Abstract
The rise of new experimental techniques, such as high-throughput combinatorial methods, and the availability of large data sets by means of the Internet have greatly increased the amount of data that must be managed by relatively small projects. Scientific data management systems developed for large projects are often not available, suitable, nor affordable for projects with lesser resources. Increasing numbers of open-source frameworks have made available numerous options for smaller facilities to build for themselves effective and robust data management solutions. We will present considerations of these options and a case study.
Introduction
Historically, scientific computing has created many of the greatest demands for high-performance computing and data storage. For projects running on large clusters and grids, this remains true. Many custom systems have been built to handle the huge quantities of data generated by endeavors, such as particle physics or gene sequencing. These were often built in the absence of alternatives: commercial systems were often found to be incapable of scaling to meet the demands of these projects or too rigid in their design to be adapted as necessary, and there was, sometimes, a perception that the nature of the scientific data required different kinds of tools to handle its characteristic scale and complexity. These large projects often have teams of software developers at work, and articles describing them are not uncommon (e.g., Ref. 1).
Examples of smaller-scale scientific data management systems are harder to find in the literature, despite a growing need for such systems. Although large scientific projects continue to stretch the envelope of computing possibilities, an interesting expansion of scientific data management needs is happening in smaller laboratories or groups within laboratories. Advances both in technologies (e.g., measurement devices) and techniques (high-throughput and combinatorial methods) have led to exponential increases in the amount of raw measurement data to manage. Besides experimental data, online data sources, such as the Kyoto Encyclopedia of Genes and Genomes 2 and the Virtual Solar Terrestrial Observatory, 3 have made it possible to easily and programmatically access large public data sets. Managing these data sets involves new kinds of problems. The statement, “Scientific work flows usually consist of many distributed computationally intensive tasks that can be executed in parallel,” 4 does not address the issues faced by many research teams being swamped by data. There is some literature in this regard, 5,6 but most articles deal with very large data sets and computationally intensive tasks, perhaps because these large projects are the only ones with computer scientists as staff.
Many of the issues discussed in this article overlap with the tasks accomplished by LIMS. The term “LIMS,” however, has come to mean commercial systems addressing the needs of laboratories conforming to strict government rules, in particular, the United States Food and Drug Administration's regulation Title 21 CFR Part 11. In this article, we discuss a more general issue: the viability of adapting open-source software (OSS) to the end goal of scientific data management. We include a case study of such an application in the section Case Study.
Scientific Data Management
The term “data management” has a broad meaning that we narrow somewhat by identifying some end goals. These relate to the human activities of doing research and include the following:
Cleaning; reducing; validating; reformatting; annotating; or, in some other way, preparing data for storage or restoration. Making the data safe by ensuring that data is not lost, corrupted, or accessed by unauthorized means. Making the data accessible, in particular, over a network. This goal also includes visualization and authorization issues.
The term “curation” is often used for these activities, especially when discussing long-term archiving in digital repositories. 7 We use the term “data management” to comprise a broader range of tasks that include curation. The word “data” can have a broad meaning: it can mean measurements, analysis documents, calculated results, procedures, settings, provenance information, or folklore. By “folklore,” we mean information that everybody in a laboratory might know but that no one has formally recorded. Data management tasks include data identification, categorization, collection, preparation, long- and short-term data storage, workflow integration, and providing appropriate access.
Identification and Categorization
The introduction of a data management system requires the examination of what data is currently considered desirable to archive and may lead to a reevaluation of what should be saved. Conducting a formal data inventory often leads to internal discussions about data quality, standards, and workflows, and what should be available to whom. The notions of metadata, provenance, and data life span are part of these discussions.
Collection
Data must be collected from its sources and converted into digital formats when necessary. This might be as simple as transcribing notebooks into digital formats or pulling an old archive compact disc (CD) collection off the shelf, but it might require changes in physical networking, instrument configuration, introduction of data transfer applications, and gaining access to secured data. The largest quantity of data will be in digital form—accumulated measurements on hard drives, images, electronic documents, and others—but the nondigital data are often equally important and harder to collect.
Preparation
This often includes parsing, translating, and reformatting, but might include fine-grained classification based on various criteria, such as security, quality, and priority. Preparation often requires human intervention, and this must be accounted for in workflows.
Archiving and Storage
We distinguish long-term archival of data from whatever storage enables quick access for searching. The long-term value of the data should be evaluated, and appropriate storage facilities should be identified. Data, especially metadata, may need to be organized so that it can be easily searched and retrieved, say with a relational database management system. It might be desirable to encrypt sensitive data.
Workflow Integration
Workflows and data are intrinsically linked: a new workflow will change how data are managed, but the new data may lead to conclusions that the workflow must be changed. Integrating data management with workflows is an ongoing task in scientific laboratories where techniques and analysis are continually evolving.
Accessing
Access to the data is provided in the form of human user interfaces or machine interfaces. Web interfaces are usually the expected form of access for humans, and their implementation requires the development of graphical user interfaces (GUI) that are particular to each data management problem (satisfactory off-the-shelf GUIs being rare). A machine interface may be services using common network protocols or it might be an application programming interface (API) exposed to software developers. All access mechanisms are subject to both security and ease-of-use considerations.
It should be clear that software is only a part of the data management process and that implementing a scientific data management system will affect many parts of an organization's scientific process. The challenges of bringing data management to a community and managing scientific data on the scale described in this article are described in Graybeal et al. 8 and Deelman and Chervenak. 9
The tasks mentioned earlier are not unique to scientific endeavors but common to many organizations for which data management is critical. Thus, it is not surprising that, for all of these tasks, there exists a variety of OSS technologies aimed at accomplishing them. Most of these have been designed with the business community in mind (the term “enterprise” occurs often in their descriptions), but to meet a wide variety of business needs, they are often designed in a general way.
Suitability of Enterprise Software for Scientific Data Management
There may still be some perception in the scientific community that software developed for the business community is inadequate for most research purposes. For example, in Critchlow et al., 10 we read that, “Scientific applications face problems not being addressed by commercial tools.” Although this might be true for some large projects, this conclusion is difficult to justify for small- and medium-size projects. (We wish to define “small and medium” no more precisely than the common and analogous “small and medium-size business,” but, generally, we mean projects that do not have the equivalent of a full-time software engineer as staff.)
We consider the scale, complexity, metadata issues, and workflow demands of such projects in light of the capabilities of some well-known open-source systems nominally designed for commercial application.
Scale
The data generated by many projects are usually in the gigabyte to low terabyte range. The number of people accessing the data are typically in the dozens or hundreds. These sorts of demands are well within the operating parameters of large open-source projects, such as the MySQL and PostgreSQL databases and the Tomcat and JBoss application servers.
Complexity
It is sometimes suggested that scientific data are somehow more complex than those encountered in the business world. This claim seems to be undocumented. In the authors' experience, the opposite is true, as measured by the number of relational tables in the data models. Scientific data for a single project rarely requires even a dozen tables, whereas commercial applications often have more than a hundred. Furthermore, scientific data models (and workflows) are greatly simplified by being based on data that are almost entirely “read-only.” Regardless, tools to handle complexities in data are more prevalent as well. At the same time that the scale of data has increased, there has been an exponential increase in the number of OSS projects. 11 Although few are specifically directed at laboratory information management problems, many are general-purpose tools that have evolved to the point where they are suitable for adaptation to scientific purposes.
Metadata
Scientific data management, compared with business data management, often requires a greater emphasis on metadata management. We include the notion of “provenance” as part of metadata here (see Deelman and Chervenak 9 for a discussion of provenance). However, handling scientific metadata issues is usually more a matter of software development emphasis than intrinsic problems with scientific metadata itself.
Workflows
Scientific workflows are often characterized by lengthy intermediate steps, for example, chemical treatments, lengthy calculations, equipment reconfiguration and recalibration, seasonal changes, and by a variety of steps, perhaps, unique to each project. However, workflows are part of every commercial enterprise, and there are many software tools addressing those needs, including open-source tools dedicated to scientific workflows, such as Kepler 12 and Taverna. 13 One may note that the major problem with the implementation of workflows is often not the software, but rather, cultural resistance to change and the training and discipline necessary in the human environment for the new workflow to be successful.
Commercial and Open-Source Data Management Systems
Whether implementing a new scientific data management system or enhancing an existing one with OSS, the process differs from buying off-the-shelf products from vendors.
Requirements and Architecture
Data management affects many aspects of a project. The high-level design of the data management system and its architecture can make the project workflows easier or more difficult. Every sizable software project has an architecture, even if it is not recognized as such. Commercial or OSS-based systems must provide an architecture that meets project goals. Vendors hired to do the entire project will create the high-level architecture themselves and aid in creating a requirements document that will be the basis of a contract.
Those who choose to assemble their system from OSS tools will need to follow good software engineering practice in creating a requirements document that includes particular laboratory needs and addresses the issues mentioned in the section Scientific Data Management. The architecture might arise more or less naturally from these requirements. The requirement-gathering process might necessarily be long and formal, or it may be quick and casual, but the end result should be a document that can be used to create an architecture meeting those requirements.
In addition to studying requirements, those creating their own architecture will need to understand how to choose a set of components that will work together to meet the project goals. The components chosen may include a mix of open-source and commercial products (in fact, many commercial projects are assembled this way). The choices of these components may be critical to the project's success.
Evaluating Open Source
There are costs and risks associated with committing to any software framework: hence, the choice must be made carefully. Gathering information about data management tools from vendors is not difficult, as vendors of data management systems are usually very willing to provide demonstrations and literature about their products. For OSS frameworks, gathering information may require more effort, especially for those not well versed in computer software, but it is usually possible, in principle, to obtain unbiased information. Some criteria are mentioned here.
Suitability
Is the tool suited in purpose and scale to the task? For popular open-source projects, this may be easily judged. For lesser-known projects, this may require inquiry and testing. The ease and degree to which a product may be configured might be a deciding factor in its suitability for adoption. If the product will be modified, then the programming language and version may be relevant (cf. section Implementing Open Source).
Quality
Is the software of good quality? Is it robust and reliable? Does it allow for modular augmentation? If the software must be modified, then it should be designed so that modifications are not too difficult and so that they can be built and deployed easily.
Standards and Interoperability
Does the product adhere to open protocols and standards? This is critically important if it is to mesh with other products. Generally, open-source products are good in this regard, but not all standards are compatible with one another.
Codebase Stability
Projects aimed at dynamic, rapidly emerging markets may evolve quickly, leading to many version changes and corresponding implementation changes. The adoption of software from such a project will likely mean higher maintenance costs that must be weighed against the advantages of the latest features.
Project Stability
Open-source projects come and go, but some are lasting. Some are started with good intent but go nowhere. The project's programmers could all be volunteers or it might be partially or completely subsidized; there might be a risk that the project could be forked to a proprietary license in the latter case. Some projects are centered around a charismatic leader whose loss would mean the end of the project; in this case, the community is critical. Assessing a project's stability will necessarily be something of a judgment call but one that can be based partly on a project's longevity and reputation.
Community
Does this open-source project have a strong community? A strong community provides the project with vitality and is often the best source for product support. It can provide ongoing stability and be an avenue for initiating needed changes, including bug fixes and new features. However, like other communities, a project's community could be fraught with dissent, schism, and aloofness.
Third-Party Documentation
Many larger projects have third-party documentation, usually in the form of books. The existence of such books is not only a help in working with the framework but also an endorsement by the publisher of the framework's viability.
Of course, many of these considerations apply to choosing commercial software as well, but the transparency of many OSS projects may make accurate evaluations somewhat easier. Despite this transparency, as with creating the data management system architecture (section Requirements and Architecture), this is an area where expert advice may be desirable. Increasing specialization has branched information technology so that expertise in storage, networking, and web technologies cannot be expected in every laboratory or even on every software team.
Implementing Open Source
Although OSS is downloaded for free, there may still be significant work to adapt or configure the product to be a part of a scientific data management system. The difficulty of configuring a large application should not be underestimated: entire books are written about configuring some applications. However, the presence of books, online articles, and mailing lists makes it easier. Scientific laboratories are often populated by technically minded quick learners who can be drafted to do minor tasks, perhaps, under the supervision of consultants.
If the software is to be modified or augmented, then programming skills will need to be applied, whether by an in-house programmer or by hired software contractors or consultants. The laboratory manager must decide on the level of commitment to software development, weighing the costs of doing nothing, purchasing a system, or building one from open-source frameworks. In the latter case, student interns or postgraduates might provide some low-cost software development skills along with valuable expertise in the science domain.
Case Study
The National Bioenergy Center (NBC) is a part of the National Renewable Energy Laboratory (NREL), with the mission to foster capability to catalyze the replacement of petroleum with transportation fuels from biomass by delivering innovative, cost-effective biofuel solutions. 14 The importance and urgency of alternative-fuel development has grown as a result of concerns regarding world petroleum fuel supplies. The NBC has, in part, reacted to this pressure by accelerating its creation of data and information, and this has increased the need for automated solutions to the problem of data management. The NBC creates data by means of data acquisition and control systems attached to pilot plant equipment, analytical laboratory instrumentation, standard business software, and written notebooks.
Analysis
An effort to develop a simple data management system for use within a research and development organization at NREL shaped our understanding of the practical knowledge necessary to successfully implement an OSS system.
Organizational and Business Context
The NREL is divided into multiple centers with missions directed at solving different parts of the energy problem (both improving energy efficiency and investigating renewable energy sources). Each of these organizations has necessarily different approaches to scientific data collection and management; the NBC is one of these centers. The cultural environment at NREL mirrors the organizational structure; most researchers work independently or in small groups and, typically, do not rely on centralized scientific data management resources. This required the development team to pay particular attention to build user acceptance for any solution.
Influence and Control
The scope of problems was restricted to a small set within the NBC. We worked with the Biomass Analysis Section (BAS), which supports much of the chemical and physical analysis performed throughout the center, to identify problems that could be solved successfully, given the available staff and resources. Adoption by BAS personnel was required, but the applications were developed with the intent of making the solutions freely available to interested personnel outside of the BAS, the NBC, and NREL.
Underlying Issues
The NBC faces problems familiar to many laboratories: we collect large volumes of data produced by scientific instruments deployed in multiple buildings in multiple laboratories. The bulk of analytical data is produced by Agilent HPLC systems using a common chromatography data system: Agilent Chemstation, version B.04.01 (Agilent, Santa Clara, CA). The NBC did not have the means to collect HPLC output into a centralized, searchable database. The data were backed up manually onto CDs at irregular intervals. Chromatography data were not formally associated with metadata, except through paper notebooks, known as technical record books. A solution was needed for collecting, searching, and safeguarding these data.
The Agilent HPLC data are typically exported in Microsoft Excel format. Analysts reformat data from these reports and paste it into other spreadsheets designed for compositional analysis. These spreadsheets contain hundreds of formulae and take a mixture of HPLC data and manually entered data to determine the components in biomass. (Examples can be found at www.nrel.gov/biomass/analytical_procedures.html.) These analyses are performed on workstations assigned to individual analysts sitting in cubicles distributed across the laboratory campus. The information is not centralized, searchable, or safeguarded. Workflow to request analyses and report results was informal and ad hoc. This situation complicated and hindered work management by making it difficult to assign and monitor analysis.
The regulatory environment at the NBC is relatively mild. We produce little hazardous waste and do not produce food or medicine; hence, we do not have an obligation regarding QC documentation beyond our own need to produce statistically sound data for internal consumption and for publication. Therefore, most commercial LIMS functionality does not apply.
The work environment is similar to an industrial research and development operation, in that analytical procedures are moderately stable but often reassessed and adjusted. Validation processes are not rigid. Many small investigations are conducted. Researchers work autonomously and are able to resist implementation of unwanted processes; hence, solutions must carefully minimize workflow changes or risk that users will not adopt them. This also means that solutions must be carefully evaluated to ensure either that they are not applied to quickly evolving systems or that they are able to contend with process change.
Proposal and Implementation
User interviews identified several problems, which were assessed for risk and were prioritized. They concerned HPLC data, compositional analysis spreadsheet data, and analytical request data.
HPLC data needed to be placed into a searchable database with metadata, and raw HPLC files needed to be copied to a centralized, backed-up file system. The upload needed to be available from computers attached to HPLC instruments running Agilent Chemstation, and searching and retrieval needed to be available from anywhere on the NREL intranet.
Compositional analysis spreadsheets needed to have relevant data ingested into a searchable database with metadata. The spreadsheets needed to be copied to a centralized, backed-up file system, with the ability to ingest, search, and download data from anywhere on the NREL intranet. Finally, an application to request analysis was required to be available from anywhere on the NREL intranet.
High-Performance Liquid Chromatography Data Management
The client identified HPLC data as of highest priority. Data insertion and data retrieval are two separate functions; hence, they were necessarily developed separately. They also required solving more fine-grained business requirements. The steps involved are summarized in Figure 1 and Table 1.
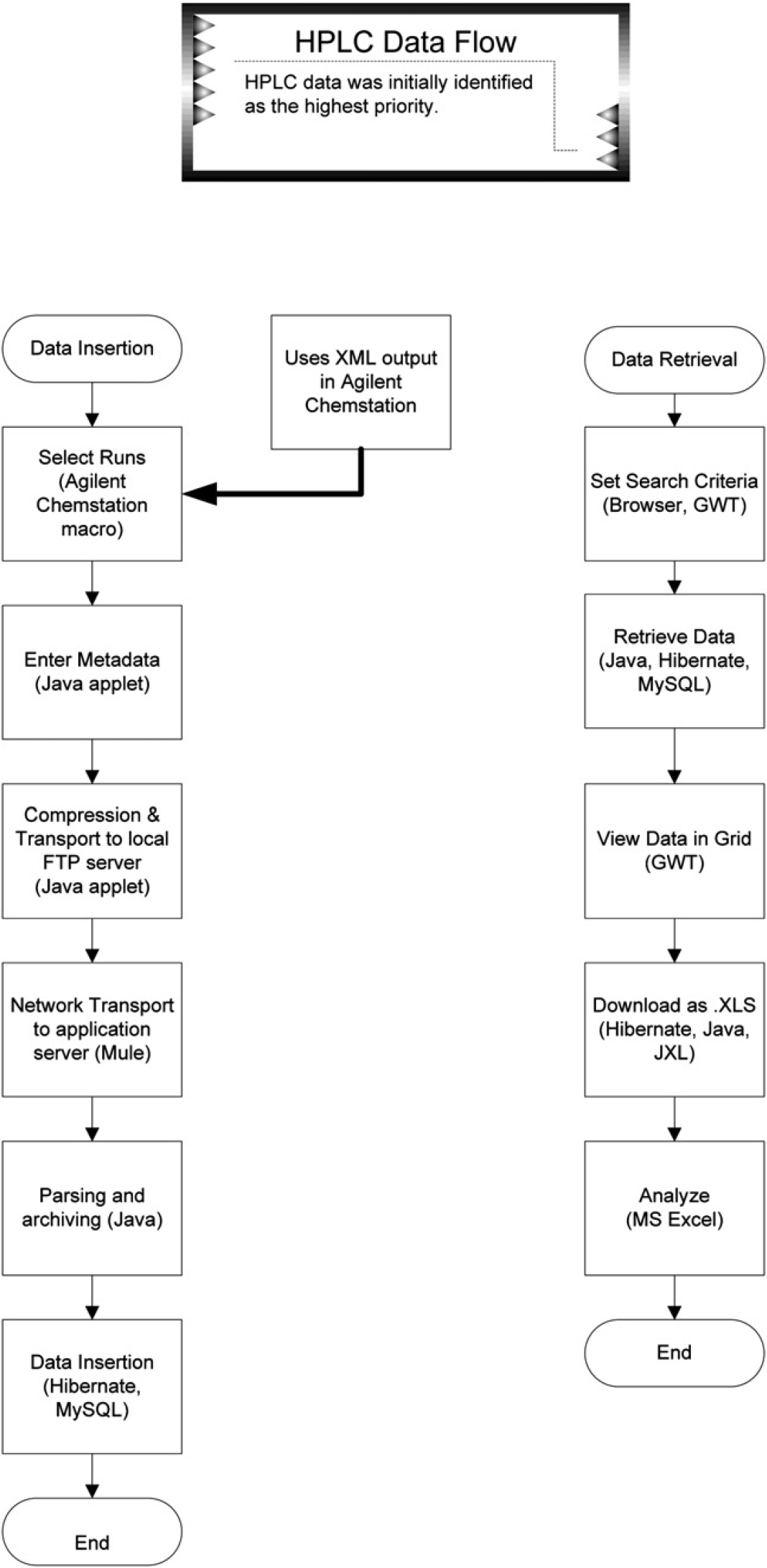
HPLC data workflow.
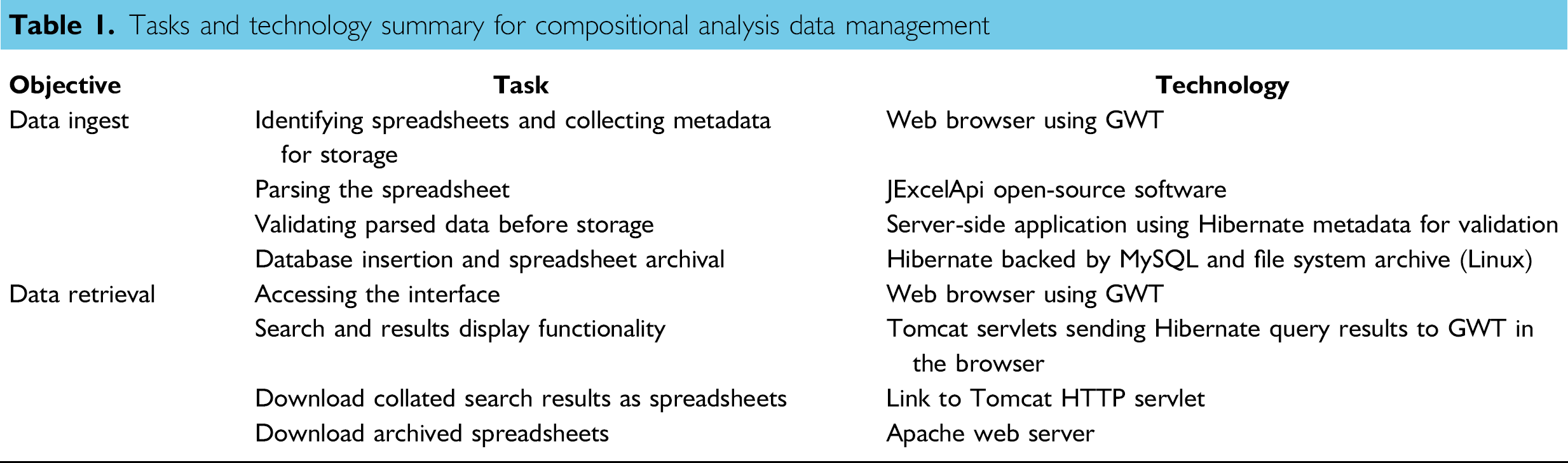
Tasks and technology summary for compositional analysis data management

HPLC data management workflow data ingestion required that users select a subset of HPLC runs from a given sequence and attach metadata for guaranteed delivery to a database and an archive. Data retrieval required an interface that would search, display, and download data and metadata.
Agilent Chemstation has a rich feature set for integration. The two features we took advantage of were the built-in macro language and the optional XML report output.
A custom menu launches a macro that generates a file containing a list of sequence data subdirectories using the Batch object, and a convenience macro opens a link to an ingestion applet. Agilent Chemstation (Agilent, Santa Clara, CA) can be configured to output data in XML format, which allowed us to parse the output using JAXP.
A signed java applet (later converted to an application) presented a GUI for entering metadata and for selecting the macro-generated file. It validates Chemstation files against Hibernate mapping files and then copies and compresses the data subdirectories, the metadata file, and the list file into a zip archive in a local FTP folder. The ESB, Mule, copies the zipped files to the server and then deletes the original. The server unpacks the file, parses the data into POJOs, and uses Hibernate to insert the data into a database. The zip archive is then moved to a repository.
The data-retrieval function presents a GUI built from Google Web Toolkit (GWT) components. The GUI design allowed for searches against metadata and HPLC data for date ranges, numeric ranges, and text expressions, and for downloading the original data and an Excel spreadsheet containing search results. The business layer interacts with the data layer through Hibernate for queries and through the JXL API for generating Excel spreadsheets for query result downloads.
Compositional Analysis Data Management
The technical problems to solve for compositional analysis ingest were to submit metadata along with a spreadsheet, validate the submission, extract relevant data from the spreadsheet, insert it into a database, and copy the original spreadsheet to an archival file system.
For search and retrieval, it was necessary to present widgets for search fields, operators, and values, a table for results, a method for retrieving copies of original spreadsheets, and a method for retrieving spreadsheets containing search results.
The interface for ingest, written in GWT, is simple and requires no host configuration. The service behind the interface was more complex; the spreadsheet copy was parsed using JXL, placed into POJOs, and ingested with Hibernate, as was the metadata, and the spreadsheet was archived. The original parser was tightly coupled to the spreadsheet architecture. Changes were later made to the spreadsheet to simplify this process.
The retrieval interface included searches against spreadsheet data and metadata. Two result sets were returned: tabular spreadsheet metadata, including download links, where at least one spreadsheet row met satisfied criteria, and tabular data containing only spreadsheet rows that satisfied criteria. The second table could also be downloaded as an Excel spreadsheet. Table 1 summarizes the tasks and technology for compositional analysis data management.
Analytical Request Data Management
NREL personnel have historically depended on ad hoc, face-to-face communication to arrange work. As the laboratory has grown, this arrangement has come under increasing pressure in response to more urgent goals and deadlines. Hence, an automated method for requesting analysis and for reviewing the requests was created (Sluiter et al., personal communication).
A satisfactory result for request management required that a custom GUI be created, necessitating tight coupling between data design and GUI. Also, a large number of choices needed to fit in the interface. A simple interface with all elements arranged on a single, long page was developed for entry and modification. Buttons to set groups of default choices were also eventually included, as was a connection to a custom sample-tracking system.
Lessons Learned
Open-source choices provide a high degree of flexibility, but this can allow a project to fail without careful oversight.
Get Your Priorities Straight
Most of the effort in the early and middle stages of the project was directed at understanding workflow and deciding on appropriate automation targets. Care must be taken to objectively enumerate priorities without prematurely attaching specific technological solutions. It is also critical to ignore the allure of a problem when deciding its priority.
We first started work on the HPLC component for two reasons: technical challenge and data volume. The HPLC problem was much more challenging technically. It was necessary to find ways to extract data from the chromatography system itself, it was necessary to move files over a network, and it required integrating several technologies. Also, the amount of data generated by the HPLC instruments was very large compared with the data from the spreadsheets; manually managing the instrument data is more cumbersome.
However, volume does not necessarily equate to importance. We failed to recognize that our compositional analysis data had much higher value to our organization. The compositional analysis data are the end product of analysis; hence, much more time and effort are invested in creating it.
The consequence of this priority is that delivery of all of the other, less-complicated, higher-value components was delayed. Although it is critical to fully investigate problems, it is also very important to judge priorities with disinterest.
Balance Necessity with Simplicity
The earliest interfaces developed in this system were unnecessarily complex. Significant effort could have been redirected by requiring that analysis tasks involving sorting, graphing, and data reduction, be performed by downloading and manipulating data in Excel or another appropriate tool.
On the other extreme, a method for placing pairs of widgets together was automated to simplify interface construction, but this method ended up being too restrictive to give good results when applied to later applications, because it forced the use of two-column grids.
In the context of this case study, the interface should have been designed to take advantage of other commonly available software tools to provide necessary functionality more rapidly and at lower cost.
Specialty Software Requires Developers
The decision to purchase or to develop specialized software does not change the need for developers. Licensing fees cover maintenance activity but not many other activities necessary for successful deployment. Extra charges apply for other essential functions, including business analysis, workflow modification, and software configuration. Vendors often charge much higher fees for this work than do other types of contract developers, because their systems are proprietary, and no other source of expertise exists. This might be acceptable when the business analysis is already complete and the product works well in a given environment without change to the “out-of-the-box” functionality. However, as the level of effort to understand the problem or to adapt the software to fit the client's needs rises, the higher cost of these specialized developers gives weight to arguments for building an in-house system using experienced open-source developers.
Conclusion
As scientific data management becomes more of a problem for small- and medium-sized projects, the use of OSS will become more attractive. Implementing successful solutions, however, still requires a significant investment in software engineering expertise. Acknowledgment of this and of the many possible pitfalls should be considered. That said, for some projects, scientific data management implementation using existing free frameworks may provide the lowest cost and best fit to growing data management needs.
Glenn Murray is a Research Professor in the Department of Chemical Engineering at Colorado School of Mines. He works as a consultant in designing and implementing scientific data management solutions, especially with OSS.
David Crocker is a Scientist II at the NREL. He has A.A., Electronics; B.S., Chemistry; and M.S., Computer Information Technology degrees.
Footnotes
Acknowledgment
Competing Interests Statement: The authors certify that they have no relevant financial interests in this manuscript.
