Abstract
The HID EVOlution—Extraction System (Tecan Group Ltd., Mannedorf, Switzerland) was developed to automate DNA extraction from biological samples using the PrepFiler Automated Forensic DNA Extraction Kit (Applied Biosystems, Foster City, CA). The system consists of a Tecan Freedom EVO 150 robot (Tecan Group Ltd., Mannedorf, Switzerland), a graphical user interface designed for use with Freedom EVOware software v 2.1 SPI (Tecan Group Ltd., Mannedorf, Switerland) as well as instrument hardware and plastic to support the PrepFiler reagents and protocol. The DNA quality and quantity obtained were comparable to that observed with the corresponding manual extraction protocol. Purified DNA was free of inhibitors and ready for downstream applications, such as real-time quantitative PCR and PCR for short tandem repeat (STR) analysis. The DNA quantity and quality obtained were consistent as demonstrated by the quantification and STR results. Our studies indicate that the HID EVOlution—Extraction System can easily be adopted in forensic laboratories to alleviate some of the bottlenecks of sample preparation in forensic laboratories.
Keywords
Introduction
The ability to detect many highly variable genetic loci simultaneously to generate an individual-specific DNA “fingerprint” for use in human genetic analysis reported by Jeffreys et al. 1 opened a new era in human identification (HID). Polymorphic short tandem repeat (STR) loci are now routinely used for HID applications, such as forensic DNA analysis, DNA databasing, paternity testing, mass disaster, and missing persons identification. 2,3 Achieving a high degree of discrimination among individuals in the population by using as many as 15 autosomal STR loci and acceptability of the results in the legal system brought forensic DNA analysis methodology to the forefront of current methodologies. 4
The STR profiling in a forensic laboratory is a multistep process that involves the extraction of DNA from a biological sample, the quantification of extracted DNA, the multiplex amplification of STR loci, the separation of the amplified products, and the analysis of data. 3 It is now possible to obtain an interpretable STR profile from as little as 100 pg of human DNA. This achievement is a result of advancements in chemistries for the isolation of DNA, realtime PCR methods for quantification, multiplex PCR systems for amplification of STR loci, separation technologies capable of resolution of fragments with 1 base difference in size, and software for data analysis.
The extraction of DNA, which is invariably the first step in STR profiling, is often challenging for a variety of reasons. Biological evidence samples often contain different body fluids or tissue, are found on a variety of substrates, can be contaminated with substances that may inhibit downstream PCR, often are present in limited quantities, and may have been exposed to environmental conditions that may degrade DNA or inhibit PCR. A variety of methods based on different principles, such as ion exchange, adsorption on silica surface, and phase separation, are available for extraction of DNA. 3,5 –9 The PrepFiler Forensic DNA Extraction Kit (Applied Biosystems, Foster City, CA) was designed specifically to support both manual and automated extraction of DNA from forensic samples. 10 The PrepFiler kit is based on a unique chemistry wherein DNA from the lysate bind to the magnetic particles by formation of a complex that remains stable during the steps for removal of inhibitors and is ultimately dissociated in the elution buffer conditions. The PrepFiler Kit enables the isolation of DNA from a variety of biological samples that contain small quantities of biological material in such a way that substances that interfere with PCR are removed and the DNA extract is of sufficiently high concentration that the volume of extract used for downstream analysis is minimal. Another challenge that forensic laboratories face is the backlog of samples to be processed. The high power of discrimination and ability to obtain STR profiles from forensic sample types increased the flow of evidence samples to laboratories for profiling. One crime scene generates multiple samples and the urgency of resolving a case requires a high-throughput sample processing workflow. Several estimates have been made regarding the overwhelming increase in the sample backlog in forensic laboratories in the United States. 11 –15 Automation, therefore, is an important tool in reducing turnaround time and the ever-increasing backlogs in the forensic laboratory, while maintaining precise and accurate sample processing. As with manual protocols, automated DNA extraction methods must generate high-quality genomic DNA with consistent yields. Some laboratories have developed automated DNA extraction protocols for internal use. 16,17 As the implementation of automated extraction methods often requires considerable resources, the availability of a validated, ready-to-adopt automated solution is desired by many small- or medium-throughput forensic laboratories.
The HID EVOlution—real-time quantitative PCR (qPCR)/STR Setup System (Tecan Group Ltd., Mannedorf, Switzerland), which automates real-time qPCR setup using the Quantifiler kits (Applied Biosystems, Foster City, CA), normalization of the DNA samples, and STR PCR setup for using the AmpFℓSTR PCR Amplification kits (Applied Biosystems, Foster City, Ca) was reported earlier. 18 We have developed the HID EVOlution—Extraction system (Tecan Group Ltd., Mannedorf, Switzerland) for automated extraction of DNA from biological samples using the PrepFiler Automated Forensic DNA Extraction Kit. The HID EVOlution—Extraction system is composed of a graphical user interface and a Tecan Freedom EVO 150 robot (Tecan Group Ltd., Mannedorf, Switzerland) developed, optimized, and validated specifically for the PrepFiler Automated Kit. The Scientific Working Group on DNA Analysis Methods (SWGDAM) has provided guidelines for validation of a method used in DNA analysis. 19 Validation studies to evaluate the performance of the PrepFiler Kit according to SWGDAM guidelines are described elsewhere. 10 This article describes the validation studies conducted to evaluate the performance of an automated DNA extraction method using the PrepFiler Automated Kit on the HID EVOlution—Extraction System.
Extraction of DNA
Materials
Biological samples of blood, saliva, and semen were purchased from Serological Research Institute (Richmond, CA). A Signature Benchtop Shaking Incubator was obtained from VWR Scientific Products (Batavia, IL). Indigo, hematin, and humic acid were purchased from Sigma-Aldrich (St. Louis, MO). The PrepFiler Automated Forensic DNA Extraction Kit, MicroAmp Clear Adhesive Film, MicroAmp Multi-Removal Tool, Quantifiler Human DNA Quantification Kit, Identifiler Kit, 7500 Real-time PCR System, 3100 Genetic Analyzer, and associated software were from Applied Biosystems (Foster City, CA). All other chemicals used in this study were of analytical grade. The stains of the body fluid on different substrates were prepared in our laboratory as described earlier. 10 The Freedom EVO 150 instrument and accompanying software and hardware are from Tecan Group Ltd. (Männedorf, Switzerland).
Lysis of Biological Material
DNA from anonymous donor samples (liquid blood, dried blood, saliva, or semen stains on cotton cloth, denim, and/or cotton swabs) was extracted using the PrepFiler Automated Kit. Cell lysis was performed either in 1.5-mL tubes or in the 96-well filter/spin plate provided in the kit. 20 Lysis of samples in 1.5-mL tubes was performed as described earlier. 10,21 The lysate obtained using the 1.5-mL tubes may contain, at times, fragments of substrate. Such insoluble visible particles were removed by centrifugation and decantation to collect clear lysate, which was then used for isolation of DNA using the robot. For more streamlined lysis and higher throughput processing, a 96-well lysis assembly composed of a filter plate and spin plate was designed (Fig. 1). Biological samples were placed in a filter plate mounted to a spin plate. Performing the lysis step in a 96-well filter/spin plate enabled the different sample types to be lysed simultaneously and the substrates removed from all samples in a single centrifugation step. To process samples in a 96-well format, the biological samples were placed in a 96-well filter plate mounted to a 96-well spin plate. Lysis in the spin plate was performed by adding 300 μL of the lysis buffer and 3 μL of 1.0 M Dithiothreitol to each well containing a biological sample. The filter plate was sealed with a MicroAmp Clear Adhesive Film. The filter/spin plate containing the lysis mixture was incubated at 70 °C for 60 min (90 min if any samples contained semen) with shaking at about 150 rpm in the Signature Benchtop Shaking Incubator. At the end of incubation, the lysate was collected in the spin plate by centrifugation at 650 × g for 10 min using a laboratory centrifuge. The filter plate was removed using the MicroAmp Multi-Removal Tool and the spin plate alone or 1.5-mL tubes containing the lysate were placed on the Freedom EVO 150 instrument for isolation of DNA.
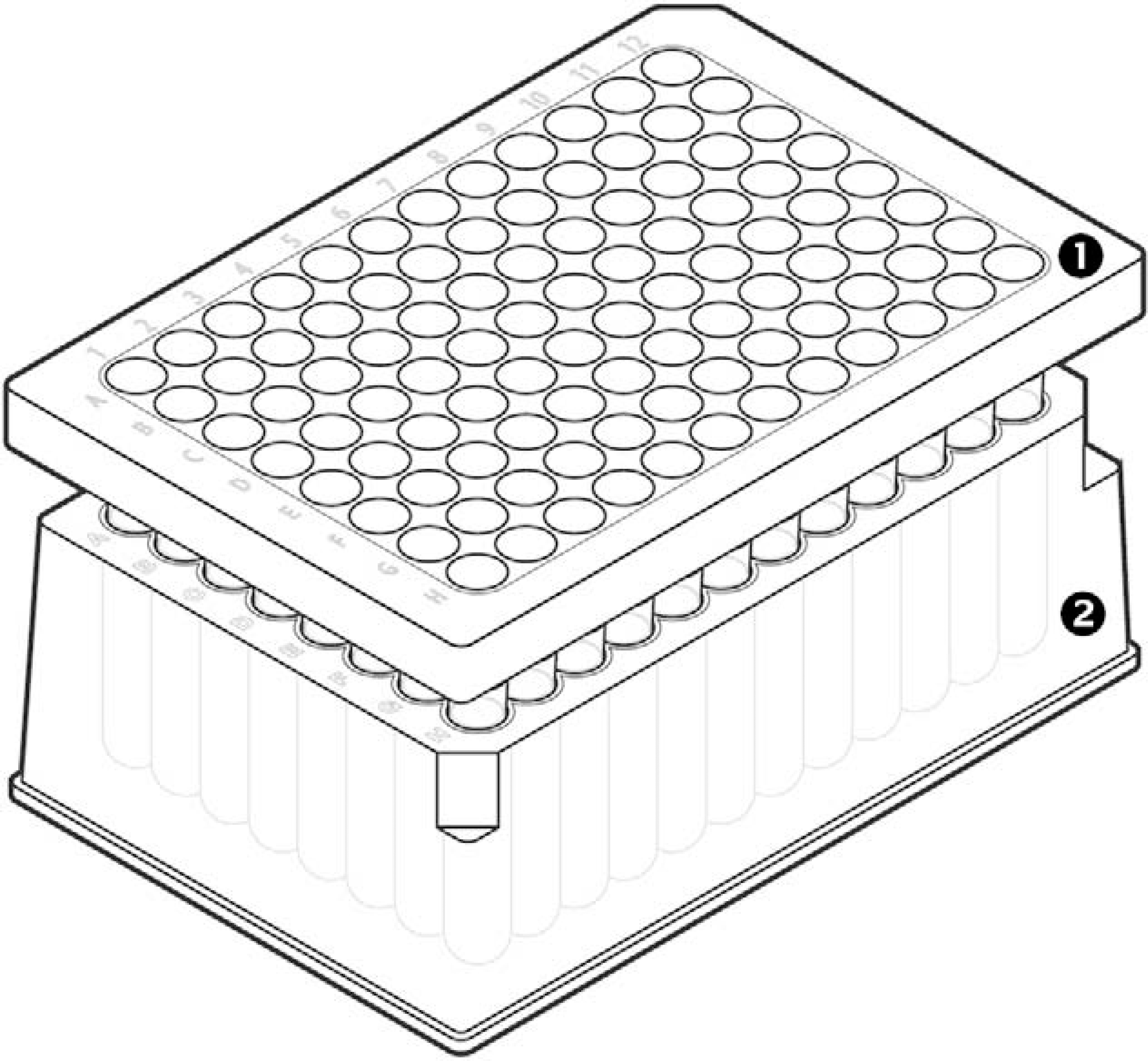
A 96-well lysis assembly filter plate and spin plate. 1. Filter plate. 2. Spin plate. Outlet of each well of the filter plate is designed as a column that forms tight seal with the respective well of spin plate. Further, the outlet extends as 7.6 mm neck for leak proof lysate delivery.
Isolation of DNA Using Freedom EVO 150
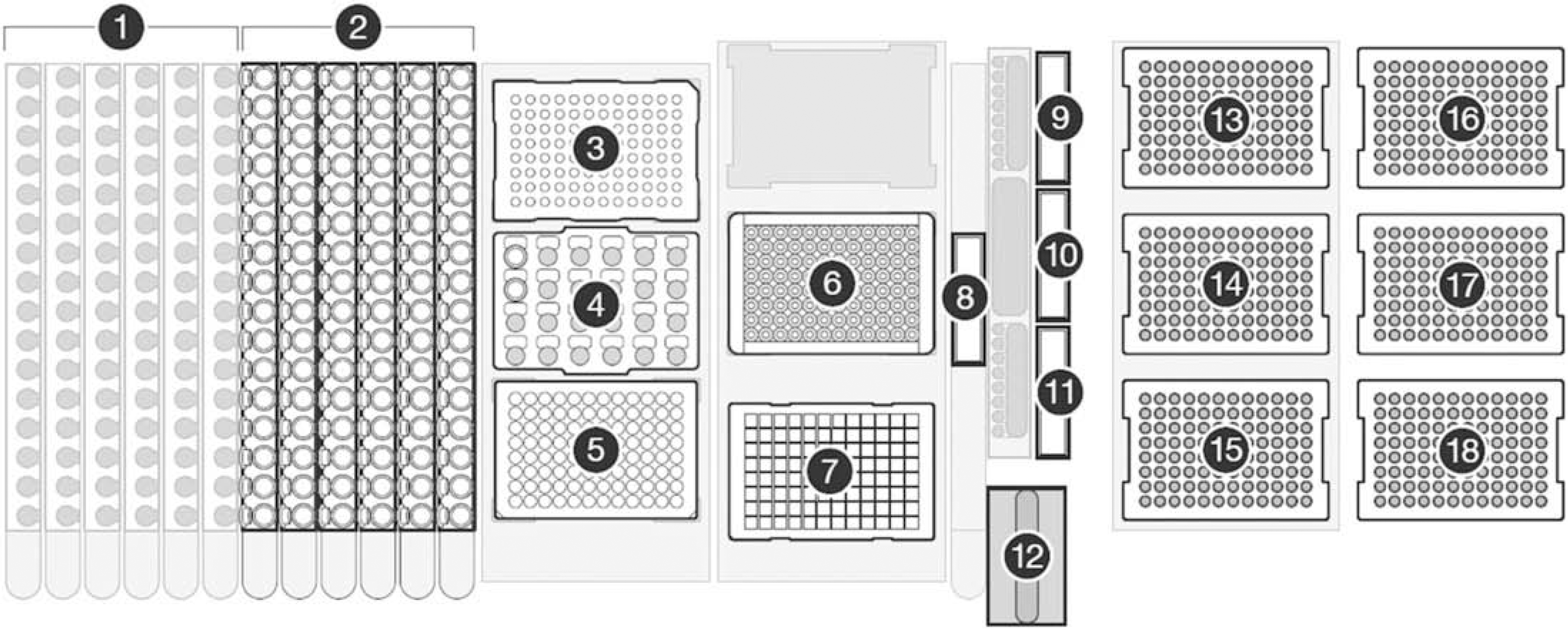
Deck layout for the operations of HID EVOlution—Extraction System. 1. Tube racks for DNA elution tubes (if eluate is collected in tubes). 2. Tube racks for sample lysate tubes (if lysis is performed in tubes). 3. 96-Well elution plate (if eluate collected in a plate). 4. Block for PrepFiler magnetic particles. 5. PrepFiler Spin Plate (if sample lysis is performed in a plate). 6. Magnetic-Ring Stand. 7. PrepFiler Processing Plate on Te-Shake. 8. Lysate waste trough. 9. Isopropanol trough. 10. Wash buffer trough. 11. Elution buffer trough. 12. Disposable tip (DiTi) waste unit. 13–15. 200-μL disposable pipette tips. 16–18. 1000-μL DiTis.
Regardless of source or elution vessels, the samples are extracted in the processing plate that is readily usable with the 96-well Magnetic-Ring Stand. The magnet used was a 96-well, o-ring magnet, which forms a donut-shaped ring of magnetic particles to optimize binding and minimize the potential loss of DNA by disturbing the magnetic particles—DNA complex. The o-ring magnet exhibited better performance compared with a pin magnet for handling of magnetic particles. The Te-Shake orbital shaker achieves vigorous shaking (without splashing) and heating required for the binding, washing, and elution steps (Fig. 3). A custom adapter for the Te-Shake was designed to hold the processing plate onto the Te-Shake and to facilitate heat transfer for effective elution of DNA from the magnetic particles (Fig. 3). Standard Tecan carriers were used to hold DiTis, troughs, 1.5-mL microfuge tubes, and 96-well PCR plates on the worktable. The spin plate, processing plate, and elution plate provided with the PrepFiler Automated kit were used.
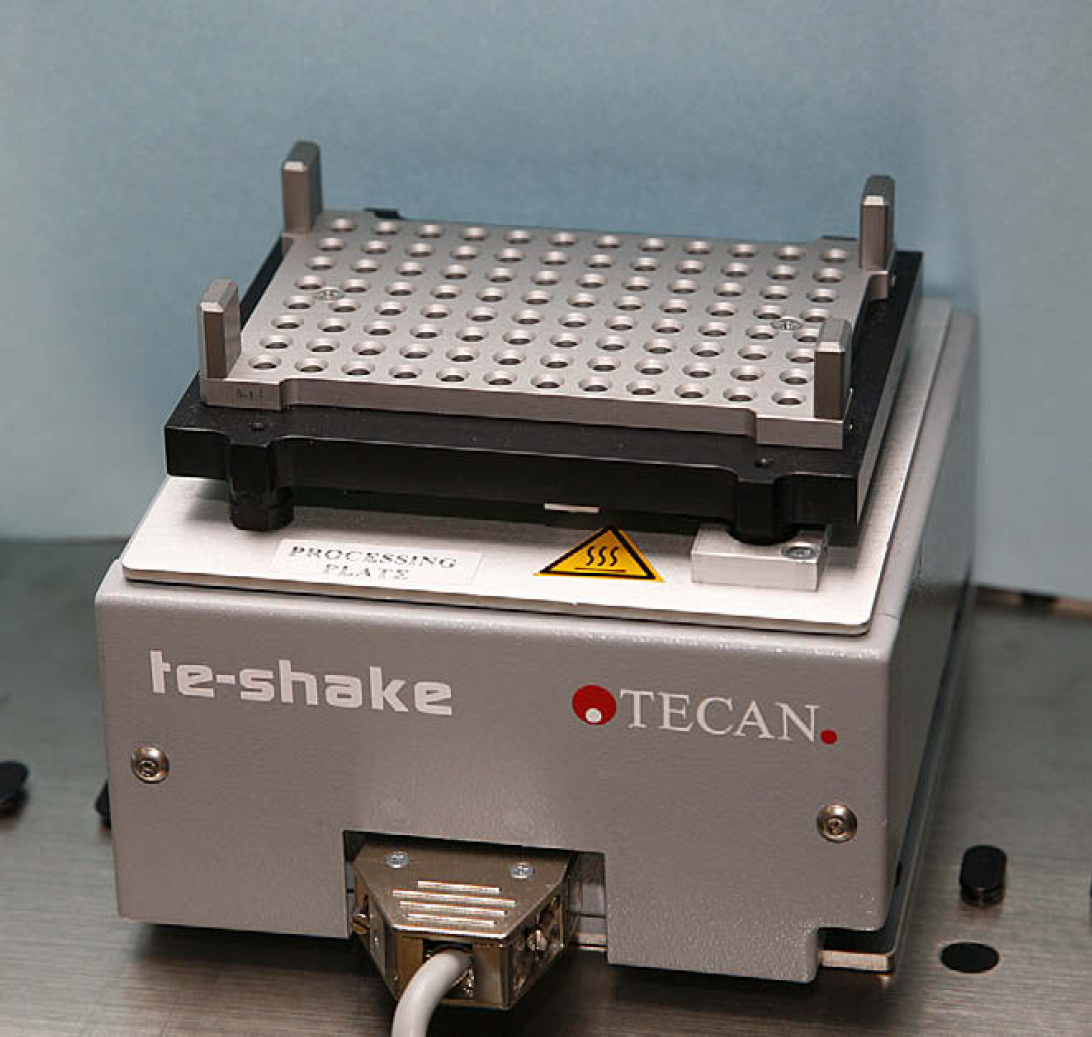

T-Shake with heating block and adapter (close-up).
The detailed workflow of the automated scripts, regardless of lysate or elution plasticware used, is as follows. A uniform suspension (15 μL) of magnetic particles is added to the lysate (280–300 μL), mixed and then the mixture is transferred to the processing plate, where 180 μL of binding solution (isopropanol) has already been added by the robot to each well. The plate is shaken on the Te-Shake (1000 rpm for 1 min, 700 rpm for 5 min), after which the processing plate is placed on the magnetic stand for 6 min to separate the magnetic particles. The resulting supernatant is transferred to the waste (a dedicated waste trough to minimize the volume of hazardous liquid waste), and the magnetic particles are resuspended by mixing with 300 μL of wash buffer. The wash is completed by placing the processing plate on the magnetic stand for 1 min to separate the magnetic particles, transferring the supernatant to waste, and repeating the wash step two more times. For elution, the processing plate is incubated at room temperature for 5 min with shaking (1000 rpm) to allow evaporation of the residual organic solvents, and then the magnetic particles are resuspended by mixing with 50 μL of elution buffer, heating the plate on the Te-Shake at 65 °C for 6 min with shaking (1000 rpm), placing the processing plate on the magnetic stand for 7 min to separate the magnetic particles, and transferring the eluate to either 1.5 mL tubes or a 96-well elution plate. The DNA extract can be stored at 4 and −20 °C for short-and long-term storages, respectively.
Custom liquid classes for magnetic particle suspension, binding solution, wash solution, and elution buffer were developed for accurate delivery of the desired volume of each reagent. Calibration curves were created for all liquid classes. The results presented in Table 1 demonstrate good precision of liquid handling by each pipetting channel.
Accuracy and precision of liquid handling for the reagents
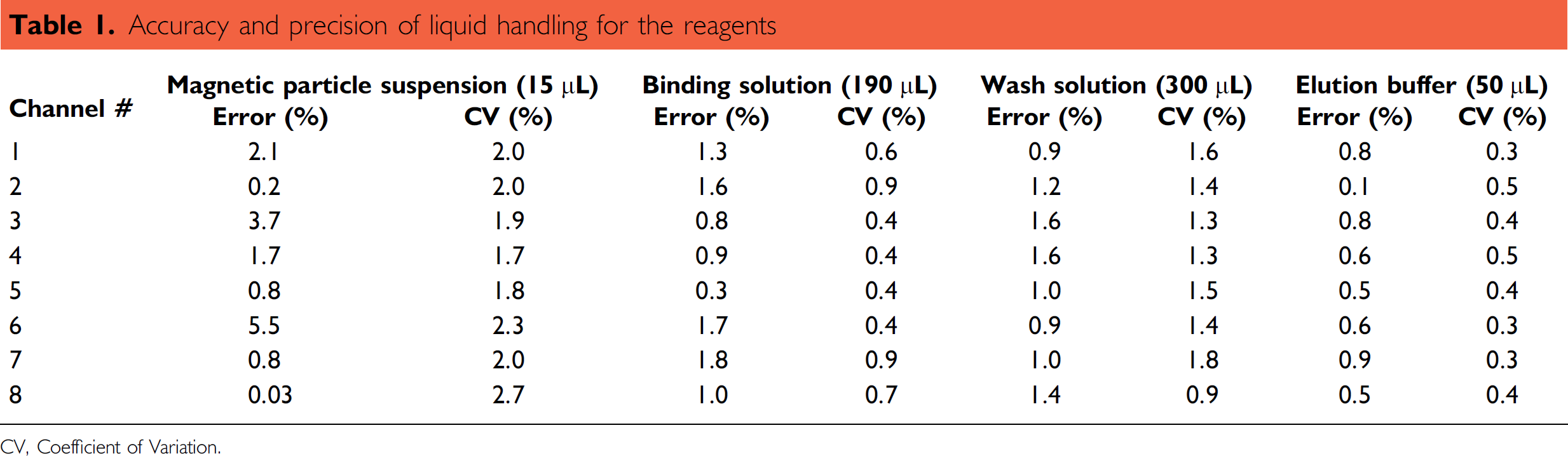
CV. Coefficient of Variation.
Total time required for the extraction of DNA from lysate using the HID EVOlution—Extraction System for 96 samples was 2.5 h. In one 8-h working shift, 288 samples can be processed without manual intervention. Manual operations take 1 h for a batch of 12 samples and 96 samples can be processed in one day. Thus, the automated workflow provides a threefold increase in sample output and allows the operator to perform other tasks in the laboratory.
Quantification of DNA and STR Analysis
To evaluate the performance of the automated extraction on the HID EVOlution–Extraction System, DNA extracts were processed for human DNA quantification using the Quantifiler Human DNA Quantification Kit and STR profiling using Identifiler and MiniFiler kits (Applied Biosystems, Foster City, CA) on the Freedom EVO 150 instrument using the HID EVOlution–qPCR/STR Setup System according to the protocol described previously. 18
Precision Study
The precision study was performed to test the precision of DNA recovery within a sample set. Eight replicates of 12 different samples were processed for extraction using the PrepFiler Automated kit. The sample set included 1, 2, 5, 10, 30, and 40 μL of liquid blood (LB) from a human donor, 5-μL human blood stain or noncolored cotton (BSC), 5-μL human blood stain on denim (BSD), stain of 5-μL human blood spiked with 1-μL inhibitor mix on noncolored cotton (BSCI), 50-μL human saliva on cotton swab (SALSw), 1-μL human semen stain on noncolored cotton (SSC), and extraction blank (XB). The samples were lysed in a 96-well filter/spin plate and eluted in a 96-well elution plate. The purified DNA extracts were processed for quantification of human DNA. The yields of DNA and standard deviation values are summarized in Table 2.
Total yield of DNA and standard deviation values for the sample process for precision study
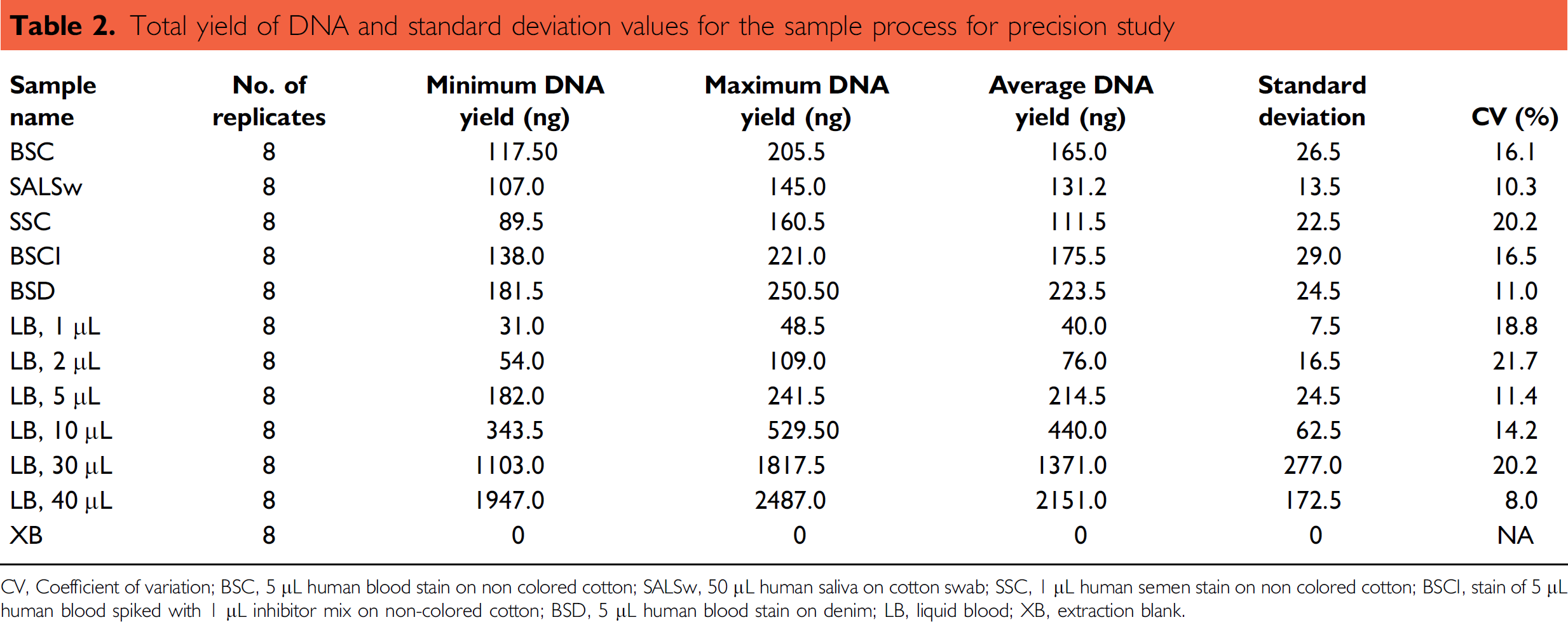
CV, Coefficient of variation; BSC, 5 μL human blood stain on non colored cotton; SALSw, 50 μL human saliva on cotton swab; SSC, 1 μL human semen stain on non colored cotton; BSCI, stain of 5 μL human blood spiked with 1 μL inhibitor mix on non-colored cotton; BSD, 5 μL human blood stain on denim; LB, liquid blood; XB, extraction blank.
The CT value of the internal PCR control (IPC) was monitored to evaluate the presence of PCR inhibitors (e.g., heme, humic acid, dyes) in the DNA extract. In general, a shift in the IPC CT value from a sample compared with the nontemplate control (NTC) of > 1.0 is considered to be an indication of the presence of PCR inhibitors. The IPC CT values for all sample types and replicates ranged between 27.3 and 27.9 and were within ±1 of the IPC CT value for the NTC (27.3). The results demonstrated that the PrepFiler Automated kit removed detectable levels of PCR inhibitors.
Similar results for the DNA yields and IPC CT values were obtained when the extraction was performed using 1.5-mL tubes as lysate and eluate containers (data not shown).
Correlation Studies
DNA yields obtained from the precision study described above were compared with the DNA yields obtained from an identical set of samples manually extracted using the PrepFiler Manual Kit. Manual extraction was performed as described previously. 10,21 The results are presented in Figures 4 and 5. Both DNA yield and IPC CT values obtained using the automated procedure on the HID EVOlution—Extraction System were similar to those obtained using the manual procedure and demonstrate that the performance of the automated method was comparable to the manual method.
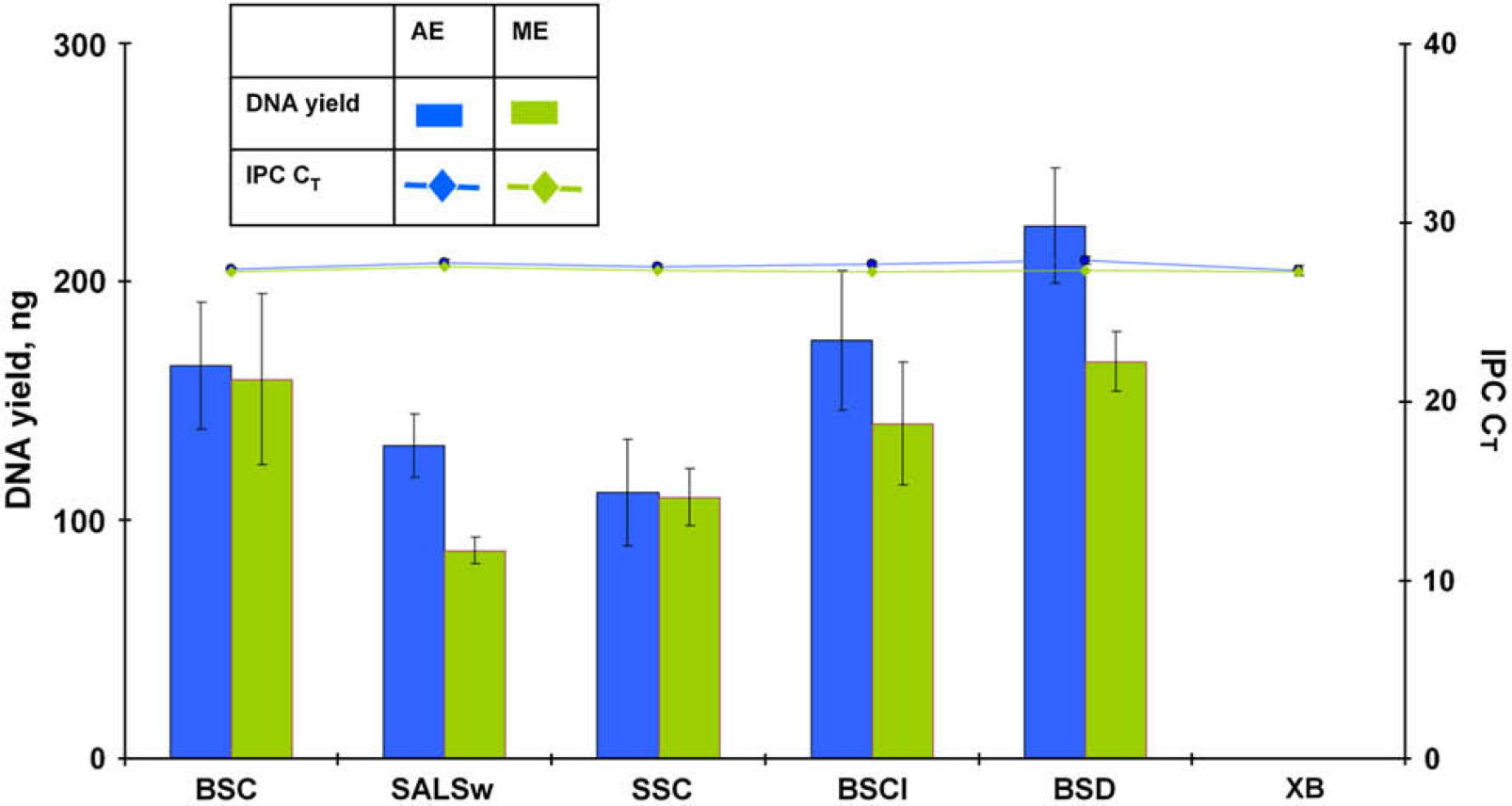
Total yield of DNA and internal PCR control (IPC) CT values for forensic type samples from automated (AE) and manual (ME) extraction experiments. BSC, 5-μL human blood stain on noncolored cotton; SALSw, 50-μL human saliva on cotton swab; SSC, 1-μL human semen stain on noncolored cotton; BSCI, stain of 5-μL human blood spiked with 1-μL inhibitor mix on noncolored cotton; BSD, 5-μL human blood stain on denim; and XB, extraction blank.
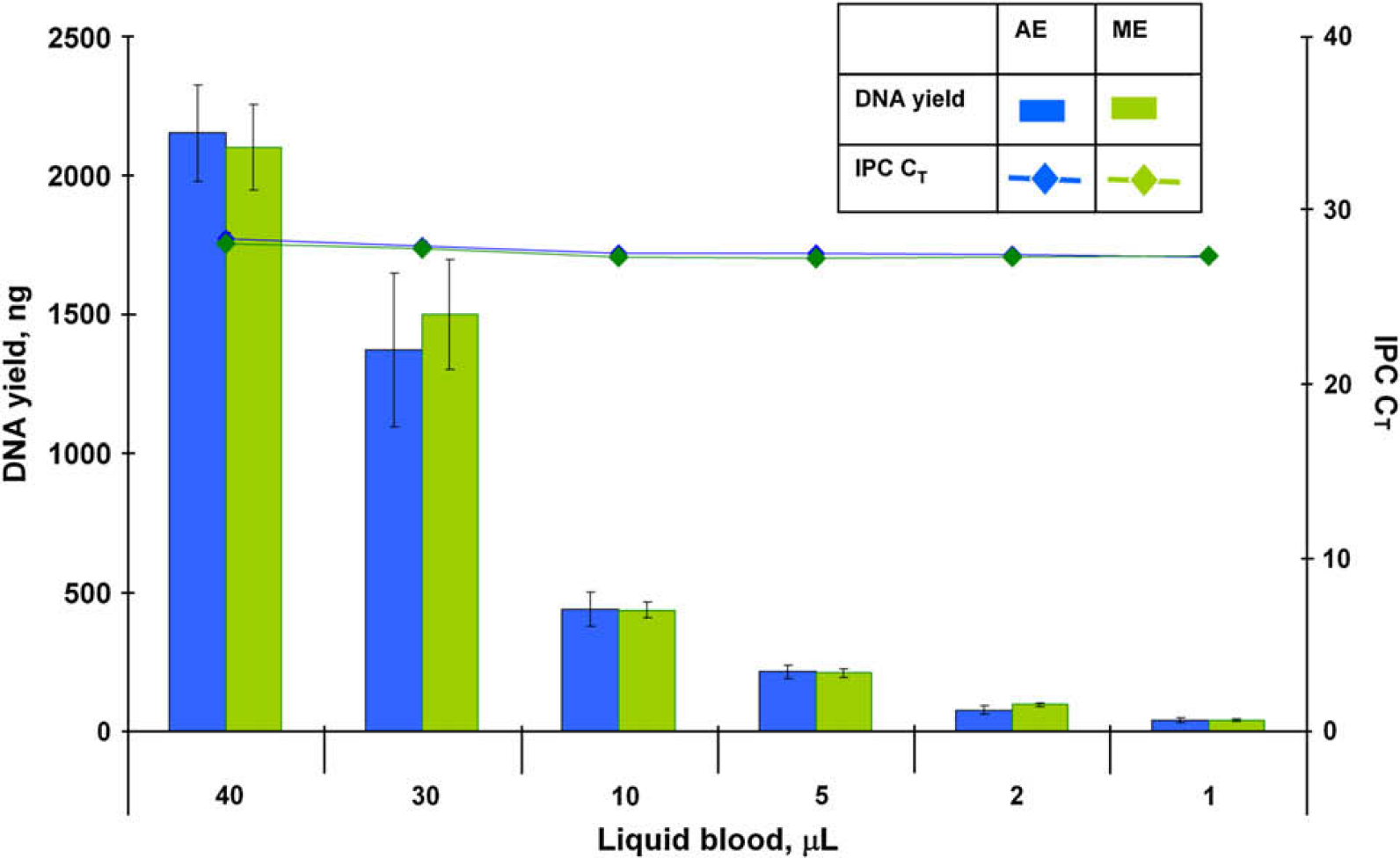
Total yield of DNA and internal PCR control (IPC) CT values for liquid blood sample from automated (AE) and manual (ME) extraction experiments.
Reproducibility Study
Replicate extractions of the same biological samples were performed in separate experiments to examine the ability to reproducibly obtain a maximum amount of high-quality DNA. The set of 12 samples described in the precision study were extracted in 8 replicates using the PrepFiler Automated kit. The extraction experiment was performed on 3 consecutive days. Lysis was performed in a 96-well filter/spin plate, and eluate was collected in a 96-well elution plate. The purified DNA extracts were evaluated based on DNA quantity and quality using the Quantifiler Human Kit. The results summarized in Figures 6 and 7 demonstrate that DNA yield and quality obtained for each sample type were reproducible. The IPC CT values for all sample types and replicates from three extraction runs ranged between 27.2 and 27.9 and were within ±1 of the IPC CT value for the NTC (27.3) indicating effective removal of PCR inhibitors.
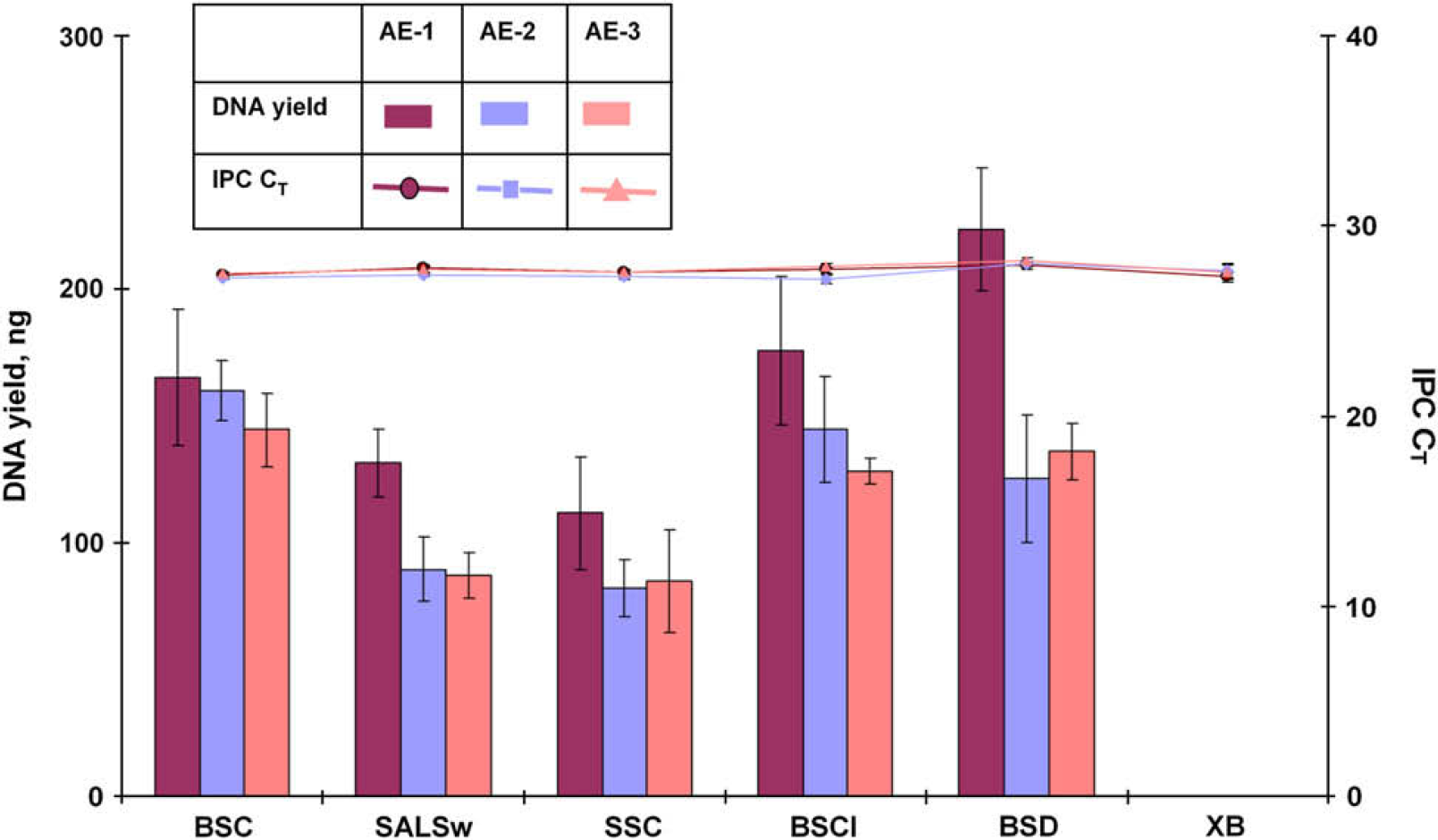
Total yield of DNA and internal PCR control (IPC) CT values for forensic type samples from three automated extraction (AE) experiments. BSC, 5-μL human blood stain on noncolored cotton; SALSw, 50-μL human saliva on cotton swab; SSC, 1-μL human semen stain on noncolored cotton; BSCI, stain of 5-μL human blood spiked with 1-μL inhibitor mix on noncolored cotton; BSD, 5-μL human blood stain on denim; and XB, extraction blank.
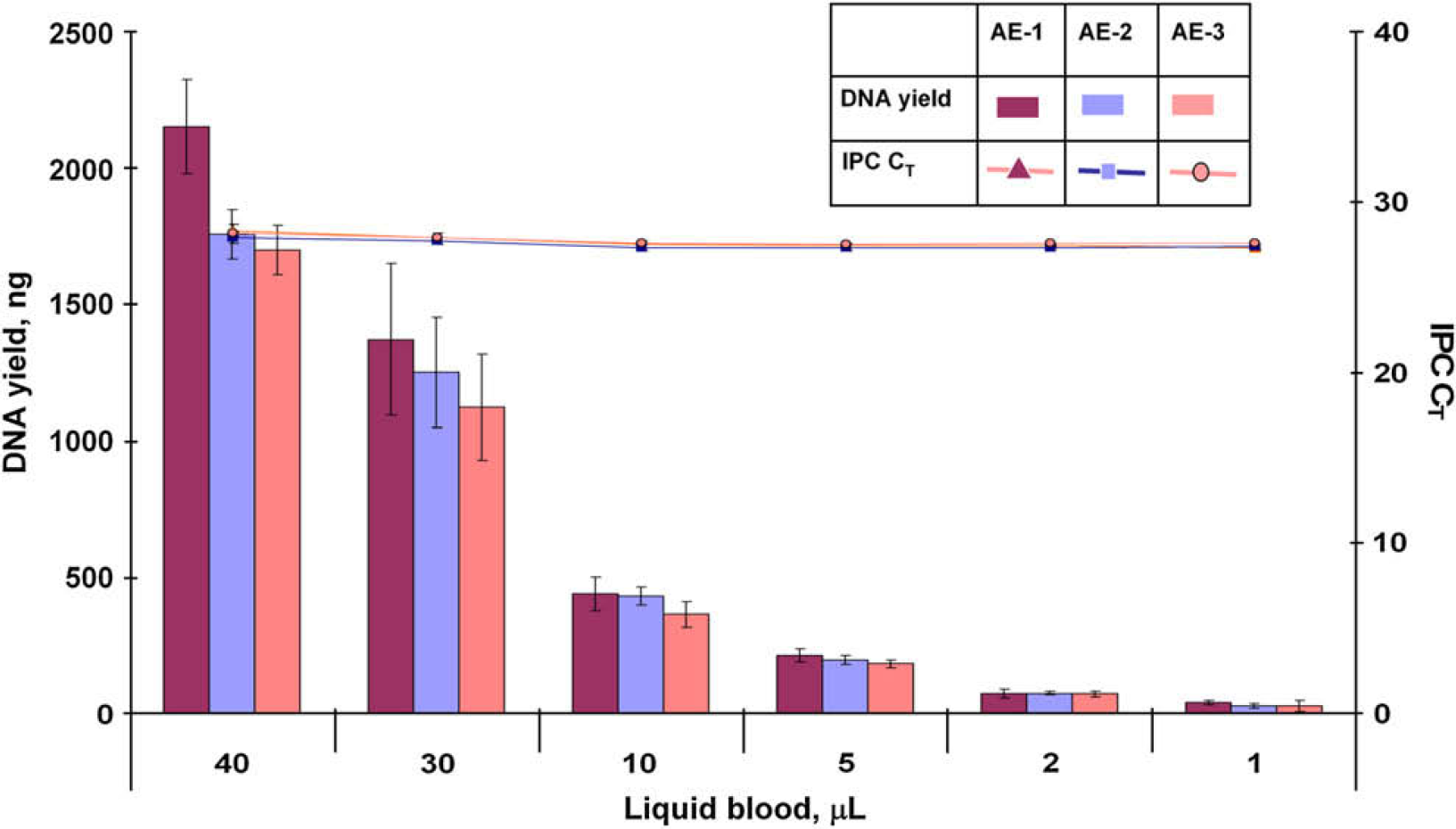
Total yield of DNA and internal PCR control (IPC) CT values for liquid blood sample from three automated extraction (AE) experiments.
Contamination Studies
The ability to minimize cross-contamination in a forensic laboratory is of utmost importance. Factors such as multiple transfer of samples and reagents, excessive foaming due to vigorous mixing or the chemical nature of the reagents, aerosol formation, and movement of the robotic arm can contribute to cross-contamination. Therefore, it is essential to optimize the liquid-handling protocols to ensure that the robotic or laboratory operations minimize the potential for contamination. To evaluate the potential for cross-contamination, 10 μL of blood from six different donors and extraction blanks were arranged in a checkerboard format in the 96-well spin plate in such a way that samples from the same donor were not in adjacent wells. Samples were dispensed manually into a spin plate one well at a time, while all remaining wells were covered and the movement of the pipette was controlled to minimize potential for cross-contamination when preparing the initial sample plate. The samples were processed for lysis and isolation of DNA using the Plate-to-Plate script. This experiment was repeated one more time. All samples and extraction blanks were analyzed with Quantifiler Human, Identifiler, and MiniFiler kits.
Of the 96 wells of extraction blanks on the two plates, only one well exhibited a CT value less than 40 (39.43). A CT value of ≥40 indicates that no human DNA was detected. Amplification of the extract from this well and all other extraction blanks did not generate STR profiles when amplified using the Identifiler and MiniFiler kits. Thus, the observance of the CT value of 39.43 is attributed to background amplification and not necessarily due to cross-contamination. The wells expected to contain DNA provided conclusive complete, well-balanced, and concordant Identifiler and MiniFiler STR profiles. The results demonstrate that HID EVOlution—Extraction System provides clean liquid handling.
Verification of Scripts
As mentioned earlier, four scripts were developed and optimized to accommodate different combinations of plastics commonly used in the forensic laboratory to store samples:
Lysis in 96-well spin plate and elution in 96-well elution plate (Plate to Plate),
Lysis in 96-well spin plate and elution in 1.5-mL tubes (Plate to Tube),
Lysis and elution in 1.5-mL tubes (Tube to Tube), and
Lysis in 1.5-mL tubes and elution in 96-well elution plate (Tube to Plate).
The core liquid-handling parameters are identical for all four scripts including DNA binding to the magnetic particles, DNA—magnetic particle complex washing, and DNA elution. The Plate-to-Plate script was the primary script used to evaluate the performance of the automated protocol as described above. The performance of the remaining scripts was confirmed by the verification studies described here. DNA was extracted from six human donors (eight donors when the lysis was performed in 1.5-mL tubes). Blood samples of 10 μL were arranged in a checkerboard pattern in combination with extraction blanks in such a way that samples from same donor were not in adjacent sample wells or tubes. The samples were evaluated based on DNA quantity and quality using the Quantifiler Human Kit. The DNA quantity and quality from the six donors used for the extraction using the four scripts were similar (Fig. 8). All but 2 extraction blanks exhibited CT values >40, and none exhibited STR profiles when amplified with the Identifiler kit. The observance of CT values <40 for these samples was attributed to background amplification and not necessarily to cross-contamination.
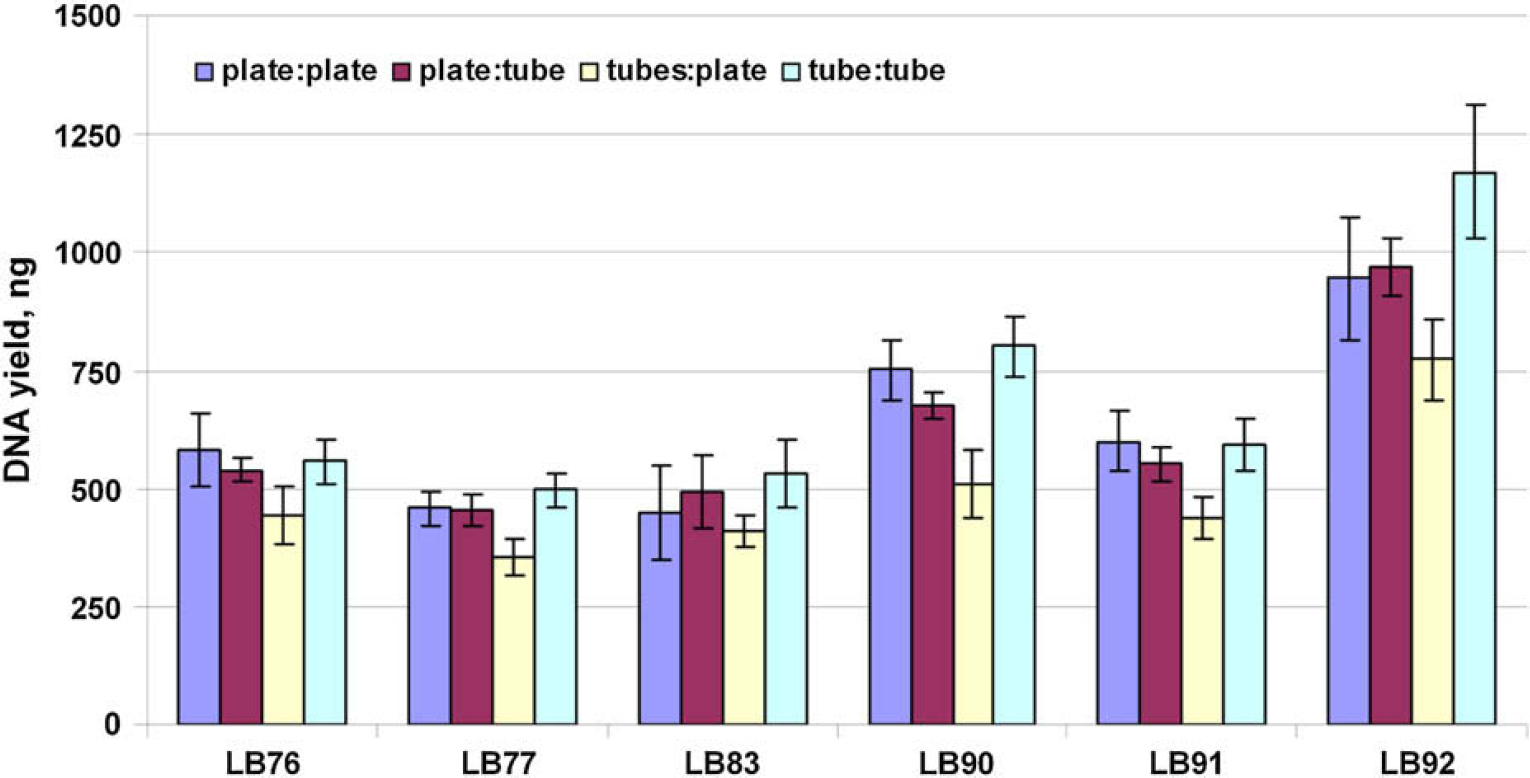
Total DNA yield for six donor samples for the four scripts. LB, liquid blood.
Str Study
The ultimate goal of the DNA extraction step in forensic DNA analysis is to obtain high-quality STR profiles. To further evaluate the performance of the automated PrepFiler kit procedure using the HID EVOlution—Extraction System, DNA extracted from 5-μL human blood stain on noncolored cotton (BSC) and denim (BSD), a stain of 5-μL human blood spiked with 1-μL inhibitor mix on noncolored cotton (BSCI), 50-μL human saliva on cotton swab (SALSw), and 1-μL human semen stain on noncolored cotton (SSC) were profiled using the Identifiler kit. Human DNA (1 ng), as determined by the Quantifiler Human Kit, was used as the template DNA. The STR profiles obtained were complete, well-balanced, and concordant with the STR profiles obtained from identical samples extracted using the PrepFiler Manual kit (Fig. 9).
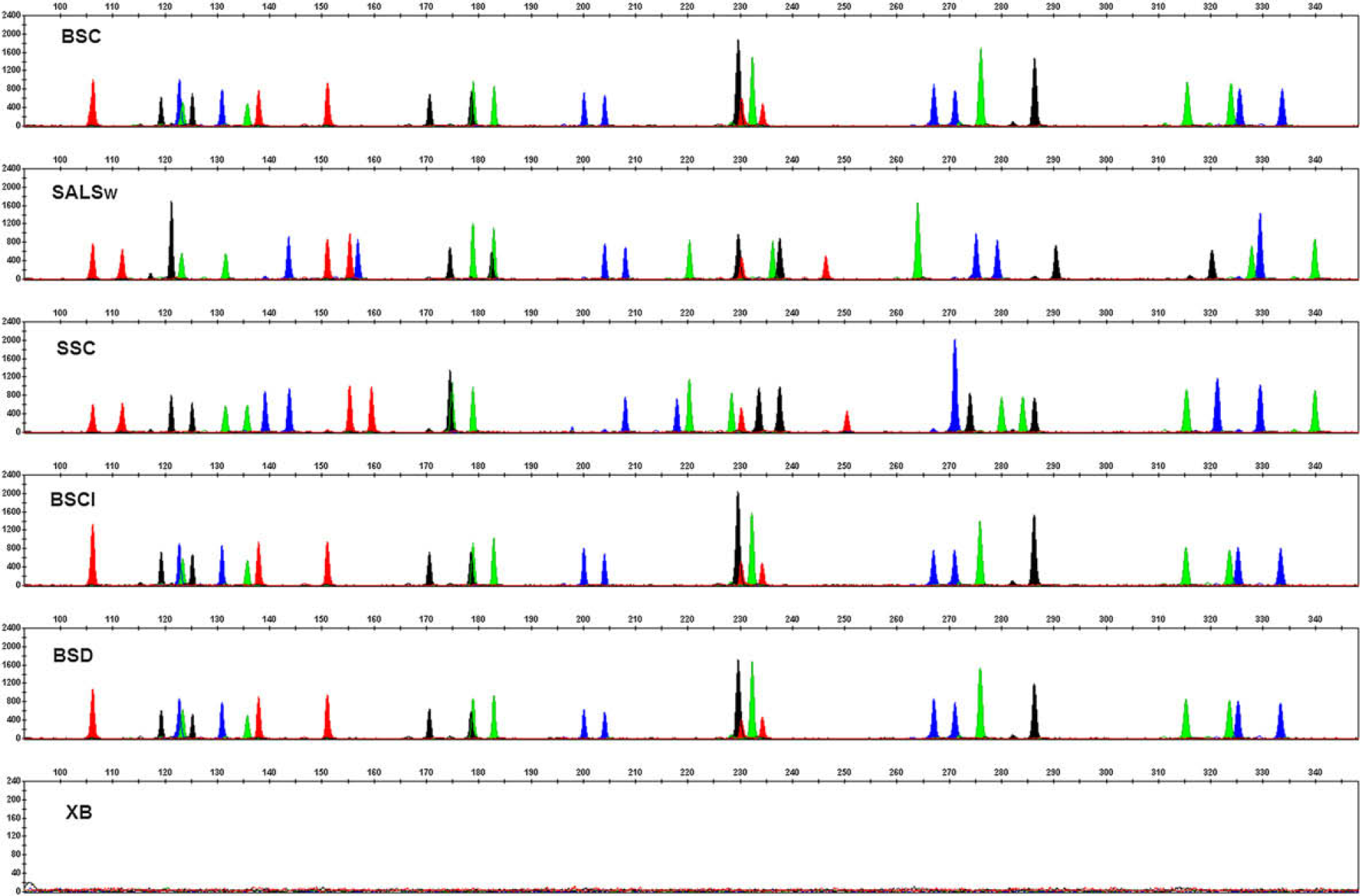
Short tandem repeat profiles of forensic type samples generated using the Identifiler kit. BSC, 5-μL human blood stain on non-colored cotton; SALSw, 50-μL human saliva on cotton swab; SSC, 1-μL human semen stain on noncolored cotton; BSCI, stain of 5-μL human blood spiked with 1-μL inhibitor mix on noncolored cotton; BSD, 5-μL human blood stain on denim; and XB, extraction blank.
Conclusions
The PrepFiler Forensic DNA Automated Extraction Kit was developed for the high-throughput isolation of genomic DNA from a variety of biological samples. The utility and performance of the kit in forensic DNA analysis was previously demonstrated to maximize the amount and purity of DNA obtained from a range of sample types, inputs, and substrates as well as from samples exposed to inhibitors. 10 The validation studies discussed here confirm that the use of the automated DNA extraction procedure using the HID EVOlution Extraction System in conjunction with the PrepFiler Forensic DNA Automated Extraction Kit is capable of clean liquid handling and produces reproducible and reliable results in terms of DNA quantity and quality for downstream applications, such as real-time qPCR and PCR for STR profiling.
Footnotes
Acknowledgments
The authors thank Heidi Kijenski and Lisa Calandro from Life Technologies and Tony Mamone from Tecan for useful discussions.
