Abstract
The culture-independent and automated detection of bacteria in the environment is a scientific and technological challenge. For detection alone, a number of sensitive methods are known (e.g., PCR, enzyme-linked immunosorbent assay [ELISA], fluorescent in situ hybridization) but a major problem remaining is the enrichment and separation of the bacteria that usually occur at low concentrations. Here, we present an automated capturing and separation system, which can easily be combined with one of the sensitive detection techniques.
We have developed a method for enrichment and detection of Legionella pneumophila in liquid media. Concentrated microorganisms were either detected by PCR or by sandwich ELISA. The limit of detection with the immunological assay was about 750 bacteria. Using PCR, the equivalent of about 2000 genomes could be detected.
The assays were then transferred to a laboratory prototype for automated processing. It was possible to automatically enrich L. pneumophila by immunomagnetic separation (IMS), and again, the bacteria were detected by sandwich ELISA and PCR amplification of the ompS gene. As a novel aspect, ompS gene was used for the first time as a target for the detection of L. pneumophila on magnetic beads.
The aim of this work was to develop an automated procedure and a device for IMS of bacteria. With Legionella as a model organism, we could show that such a novel fully automated system can be an alternative to timeconsuming conventional cultivation methods for detecting bacteria or other microorganisms.
Introduction
The standard detection methods for viable
Fluorescent in situ hybridization is a sensitive and rapid detection method for microorganisms. Water samples can be filtered, and the captured bacteria are hybridized with a fluorescence-labeled probe complementary to 16S rRNA. 6,7
For rapid enumeration of different strains of viable
However, all these detection methods are not sensitive enough to detect bacteria at very low concentrations. One solution to this problem is an enrichment step by filtering the water sample, but often, the bacteria stick to the filter membrane, and it is not possible to completely recover them from the filter surface. Micromechanical filters have successfully been used as a tool for efficient filtration and enrichment of bacteria. Because of their absolutely flat surface, bacteria can be recovered and transferred to a biodetector system. 9
Another technique for capturing microorganisms is immunomagnetic separation (IMS). IMS uses small superparamagnetic particles or beads coated with antibodies against surface antigens of pro- and eukaryotic cells. Many applications of IMS can be found in the literature (e.g., Refs. 10 and 11). This technique is also used in diagnostic applications to detect a wide spectrum of pathogens. After capturing the organisms with the beads, nearly all bioanalytical methods can be applied for detection, such as immunoassay, hybridization techniques, and PCR. 12,13 Furthermore, immunomagnetic cell sorting is also possible. For extracting mesenchymal stem cells from peripheral blood, an immunomagnetic cell sorter in combination with a mixer was developed by Inokuchi et al. 14 In the first step, the cell suspension is mixed with antibody-coated paramagnetic beads. During mixing, the paramagnetic beads interact with the stem cells by antigen—antibody reaction. In the next step, the captured cells are separated from the medium by a magnetic separator.
A different capturing technology is to bind the cells to cationic magnetic beads. These positively charged beads interact with the negatively charged lipopolysaccharides on the surface of gram-negative bacteria. With the commercial capture system Pathatrix
In this article, we present a novel automated system for capturing and enrichment of bacteria. Paramagnetic beads coated with a specific antibody against
Materials and Methods
Bacteria, Genomic DNA, and Determination of Cell Numbers
Experiments were performed using inactivated
Immunoblot Assay (Western Blot) of Legionella pneumophila
A suspension of heat-killed
Twenty microliters of biotinylated polyclonal anti-
PCR mixtures (48 μL) contained 10 mM Tris-HCl at pH 8.3, 50 mM KCl, 1.5 mM MgCl2, 2.5 mM each of the four deoxynucleotides, 100 pmol of each primer, and 0.02 U of Taq DNA polymerase (New England Biolabs, Ipswich, USA), to which 2 μL of beads with captured
PCR assays were conducted in a RoboCycler (Stratagene; Agilent Technologies, Santa Clara, USA) with the following thermal profile: initial denaturation for 5 min at 95 °C, followed by 30 cycles of denaturation at 95 °C for 20 s, annealing at 54 °C for 20 s, extension at 72 °C for 1 min, and final extension at 72 °C for 5 min. Amplified PCR products were separated in a 2% (wt/vol) agarose gel and stained with ethidium bromide. Agarose gels were prepared and run in 1 × Tris-acetate-EDTA buffer (TAE) (50x TAE: 242 g Tris-base, 57.1 mL acetic acid, 100 mL 0.5 M EDTA (Ethylenediaminetetraacetic acid), water to 1000 mL, pH adjusted to 8.5). PCR products were compared with a 1-kbp DNA molecular size ladder (New England Biolabs, Ipswich, USA) to estimate their size. Results were scored as positive or negative by visual inspection.
Results
Automated Immunomagnetic Enrichment of Legionella pneumophila
We have developed an automated system for capturing bacteria using paramagnetic beads (principle shown in Fig. 1A). Two pipettors-one for larger volumes up to 10 mL and the other for small volumes up to 300 μL-are mounted on independent
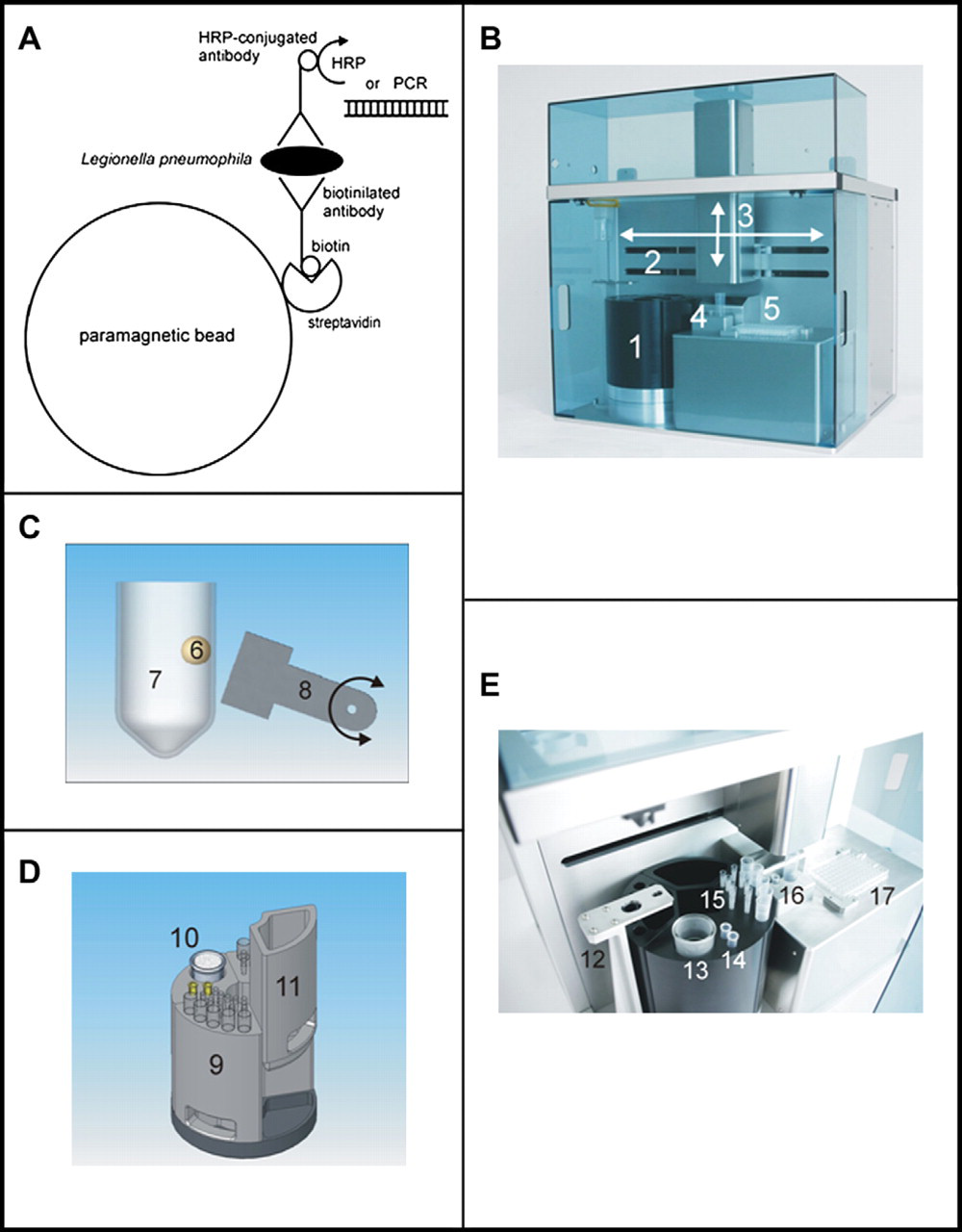
Illustration of the automated separation system and schematic diagram of capturing by IMS. A) Schematic illustration of the capturing and detection of
Using a large and a small reaction vessel, magnetic beads from a sample volume of 10 mL can be washed and concentrated into any volume between 20 μL and 1 mL. An integrated heater enables denaturation of samples. As a last step of the process, the concentrated beads are transferred into a microplate for further processing. The whole system is controlled by custom-made software with an integrated editor for structured programming.
A typical processing cycle starts with an initial sample volume of 5 mL in the large vessel. 6 × 108 beads are added to the sample and mixed with the large pipette. After an incubation time of 30 min, magnetic separation starts followed by a concentration of the beads into a volume of 1 mL. The beads are washed three times with 1 mL washing buffer (PBS with 10% BSA and 0.01% Tween-20) using the large pipette. The beads are collected again and resuspended in 1 mL washing buffer containing detection antibody conjugated with HRP (1:1000), followed by an incubation time of 20 min, including mixing every 5 min with the small pipette. The beads are collected again, resuspended in 300 μL buffer and transferred to the small reaction vessel. To remove unbound antibody, beads are washed six times with a volume of 300 μL. In the last processing step, beads are concentrated to a final volume of 100 μL and fractionated into five aliquots of 20 μL each in a microplate. These samples were either subjected to HRP reaction (immunoassay) or to PCR detection. The whole processing cycle for sandwich ELISA was about 2.5 and 2 h for processing for detection with subsequent PCR (without addition of detection antibody). This results in a total time of about 3 h for immunodetection (2.5 h washing and enrichment plus ca. 0.5 h HRP reaction and reading). In the case of PCR, about 4–5 h (including analysis of reaction products on a gel) have to be taken into account. The conventional detection of
To estimate the sampling efficiency, a small aliquot of beads was taken out after each concentration step and counted under the microscope with a Neubauer improved counting chamber. From an initial amount of 6 × 108 beads, 2.5 × 108 beads were remaining after all concentration and washing steps, that is, 45% of the magnetic beads were left over for detection by sandwich ELISA or PCR. The loss of beads of 55% seems to be very high, but it results from the high number of washing steps. We found out that the major loss (approx. 40%) comes from the washing with the large pipette. Further improvements of the system are underway to reduce this loss. In summary, the sample volume of 5000 μL was reduced down to 100 μL (50-fold concentration). From this, a fraction of only 2 μL was used for the assays (i.e., HRP reaction or PCR).
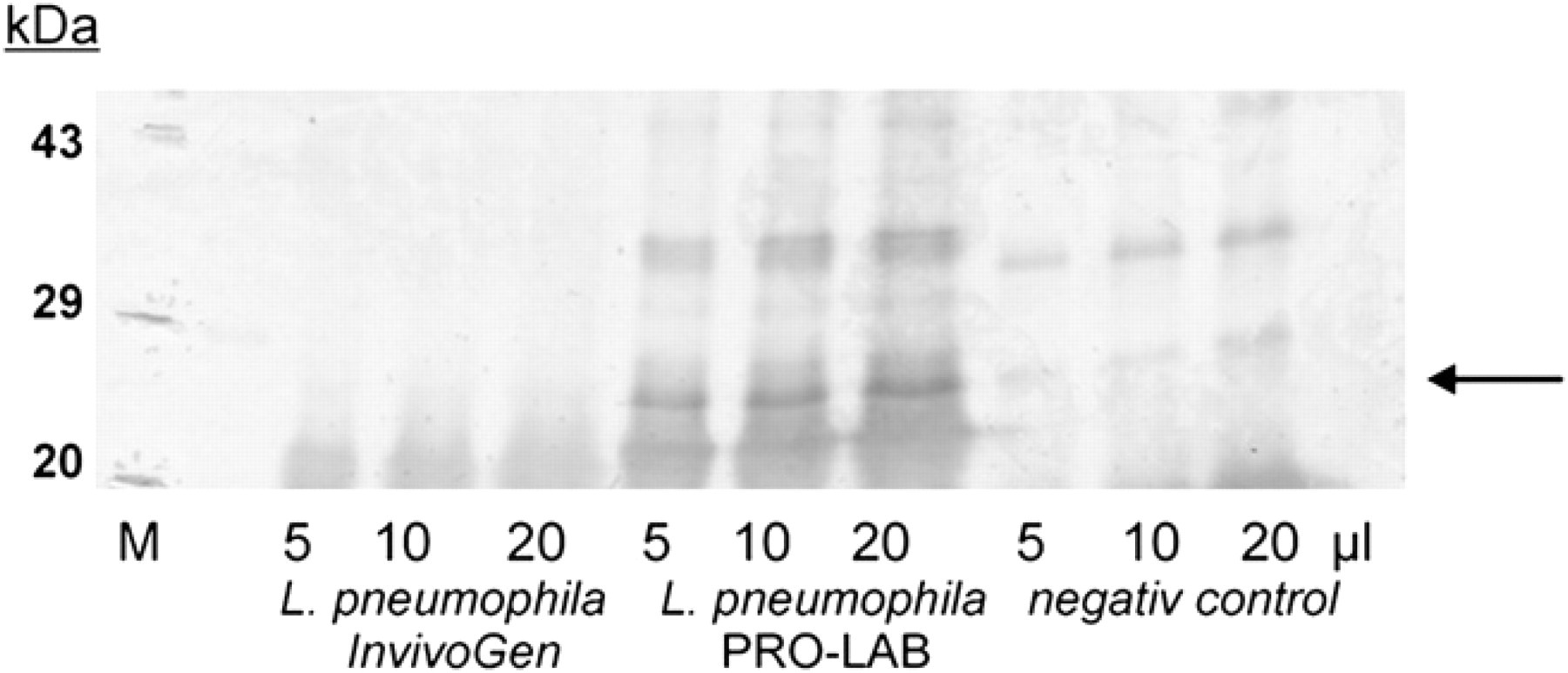
Definition of the antigen with the method of the Western blots. Immunoblot assay (Western blot) of
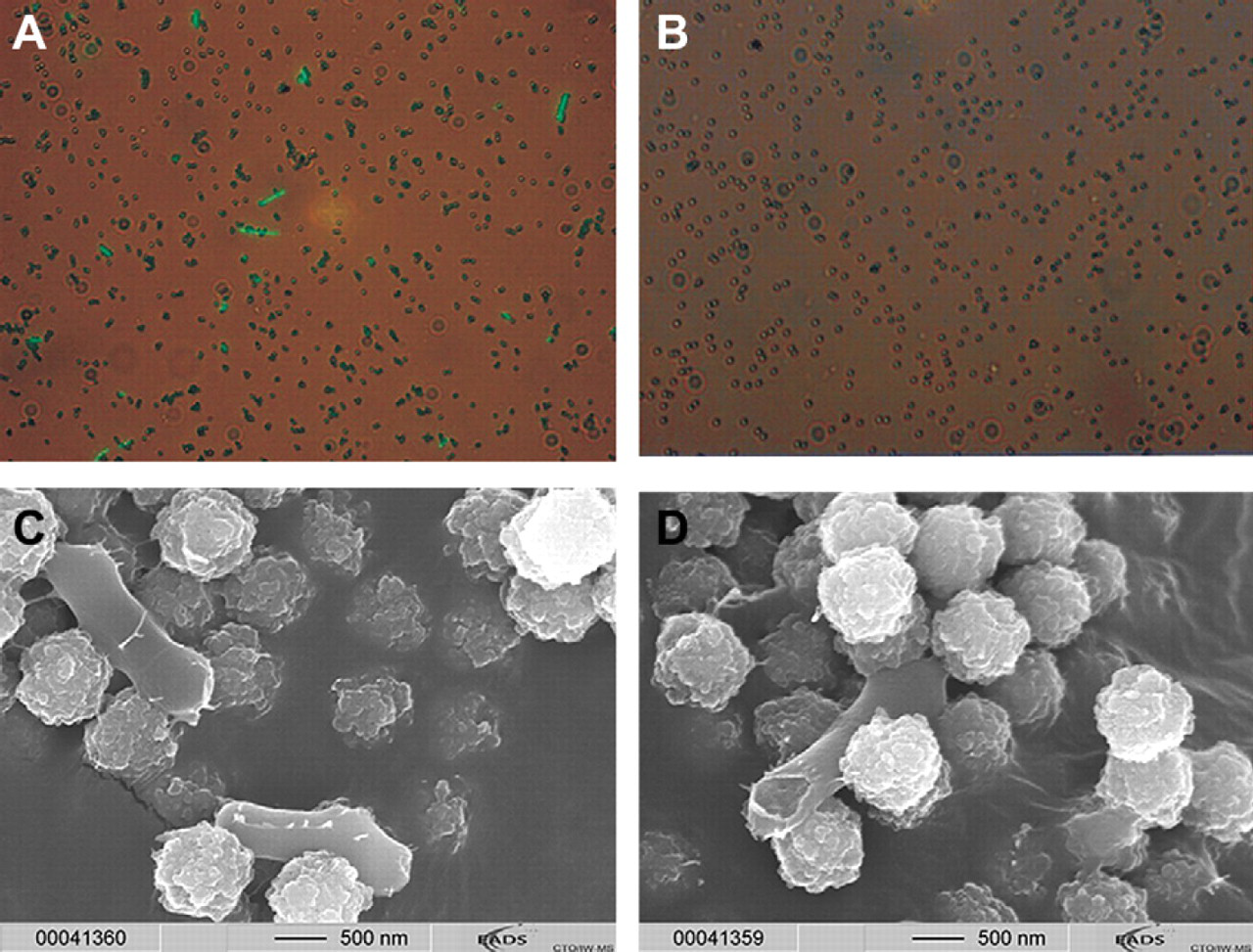
Fluorescence and electron microscopy of
In addition, electron micrographs were taken from heat-inactivated
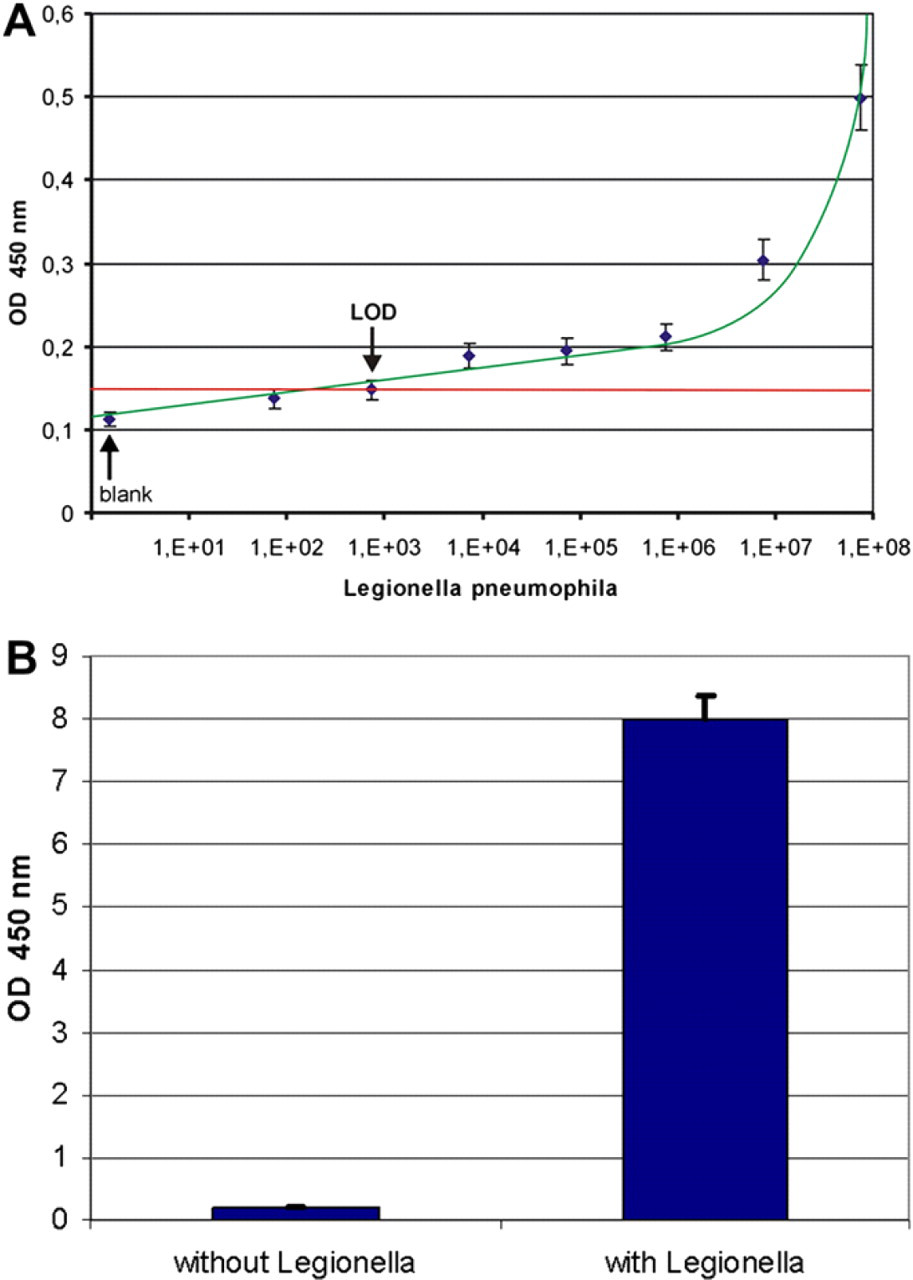
Detection of
In a next step, the assay was transferred to the automated separation system. After binding of
After staining, the bands from paramagnetic beads and from the genomic DNA dilution series were compared with each other, and the number of genomic DNA equivalents was estimated. In our PCR assay, the limit of detection was between 200 and 2000 genomes (Fig. 5A).
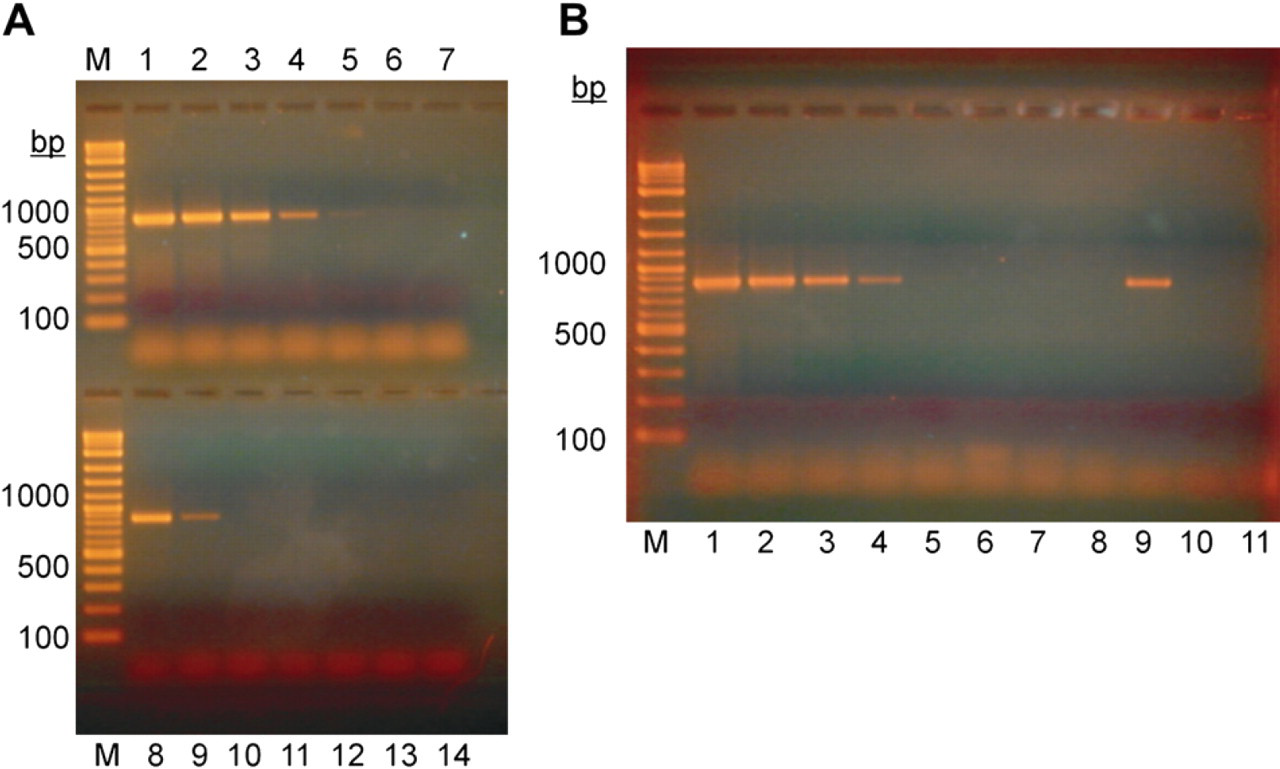
PCR Detection of
Again, we used the automated separator for the immunomagnetic capturing step. Bacteria were washed several times and concentrated from 5-mL volume down to a volume of 100 μL. For detection, 2 μL of bead suspension was transferred to the PCR reaction, and the amount of amplification product was compared with PCR products from a dilution series of the
Conclusions
We have developed an automated capturing system for microorganisms. Its practical use could be demonstrated using
We have also developed a sensitive PCR detection method for
Capturing and enrichment of bacteria and other microorganisms are necessary in environmental analysis, such as water and air monitoring. Our device is prepared for an aerosol collector being integrated. The complete system will then be able to concentrate airborne pathogens contained in several cubic meters of air into a small volume of about 20 μL of liquid.
Another advantage of using IMS is the possibility of multiplexing. By combining several types of beads coated with different antibodies, the number and type of target organisms can easily be adapted to the particular application.
The system can be regarded as a major step toward an automated standalone analyzer for environmental monitoring (water and air). It should also be possible to use it with other liquid-sample matrices, such as in food analysis (milk, beer, and others) and in diagnostics (blood, serum, urine, saliva, and others).
ACKNOWLEDGMENTS
The authors gratefully acknowledge that a part of this work has been supported within the frame of the German BMBF project ZentriLab (funding agency: VDI/VDE-IT GmbH, Germany). They appreciate the support from Dr. Müller and Dr. Waltenberger (MicroCoat Biotechnologie GmbH, Bernried, Germany) and, especially, from Dr. Hero Brahms (DRG Instruments GmbH, Marburg, Germany), who provided the HRP-conjugated polyclonal anti-Legionella antibody. The authors thank Plugit AG, HTI bio-X GmbH, and Infoteam Software GmbH for construction, hardware assembly, and software development.
Competing Interests Statement: The authors certify that they have no relevant financial interests in this manuscript, and that all financial and material support for this research and work are clearly identified in the manuscript.
