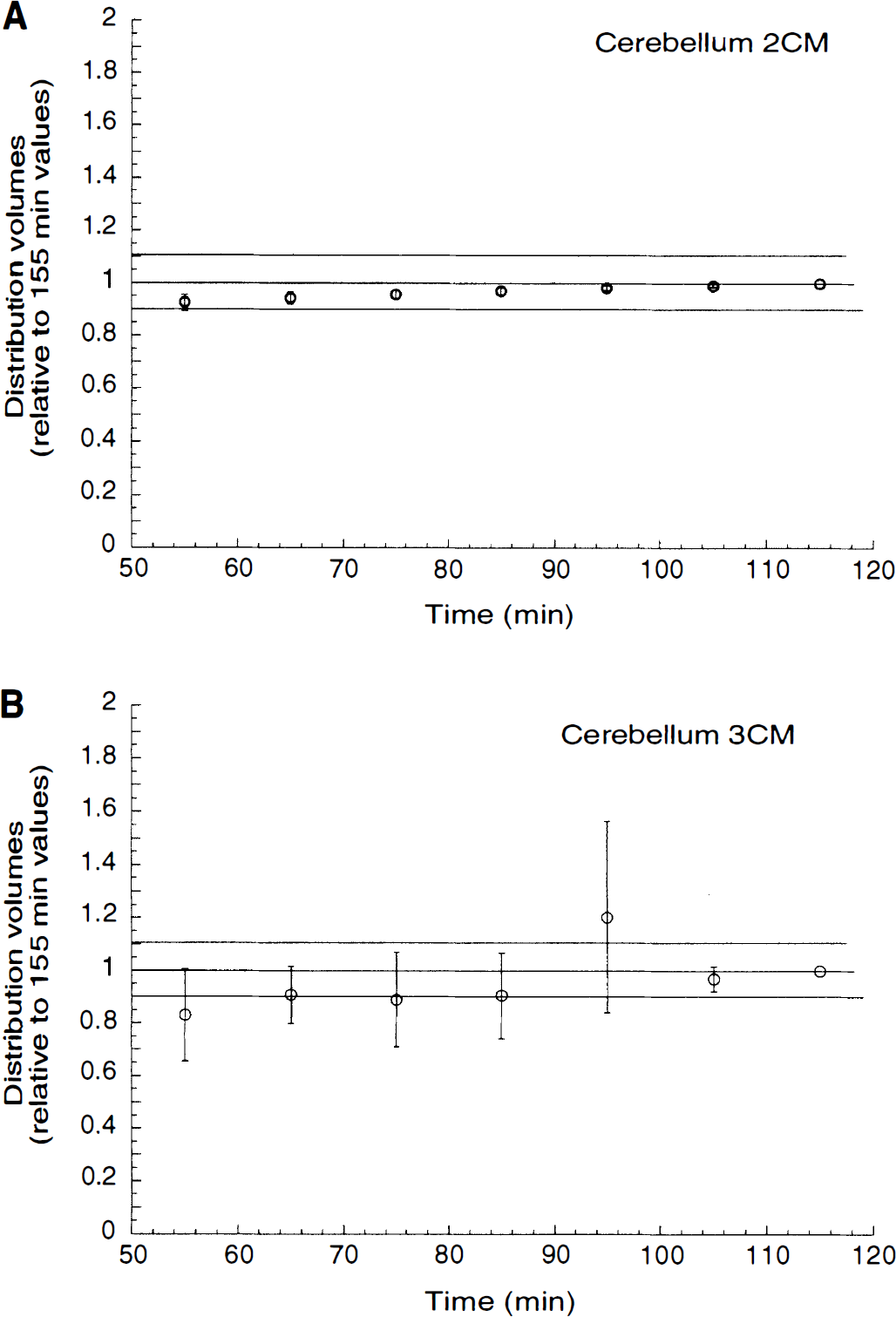
Relationship between duration of data collection and kinetic derivation of total distribution volume (
Kinetic analysis of regional [11C]NNC 112 uptake in human brain (three-compartment model constrained)
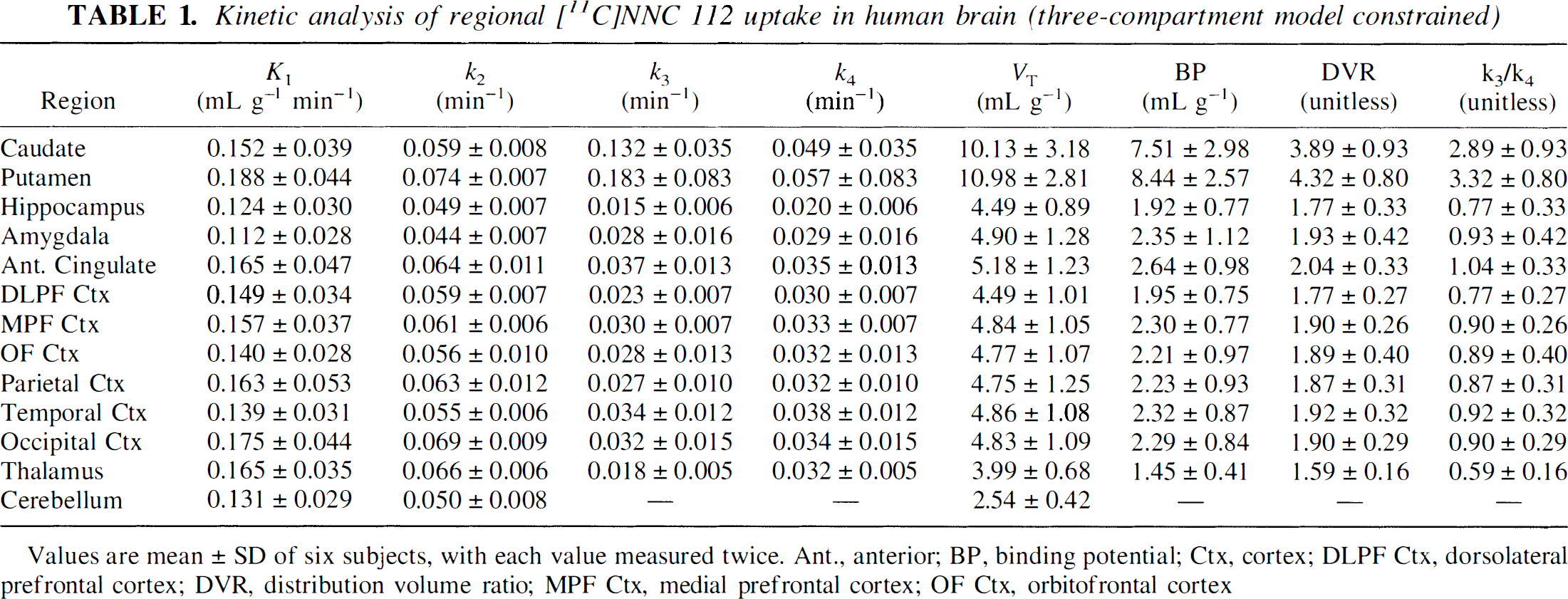
Values are mean ± SD of six subjects, with each value measured twice. Ant., anterior; BP, binding potential; Ctx, cortex; DLPF Ctx, dorsolateral prefrontal cortex; DVR, distribution volume ratio; MPF Ctx, medial prefrontal cortex; OF Ctx, orbitofrontal cortex
Regional distribution of D1 receptors in human brain:
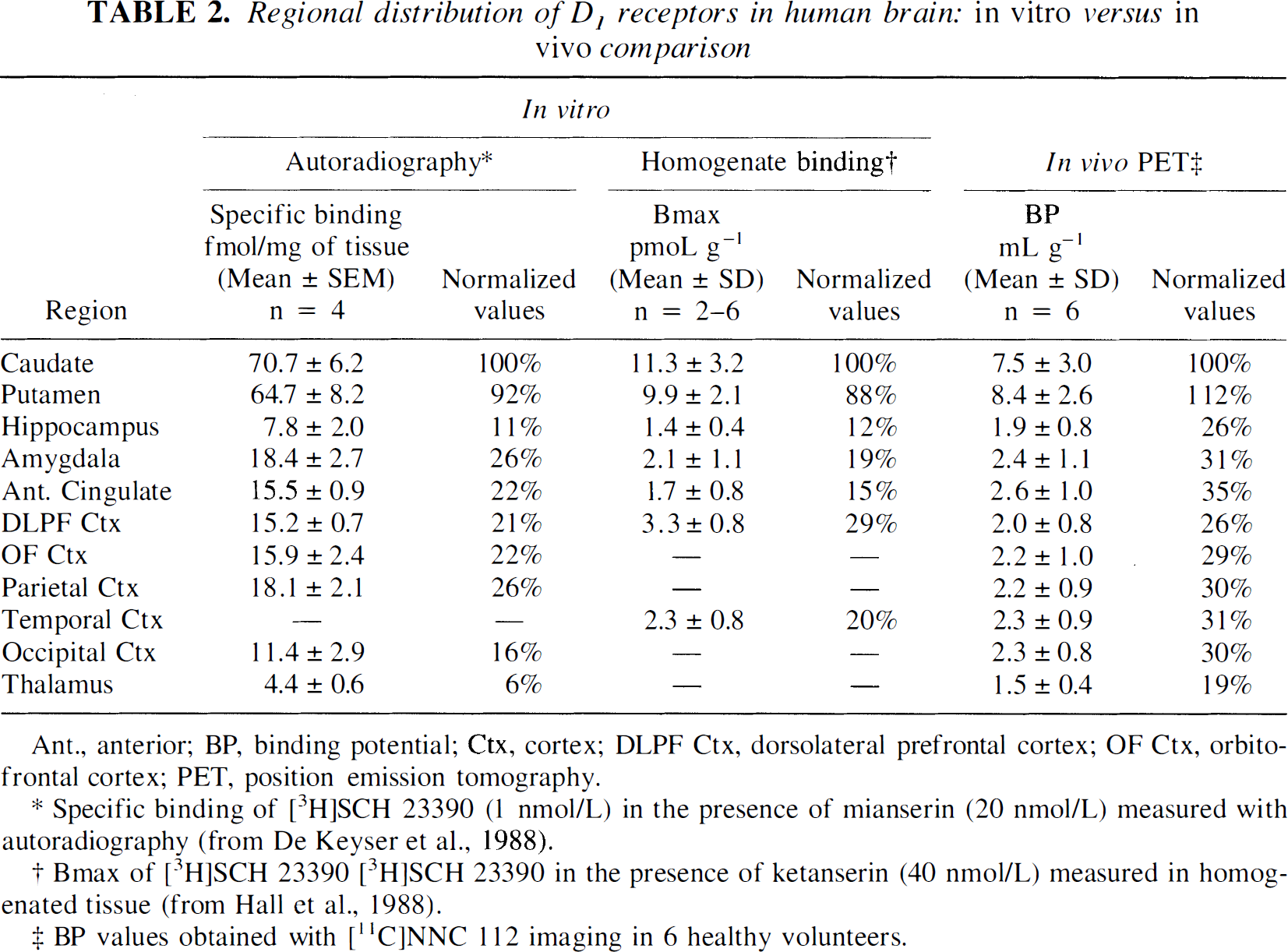
Ant., anterior; BP, binding potential; Ctx, cortex; DLPF Ctx, dorsolateral prefrontal cortex; OF Ctx, orbitofrontal cortex; PET, position emission tomography.
Specific binding of [3H]SCH 23390 (1 nmol/L) in the presence of mianserin (20 nmol/L) measured with autoradiography (from De Keyser et al., 1988).
Bmax of [3H]SCH 23390 [3H]SCH 23390 in the presence of ketanserin (40 nmol/L) measured in homogenated tissue (from Hall et al., 1988).
BP values obtained with [11C]NNC 112 imaging in 6 healthy volunteers.
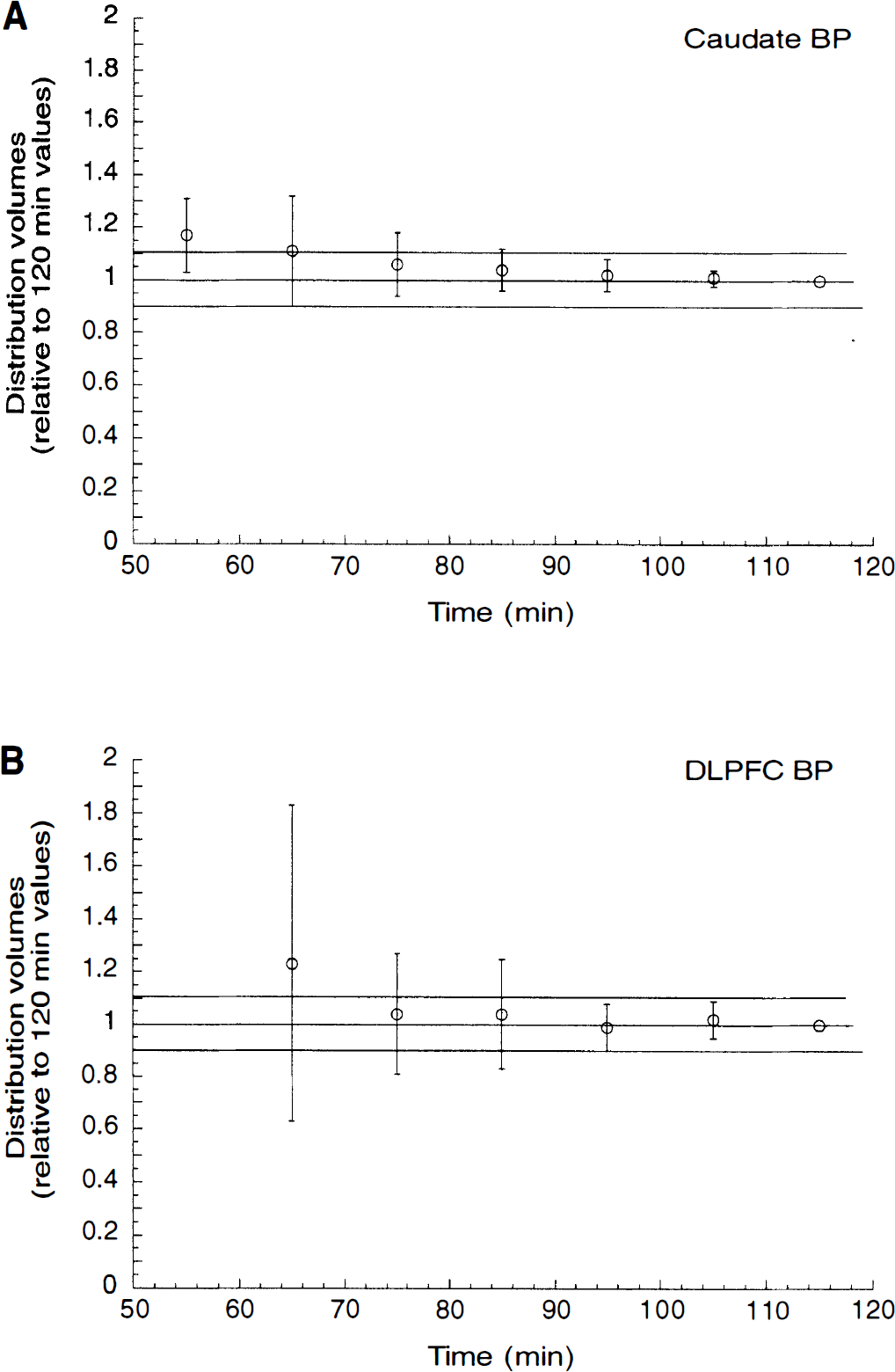
Relationship between duration of data collection and kinetic derivation of binding potential (BP) in the caudate
Reliability of
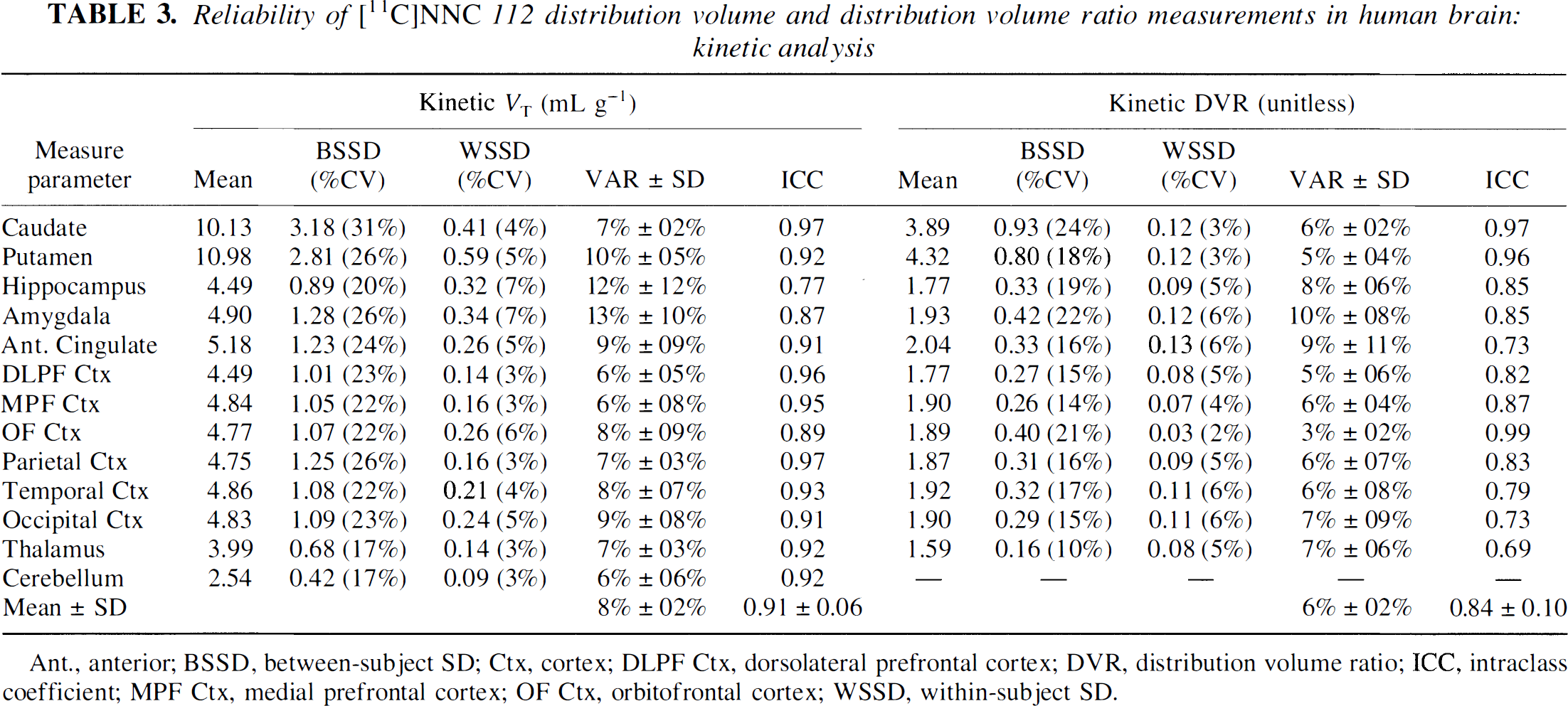
Ant., anterior; BSSD, between-subject SD; Ctx, cortex; DLPF Ctx, dorsolateral prefrontal cortex; DVR, distribution volume ratio; ICC, intraclass coefficient; MPF Ctx, medial prefrontal cortex; OF Ctx, orbitofrontal cortex; WSSD, within-subject SD.
Reliability of [11C]NNC 112 BP and
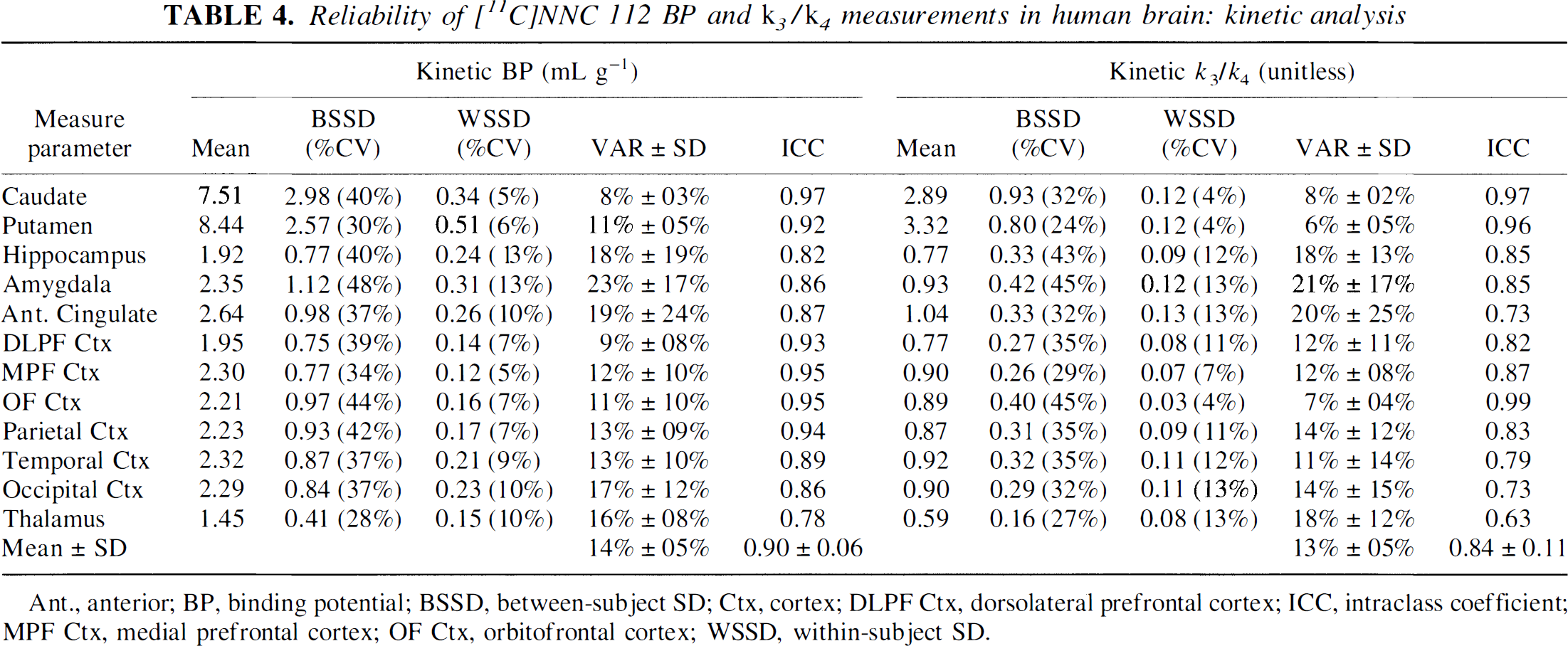
Ant., anterior; BP, binding potential; BSSD, between-subject SD; Ctx, cortex; DLPF Ctx, dorsolateral prefrontal cortex; ICC, intraclass coefficient; MPF Ctx, medial prefrontal cortex; OF Ctx, orbitofrontal cortex; WSSD, within-subject SD.
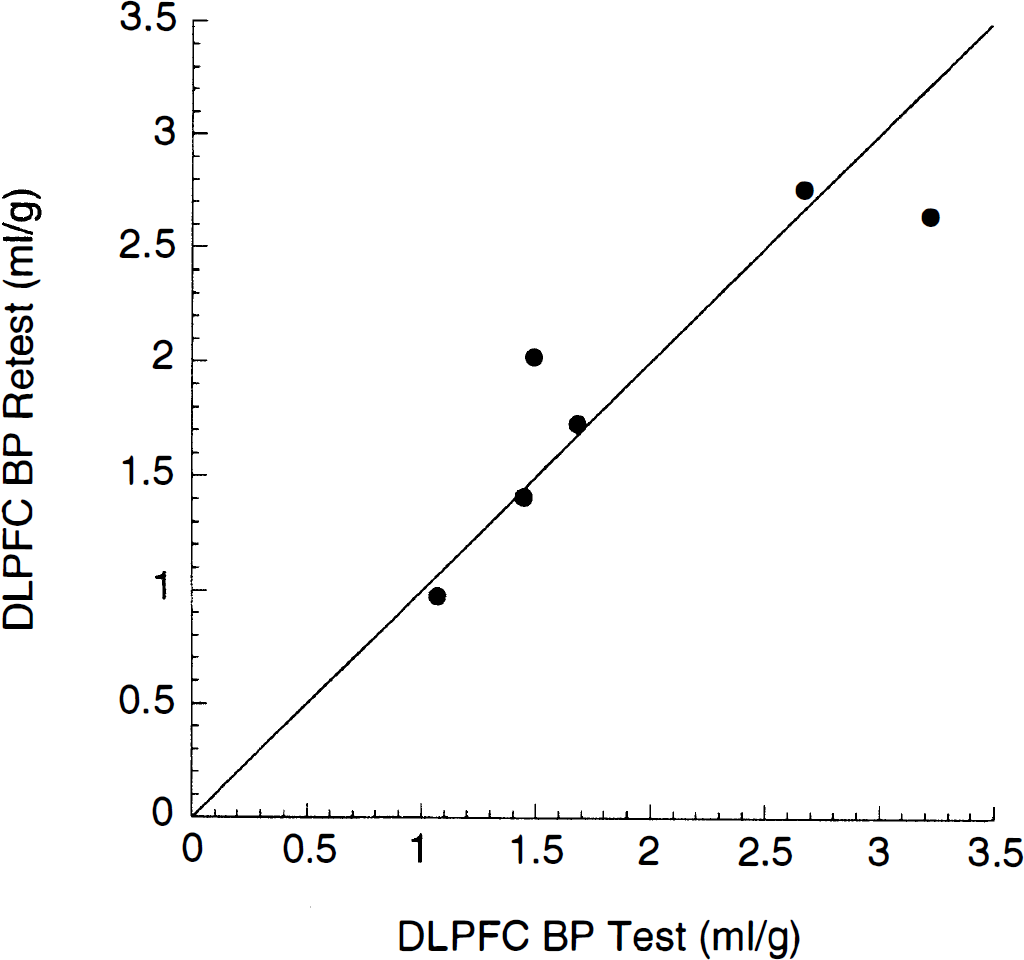
Relationship between [11C]NNC 112 binding potential (BP) in dorsolateral prefrontal cortex in test (
Graphical analysis
By visual inspection, Logan plots achieved linearity at times of >25 minutes, and a linear regression was performed on data from the 30- to 120-minute interval (Fig. 10). The
Graphical analysis of regional [11C]NNC 112 uptake in human brain
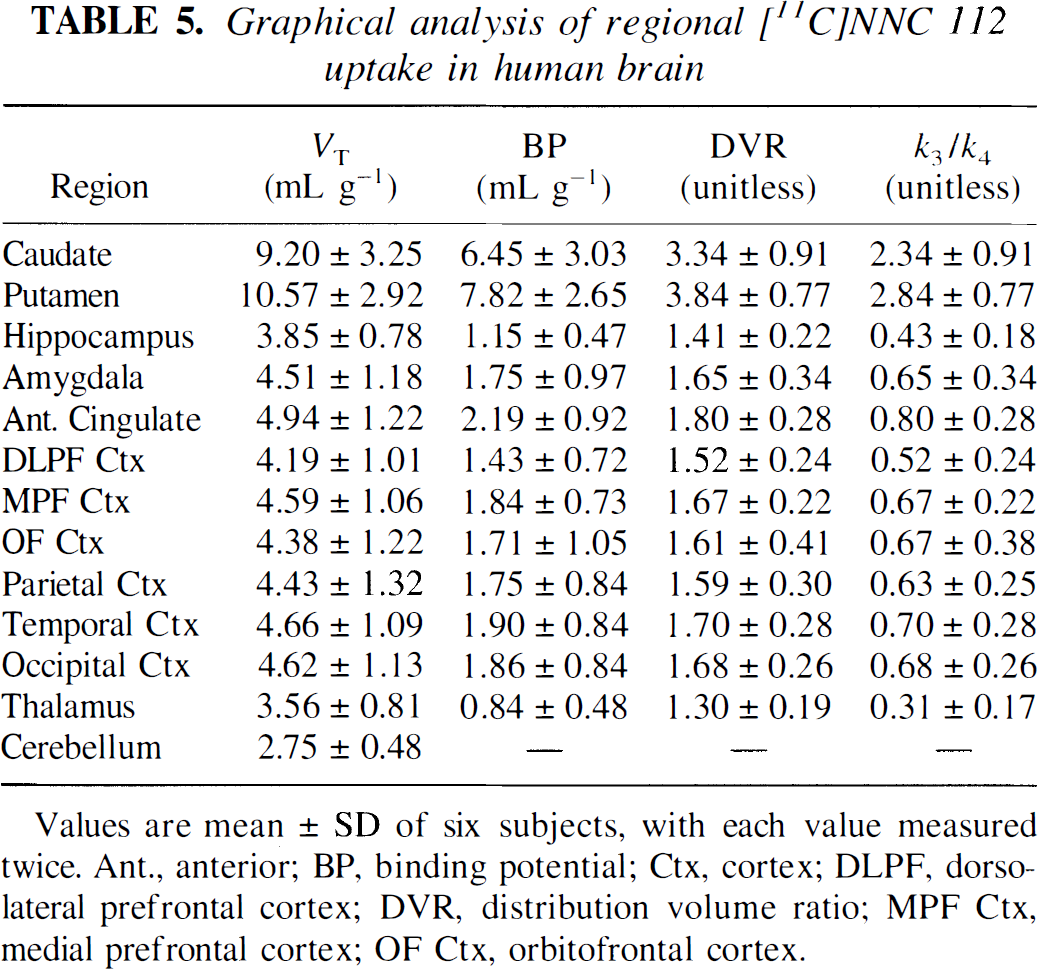
Values are mean ± SD of six subjects, with each value measured twice. Ant., anterior; BP, binding potential; Ctx, cortex; DLPF, dorsolateral prefrontal cortex; DVR, distribution volume ratio; MPF Ctx, medial prefrontal cortex; OF Ctx, orbitofrontal cortex.
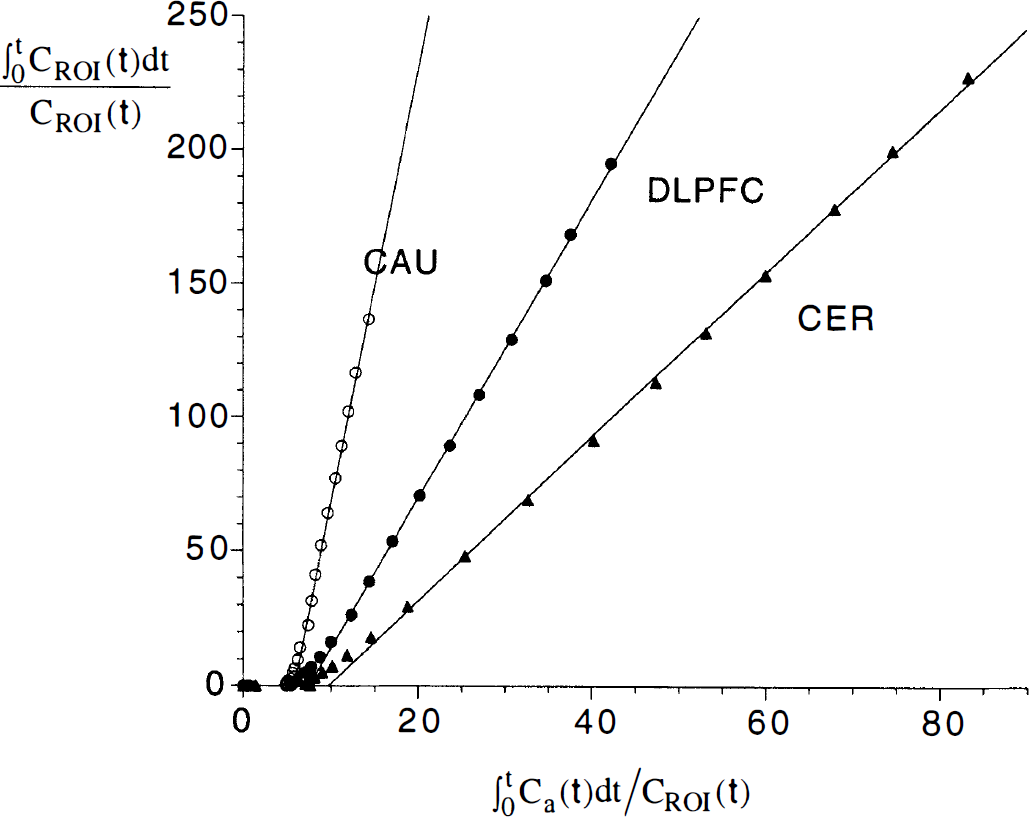
Graphical analysis of [11C]NNC 112 uptake in caudate (open circles), dorsolateral prefrontal cortex (filled circles), and cerebellum (triangles) after injection of 18 mCi of [11C]NNC 112 (same experiment as in Fig. 6). The linearity of the relationship at normalized times of >15 min (equal to 25 minutes of real time) confirms the reversibility of the binding.
Tables 6 and 7 display the reproducibility parameters of the graphical method. As observed for kinetic analysis, the test/retest variability of
Reliability of [11C]NNC 112 distribution volumes and distribution volume ratio measurements in human brain: graphical analysis
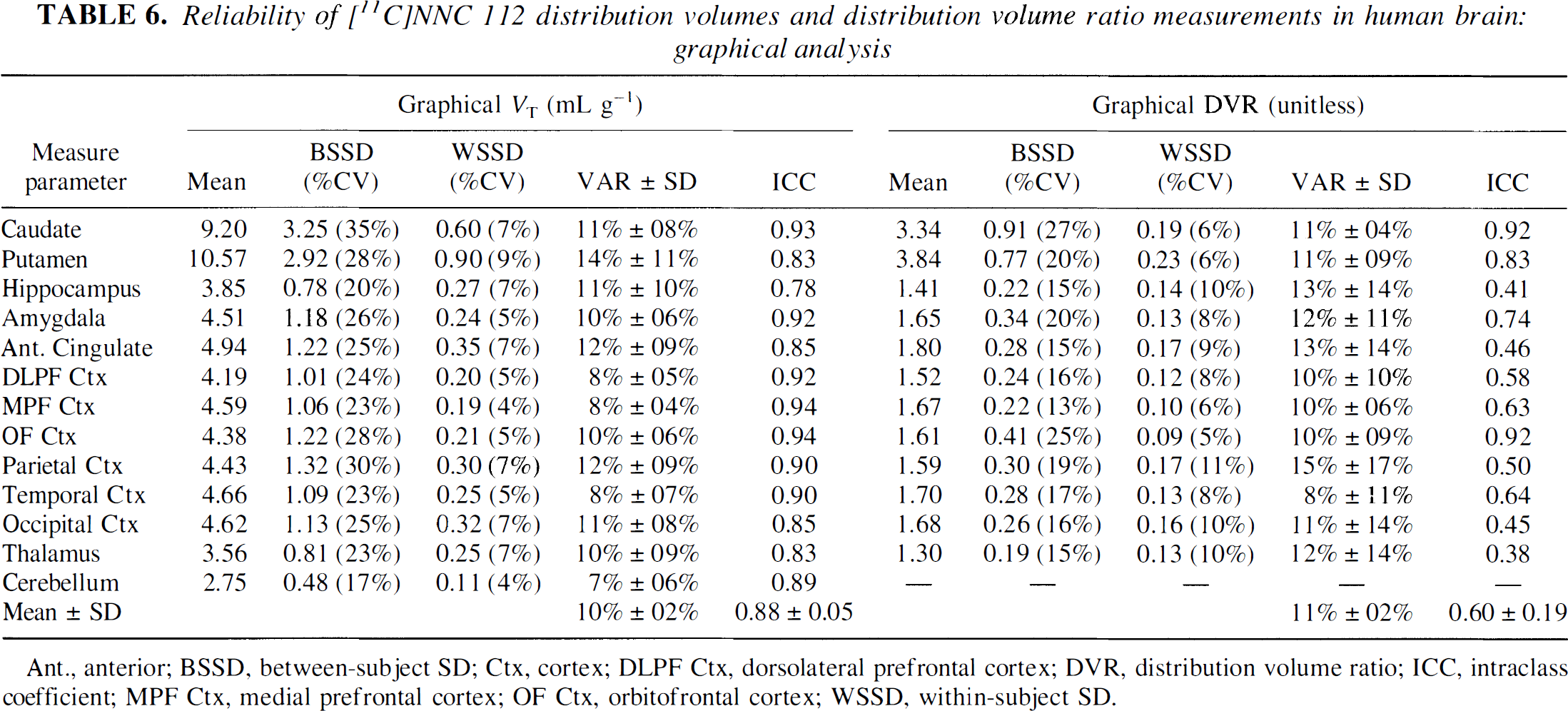
Ant., anterior; BSSD, between-subject SD; Ctx, cortex; DLPF Ctx, dorsolateral prefrontal cortex; DVR, distribution volume ratio; ICC, intraclass coefficient; MPF Ctx, medial prefrontal cortex; OF Ctx, orbitofrontal cortex; WSSD, within-subject SD.
Reliability of [11C]NNC 112 BP and
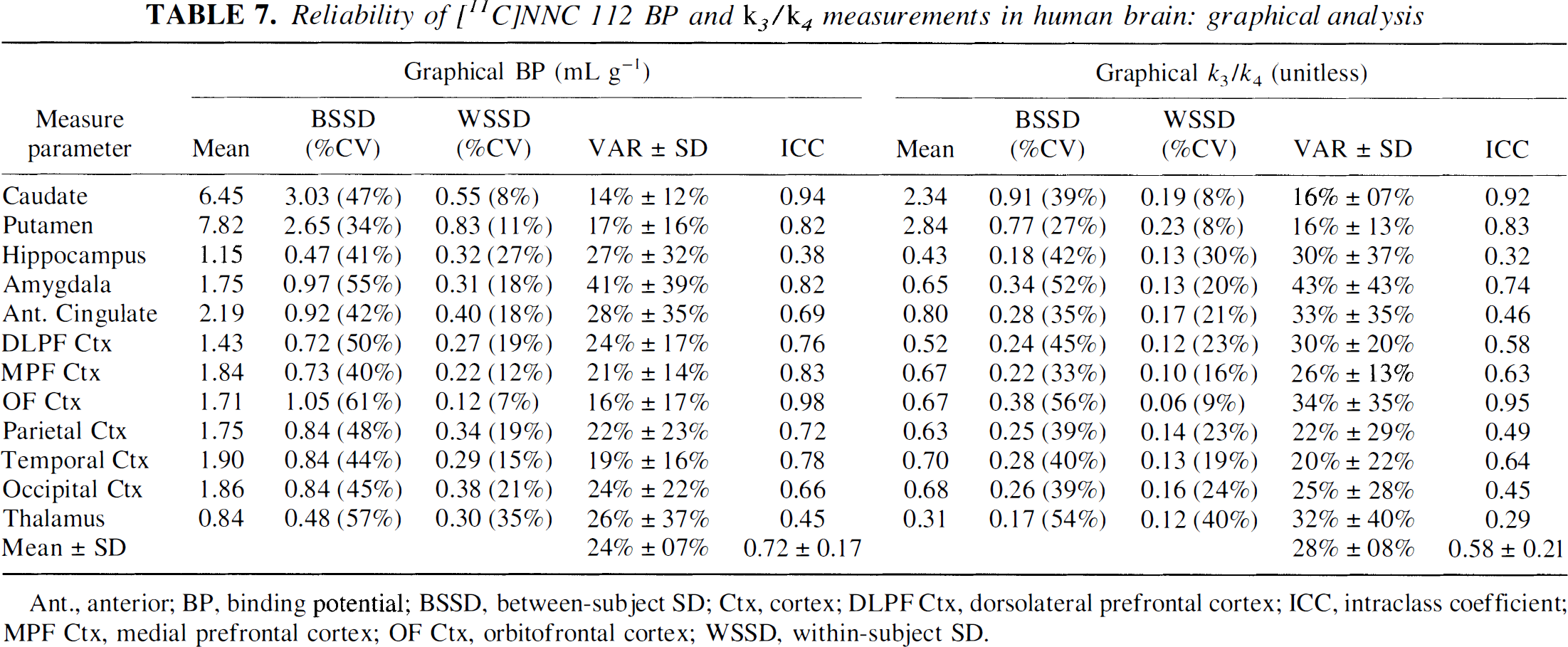
Ant., anterior; BP, binding potential; BSSD, between-subject SD; Ctx, cortex; DLPF Ctx, dorsolateral prefrontal cortex; ICC, intraclass coefficient; MPF Ctx, medial prefrontal cortex; OF Ctx, orbitofrontal cortex; WSSD, within-subject SD.
Noise analysis
Noise analysis was done to compare the effect of realistic noise in the images on results from the kinetic and graphical analyses (Fig. 11). A typical experiment was used to generate a simulated data set that included one cerebellar curve (two-compartment model,
Noise analysis of cerebellum and caudate: comparison of kinetic and graphical methods
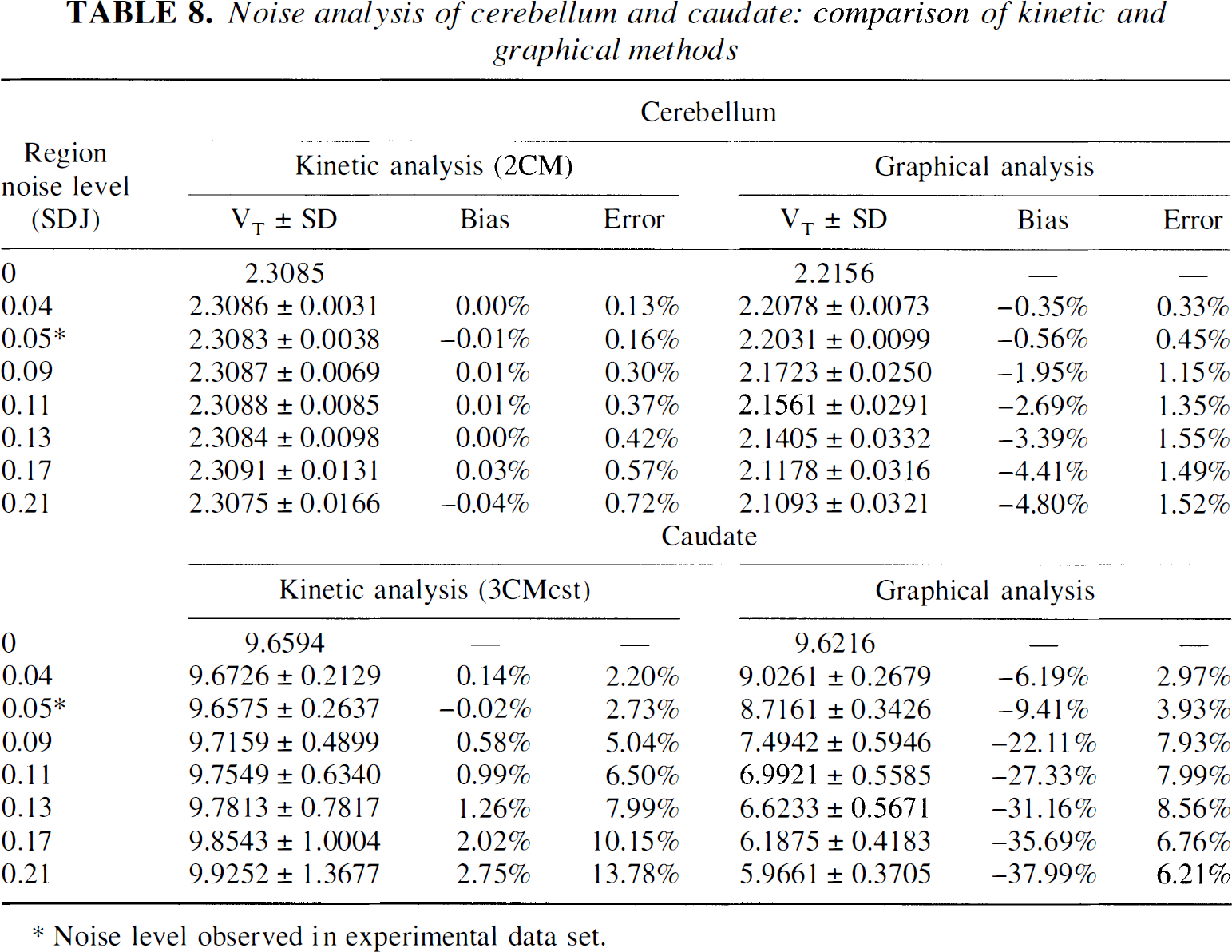
Noise level observed in experimental data set.
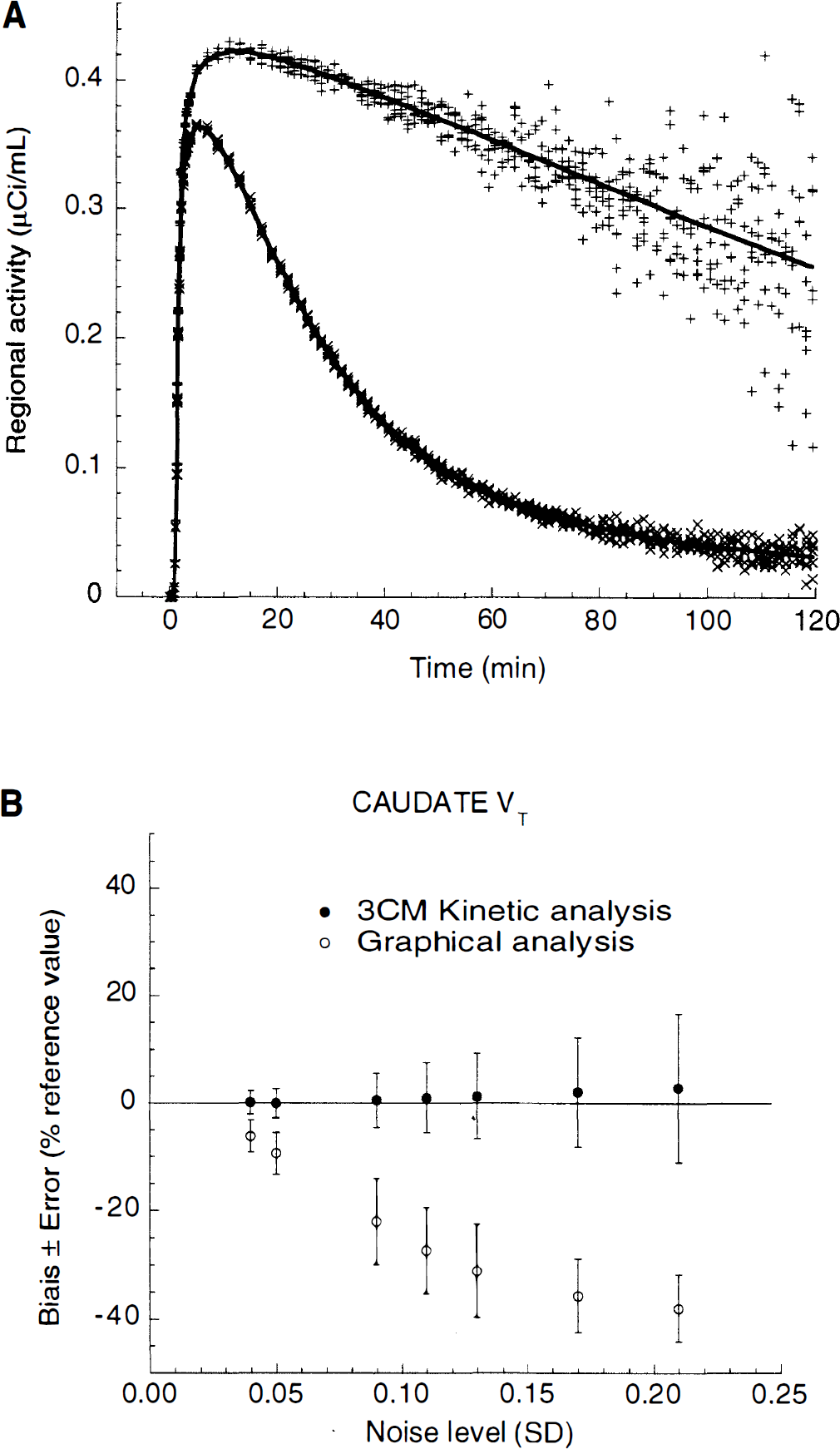
Noise analysis.
Both analyses were very robust as far as the cerebellum was concerned. For cerebellar kinetic analysis, the bias was negligible, and the error was very small (both bias and error < 1%). Cerebellar graphical analysis showed higher bias and errors than the kinetic analysis but remained within acceptable levels at each noise level.
Analysis of caudate curves was more sensitive to noise. Kinetic analysis showed a small bias (+2.7% at higher noise level), and the error increased to ±14% at the highest noise level. Graphical analysis was characterized by a pronounced bias. As noise increased, graphical
DISCUSSION
This study confirms the excellent imaging properties of [11C]NNC 112 previously reported in two healthy volunteers (Halldin et al., 1998). In addition, model-based analysis demonstrates that kinetic analysis of brain regional uptake of [11C]NNC 112 provides reliable estimates of D1 receptor BP not only in the striatum but also in extrastriatal areas such as the limbic and neocortical regions. To our knowledge, this study represents the first demonstration of the reliability of the measurement of D1 receptor BP in extrastriatal areas. In a recent study of the reproducibility of D1 receptor quantification with [11C]SCH 23390, only measurements in the striatum were reported (Chan et al., 1998).
Comparison with [11C]SCH 23390
Results of this study indicate that [11C]NNC 112 achieves higher specific/nonspecific DVRs at equilibrium than [11C]SCH 23390. In the putamen, [11C]SCH 23390
Neocortical region-of-interest sampling
In this study, an original method was developed for sampling of activity in neocortical regions. Measurement of activity in large subcortical regions such as the striatum is generally straightforward, inasmuch as the boundaries of activity can be well defined anatomically. In the striatum, MRI-guided ROI definition does not substantially improve the reliability of the measurement compared with direct definition of the ROI on the PET image based on activity distribution (Wang et al., 1996). In contrast, measurement of activity distribution in GM cortical areas is a difficult problem due to the extent and complexity of cortical gray geometry (see discussion in Koepp et al., 1997). The simplest method is to draw geometrically simple structures (spheres, rectangles) directly on the PET scan, but this method is unreliable and influenced by the activity distribution on the PET image. Thus, defining the ROI on a coregistered MR image is preferable. However, tracing contours of a cortical region manually around the GM on high-resolution 128-slice MRI is time consuming and subject to rater bias and errors. In this study, we used a method based on coregistration of the PET data set to a segmented MRI data set, that is, a data set in which each pixel has been designated as GM, WM, or CSF value. Only the activity located in the pixels classified as GM is sampled. The advantage of constraining ROI on GM is that the segmentation procedure is operator independent and much faster and reliable than manual tracing of GM/WM/CSF boundaries. The only operator-driven choice is the choice of the limits between cortical regions, which follows well-defined criteria. The excellent reliability of the cortical measurement of [11C]NNC 112 BP reported in this study indirectly supports the robustness of the method.
It is, however, important to keep in mind that this method has several limitations. Like all MRI-based ROI definitions, this method is sensitive to errors in PET-MRI coregistration. In this study, we used the minimization procedure developed by Woods et al. (1993). This procedure has a reported average three-dimensional error of <2 mm for between-modality registration, which is acceptable, given a camera resolution of 6 mm. Second, this sampling method results in inherent subsampling, inasmuch as the activity spilling over the WM is not measured. This subsampling is desirable from an accuracy point of view but results in a loss of information that might translate into increased noise. Finally, this method does not correct for partial voluming, which is considerable in the cortex. Yet, it provides a necessary first step for implementation of ROI-based partial volume correction (Rousset et al., 1998). In fact, partial volume correction would result in a recovery of the spilled-over information. Thus, partial volume correction has the potential not only to increase the accuracy of the measurement but also to reduce the error by using the information that is rejected in the present method. We are currently developing and testing a partial volume correction method that would address these issues.
Evaluation of kinetic approach
Evaluation of model-based methods for quantification of receptor parameters includes assessment of various criteria, namely, identifiability, validity, stability, and reliability. The identifiability is a mathematical dimension referring to the confidence in the value of the parameter as derived by the fitting process. The validity refers to the physiological plausibility of the parameters and is usually determined in relationship to
Kinetic analysis of the cerebellum presented an interesting problem. The three-compartment model provided a slightly but significantly superior fit to the data compared with the two-compartment model. The observation of a better fit by the three- compared with the two-compartment model for a region of a reference is the rule rather than the exception in neuroreceptor imaging (Frost et al., 1989; Logan et al., 1990; Laruelle et al., 1994b, 1994c; Lammertsma et al., 1996; Abi-Dargham et al., 1997; Ito et al., 1998). The absence of specific binding in the cerebellum is confirmed by the absence of displacement of [11C]NNC 112 uptake in the cerebellum after injection of SCH 23390 (2.5 mg/kg) (Halldin et al., 1998). Thus, the small second tissue compartment revealed by the three-compartment model's superior fit might reflect a slow buildup of radiolabeled metabolites in the brain or the existence of a nonspecific binding component with slow kinetics. The presence of a statistically superior fit with the three-compartment model would usually suggest the use of a three-compartment model to derive cerebellar
For ROIs, we used a three-compartment model with
A significant relationship was observed between regional BP and regional density of D1 receptor measured
Results of kinetic modeling were essentially stable from 90 minutes to the end of the experiment. The stability over time of the solution means that the model describes the data appropriately and that enough data were collected to inform the model. Remarkably, this was true for both striatal and extrastriatal areas. It is often difficult to accurately measure receptor parameters with the same radiotracer both in regions with high and low receptor densities. High affinity is required for a high signal/noise ratio in regions with low receptor density, but high affinity might preclude favorable kinetics in regions with high receptor density. [11C]NNC 112 achieves an effective balance between these competing requirements. However, 90 minutes of data was needed to achieve stability of the outcome measure. The three-dimensional acquisition mode, with its higher sensitivity, is crucial to enable data collection up to 120 minutes. The use of the polyurethane head mold was effective, as only minimal head movement was recorded during this prolonged time frame. Head movement was measured by comparing the spatial location of the 11C markers on each frame and was <2.5 mm per frame in ∼90% of the cases. Thus, the combination of the high sensitivity of the three-dimensional mode and the effective headholder allowed collection of enough data to achieve stability of the results.
Reproducibility and outcome measures
Several statistical parameters and outcome measures are used in the PET/single photon emission computed tomography neuroreceptor literature to report the reproducibility of the measurements. The most straightforward reproducibility parameter is the absolute test/retest variability, calculated as the absolute difference between test and retest values divided by their average (Abi-Dargham et al., 1995; Seibyl et al., 1995; Smith et al., 1998). A second method consists of reporting the within-subject SD (or %CV) of the measurement (Chan et al., 1998). When the number of repeat scans is 2, the within-subject %CV is smaller than the variability by a factor of 21/2. Finally, the ICC provides a measure of the reproducibility of the measurement relative to between-subject differences and is the most useful measure to compare results of various methods.
In this study, four outcome measures (
In contrast to
BP versus k 3/k 4
As a region of reference can be measured for [11C]NNC 112 (i.e., cerebellum), the only meaningful choices of outcome measures for clinical studies are either BP or
The BP as expressed in Eq. 10 is the only expression of BP that is exclusively dependent on receptor parameters and unbiased by factors unrelated to receptors. However, in this study, BP was not corrected for
Kinetic versus graphical analysis
Distribution volumes derived by graphical and kinetic analyses were strongly correlated (
Analysis of the effect of noise on the outcome measure derived by both methods was performed to better understand this phenomenon. Realistic noisy data sets were generated in which the noise increased with the duration of the experiment proportionally to the decay of 11C. The impact of the noise on the outcome measure was very different for kinetic and graphical analyses. As noise increased, both methods were associated with a similar variability, but
At the highest noise level (SDJ = 0.21), this bias resulted in a 38% underestimation of
A large range of noise was used to generate these simulations (SDJ from 0.04 to 0.21). To determine which level of noise corresponded to the experimental noise, we compared the mean sum of squares of the residuals as well as the identifiability (percent error) of
At this noise level, caudate kinetic analysis had no bias (−0.02%), but the graphical analysis had a significant bias (−9.41%). The bias associated with graphical analysis at this noise level might account for the lower
CONCLUSION
This study confirms that [11C]NNC 112 is a superior radiotracer with which to measure D1 receptors in both the striatal and the extrastriatal areas. The main limitation of this radiotracer is that 90 minutes of data collection is needed to yield stable outcome measures. Simplified approaches that do not require input function measurement still need to be evaluated. Because it allows reliable estimation of D1 receptor parameters in extrastriatal areas, [11C]NNC 112 might play an important role in elucidating the role of D1 receptors in cognition in health and disease and in psychopathology.
Footnotes
Abbreviations used
Acknowledgements
The authors thank Suehee Chung, Bryan Bergert, Ann Shinn, Alexei Kartachov, Richard Weiss, Analia Arevaldo, Julie Montoya, Daniel Schneider, and the technologists of the Columbia PET Center for excellent technical assistance.
