Abstract
Hairs are commonly submitted as evidence to forensic laboratories, but standard nuclear DNA analysis is not always possible. Mitochondria (mt) provide another source of genetic material; however, manual isolation is laborious. In a proof-of-concept study, we assessed pressure cycling technology (PCT; an automated approach that subjects samples to varying cycles of high and low pressure) for extracting mtDNA from single, short hairs without roots. Using three microscopically similar donors, we determined the ideal PCT conditions and compared those yields to those obtained using the traditional manual micro-tissue grinder method. Higher yields were recovered from grinder extracts, but yields from PCT extracts exceeded the requirements for forensic analysis, with the DNA quality confirmed through sequencing. Automated extraction of mtDNA from hairs without roots using PCT could be useful for forensic laboratories processing numerous samples.
Keywords
Introduction
The National Institute of Justice (NIJ) defines a backlogged case as one that has not been completed within 30 days of receipt at the laboratory. 1 With the ever-increasing number of cases submitted for testing to both national and international forensic laboratories, it is imperative to implement automated, efficient, and accurate methods to streamline workflows. Hairs are commonly collected from crime scenes and submitted for forensic analysis. In many cases, nuclear DNA may not be available, and thus analysis of the mitochondrial DNA (mtDNA) may be the only possibility. A common practice for the isolation of mtDNA from hair requires manual mechanical disruption using micro-tissue grinders. As this method is laborious, developing an automated extraction approach that facilitates the recovery of sufficient copies of high-quality mtDNA would be of great benefit.
Pressure cycling technology, or PCT, uses cyclic hydrostatic pressure changes to lyse tissues and cells, resulting in the release of intracellular contents such as DNA, RNA, and proteins. The sample is placed in a single-use tube and is subjected to alternating cycles of high pressure (up to 35 KPSI) and ambient pressure in a Barocycler (Pressure BioSciences, South Easton, MA). The sample must be contained in a liquid, as liquid compression provides the force that helps disrupt membranes and the intracellular contents. The number of cycles, the minimum and maximum pressure, and the time at each pressure can be adjusted to improve yields of the desired material. The temperature of the reaction chamber can be controlled using an external water bath, so the samples are not subjected to high temperatures. This is advantageous when working with nucleic acids, since bases can hydrolyze at elevated temperatures, preventing subsequent hybridization, 2 a potentially complicating factor with subsequent PCR amplification.
PCT has been used to isolate DNA from pathogens that are very hard to lyse 3 ; a 16-fold improvement in DNA yield was obtained with PCT over the author’s standard mechanical grinding method. Gross et al. 4 reported using PCT for isolation of mitochondria from cells grown in suspension. The authors pointed out that gentle pressure cycling could be used to isolate mitochondria for functional studies and suggested intense pressure cycling might be capable of releasing constituents of the organelle, such as the DNA. PCT has also been used to recover unfragmented RNA from epithelial cells 5 and reduce the effect of inhibitors in PCR.6,7 Several groups have shown that PCT increased both the amount and number of extracted proteins from cells.8–11
The utility of PCT for extracting DNA from low-template forensic samples,12,13 including hair,14,15 bone fragments, 16 skin and blood stains, 15 sexual assault samples, 17 and fecal matter, 18 has also received some attention, due to the foreseeable advantages of an automated approach for reducing casework backlogs and potentially improving DNA recovery. In addition to reducing backlogs, using PCT may decrease the risk of cross-contamination and sample mix-ups, especially in cases where hairs submitted for DNA analysis arrive at the laboratory on an item of clothing. In these cases, the hairs are generally picked off the item and put into a plastic container until further analysis. However, it may be possible for these hairs to be placed directly into a PCT MicroTube (Pressure BioSciences). The main foreseeable advantage to using PCT is that sample preparation is automated using the Barocycler, so results should be more reproducible and less influenced by the extent of manual grinding. In addition, the Barocycler can simultaneously extract up to 48 samples, reducing sample processing and handling times for analysts.
To date, the studies that have examined the utility of PCT for extracting DNA from forensic-type samples have only reported preliminary data on the impact of varying PCT operational conditions on DNA yield.12 –18 Given that these studies largely remain unpublished or were completed by the manufacturer of the Barocycler (Pressure BioSciences), this proof-of-concept study was focused on performing a systematic assessment of PCT for the extraction of mtDNA from single short hairs without roots. Specifically, we examined (1) the ideal extraction conditions to ensure the highest mtDNA yield using PCT, specifically focusing on the extraction buffer, incubation, and cycling conditions; (2) the reproducibility of PCT cycling; and (3) the yields of PCT compared to traditional mtDNA extraction methods.
Materials and Methods
Hair Collection, Characterization, and Preparation
Shed fresh hairs were collected from eight volunteers in accordance with an FBI Institutional Review Board–approved project plan. Hairs were mounted with Permount (Electron Microscopy Sciences, Hatfield, PA) and characterized by a qualified forensic examiner. To ensure high similarity among donors but also to preferentially select donors likely to have larger populations of mitochondria in their hair shafts, four main characteristics were examined ( Table 1 ): (1) color/pigment, which indicates the presence of active melanocytes that produce new mitochondria and pigment granules (e.g., in gray hairs, the melanocytes are mainly dormant, thus not producing as many mitochondria); (2) the presence/absence of medulla, as shafts with medullas contain less mitochondria; (3) the cortical cells, as they are recipients of the pigment granules and likely the mitochondria; and (4) the diameter of the hair shaft, as hairs with a larger diameter with many cortical cells are likely to contain more mitochondria than hairs with a smaller diameter. 19 Following donor selection, hairs for extractions were individually affixed to a sheet of Scotch (Staples, Stafford, VA) lint rolled paper. To ensure no root tissue was present in extractions, 5 mm was trimmed off from the proximal root end. The adjacent hair was then sectioned into four 5-mm segments. All hairs were washed as follows prior to extraction, to clean the hair and remove any extraneous DNA 20 : (1) sonicated for 20 min in 1 mL Shandon Xylene Substitute (Thermo Scientific, Waltham, MA), (2) sonicated for 20 min in 1 mL 5% Tergazyme (Alconox, White Plains, NY), (3) rinsed in 1 mL of 100% ethanol (Sigma-Aldrich, St. Louis, MO), and (4) rinsed in 1 mL of 18 MΩ cm MilliQ water (Thermo Scientific). To minimize the inconsistency in mtDNA yields due to slight variations among individual hairs, fragments from multiple hairs from each donor were initially pooled together. From each donor pool, four random fragments were chosen (total of 2 cm of hair) and were used for each extraction. Following the current standard operating procedure (SOP), we did not weigh the hair fragments prior to extraction. Thus, it is likely that even with our attempts to minimize variation in the starting material among extracts, some extracts likely had more tissue than others.
Similarities in Hair Characteristics among the Three Donors Used in This Study.
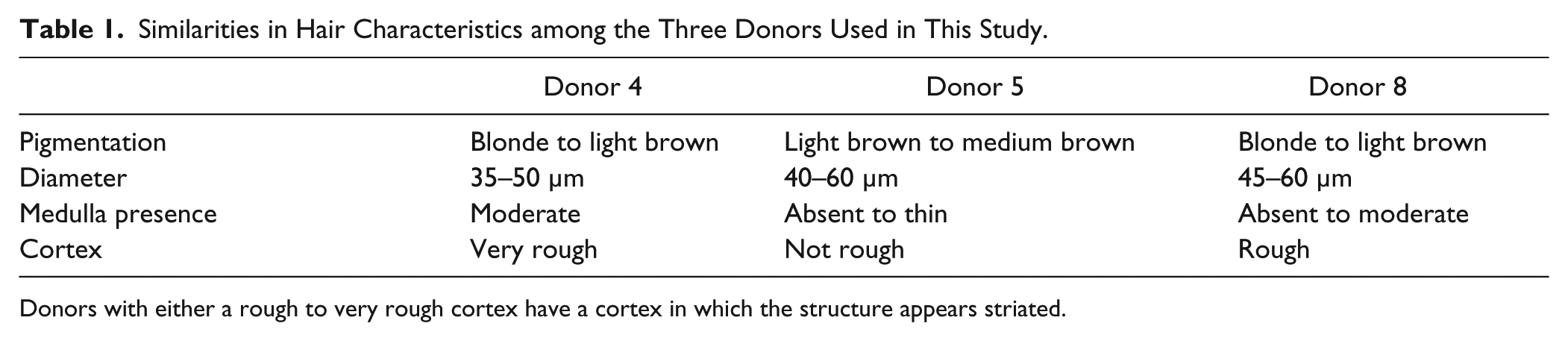

Donors with either a rough to very rough cortex have a cortex in which the structure appears striated.
PCT
Pressure cycling was performed using a Barocycler NEP3229 and a PCT MicroTube Adapter Kit (Pressure Biosciences). To determine the best operating conditions/parameters for maximum mtDNA yield, PCT was performed using varying (1) extraction buffers, incubation steps, and reaction volumes; (2) number of pressure cycles; (3) maximum pressures during cycling; and (4) times at maximum pressure. These variables were examined for each donor in the order listed above ( Figure 1 ). The ideal conditions were identified and used in subsequent extractions. All extractions were performed in triplicate unless otherwise noted. Reagent blanks were prepared and carried through each extraction (i.e., also in triplicate) and were found to be free of mtDNA.
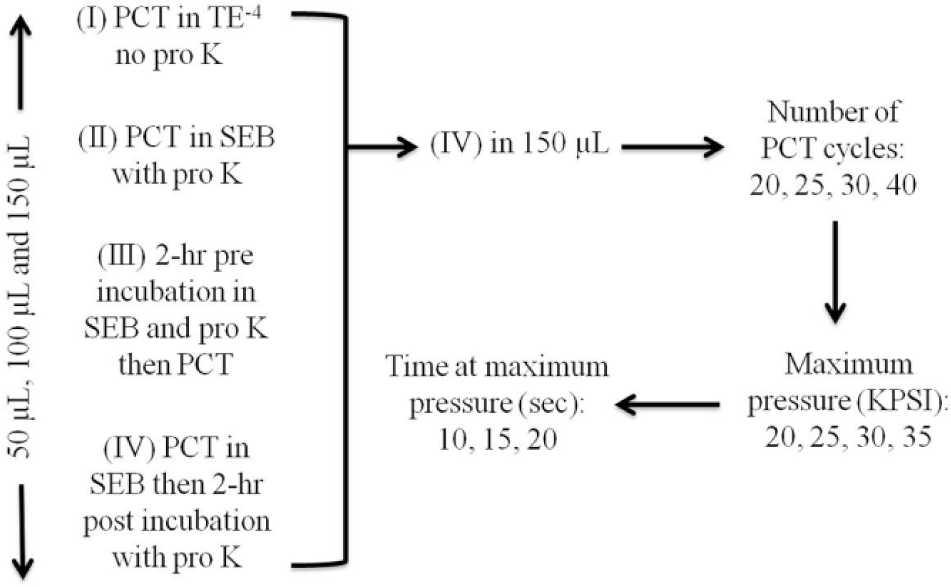
Experimental design for determining the ideal pressure cycling technology (PCT) conditions. First, a range of incubation conditions and reaction buffers was assessed, then (IV) was chosen and used at 150 µL for all subsequent reactions. Following this, the number of PCT cycles, the maximum pressure and the time at maximum pressure were examined. SEB, stain extraction buffer.
Evaluating Reaction Volumes and Incubation Conditions
Initial PCT extractions were performed in three reaction volumes with different incubation conditions: (1) PCT in TE–4 (10 mM Tris-HCl, 0.1 mM EDTA, pH 8.0) without proteinase K (pro K; 0.02 µg; Amresco, Solon, OH), (2) PCT in stain extraction buffer (SEB; 10 mM Tris, 100 mM NaCl, 29 mM dithiothreitol, 10 mM EDTA, 2% sodium dodecyl sulfate [SDS]) with pro K (0.02 µg) but no incubation, (3) 2-h preincubation in SEB with pro K (0.02 µg) at 56 °C in a 1.5-mL screw-cap tube prior to transferring the reaction mix to a MicroTube and performing PCT, and (4) PCT in SEB, followed by a 2-h postincubation with pro K (0.02 µg) at 56 °C (contents were transferred to a 1.5-mL screw-cap tube as MicroTubes could not be heated to 56 °C). For each of the above chemistries, hair extractions were performed separately in 50-µL, 100-µL, and 150-µL reaction volumes, using MicroTubes (part MTWS-MT-01), which used different-sized caps to create the tube volume (part MTWS-MC50-01, MTWS-MC100-01, MTWS-MC150-01, or MTWS-MC150-RK, respectively). Three different lots of MicroTubes and caps were obtained from the manufacturer, and none were contaminated with human mtDNA, as determined by the blanks run with each experiment. For these initial PCT extractions, the cycling conditions used were the same as those implemented by Feller et al. 14 : 30 cycles of 35 KPSI for 20 s and ambient pressure for 10 s.
Examining the Number of Cycles, Maximum Pressure, and Time at Maximum Pressure
First, to identify the ideal number of PCT cycles, 20, 25, 30, and 40 cycles were tested across donors (35 KPSI for 20 s, ambient pressure for 10 s). Following this, we examined the impact of varying the maximum pressure during the cycling, with the time samples subjected to maximum and ambient pressures kept constant (20 s and 10 s, respectively). Considering the Barocycler NEP3229 has a pressure limit of 35 KPSI, we tested cycling with a maximum pressure at 20, 25, 30, and 35 KPSI. The final variable tested was the length of time at the ideal maximum pressure. Holds of 10, 15, and 20 s were tested, with the length of time at ambient pressure remaining constant (10 s).
Following PCT, the homogenate volumes were adjusted to 200 µL in the same buffer in which the PCT was performed (i.e., low TE–4 buffer or SEB). Each homogenate was purified using phenol/chloroform/isoamyl alcohol (25:24:1) (Sigma-Aldrich) and concentrated by Microcon YM-100 (Millipore, Billerica, MA). To ensure the efficient recovery of DNA from the Microcon YM-100, hot 18 MΩ cm MilliQ water (Thermo Scientific) was used to elute the DNA. Extracts were stored at 4 °C until quantification.
Micro-Tissue Grinder DNA Extraction
To allow for a comparison of yields obtained using PCT to the standard mechanical disruption approach, additional mtDNA extractions were performed using paired mortar and pestle Kontes micro-tissue grinders (Kimble Chase, Vineland, NJ). Micro-tissue grinders were cleaned prior to extraction using 5% Tergazyme (Alconox), 4N H2SO4, and UV exposure (Spectrolinker XL-1500 UV Crosslinker; Spectronics Corporation, Westbury, NY). Similar to the PCT extracts, four randomly pooled 5-mm hair fragments were homogenized in 200 µL SEB until there were no visible pieces. Prior to a 2-h incubation at 56 °C and 500 rpm in a Thermomixer (Eppendorf, Hauppauge, NY), 1 µL of 20 mg/mL pro K solution was added to each sample. Phenol/chloroform/isoamyl alcohol (24:25:1) purification and Microcon YM-100 cleanup and concentration were performed as outlined above. Ten extractions were completed for each donor using the tissue grinder approach. A reagent blank was prepared and carried through extraction for each grinder by simulating grinding in 200 µL SEB, prior to processing each hair sample. An additional 10 PCT extractions (and reagent blanks) were also completed using the ideal PCT conditions. All reagent blanks were later observed to contain no mtDNA. Extracts were stored at 4 °C until quantification.
Incubation-Only Extractions
Ten samples from each donor were extracted following only incubation, no PCT, or grinding to determine the baseline yield when neither of these processes is implemented. For these samples, four randomly pooled 5-mm hair fragments were incubated for 2 h at 56 °C in 200 µL SEB with 1 µL pro K solution (20 mg/mL). Following incubation, samples were purified, cleaned, and concentrated as described above.
DNA Quantification
Extraction quantification was performed using a duplex TaqMan qPCR assay 21 that has been validated for forensic casework, 22 which included a human mtDNA-specific high performance liquid chromatography (HPLC)–purified probe (QRL8: 5′-6FAM CAT TCC TGC ACA TCT G MBGNFQ-3′) (Applied Biosystems, Foster City, CA) and TaqMan Exogenous Internal Positive Control Reagents (Applied Biosystems). Amplification was performed in 25-µL reactions: 12.5 µL TaqMan 2× Fast Universal Master Mix (without UNG) (Applied Biosystems), 900 nM forward primer (5′-GGC ATC AAC CAA CCA CAC CTA-3′), 900 nM reverse primer (5′-ATT GTT AAG GTT GTG GAT GAT GGA-3′), 250 nM QRL8 probe, Exogenous IPC Reagents (2.5 µL of 10× IPC primer and probe, 0.5 µL of 50× IPC DNA), and 2 µL of sample, standard, or control (water, TE–4, or IPC Block [0.8×]). This primer set allows the amplification of a 105–base pair (bp) target (positions 12,288–12,392 of the revised Cambridge Reference Sequence [rCRS]).
A polyacrylamide gel electrophoresis (PAGE)–purified synthetic quantitative PCR (qPCR) standard (Ultramers; Integrated DNA Technologies, Coralville, IA) composed of two complementary oligonucleotides, Tfor8sig and Trev8sig, 21 was used in the qPCR assay. Quantitative PCRs were performed in duplicate on a 7900HT Fast Real-Time PCR System (Applied Biosystems) in “Fast” mode: 95 °C for 20 s, 40 cycles of 95 °C for 1 s, and 60 °C for 20 s. Data analysis was performed using Sequence Detection Software version 2.4 (Applied Biosystems) with the cycle threshold (Ct) set to 0.2 and an automatic baseline. The efficiency of each run was determined based on the slope of the standard curve plot: efficiency = 10(–1/slope) – 1. The qPCR data were imported into Microsoft Office Excel 2007 (Microsoft Corporation, Redmond, WA) to calculate the average among duplicates and the standard deviation among extracts. Microsoft Office Excel 2007 was also used to identify any potential outliers by calculating the lower and upper outlier boundaries: 1.5 times the interquartile range was either subtracted or added to the value of quartiles 1 and 3, respectively. 23
Amplification and Sequencing of Hypervariable Regions 1 and 2
To assess mtDNA quality, hypervariable region 1 (HV1) and hypervariable region 2 (HV2) were amplified and sequenced from a subset of PCT extracts from all donors, using the primers outlined in Wilson et al. 24 PCRs were performed in 25-µL reactions: 2.5 µL GeneAmp 10× PCR Buffer (Applied Biosystems), 2.5 µL bovine serum albumin (BSA, 1.6 µg/µL; Invitrogen, Carlsbad, CA), 2.0 µL dNTP mix (10 mM; Applied Biosystems), 0.5 µL forward primer (30 µM), 0.5 µL reverse primer (30 µM), 1.0 µL AmpliTaq Gold DNA Polymerase (5 U/µL; Applied Biosystems), 6.0 µL high-purity water, and 10 µL sample or control (water or positive control DNA). Reactions were amplified on a 9700 GeneAmp PCR System (Applied Biosystems) under the following conditions: 95 °C for 9 min, 36 cycles of 95 °C for 10 s, 60 °C for 30 s, 72 °C for 30 s, and hold at 4 °C. Following PCR, amplicons were treated by incubating the entire volume with 5 µL ExoSAP-IT (Affymetrix, Santa Clara, CA) at 37 °C for 15 min and 80 °C for 15 min, before bringing to 4 °C (using a 9700 GeneAmp PCR System). Samples were quantified using a 2100 Bioanalyzer (Agilent Technologies, Santa Clara, CA) and a DNA 1000 Series II LabChip Kit (Agilent Technologies) with 2100 Expert Software (Agilent Technologies). Cycle sequencing was performed using the BigDye Terminator v1.1 Cycle Sequencing Kit (Applied Biosystems). Each sequencing reaction contained 3.5 µL of primer (1 µM), 9.5 µL of diluted BigDye (0.85×), and ~10 ng of template, with the final volume of 20 µL made up with nuclease-free water. Reactions were amplified on a 9700 GeneAmp PCR System (Applied Biosystems) under the following conditions: 96 °C for 1 min, 25 cycles of 96 °C for 15 s, 50 °C for 1 s, 60 °C for 1 min, and hold at 4 °C. Sequencing products were separated using a 3130xl Genetic Analyzer with POP-6 polymer (Applied Biosystems) and data analyzed using Sequence Analysis 5.2 (Applied Biosystems) and ChromasPro 2.4.3 (Technelysium Pty Ltd, South Brisbane, Qld, Australia).
Results and Discussion
Similarity between Donors
Multiple hairs from eight donors were microscopically examined by a qualified forensic examiner to determine the three most similar donors. Of the three donors selected (donors 4, 5, and 8), all were Caucasian, had blonde to medium brown hair pigmentation, had similar average hair shaft diameter, and had a discontinuous medulla when present ( Table 1 ).
Ideal Extraction Volume and Incubation Conditions
Initial extractions were performed using two different buffers, three buffer volumes, and several incubation conditions. Irrespective of the reaction volume and donor, none or very few copies of mtDNA were obtained when hairs were subjected to PCT only in TE–4 or in SEB that contained pro K ( Fig. 2 , I and II, respectively). A preincubation of the hairs in SEB with pro K at 56 °C resulted in a good yield of mtDNA ( Fig. 2 , III). However, the highest mtDNA yields were obtained when PCT was performed in SEB, followed by 2-h postincubation with pro K at 56 °C ( Fig. 2 , IV). This method and extraction buffer was used for all subsequent optimization extractions and most closely mimics the traditional method (samples are homogenized in SEB buffer and subsequently incubated with pro K). Substantial deformity of the MicroTubes was noted after PCT in 50- and 100-µL reaction volumes but not as much in 150 µL (complete tube destruction was not observed). Comparable yields were obtained among the two highest extraction volumes ( Fig. 2 ); therefore, all subsequent homogenizations were performed in 150 µL. These results highlight the importance of assessing the impact of varying reaction and incubation conditions on mtDNA yield.
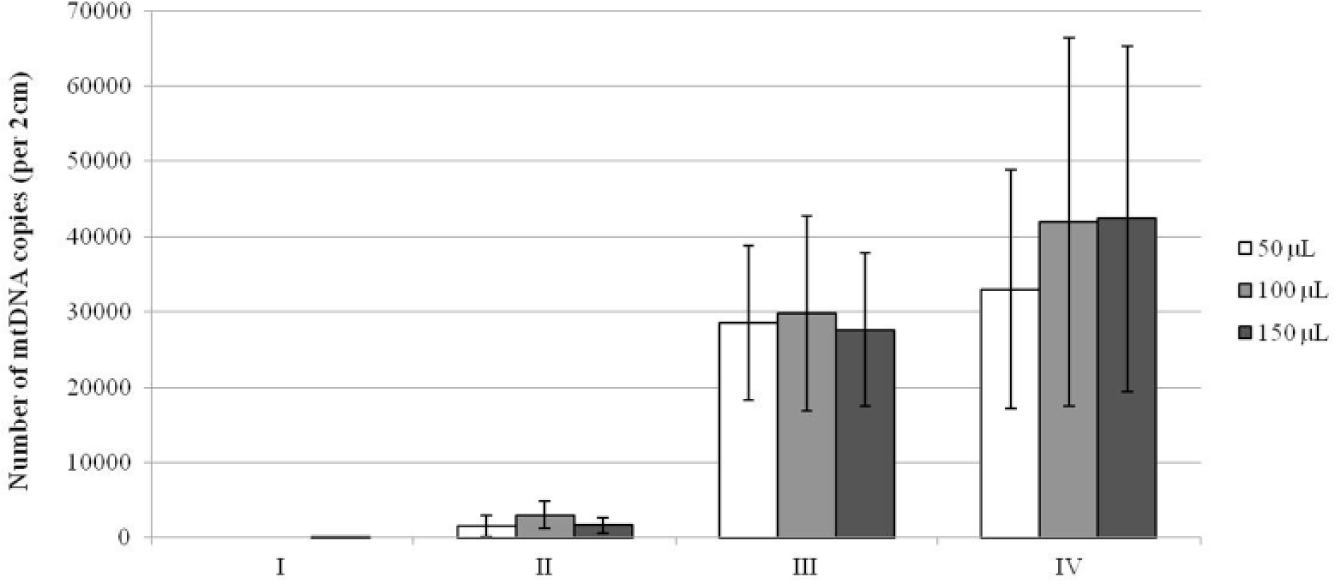
Comparison of mitochondrial DNA (mtDNA) yield from pressure cycling technology (PCT) extractions in a variety of reaction volumes and incubation conditions. Incubation conditions as follows: (I) PCT in TE–4 without proteinase K, (II) PCT in stain extraction buffer (SEB) with proteinase K but no incubation, (III) 2-h preincubation in SEB with proteinase K at 56 °C prior to PCT, and (IV) PCT in SEB, followed by a 2-h incubation with proteinase K at 56 °C. Average yield across the three donors is given (total N = 9, as extracts were performed in triplicate for each donor). The bars represent the standard deviation among all extracts, which was calculated from the duplicate average of each extract.
Ideal PCT Conditions
To comprehensively evaluate PCT as an automated approach for extracting mtDNA from hair shafts, we attempted to find the ideal cycling parameters. As the mtDNA yields for the three conditions examined (number of cycles, maximum pressure, and time at maximum pressure) were largely overlapping for all donors, it was difficult to conclusively determine the ideal PCT conditions ( Fig. 3a–c ). The donor dependency of mtDNA yields appeared to vary between the type of parameters investigated. For example, donors 4 and 8 showed the lowest yields when examining the number of cycles ( Fig. 3a ), but donor 8 showed the highest yields when different times at maximum pressure were assessed ( Fig. 3c ). For donors 4 and 5, the number of cycles that gave the highest yield was used in subsequent extractions to determine the ideal parameters (20 and 40 cycles, respectively; Fig. 3a ). However, for donor 8, 40 cycles were chosen as the variation among the number of cycles was largely overlapping ( Fig. 3a ), and other studies have shown that performing increased cycles (up to 60) does not negatively affect yield. 17 It was difficult to determine an optimal maximum pressure for cycling but also the time at that maximum pressure from our results. Although the lowest yields were obtained at 20 KPSI, when the variation is taken into account for a given donor, the yields were comparable across 25, 30, and 35 KPSI ( Fig. 3b ). For each donor, the mtDNA yields were very consistent when maximum pressure was held for 10, 15, or 20 s ( Fig. 3c ). For all subsequent extractions, we chose cycling to reach a maximum pressure of 35 KPSI for 20 s. These settings have been used by others with a broad range of tissues.6,25,26 In summary, we chose the following ideal cycling conditions for subsequent extractions of mtDNA from hair shafts using PCT: 40 cycles of 35 KPSI for 20 s.
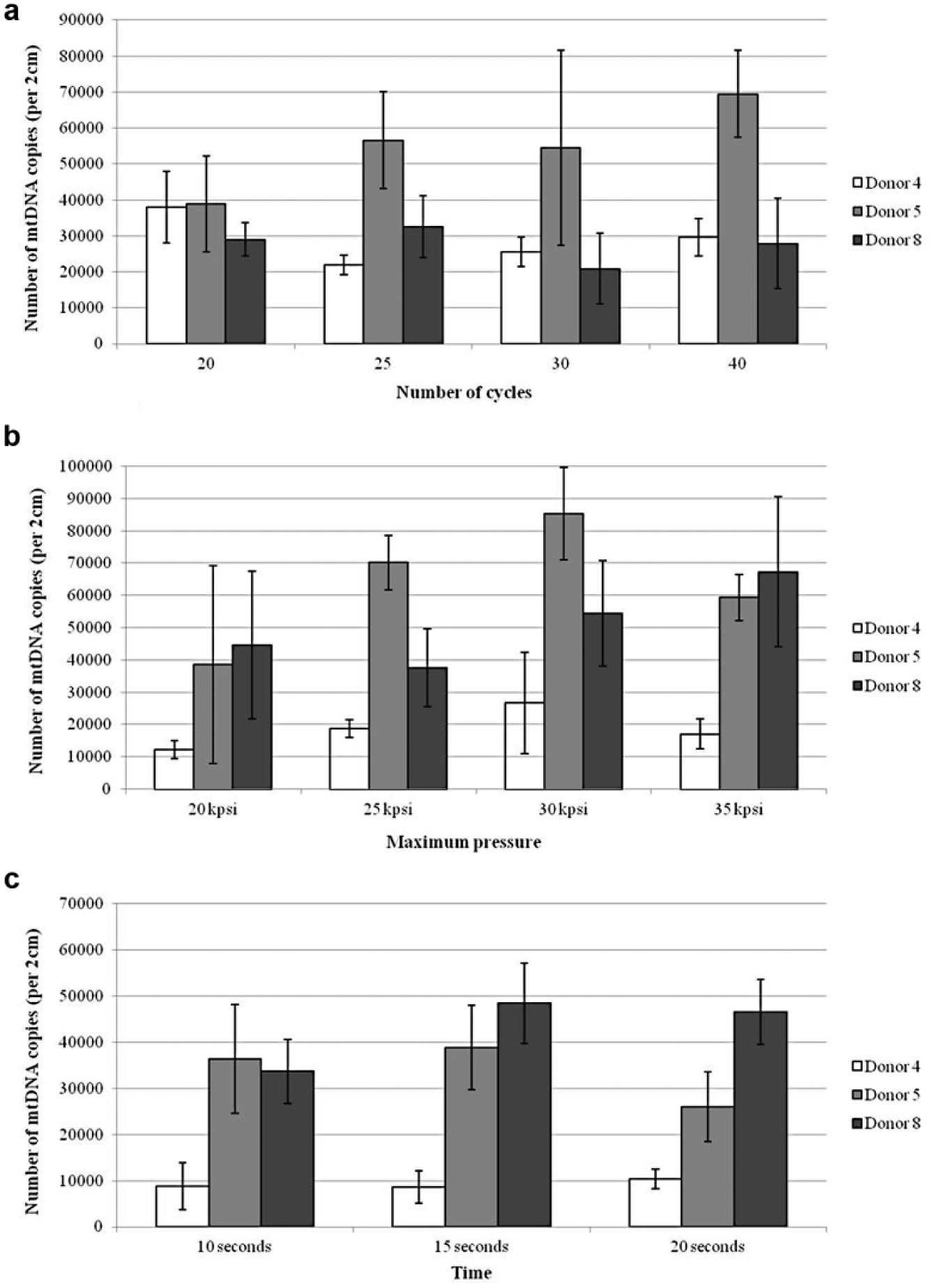
Comparison of mitochondrial DNA (mtDNA) yield obtained when pressure cycling technology (PCT) parameters are varied. Average mtDNA yields for donors 4, 5, and 8, with bars representing the standard deviation among extracts for a given donor (n = 3). Examination of the ideal: (
Comparison of PCT to the Tissue-Grinding Procedure
We performed parallel extractions from each donor using (1) only an incubation (with neither grinding nor PCT), (2) the ideal PCT conditions, and (3) the micro-tissue grinder method, to allow for a comparison between approaches. As expected, the incubation-only extractions yielded a relatively low yield of mtDNA ( Fig. 4 ). For donors 4 and 8, the micro-tissue grinder extracts had higher average yields than the PCT extracts, whereas for donor 5, the yields from PCT and tissue grinding were highly comparable ( Fig. 4 ). The lower apparent performance of PCT for extracting mtDNA may be attributed to some deformity in the MicroTubes after PCT (even when using the 150-µL extraction volume), along with incomplete dissolution of the hairs. Both of these factors made it difficult to ensure the complete transfer of the homogenate, along with any hair fragments, to a new tube after PCT for the 2-h postincubation at 56 °C. In addition to this, it was possible that the presence of hair fragments may have negatively affected the subsequent phenol/chloroform/isoamyl alcohol extraction. We observed that when hair fragments were present, they were located in the interface, which may have influenced the complete recovery of the aqueous layer, likely decreasing yields. The inclusion of partially fragmented hairs did not occur when using the manual tissue grinder method, as all hairs were completely homogenized.
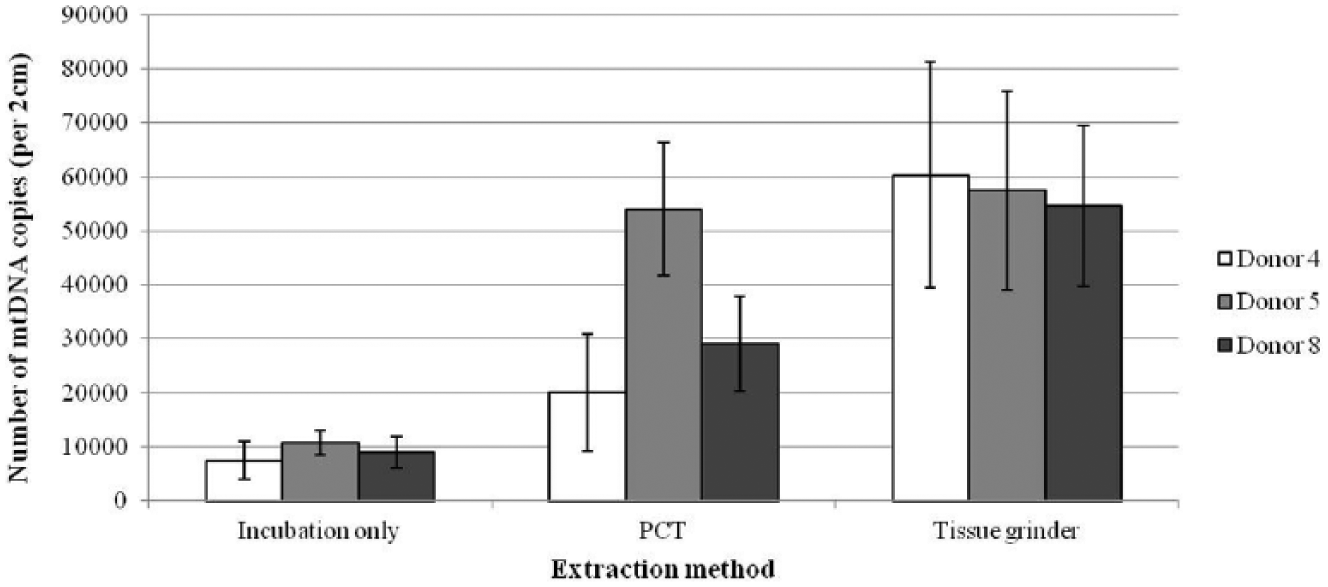
Comparison of mitochondrial DNA (mtDNA) yield obtained from three different extraction methods. Extraction methods as follows: incubation only, pressure cycling technology (PCT) followed by 2-h postincubation, and the traditional tissue grinding method. All reactions are in stain extraction buffer (SEB) with proteinase K. Yields given are the average of 10 extracts for a given donor, and the bars represent the standard deviation among extracts.
It is plausible to suggest the implementation of PCT as an automated approach for the isolation of mtDNA from hair shafts. If the MicroTube were improved, such that it could be heated to 56 °C and obviating the transfer of homogenate to a new tube, it is likely that the yields between PCT and micro-tissue grinders would be more comparable. The mtDNA yields from all three extraction methods exceed the required number of copies for forensic analysis (minimum of 100–760 copies required22,27). Even though the incubation-only extractions met the minimum copy threshold, it is always preferable to maximize DNA yield from vital casework samples, not only to obtain what is sufficient for analysis. It is important to reemphasize that the yields shown here were obtained from pristine hair shaft samples. As such, it is not expected that similar mtDNA yields would be obtained using either PCT or the micro-tissue grinder approach with typical casework hair samples. After completion of this work, a report that identified an automatable approach for mtDNA extraction from hair shafts using commercial kits was published. 28 Although this approach produced high yields of good-quality mtDNA, using PCT is still an attractive automated alternative.
Reproducibility of Extractions
Substantial inconsistency in the number of mtDNA copies obtained from extracts both within and among donors, but also across all extractions, was noted ( Figs. 2 , 3 ). To reduce variation among extracts, the starting samples for each extract were randomized; fragments from multiple hairs from a single donor were initially pooled together prior to random selection. However, given that we anticipated some level of consistency among hairs from a single donor, only enough fragments needed for a set of optimization extractions were pooled: if 10 extractions were to be completed on a given day, fragments from 10 random hairs were pooled, even though ~100 hairs were initially collected per donor. Thus, the difference in the average yield within a donor but across different conditions could be a reflection of the difference among sets of pooled hairs. The subtle differences in presence of medulla among donors ( Table 1 ), which is known to contain low amounts of mtDNA, 19 may have attributed to the interindividual variation noted. Comparatively high levels of both intra- and interindividual variation in mtDNA yields from hair shafts have been noted in other studies.27,29
Quality of PCT-Extracted DNA
Quantitative PCR validated that PCT can be used to extract mtDNA from hair shafts; however, the quality of this PCT-extracted DNA needed to be assessed. For example, the rapid changes in pressure could cause a whiplash movement of the molecule and break the DNA into fragments too small to sequence effectively but still long enough to be quantitated by qPCR (105-bp target). By analyzing the total genomic DNA by gel electrophoresis, one can determine whether the DNA has been fragmented or if a change in molecular weight has occurred after PCT. Using this approach, studies have shown that PCT extracts contain high molecular weight and unfragmented DNA.3,14 In addition, it has been reported that PCT extracts have routinely yielded PCR amplicons, ranging in size up to 700 bp.3,6,14
In this study, we amplified the HV1 and HV2 regions in a total of four fragments, each ~280 bp in length. Amplifications of these smaller fragments were successful for all PCT extracts, as determined by gel electrophoresis with the 2100 Bioanalyzer chip ( Fig. 5a ). The sequence data showed no base ambiguities, and peaks were well defined with little to no background interference at the baseline ( Fig. 5b ).
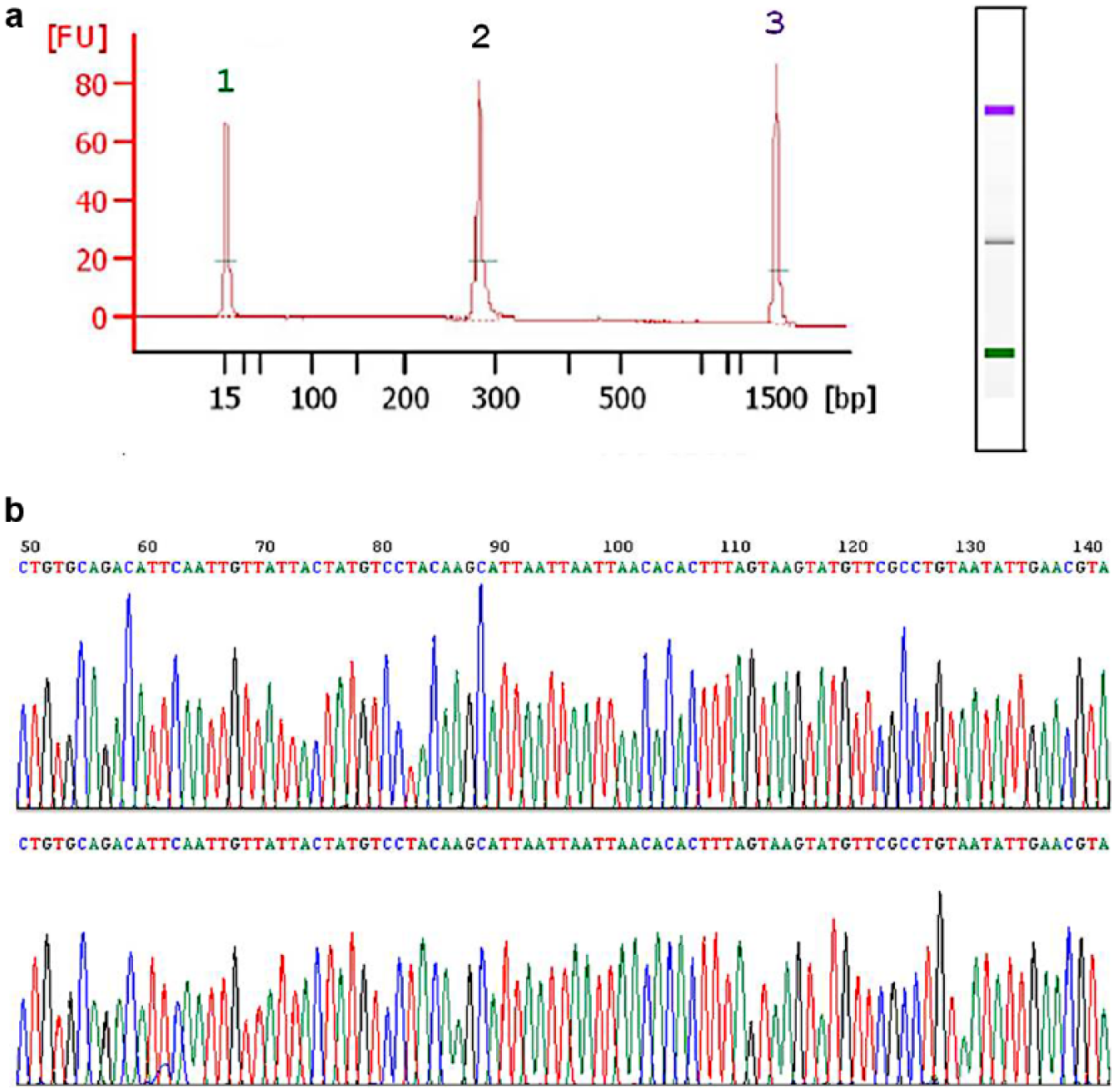
Qualitative evaluation of mitochondrial DNA (mtDNA) from pressure cycling technology (PCT) extracts. (a) Example electropherogram from the 2100 Bioanalyzer of hypervariable region 1A (HV1A) amplicon from a PCT extract. The single large peak (denoted by 2) correlates to the HV1A amplicon (~280 bp). Peaks at (1) and (3) are size standards (15 and 1500 bp, respectively). The X-axis is not linear. The concentration calculated by the 2100 Expert Software for the HV1A amplicon was 12.85 ng/µL. (b) A portion of the electropherogram for both the forward and reverse sequencing reads of HV1A from a PCT extract.
In conclusion, this study demonstrated the feasibility of PCT as an automated method for extracting mtDNA from single, short hairs without roots. Although the yields from PCT extracts were lower than those obtained using the micro-tissue grinding approach, the number of copies obtained using this automated approach exceeded the requirements for forensic analysis. The quality of the DNA from PCT extracts was validated by successful amplification and sequencing of the HV1 and HV2 regions. As this was a proof-of-concept study, we only focused on three donors and performed extractions in triplicate for most experiments. We observed a high degree of variance in some cases, which is not atypical of multiple hair extractions from a single donor. The removal of potential outliers would have decreased the variation; however, as no “true” statistical outliers were identified, all data were used in comparisons, allowing for an unbiased evaluation of PCT for extracting mtDNA from hairs shafts. To get a better understanding of the variation, we suggest a larger number of donors be examined in future studies. Hairs of various thicknesses also need to be evaluated because some hairs might yield more mtDNA under the PCT treatment than without it. To accomplish this, hairs from different biogeographic ancestries could be subjected to PCT treatment. In addition, PCT should be evaluated for hairs representing varying chemical treatments, as they can be resistant to chemical and enzymatic digestion. 24 Wilson and colleagues 24 have also reported that melanin, a hair pigment that can be coextracted with DNA, can inhibit PCR but may be removed with pressure cycling.6,7 As PCT technology provides an automated approach for processing numerous samples in a time-effective and consistent manner, it could be implemented for mtDNA casework to help reduce backlog.
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported in part by an appointment to the Visiting Scientist Program at the FBI Laboratory Division, administered by the Oak Ridge Institute of Science and Education, through an interagency agreement between the US Department of Energy and the FBI.
Disclaimer
This is publication number 14-23 of the Laboratory Division of the Federal Bureau of Investigation. Names of commercial manufacturers are provided for information only, and inclusion does not imply endorsement by the FBI or the US government. The views expressed are those of the authors and do not necessarily reflect the official policy or position of the FBI or the US government.
