Abstract
Introduction
Human activity in wilderness areas has the potential to affect aquatic ecosystems, including through the introduction of microorganisms associated with fecal contamination. We examined fecal microorganism contamination in water sources (lake outlets, snowmelt streams) in the popular Absaroka Beartooth Wilderness in the United States. Although the region is remote, increasing human visitation has the potential to negatively affect water quality, with particular concern about human-derived microorganism fecal contaminants.
Methods
We used standard fecal indicator bacterial assays that quantified total coliform bacteria and
Results
Total coliforms were detected at all lake outlets (21 of 21 sites), and
Conclusions
Our results suggest low prevalence of water-borne pathogens (specifically
Keywords
Introduction
Designated wilderness areas in the United States are unique in that there are restrictions on permanent facilities to manage visitor impacts and human waste. 1 However, human visitation to these areas continues to increase, 2 creating challenges for managers seeking to minimize impacts of human activities on wilderness natural resources. 2 -4 In particular, fecal contamination of wilderness water sources presents a growing problem, requiring managers to identify areas of potential contamination to protect public health. 5 Outbreaks of illnesses associated with fecal bacteria (eg, diarrhea) have been documented in congested recreational sites in wilderness areas,6,7 with possible sources of fecal contaminants including livestock, dogs, wild animals, and humans.
Fecal indicator bacteria (FIB) assays are often used to identify the presence of total coliform bacteria, fecal coliforms, and
Bacteria belonging to the phylum Bacteroidetes are often used as targets in MST assays because of source specificity to mammalian guts and feces.
12
-15 Members of
Our study focused on potential fecal contamination of waters in the Absaroka Beartooth Wilderness (ABW) in south central Montana. The ABW is a mountainous wilderness area northeast of Yellowstone National Park. The region ranges in elevation from approximately 1.6 km to over 4.1 km and contains a network of subalpine forests and alpine tundra. With over 1100 km of trails, the area receives high recreational use, including backpacking and stock packing, but no livestock grazing. Most human visitation is concentrated around lakes and at water sources near popular summits. Backcountry campsites are required to be >60 m from lakeshores, and popular campsite locations have been cataloged in a geospatial database maintained by the US Forest Service for over 20 y. We sought to examine the occurrence of total coliform and
Methods
Study Area and sampling sites
We selected a total of 23 sampling locations within the ABW (21 remote alpine lake outlets and 2 snowmelt streams) using geospatial information on campsite condition and popular recreational sites. Sampling sites were identified using ArcMap (version 10.5.1), overlaying campsite point locations with a wilderness-wide opportunity class layer representing 3 zones of recreational use: 1) pristine, 2) primitive, and 3) transition zones. Pristine zones were those with negligible anthropogenic influence, primitive zones were those with measurable anthropogenic influence, and transition zones were those where human impacts were moderate to substantial. All of our sampling was done within the transition zone, between 4.5 and 24 km from main trailheads (Figure 1).
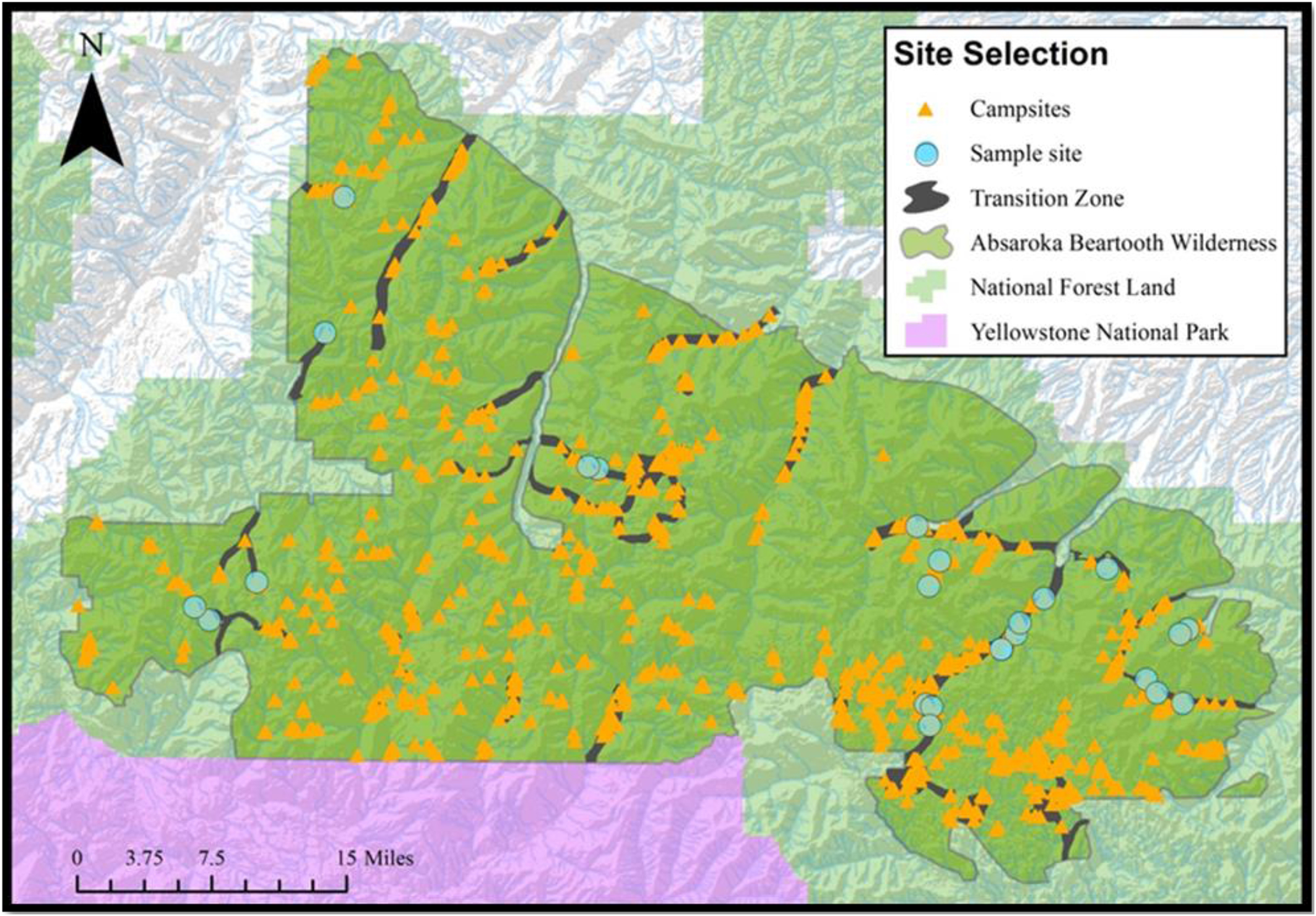
Sampling sites in the Absaroka Beartooth Wilderness. Backcountry campsites are indicated with orange triangles, and sampling sites for this study are indicated with blue circles.
Water sampling, FIB detection, and DNA extraction
Water samples from 21 different alpine lake outlets and 2 snowmelt streams were collected in triplicate over 13 d in July and August 2018 (Table 1). Water samples were placed into sterilized 250-mL polypropylene bottles and transported to the laboratory in the dark and on ice. In the laboratory, samples were stored at 4°C until processing (within 48 h of sample collection). Two 100-mL aliquots of sample water from each of the triplicate 250-mL sample bottles were vacuum filtered onto separate 47-mm diameter, 0.45-μm pore size (mixed cellulous ester) gridded, pre-sterilized filters (Millipore Sigma
Elevation, distance to trailhead, number of campsites, groundcover, and types of use for all sites surveyed in this study
The US Forest Service does not designate campsites in this area, but it monitors well-established but illegal camping sites.
Ground cover designations defined by US Forest Service monitoring.
Use-type: Foot refers to human foot traffic; foot and stock refers to both human and stock animal traffic.
Filters for FIB analyses were placed in M-ColiBlue24 broth petri dishes and incubated at 35°C for 24 h (Millipore Sigma). Colony-forming units (CFUs) were quantified by counting and recording the number of red (non-
DNA was extracted from filters using the MasterPure DNA Purification kit (Lucigen Corporation, Middleton, WI). Triplicate blank filters (ie, no sample filtered onto them) were processed alongside samples. Filters were transferred from 15-mL centrifuge tubes to 2-mL microcentrifuge tubes containing 600 μL of a cell lysis solution and 100 μL of 0.1-mm and 100 μL of 0.5-mm glass beads. The tubes containing filters were frozen at -80°C, thawed, and placed into a mechanical bead beater for 2 min, followed by the addition of proteinase-K (50 μg·μL-1 final concentration). Samples were incubated at 65°C for 15 min and placed on ice for 3 to 5 min, and DNA was extracted following the MasterPure DNA Purification kit protocols. DNA was resuspended in 100 μL of nuclease-free water and stored at -80°C.
Droplet digital PCR for detection of universal and human-specific Bacteroides gene markers
We used 2 different droplet digital PCR (ddPCR) assays: one targeting bacteria belonging to the genus
Polymerase chain reaction primers used for microbial source tracking analyses
Triplicate filter extracts from the lake outlets and snowmelt streams were analyzed by ddPCR using a QX200 Droplet Digital PCR System (Bio-Rad Laboratories, Hercules, CA). Each ddPCR reaction (20 μL total) contained EvaGreen Supermix (Bio-Rad Laboratories), nuclease-free water, DNA extract, and 0.18 μM (final concentration) of each primer. Triplicate controls (with no added DNA) were included as negative controls. Filter blank DNA extracts were also analyzed in triplicate to estimate the detection limits associated with both ddPCR assays (see description in Statistics subsection). PCR reaction mixes were combined with droplet generation oil specific for EvaGreen, and droplets were generated using the droplet generator (Bio-Rad Laboratories). Droplets (40 μL total, including PCR reaction mix and oil) were transferred by multichannel pipettor into 0.2-mL 96-well PCR plates. Plates were heat sealed and placed in a Bio-Rad C1000 thermal cycler.
Amplification conditions were as follows: 95°C for 6 min, followed by 40 cycles of 95°C for 30 s, 61°C for 30 s, and 72°C for 45 s. 13 A QX200 Droplet Reader (Bio-Rad) was used to quantify droplet specific fluorescence. The fluorescence amplitude threshold was manually adjusted to distinguish positive droplets from those that demonstrated no amplification (negative droplets). We excluded reactions with <10,000 accepted droplets from subsequent analyses (n=3). Gene abundances (copies per milliliter of sample water) were quantified for each sample using the manufacturer’s software (QuantaSoft, Bio-Rad Laboratories).
Statistics
The method detection limit (MDL) of the ddPCR assays was defined as the lowest number of genes that could be distinguished based on replicate (n=4) amplification of the filter blank DNA extracts. We estimated the MDL for each assay as
We defined the target gene as “present” if 1 or more of the triplicate ddPCR reactions amplified above the detection limits after 40 cycles. We quantified gene abundances only for those samples for which all triplicate ddPCR reactions amplified above the detection limits. Data are presented as mean±SD with range.
Least-squares linear regression analyses of square root transformed count data were used to examine relationships between CFUs of coliform and 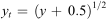
Results
Total coliform and E coli occurrences
Total coliforms were found in all 21 of the lake outlets sampled, and
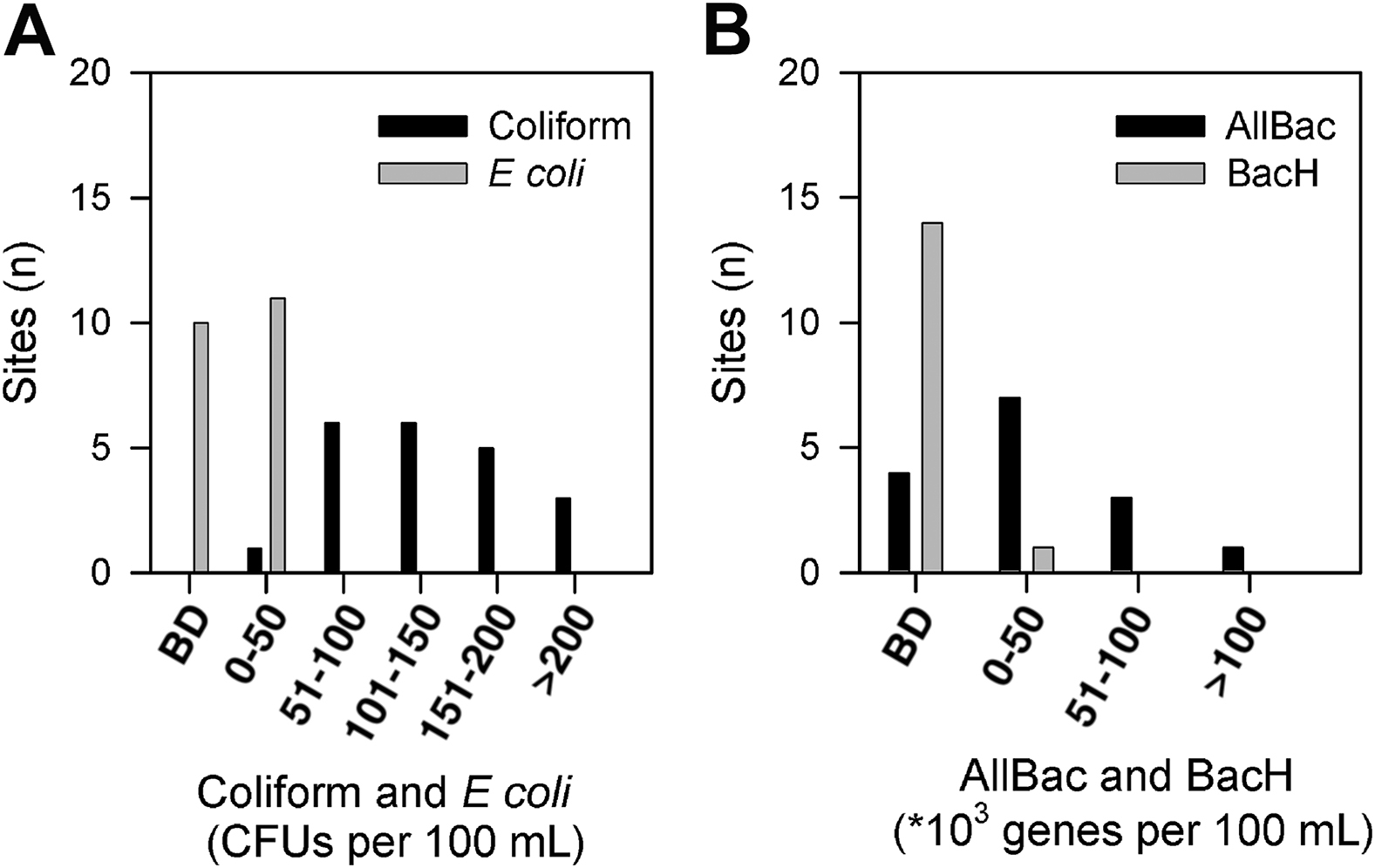
Histograms of colony-forming units (per 100 mL of water) for total coliforms and
Occurrence of total coliform and
Presence of target gene indicates detectable but not quantifiable genes (ie, at least 1 sample from the triplicate ddPCR reactions amplified; “No” indicates none of the triplicate polymerase chain reaction reactions amplified above the lower limit of detection). Mean±SD of triplicate analyses from each lake outlet or snowmelt stream. BD indicates gene abundances below detection (<900 gene copies per 100 mL of lake water for BacH and <2100 gene copies per 100 mL of lake water for AllBac). NA indicates sample not analyzed.
Universal and host-specific Bacteroides MST
The amplitudes of fluorescence (in relative fluorescence units [RFUs]) for droplets binned as positive for the AllBac and BacH assays were 1.7- and 1.6-fold greater, respectively, than fluorescence amplitudes of the negative droplets. The amplitude of fluorescence for positive droplets in the AllBac assay was 15,571±1195 (11,628–17,579) RFUs, whereas the fluorescence amplitude of negative droplets was 9247±773 (8044–11,026) RFUs. The fluorescence amplitude of the positive droplets for the BacH assay was somewhat lower at 9206±2849 (6270–19,885) RFUs, with negative droplets at 5799±1030 (4129–9971) RFUs. The number of accepted droplets for both assays was 15,043±1877 (10,404–18,558). The MDL for the BacH ddPCR assay was 18 copies per ddPCR reaction, equivalent to 9 copies per milliliter of lake water, whereas the MDL for the AllBac ddPCR assay was 42 copies per reaction, equivalent to approximately 21 copies per milliliter of lake water.
AllBac genes were present in all of the sites examined by ddPCR, with gene abundances quantifiable in 11 of 15 sites (Table 3). For those sites where gene abundances were quantifiable, AllBac gene abundances ranged from 5×103 to 131×103 genes per 100 mL of sampled water, with peak abundances observed in the same 2 lakes where
Human-associated
Based on least-squares linear regressions of the square root transformed data, total coliform CFUs and the AllBac gene abundances were correlated (R2=0.57,
Discussion
By leveraging culture-based FIB assays and culture-independent MST assays, we evaluated the spatial distribution and potential sources of fecal bacteria in selected ABW waters. Total coliforms were present in all of the lake outlets sampled, with positive occurrence of
Our study highlights the utility of combining FIB and MST methods for discerning possible sources of feces-associated microorganism contaminants. For example, we did observe a weak but significant positive relationship between the FIB-derived total coliform CFUs and AllBac gene abundances, suggesting the presence of mammalian
Use of the AllBac ddPCR assay revealed widespread occurrence of members of
We also sampled 2 snowmelt streams near Granite Peak, Montana’s highest point and a popular summit within the ABW. The scarcity of water around Granite Peak creates congested basecamp areas that place disproportionate pressures on adjacent water resources. Prior studies on wilderness waters noted that both total coliform and
Although use of culture-based FIB- and PCR-based MST methods revealed that several lakes in the ABW appear to be influenced by fecal microorganism contaminants, for most lakes we were unable to identify sources of these potential contaminants. The human-specific fecal indicator MST assay indicated that most of these lakes received little fecal input from humans. These results highlight the utility of combining more traditional FIB assays with emerging MST methods for information on both the occurrence and specific sources of microorganism contaminants to aquatic systems.
Limitations
A limitation in this study was the relatively high MDLs estimated for both ddPCR assays. Detection limits for both assays were somewhat higher than has been previously reported for quantitative PCR-based MST methods 27 ; the detection limits for the ddPCR assay derive from any amplification in the blanks, together with the volume of water filtered to concentrate microorganism biomass. We processed relatively small volumes of water for subsequent extraction of DNA, constrained in part by the remote locations and the need to transport water back to the laboratory. We also did not include PCR inhibition controls as part of our ddPCR analyses; hence, we do not have information on the extent to which the presence of inhibitors may have resulted in underestimation of the target gene abundances. In addition, our sampling was restricted to a single summer season; hence, we do not have information on how seasonal- to episodic-scale changes in the local environment (eg, rainfall, snowfall) might affect the introduction of fecal contaminants to these waters.
Conclusions
Our results suggest relatively low prevalence of water-borne pathogens (specifically
Footnotes
Acknowledgements
Acknowledgments: This study benefited by input from A. Wood, K. Crootof, K. Barker, J. Zimmer, C. Neppl, L. Oswald, E. Bilbrey, and the US Forest Service. We thank the Franke College of Forestry and Conservation, the Human Dimensions Lab, L. Gerloff, and the Wilderness Institute at the University of Montana; W.A. Sigler and the Zabinski Lab at Montana State University; and Dr. R.O. Hall and T. Bansak at the Flathead Biological Station. The comments of 2 reviewers and the editor substantially improved this work.
Author Contributions: Study concept and design (DPP, MJC, ALM, THD); acquired data (DPP, JR, LE, AB); analyzed the data (DPP, JR, AB, MJC); wrote and revised the manuscript (DPP, MJC); all authors approved the final version of the manuscript.
Financial/Material Support: This study was funded by the Aldo Leopold Wilderness Institute, Montana Institute on Ecosystems, and the University of Montana. MJC acknowledges support from the Simons Foundation (award 329108).
Disclosures: None.
