Abstract
Fully automating laboratory processes is a nontrivial task made more difficult by a lack of standardization between components. Each device in the system requires custom interfaces and translators to permit sharing of data and remote control by a central protocol. The BioCube System from Protedyne features the LILY data management software that enables communication between external databases, the robot controller, and integrated third-party devices. Data is formatted and passed internally as XML files to provide a generic interface for all the components. The resulting system allows the process to begin with the input of samples, and finish with the automatic export of data to a variety of data management software.
Introduction
Automating laboratory processes can provide great benefits for throughput and consistency. However, to realize the full potential of automation, it is important to integrate the devices involved in the process, as well as the data storage and processing. While in certain situations there may be good reasons for adopting an “islands of automation” philosophy, production environments typically are more suited to fully integrated solutions. Implementing a system in which peripheral devices such as incubators, thermal cyclers, plate washers, imagers, and detectors are controlled by the automation prevents errors arising from operator interaction. Additionally, centralized control of the process allows for easier operation and simplified user training. Integrating data tracking and processing, as well as the ability to use data-driven protocols, requires a dedicated robotic Laboratory Information Management System (LIMS).
Existing enterprise LIMS include products by IDBS 1 (Guildford, Surrey, UK) and LabVantage Solutions, Inc. 2 (Bridgewater, NJ). Other options for coordinating multiple devices in a lab include Peer-to-Peer 3 and Service-Oriented 4 architectures. All of these solutions enable laboratories to integrate multiple instruments and coordinate and data track samples as they are processed. The BioCube System, however, operates essentially as an automated laboratory, so it requires an additional level of data management that runs internally. The Protedyne Corporation has developed the LILY data management software, a LIMS system dedicated to the BioCube System, for that purpose. The BioCube System generates a large amount of data, which are stored internally, as a consequence of normal operation, and it coordinates the use of integrated devices during protocol execution. LILY performs similar functions as the LIMS solutions mentioned above; however, it does so to handle the internal data rather than to coordinate the entire laboratory. LILY works in coordination with those other solutions by distilling its internal data down to the parts that are critical for interfacing to external LIMS. Because the type of data critical to a user varies from protocol to protocol, LILY allows the user to define what data will be exported. Such a dynamic data management system allows more functionality than the traditional log file approach.
Integrating all the components of a process in a single system tied together with a dedicated LIMS offers several benefits to the end user. These include: a reduction in the number of errors caused by manual intervention, the simplified operation provided by a single user interface, and the increased efficiency produced by enabling reactions to be performed in higher density plates and at lower volumes. Fully automated systems can also be schedule-optimized for higher daily throughput with batch size and batch interleave timing adjusted to maximize resource use. Finally, integrated systems can automatically track sample data throughout the process and export the data into an enterprise LIMS, or custom database, or as text-based reports.
As an example, consider a system to perform Enzyme-Linked ImmunoSorbant Assay (ELISA) testing (Fig. 1). In Protedyne's case, the BioCube System is capable of processing samples from bar code-labeled patient tubes or bar codelabeled microtiter plates. The system contains one or more automatic plate washers and absorbance readers that the robot controls through standard protocol statements. Since the system tracks all the sample information as well as the detector data, it can easily link the results to the original sample. LILY allows the system to export absorbance data in a variety of formats linked to the original sample tube or well.
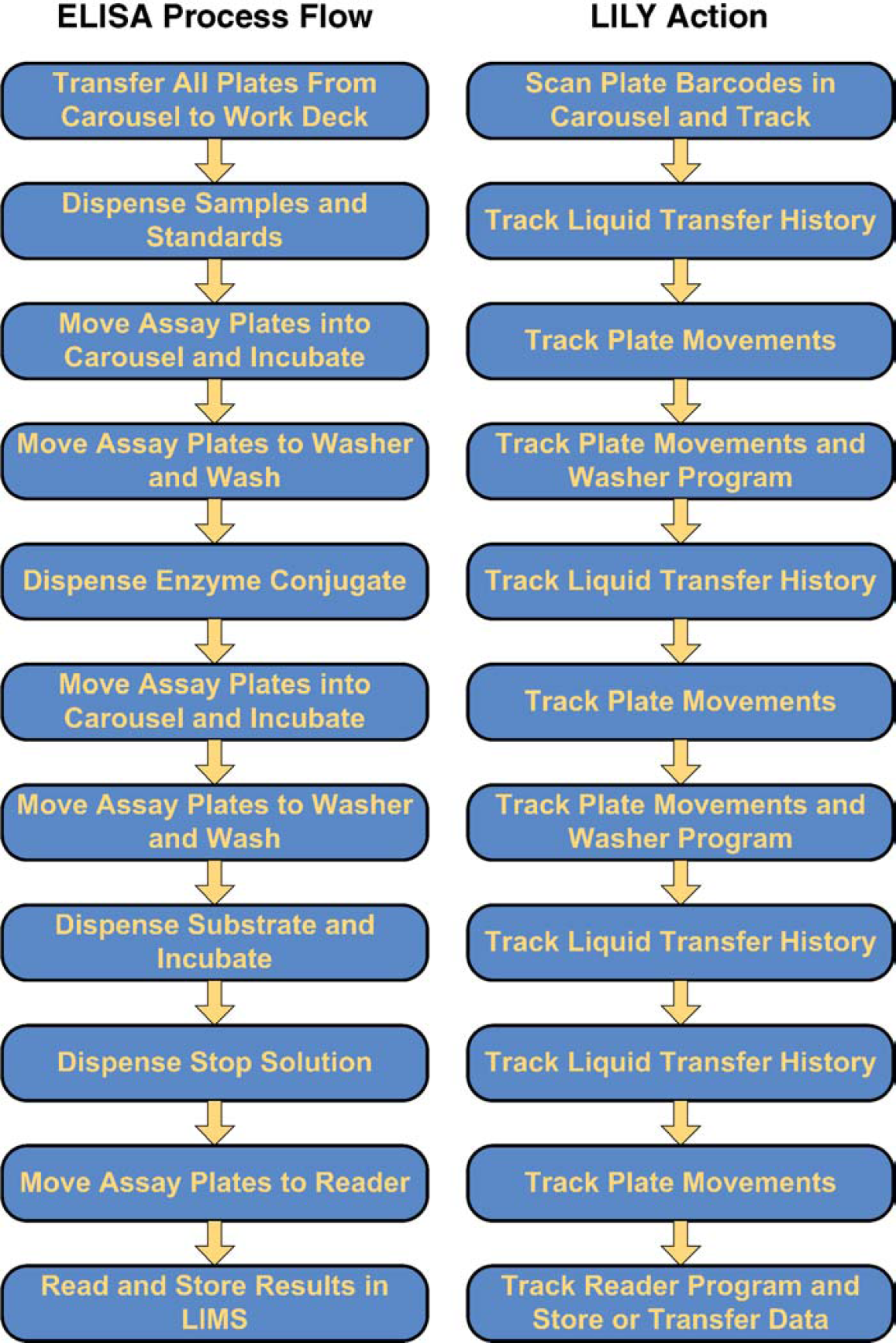
The generalized protocol for automated ELISA-type assays involves binding samples, antibody conjugates, and reagents to wells in microtiter plates. Integrated incubators, plate washers, and plate readers enable parallel processing of several batches on one system.
In the past, transferring data between devices and databases involved manually exporting and importing text files. Such a system often produced problems when each device typically exported data in a different format, and each database often required data files to match yet another format. Today, Extensible Markup Language 5 (XML) allows data to be stored in a more generic format 6 . XML was designed specifically as a data transfer language, independent of operating system, software, and hardware. It allows custom definitions of data types, and compatibility with HTML makes it easy to display the data on web pages. XML files are essentially plain text files in which the data are associated with custom tags. These custom tags can be shared across applications, a feature that makes it possible for different software to access the same information, for example, with common tags (plate_id, sample_id, temperature, etc.). XML tags can also be interpreted as commands, for example, get_carrier_status. An XML example might look like:
In the example above, “aaaa”, “bbbb”, “cccc”, and “dddd” represent the data, while the tags (indicated by text enclosed with <>) define the start and end of the data records and data fields. The tag <sample> indicates the start of the sample data, which is ”dddd”, and the closing tag </sample> indicates that the data record is finished.
Currently, there are several hurdles to establishing full system integration. A central issue is the lack of a consistent set of standards for data transfer. This means that each device must be dealt with on an individual basis, often through serial commands. A standard XML command and data set would make it easier for manufactures to integrate with one another's devices, and would also make it easier for users to perform their own integrations when modifying existing automation systems.
The immense amounts of data generated by Protedyne's customers are generally stored using methods ranging from simple network locations and files, such as Excel (Microsoft Corporation, Redmond, WA) spreadsheets, to a large distributed LIMS. To integrate the BioCube System with this wide variety of information systems, Protedyne employs its LILY data management software that is based on a Structured Query Language (SQL) relational database. LILY communicates to external systems and devices via XML, and runs in real time to ensure that both the BioCube System and external information systems receive a rapid response when making requests or posts. LILY communicates bidirectionally to allow external data to drive the BioCube System and to track sample data and results. Finally, LILY can access the Internet to enable web-based and wireless access for monitoring the automation and notification of status or errors. As a part of the BioCube System, LILY provides multiple benefits to high-throughput laboratories, such as easier sample tracking, real-time sample validation, and automated hit picking.
Technical Details
CILA
CILA System Software, a database-driven real-time control system, manages the operation of the BioCube System. CILA is an extension of the Adept AIM application information manager and Adept V+ operating system 7 that runs on the Adept controller (Adept Technology Inc, Livermore, CA) and executes commands on the BioCube System. During protocol execution, CILA retrieves front-end data (e.g., hit-picking maps) from LILY in real time and sends a constant stream of tracking data (e.g., sample transfer data) back to LILY (Fig. 2).
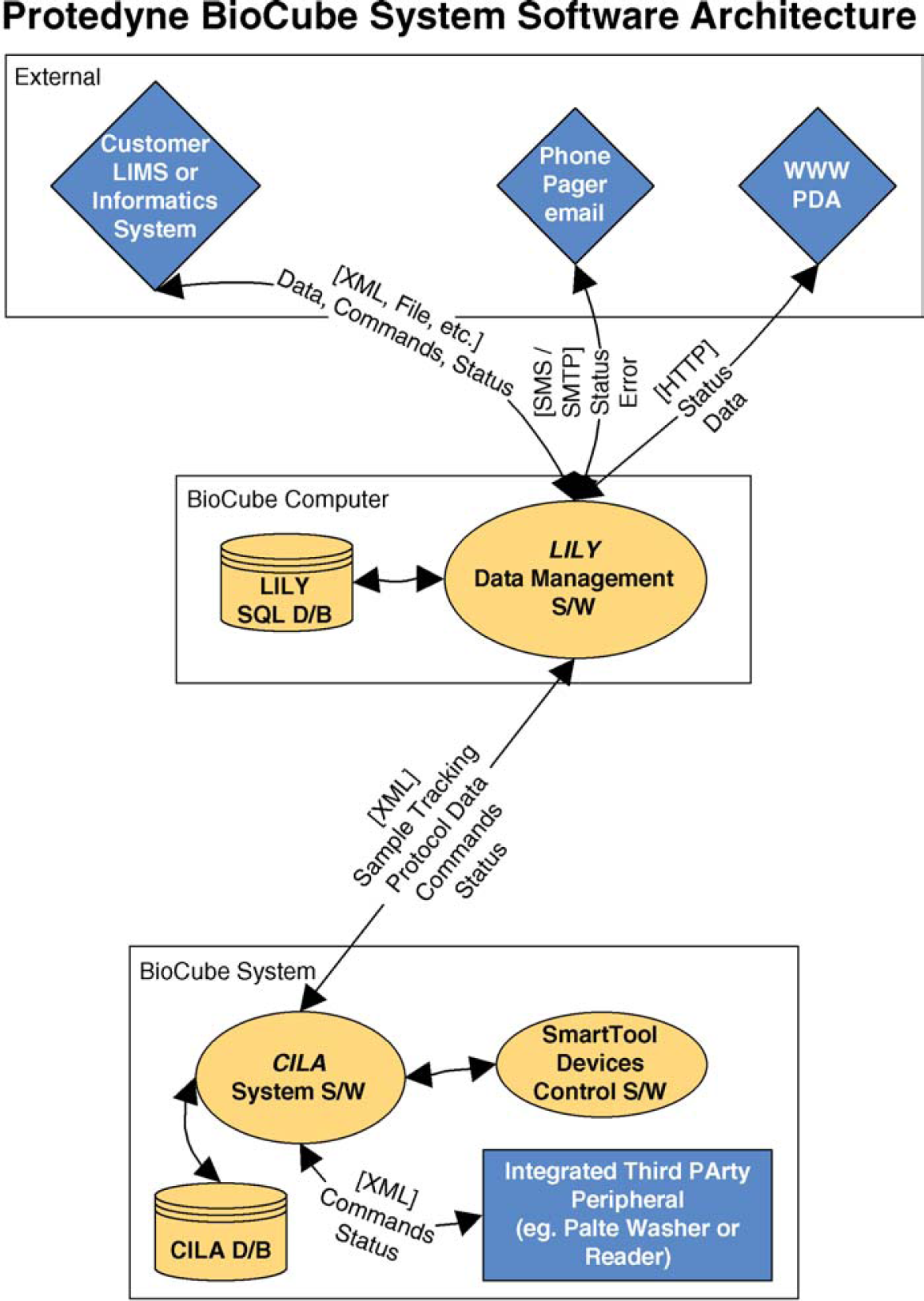
On the BioCube System, the CILA system software interacts with an internal database, SmartTool devices (e.g., plate grippers, pipettors, etc.), and integrated third party devices. It communicates with LILY, which resides on a Windows-based PC in the robot, through XML files. LILY then communicates with external systems, including customer LIMS and Web sites, by a variety of methods as required by the customer.
LILY
LILY is a dedicated, web-based LIMS designed specifically for the BioCube System, which handles both front- and backend sample management. Users can access LILY through (1) a browser interface that provides real-time BioCube System status information, (2) data files (text, spreadsheet, etc.) that can take the form of LILY-generated reports for export or LILY-interpreted information files for import, and (3) a standard XML command set that allows simple real-time integration of the BioCube System with a preexisting LIMS.
Internally, a web application resides on the server to interpret a proprietary set of XML commands for database access. This application is WebService-enabled 8 to provide a subset of the available functionality via Simple Object Access Protocol (SOAP). 9 A Web Service Description Language (WSDL) 10 definition of the WebService capabilities is available to Protedyne customers. LILY also incorporates an Extensible Stylesheet Language (XSL) 11 translator layer that allows data to be imported and exported through a variety of formats. The XSL translator can alter elements in XML documents, as well as change how the XML documents are displayed (e.g., in HTML). Using the two functions, the translator reformats XML documents that are sent out of LILY either as HTML for display or XML for communicating with other instruments and LIMS (Fig. 3).
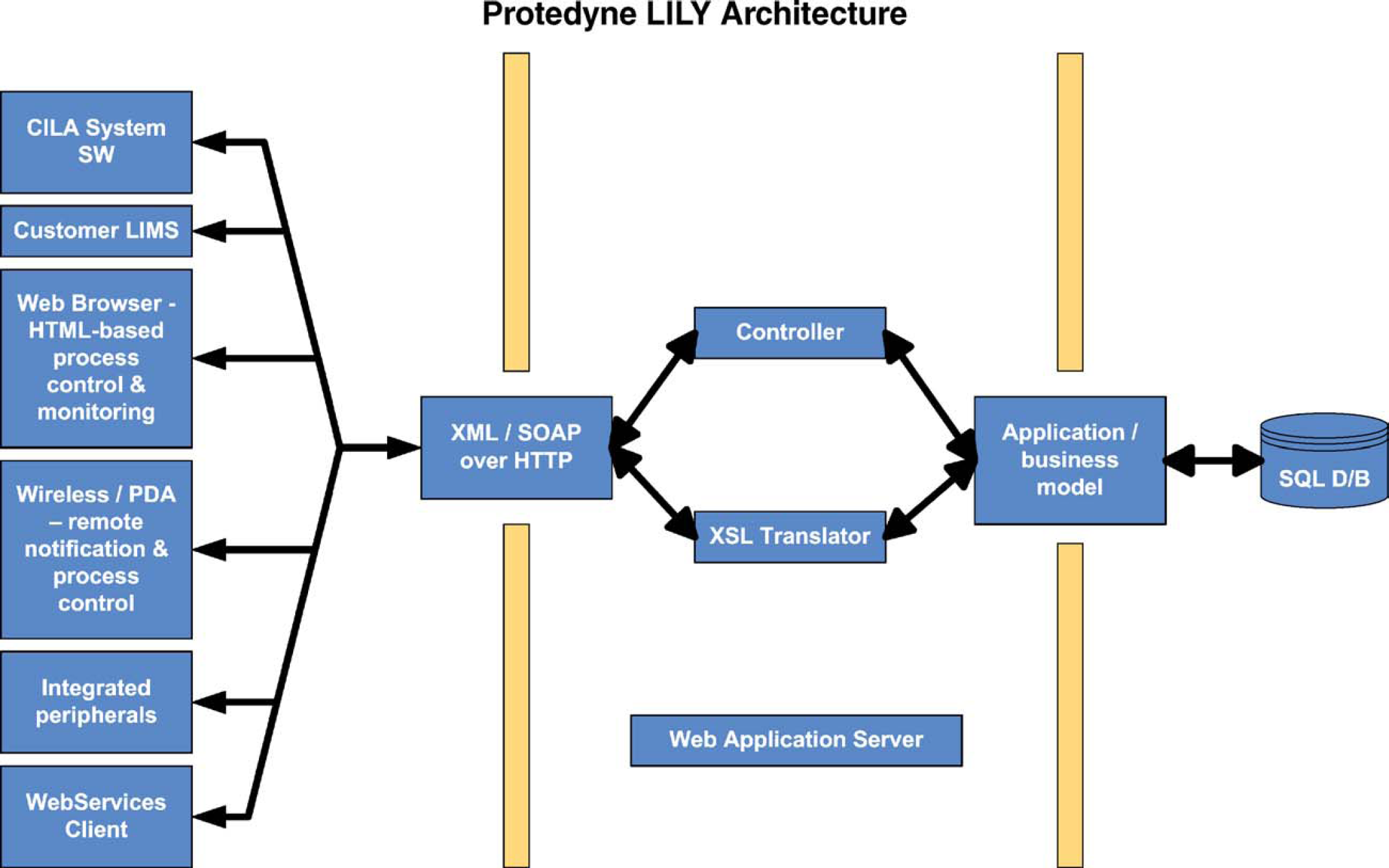
The various clients and devices shown on the left communicate with LILY through XML files. LILY interprets the files and acts as an interface for the internal SQL database. Depending on the application, XML-based commands can either modify the database (i.e., store data) or pass a request (i.e., to retrieve data and format it for export).
LILY allows multiple levels of sample management: offline, self-contained operation; online data tracking; sample validation; and recipe-based, data-driven protocols.
System Components
Discussion
In the example ELISA assay, samples enter the system in bar-coded microplates. Information, such as sample bar code, reagent bar code, volume pipetted, and current protocol step, is stored as the protocol executes. After the samples are transferred to assay plates, which are also bar code labeled and tracked, the assay plates are sent through rounds of reagent and buffer addition, followed by washing steps in integrated plate washing devices. At the end of the protocol, the assay plates are sent to an integrated microplate reader, where the absorbance data is stored in the database and linked to the original sample plate.
The BioCube System is designed to interact with these integrated devices (Fig. 4) through several methods. In the simplest method, using a microplate reader as an example, the reader is connected via an RS232 cable and serial communication. Individual commands are sent through CILA statements to load plates, set wavelength, measure absorbance, etc. The BioCube System can also interact through the device's native software by calling programs already installed on the system. For example, an integrated plate washer might have several programs for different wash cycles. CILA statements would call individual programs as needed in the protocol by passing the program name as a variable.
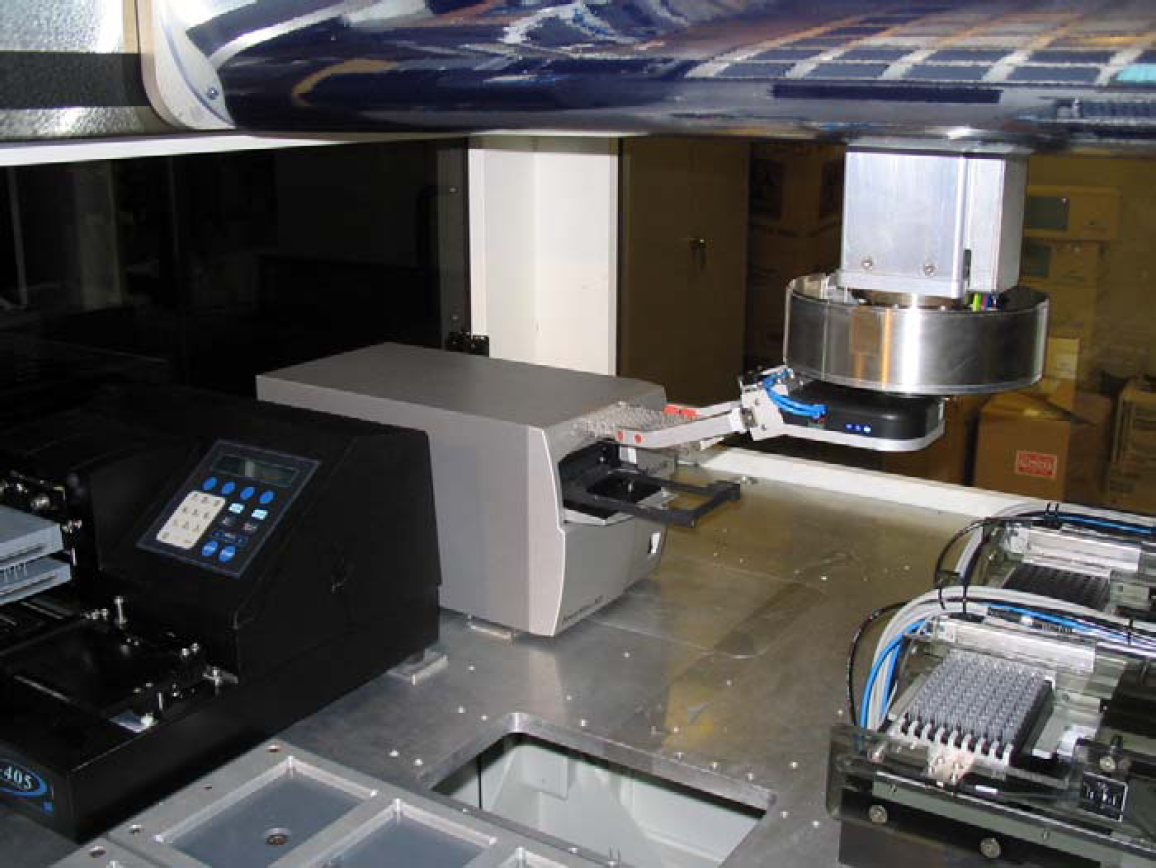

The BioCube System solution often includes integrated peripheral devices. In the case of the ELISA assay (see Fig. 1), a third party plate washer (left side) and reader (center) are mounted on the deck where they are loaded by the robot. The devices are controlled by the protocol through the CILA system software.
Since LILY tracks the system status and progress during protocol execution, a user can submit a query and have the results exported as an HTML file (Fig. 5). Similar queries performed through LILY's web interface can show the history of a particular well (Fig. 6). For example, an operator may use the reagent history information for a specific sample's results in order to interpret the data or debug failed experiments.
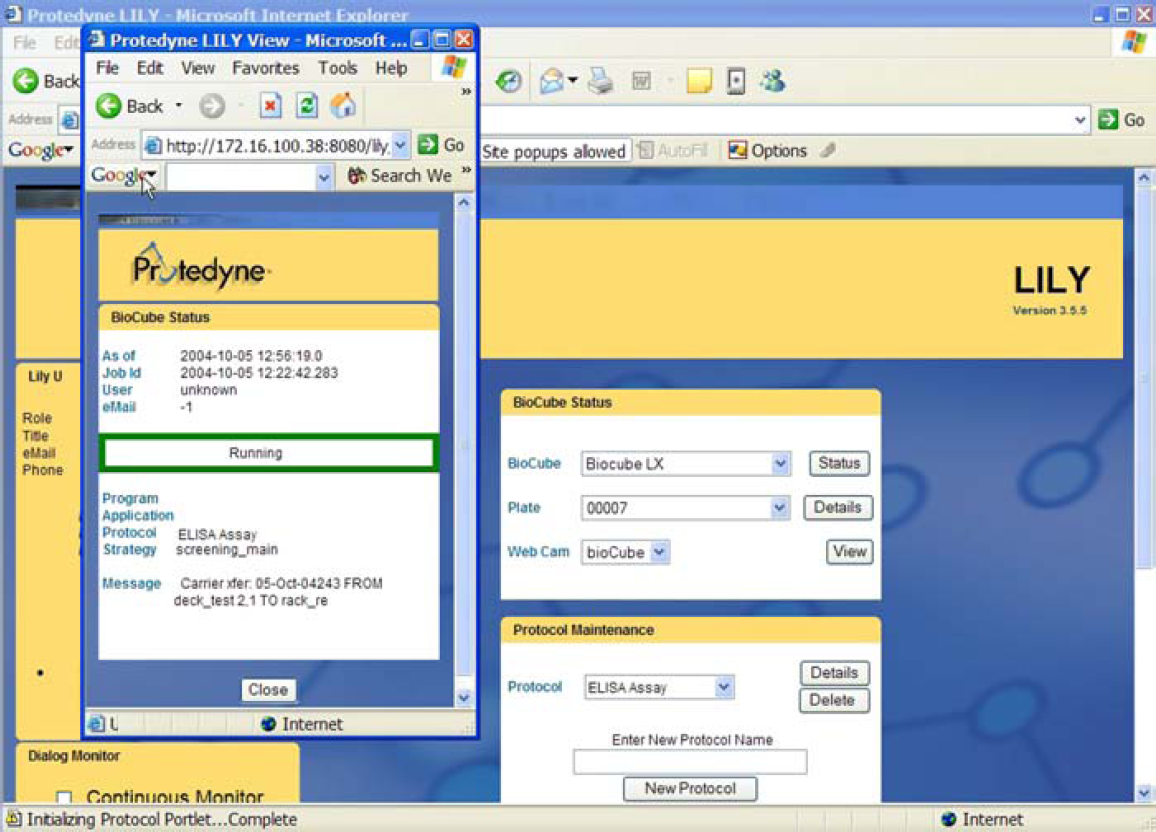
LILY's web-based interface allows users to easily monitor system status from any location, and can be customized to provide additional information.
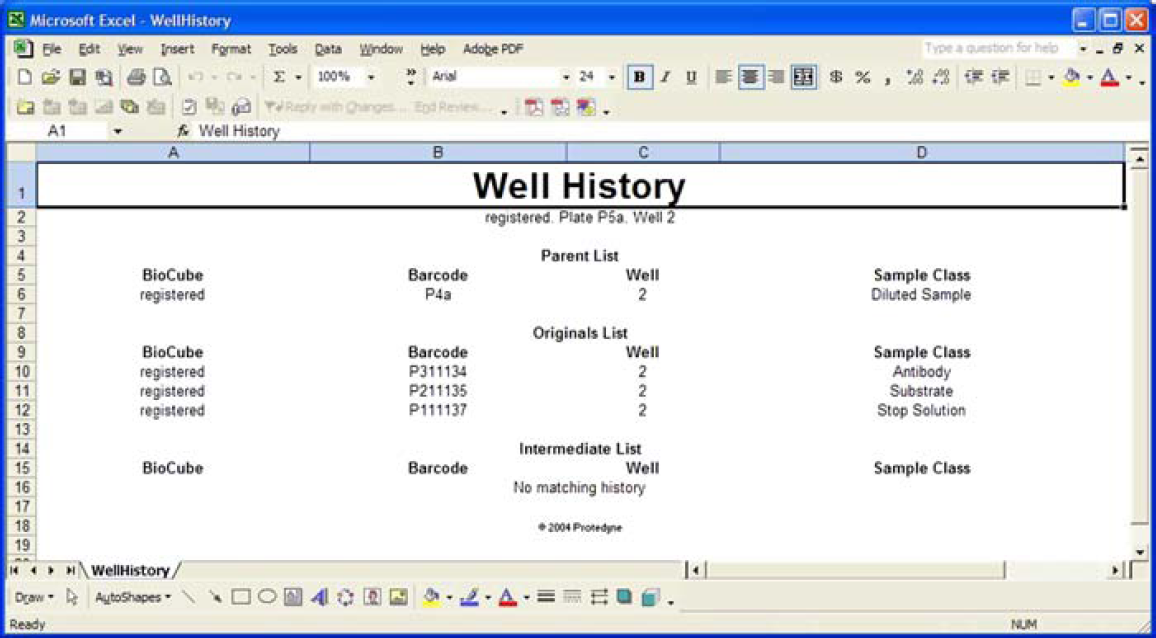
LILY's web-based interface also allows users to generate reports of tracked data in multiple formats, for example, the history of a particular plate and well in a Microsoft Excel spreadsheet format. Reports can also be generated as HTML files for platform-independent viewing.
Information doesn't just flow into the internal database and get pulled out manually; the system can be set up to automatically send XML-formatted data out to external LIMS for offline storage and processing, to users as alerts, and to control protocols running in data-driving mode. XSL translation allows LILY to format the data to be readable by external LIMS, spreadsheets, and so on, but some users find it most convenient for their LIMS to query LILY and transfer data automatically. LILY's WebServices capabilities can send real-time alerts via electronic notification such as e-mail, pager, and text messaging, to alert operators of errors or required interaction.
Finally, the combination of integrated devices and dedicated LIMS allows the BioCube System to generate data, reduce it in real time, and use the results to drive subsequent protocols such as closed-loop hit-picking. 16 Protocols that are either data-driven or that make use of run-time LILY access also include LIMS (internal or external)-supplied recipes, such as DNA concentration normalization, and sample validation at run time, such as comparing clinical sample bar codes to the accessioning database before performing requested assays.
Although integrating automation platforms, LIMS, and scientific instrumentation has been possible in the past, the various parts of the system were not designed to share data easily. The BioCube System serves as a model to demonstrate how data and control statements can be packaged as XML files to facilitate system integration. Even if the various components of a large system use different data tags, industry-wide standardization on the XML format will make complex projects easier to design, build, and maintain. While our current methods of intrasystem communication allow such integration, Protedyne is continually improving the interface to ease the process of linking to an external LIMS. Even while wrapping the XML interface into .Net 17 protocols, LILY still preserves the underlying universal XML interface.
Conclusions
In developing the BioCube System for customers, Protedyne has found that fully integrated solutions are often the most efficient choice. To ease the integration process, we have developed an internal method of sending data back and forth between the various parts of the system in XML format, whenever possible. However, we often find it necessary to write custom interfaces for each new device we integrate. The adoption of an industry-wide XML standard would make systems integration more generic for the industry as a whole.
