Abstract
Progress in laboratory automation depends not only on automating the physical aspects of scientific experimentation, but also on the intellectual aspects. We present the conceptual design, implementation, and our user-experience of “Adam,” which uses machine intelligence to autonomously investigate the function of genes in the yeast Saccharomyces cerevisiae. These investigations involve cycles of hypothesis formation, design of experiments to test these hypotheses, physical execution of the experiments using laboratory automation, and the analysis of the results. The physical execution of the experiments involves growing specific yeast strains in specific media and measuring growth curves. Hundreds of such experiments can be executed daily without human intervention. We believe Adam to be the first machine to have autonomously discovered novel scientific knowledge.
Introduction
We wish to automate all aspects of laboratory science, not just the physical experimentation, but also the intellectual aspects of hypothesis formation, experiment planning, and results analysis. A “Robot Scientist” is a physically implemented robotic system that applies techniques from artificial intelligence (AI) to execute cycles of automated scientific experimentation. 1 This contrasts with standard laboratory automation that normally focuses on just the physical aspects of experimentation. Our Robot Scientist “Adam” executes, with minimal human intervention, a complex combination of operations on yeast cell cultures at medium to high throughput and, moreover, is capable of modifying those operations according to the behavior of the organisms. 2 Again, this contrasts with standard laboratory automation that is normally characterized by medium- or high-throughput execution of a linear sequence of a relatively small number of different operations. 3 5
Automating Scientific Discovery
Automation has been integral to many of the changes in human society since the 19th century. The advent of computer science in the mid-20th century has made practical the idea of automating aspects of scientific discovery. Computers were initially used to automate simple linear processes, for example, to collect and process laboratory instrument data, perform astronomical calculations, and create ballistic tables. Later, AI began to be used to automate aspects of planning experiments and analyzing results. Meta-DENDRAL, developed in the 1960s, was the first automated system for scientific hypothesis generation. It used inference to suggest chemical structures responsible for mass spectrometry data. 6 BACON, 7 ABACUS, 8 FAHRENHEIT, 9 and IDS 10 were all impressive examples of automated data-driven discovery systems that could discover scientific laws as algebraic equations. A more recent example uses iterative cycles of algorithmic correlation to distil natural laws of geometric and momentum conservation, using data captured from the motion-tracking studies of a range of simple and complex oscillators and pendula. 11
However, none of the systems described fully “closes the loop;” they either do not collect their own data, or do not use analyzed results to update their knowledge base, or do not automatically perform cycles of scientific discovery.
Our Robot Scientist “Adam” goes a step further than these other systems and also automatically performs experiments. Adam uses a detailed logical model of yeast metabolic pathways, 12,13 along with bioinformatic and AI methods to form sets of hypotheses about yeast functional genomics. Adam then designs and executes physical experiments to test these hypotheses. The experimental observations are optical density (OD) readings, from which growth curve parameters are derived and automatically analyzed by statistical and machine learning methods, and the results are used to confirm or refute the hypotheses. Adam then decides whether to execute further cycles of hypotheses generation and experimentation, or whether it can infer new knowledge and update its model (see Fig. 1). Human intervention is limited to creating yeast library stocks and supplying consumables.
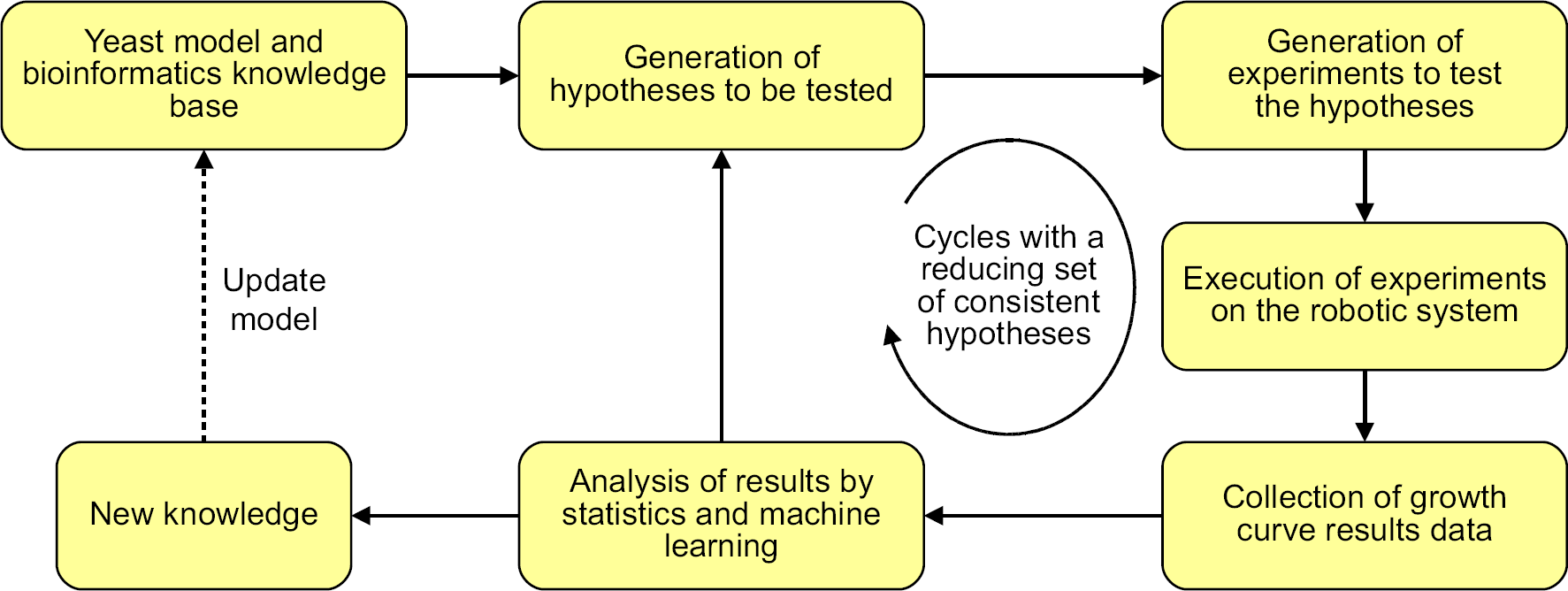
Hypothesis generation and elimination.
Why Yeast?
The focus of Adam is on generating novel biological knowledge. We therefore deigned Adam to work with the model organism Saccharomyces cerevisiae (baker's yeast). It has a relatively small genome, and it is the best-understood eukaryotic organism. It grows fast and is ideal for experimentation in microtiter plates. It also has an unrivaled collection of mutant strains. The genetically modified haploid strains used in Adam's experiments each have a single gene deleted, and Adam's library contains over 4500 such strains. 14 More than 900 yeast genes still have unidentified functions 15 and there are reactions in the metabolic pathways that are catalyzed by enzymes for which related genes are unidentified (referred to as “locally orphan”).
System Description: Technologies and Methods
The robotic system executes iterative cycles of sample preparation and monitoring (see Fig. 2) and comprises numerous pieces of automated equipment (see Fig. 3) arranged as three subsystems (see the next section). AI software creates plate layouts that specify yeast strains, the required volumes and concentrations of growth media and metabolites, and the layout of the experiments on the microtiter plates. The plates are then created and processed, with growth data automatically recorded, and then analyzed to confirm or refute hypotheses. Finally AI software decides whether to perform further cycles or whether new knowledge has been inferred.
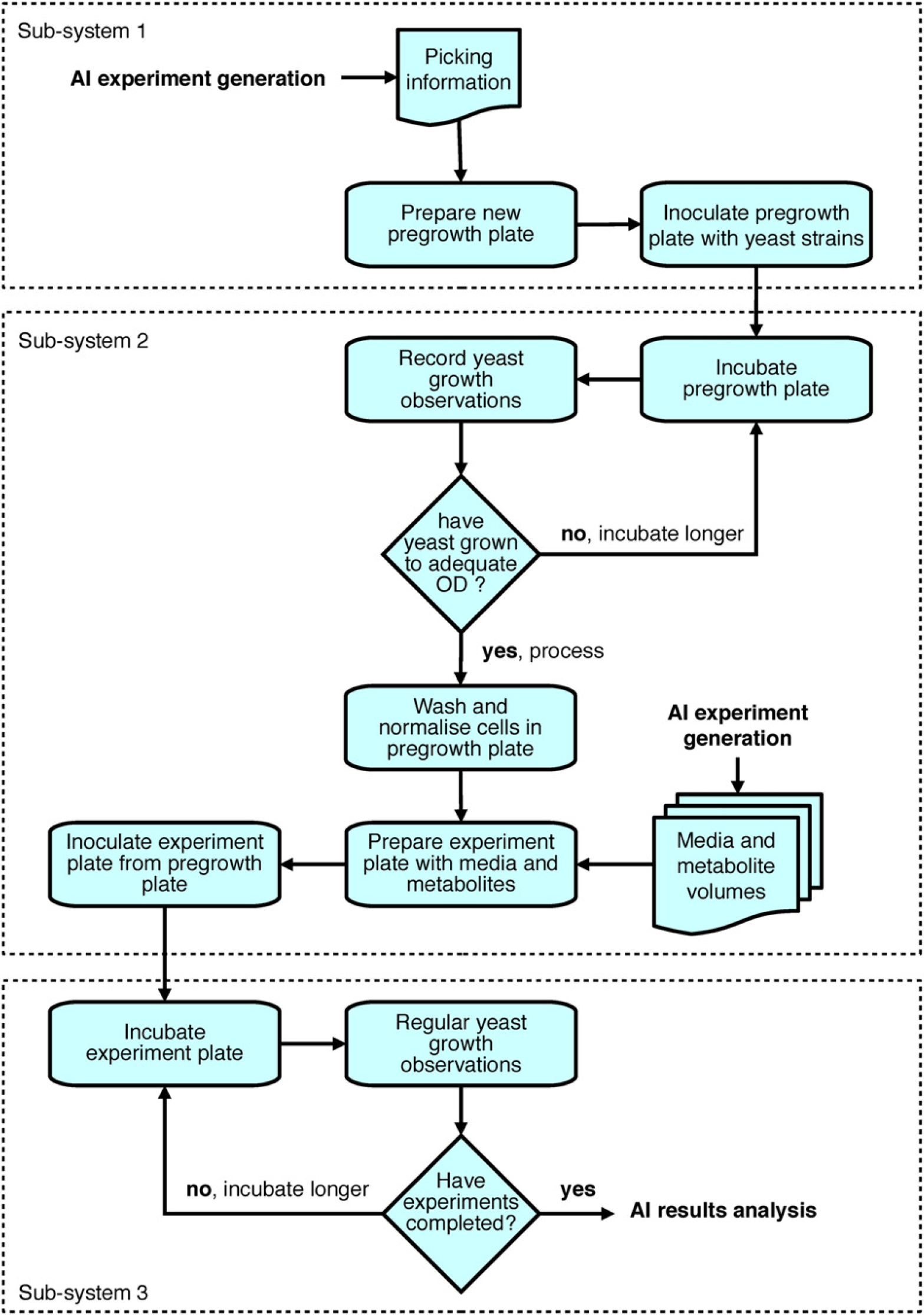
One cycle of automated experimentation for a single microtiter plate.
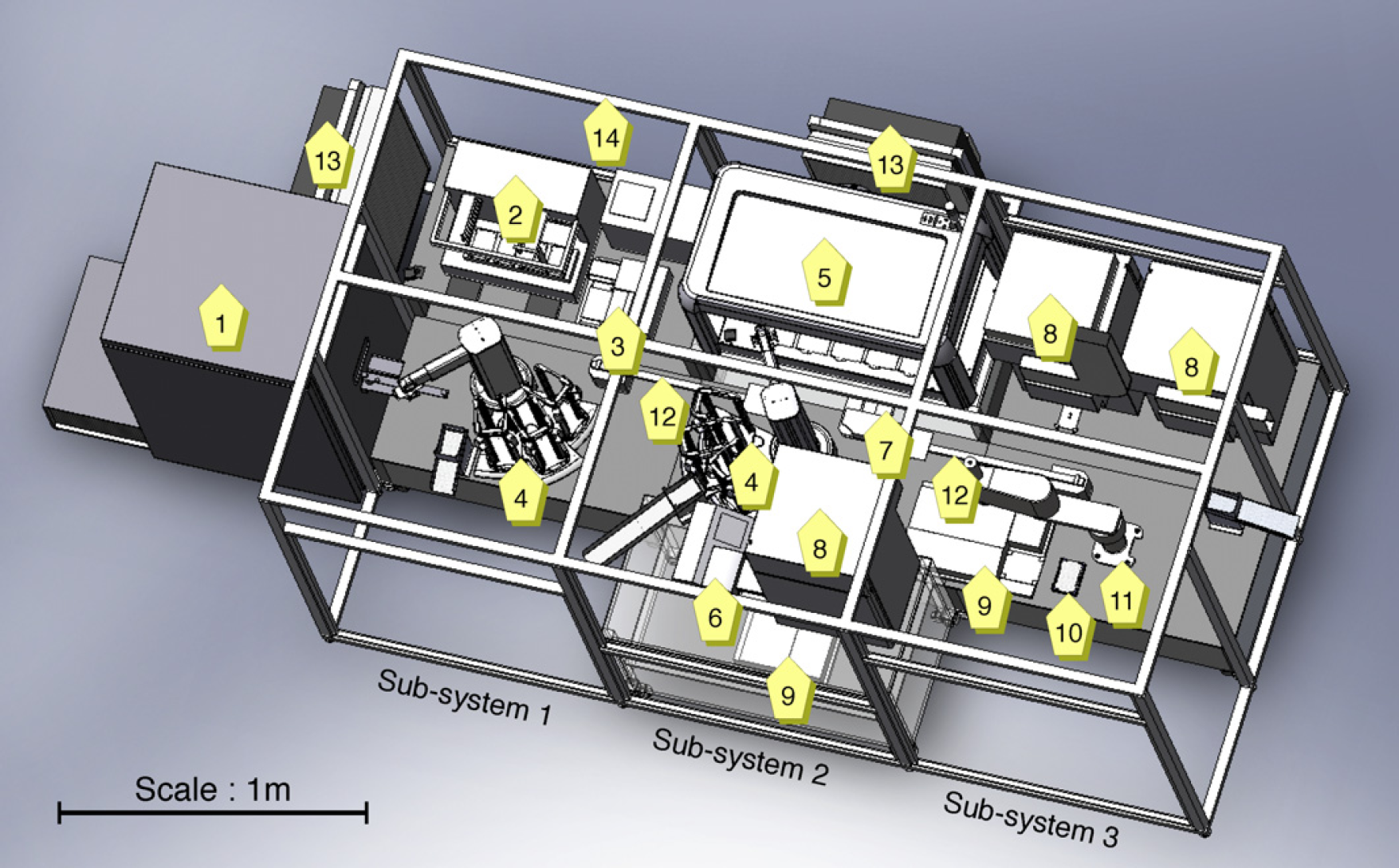
Layout of the robotics and instrumentation in the Adam Robot Scientist: (1) Liconic STR602 freezer, (2) Zymark Presto liquid handler, (3) Thermo 384 multidrop, (4) two Caliper Twister 2 robot arms, (5) Caliper Sciclone i1000 liquid handler, (6) Bio-Tek 405 plate washer, (7) Velocity 11 VSpin plate centrifuge, (8) three Liconic STX40 incubators, (9) two Molecular Devices Spectramax 190 plate readers, (10) Variomag plate shaker, (11) IAI Corporation Scara robot arm, (12) two pneumatically actuated plate slides, (13) two high efficiency particulate air (HEPA) filters, and (14) aluminum and rigid transparent plastic enclosure.
Process Description
The robotic system comprises three subsystems. Subsystem 1 picks selected yeast strains from frozen library microplate stocks and inoculates pregrowth microplates containing YPD-rich growth medium, a according to a “picking” file generated at the plate layout design stage. Subsystem 2 incubates the pregrowth plates and monitors cell growth, allowing the yeast to multiply until cell concentrations in most of the wells have reached a predefined OD measured at a wavelength of 595 nm. It then harvests and normalizes the yeast cell concentrations, dispenses the required media and metabolite solutions into the experiment plate according to the relevant “volume” files generated at the plate layout design stage, and inoculates the experiment plate with the yeast strains from the pregrowth plate. This starts the experiments that will be monitored by subsystem 3. The pregrowth plate is then removed as waste. Subsystem 3 incubates the experiment plates and monitors cell growth via regular OD readings; after 72 h the microplate is sent to waste and growth curve analysis triggered for both the pregrowth and experiment plate results.
Major Software Components
The availability of the required metabolites and yeast strains were ascertained and then Adam designs the physical experiments. Not all hypotheses in each hypotheses set can necessarily be translated into experiments. The end result of this process is a set of viable “trial” groups of individual “tests,” where a trial is related to a specific hypotheses set. A typical trial is made up of four “test” conditions: deletant yeast strain with metabolite, deletant yeast strain without metabolite, wild-type yeast strain (BY4741) with metabolite, and wild-type yeast strain without metabolite. This experimental design enables Adam to compare the growth of the deletant strain in the presence of the relevant metabolite with the growth of the standard wild-type. Each test also has a number of “test instances,” which is the requested number of replicates to be run to give statistically meaningful results. An evolutionary multiobjective optimizer 19 can then be used to determine which trials can be run together, without changing the metabolite solutions (the system can dispense no more than eight different metabolites or medium components), using factors, such as trial priority, number and identity of metabolites required, availability of metabolites, their cost, and suggested size of the “trial set.”
Parametric and nonparametric statistical tests, random forests, and Monte Carlo resampling (used to establish significance) are then applied to determine whether there is a significant difference observed when adding the metabolite(s) to the deletant strains compared with the wild-type. This enables Adam to decide which hypotheses have evidence consistent with them and which do not.
Control of Robotics and Instrumentation
The equipment control software was provided by the system integrators, Caliper Life Sciences. Three instances of the Zymark Control Supervisor (ZCS) software using “iLink” protocols are executed (one for each subsystem), using separate PCs running Windows XP. Each iLink method interacts with an MS Excel spreadsheet that stores system control and plate tracking data, and also with the instrument control programs (ICPs) that control individual instruments and robots. Some ICPs are as supplied by their manufacturers but most were developed by Caliper. A fourth PC coordinates the three independent ZCS instances via Caliper software written in MS Excel Visual Basic, providing the nondeterministic action of subsystems required by unpredictable growth and a simple graphical user interface for user interaction, error handling, and status monitoring. Experiment data are output in the form of text files containing the OD readings from each Spectramax plate reader (SoftMax Pro software v. 4.8; MDS Analytical Technologies, Concord, Ontario, Cananda).
Results
We believe Adam to be the first machine demonstrated to be able to discover novel scientific knowledge. Adam has hypothesized and experimentally confirmed the function of 12 genes encoding locally orphan enzymes in yeast (see reference 2 ). For three of these genes we manually confirmed that Adam was correct by purifying the protein product and demonstrating enzymic function. For six more of the genes we discovered that strong empirical evidence already existed in the scientific literature that supported Adam's conclusions, that is, that they were not in fact true local orphans, and that Adam only identified them as such because its bioinformatic database was incomplete. This inadvertently provided us with positive controls showing Adam's methodology does work. For one hypothesis Adam came to an inaccurate conclusion; this was because the hypothesized gene candidate YIL033C (BCY1) was predicted to be a glutaminase (enzyme class 3.5.1.2), and Adam experimentally confirmed this using 11 metabolites predicted to have differential effect on a glutaminase deletant. However, YIL033C has a cAMP-dependent protein kinase subunit that is involved in regulating metabolism, so this may be sufficient to explain the same phenotype. This exposes a weakness in Adam's current metabolic model that does not include control mechanisms. However, it is possible that YIL033C is both a kinase and a glutaminase, and scientific evidence also exists to support this theory (see reference 23 ). See our website for more details on Adam's results, its hardware, and its software. c
Discussion
Procuring Laboratory Automation
For the benefit of those new to laboratory automation we give here a brief summary of the procurement process.
It is important for the user to carefully research and define the proposed biological application, manually testing key experimental aspects where possible. Then he or she should investigate the availability, suitability, and capital, running, and maintenance costs of any potentially appropriate equipment. An outline design will help clarify system requirements, including nonfunctional aspects such as access for cleaning and maintenance, capacities for consumables and wastes, and ease of their replenishment and removal.
Integration companies should then be invited to submit proposals against detailed requirements documentation, in accordance with relevant procurement legislation. The selected integrator should clearly understand the requirements, be happy to work closely with the end-user during the design and build, and afterward to refine the system. Proposed control software should be reliable, user-friendly, flexible (to allow later refinements and improvements to the application), expandable (to allow additional instruments or functionality to be added subsequently, if required), have good error reporting and recovery, good event tracking, and be fully maintainable.
Subsequent stages involve the build of the full system, integration of the control software, factory acceptance tests (FAT), delivery to end location (preferably by a specialist equipment removals company), site acceptance tests (SAT), commissioning, refinement of biological application, and integration with the laboratory software. The user should detail the FAT specification, for agreement with the integrator, and this should specify a comprehensive set of tests to exercise all the basic capabilities of the system and associated equipment but normally without organisms and chemicals. The SAT should repeat the FAT and then test the ability of the system to execute the required biological protocols to an acceptable standard of reproducibility in the laboratory environment, with actual organisms and chemicals where required.
Practical Experience of Adam
Here we discuss some of the more important issues that arose with the use of Adam.
The design of plate trays or drawers, varies widely between different items of equipment. They are often deep relative to the height of a microplate and are a “tight fit,” requiring plates to be grasped close to their upper rim, potentially reducing grasp reliability. Ideally a plate handling robot should be presented with an area larger than the microplate, onto which plates may be placed with wide tolerance; when the plate is taken into the instrument a locating mechanism should press the plate into one corner of the tray to provide precise positioning, which provides a precise location from which the plate is subsequently picked. Increasingly, manufacturers are incorporating such mechanisms but in our system only the ELx405 washer from Bio-Tek has such an arrangement and this provides very high pick–place reliability.
Originally, movement of microtiter plates from our Liconic incubators to the Molecular Devices readers was inadequately reliable. There are three such moves: one involving a Twister II arm and the subsystem 2 incubator and reader, the other two involving the IX Scara arm and the two subsystem 3 incubators and reader. Even with the excellent repeatability of the IX Scara robot arm (±0.01 mm) plates would often fail to locate correctly in the reader drawer, leading to jamming. Plates are a close fit in this drawer and although it has beveled guides, the lack of compliance in the Scara renders these ineffective. Consequently, plates have to be placed with sufficient accuracy to seat directly into the drawer. Investigation revealed that the incubator pick zones were slightly longer than the recess in the reader drawer, leading to variation in the grasped position which, in turn, led to inaccurate placement into the reader. End stops on the incubator unload trays were machined so that the length of the pick zone became equal to that of the recess in the reader drawer, leading to much higher reliability.
Many plate handling robots can pick and place reliably only where plate locations are horizontal, but few items of automation equipment have leveling screws. Leveling errors of a millimeter or two can be cumulative across a pick and place move and can reduce grasp reliability or “jolt” the plate and disturb its contents.
Conclusions
Few laboratory automation systems are as flexible or functionally complex as Adam. This level of diversity and flexibility is challenging from the point of view of achieving the essential high level of reliability. Nevertheless, Adam achieved its design goal of being the first machine to discover novel scientific knowledge. 2
Acknowledgments
The authors thank the reviewers for their helpful suggestions and comments. This work was funded by the U.K. Biotechnology and Biological Sciences Research Council (grant BBD00425X1), by a Society for Biomolecular Screening 2 award from the Higher Education Funding Council for Wales, and by fellowships from the Royal Academy of Engineering/Engineering and Physical Sciences Research Council, and by the Royal Commission for the Great Exhibition of 1851 for Dr. Amanda Clare.
Competing Interests Statement: The authors certify that they have no relevant financial interests in this manuscript.
