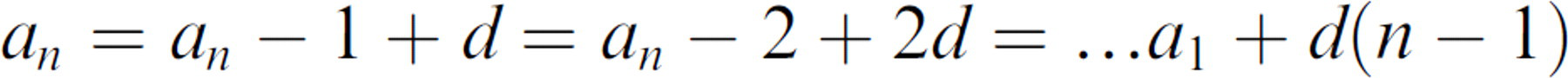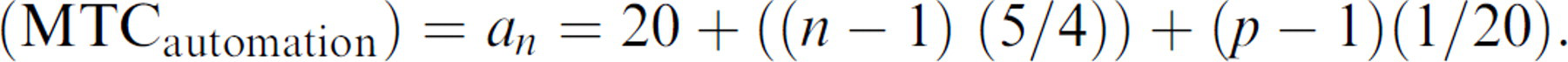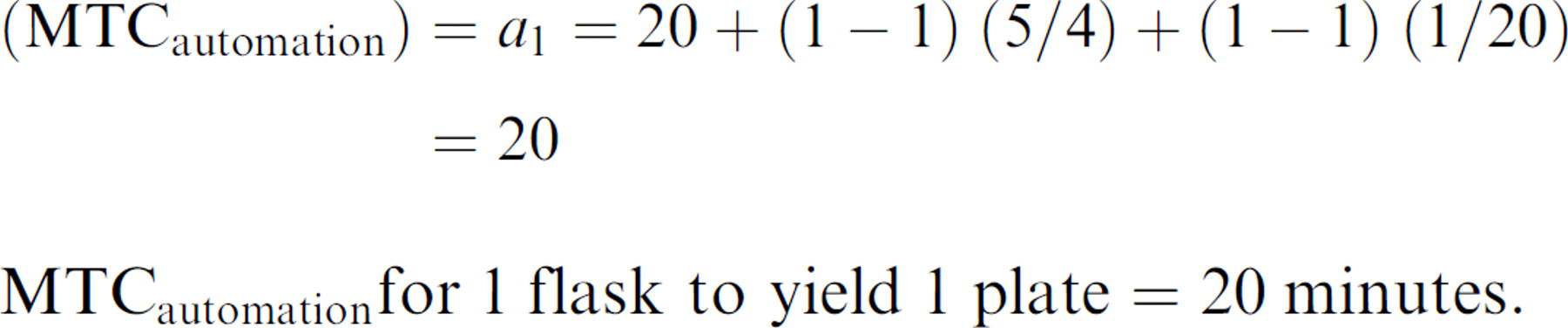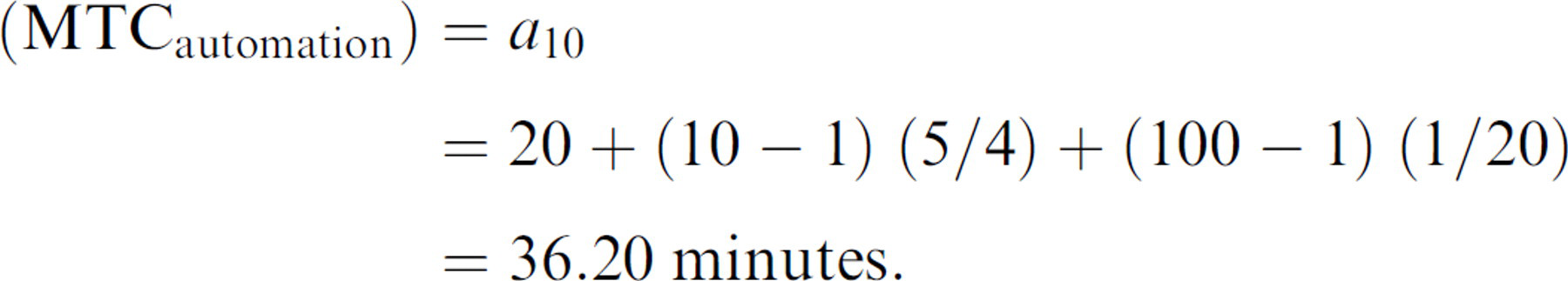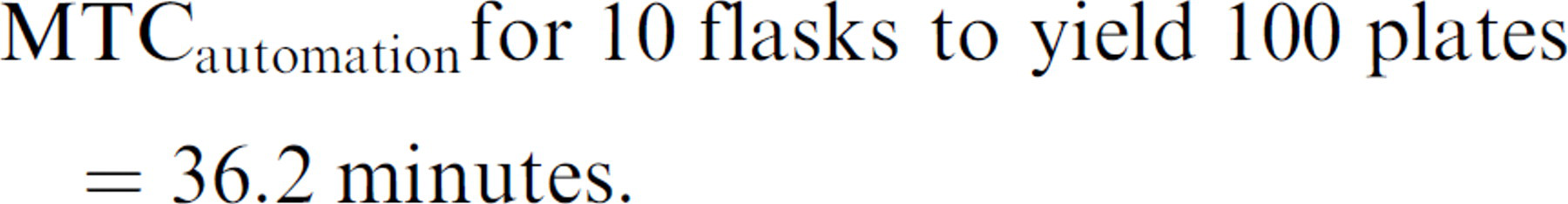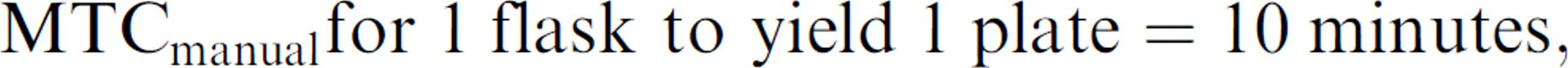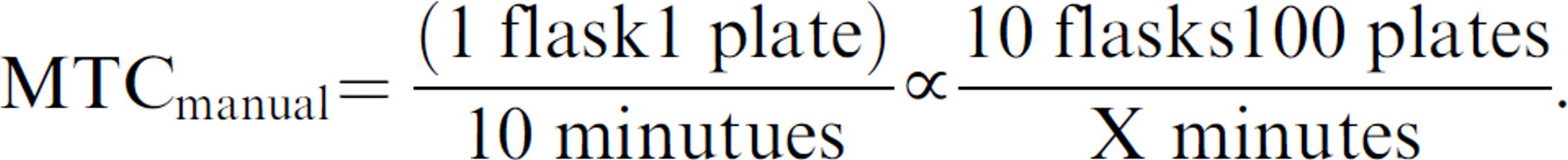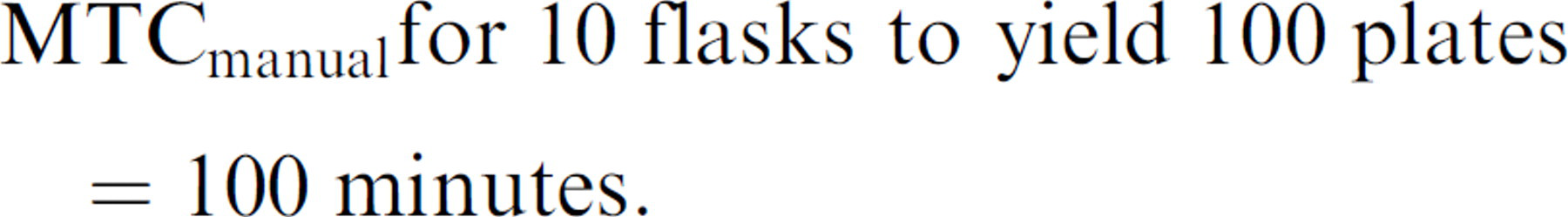Abstract
An automated cell-culture platform becomes the nucleus of an organization performing cell-based research. However, every cell-based project placed on the system brings unique challenges. With each cell line comes millions of years of evolutionary encumbrance and a genetic inclination driving unique phenotypic peculiarities. In vivo, diverse eukaryotic cells rely on their “mammalian host” for survival. An automated system must perform in vitro, the myriad actions needed to sustain multiple cell lines as well, hence becoming an “automated host.” Cells invariably, will endeavor to do as they please. Molding these cells into the operational bounds of a man-made system requires insight into the relationship between cell and machine.
Citing our own experiences, we will describe herein the use of the SelecT automated cell-culture platform (The Automation Partnership, Hertfordshire, England) in our discovery and preclinical profiling programs at Novartis. Achieving the balance between cells and the automated environment, and accommodating variable cell dynamics are discussed.
Keywords
Introduction
Expanding Needs of Cell-Based Assays
In recent years, the scientific community has entered a new phase in its ability to study diverse compound libraries in the context of living organisms. Crude tissues and the immortal HeLa cell line once embodying much early in vitro eukaryotic cell modeling have given way to diverse cell lineages for studying the effects of compounds on a living organism. Today, the availability of cells to researchers encompasses lineages representing nearly every tissue, of every organ, of myriad different species. The American Type Culture Collection (Bethesda, MD) for example, lists over 3600 phenotypically distinct cell lines from over 80 different species. These lineages are derived from keratinocytes, osteocytes, tumors, neuron, lymphocytes, stem cells, etc. The basic components of an entire being, sans sentience and a hominid form, can be modeled in living cells. Specialized cell lines, introducing reporter genes, protein tags, response elements, and allele knockouts 1 –3 expand the repertoire for modeling cell-compound interactions.
As the size of compound libraries has grown over the years, 4 the size of the reaction volume in which specific cell assays are performed has decreased. 5 Cell-based assays have shrunk from the Petri dish, to 96-well, to 384-well, and on to the 1536-well format, where a mere few hundred cells per assay may be analyzed. The high content analysis assays of the present now promise detailed analysis in a single cell. 6 The need to miniaturize, increase speed, precision, and reduce costs while permitting increased multitasking and flexibility has historic precedence in many technologies. The evolving trend from single readout analysis of pooled cells in Petri dishes to multiplexed readout and analysis of a single cell 7 resembles the well-known evolution of computer components—a process whereby the vacuum tube gave way to transistors, and later to silicone chips. The sophisticated cell handling and imaging of today's 1536-well formats bear as much resemblance to the manual cuvette readers of a few decades ago, as a modern microprocessor bears resemblance to the Charles Babbage's hulking “difference engine” of the 19th century, which at 15 tons, and a height of 8 feet, could automatically perform simple calculations on tabular data (Fig. 1).
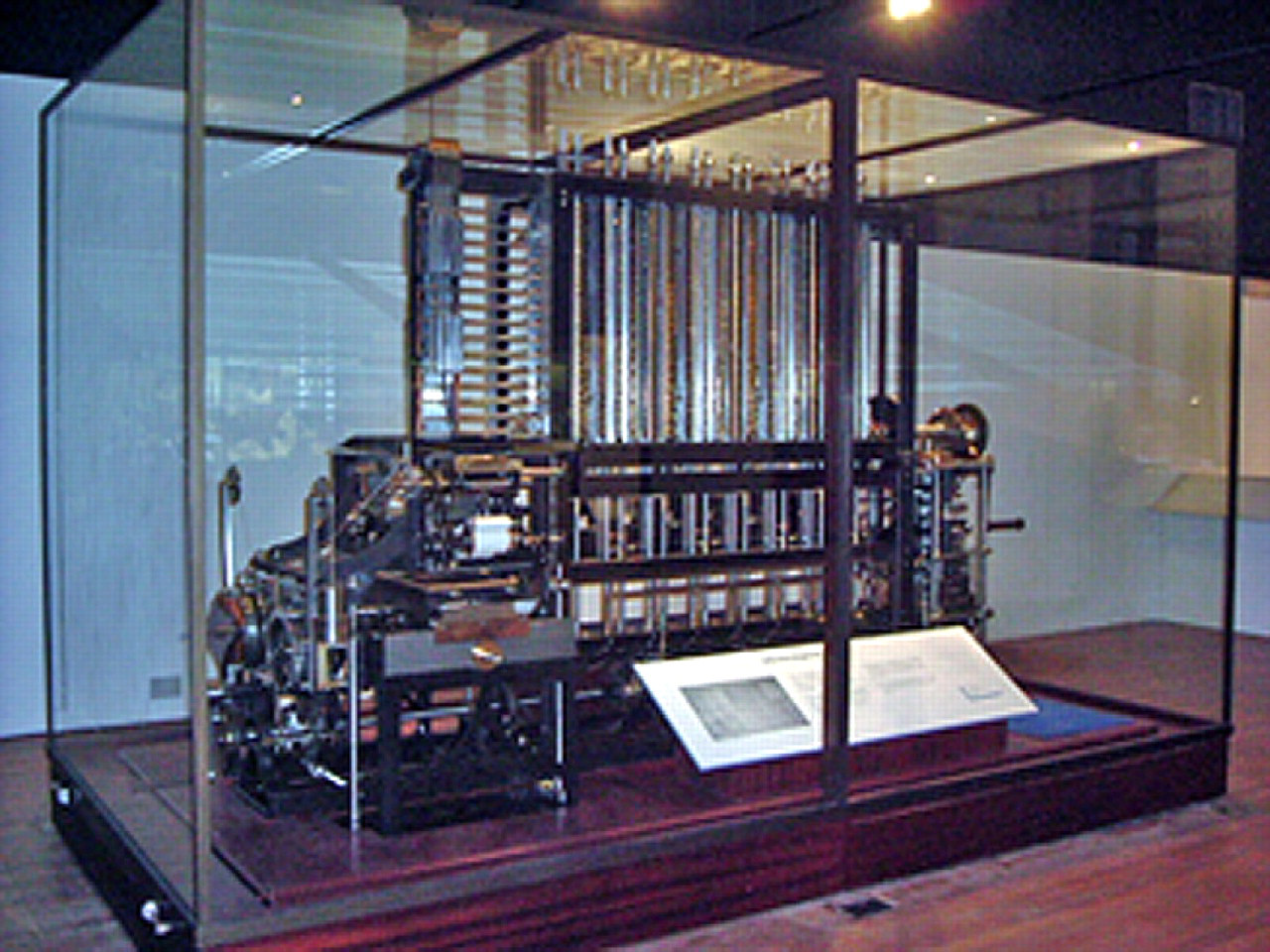

Babbages difference engine.
The use of mammalian cells as a physiological barometer of the drug/target interaction in HTS and early clinical profiling has assumed a larger role in the drug discovery efforts of pharmaceutical companies, including Novartis.
According to a mid-decade analysis, the global cell-based assay market in 2004 was pegged at approximately $475 million dollars, with an expected increase to $720 million dollars by 2009. 8 In our laboratory, technology is trending toward miniaturization. Even so, the proliferation of new target classes and demands for more early clinical toxicology profiling models places a net trend in demand toward manipulation of more cells in increasingly diverse cellular models.
We are among a growing number of cellular automation biologists who are, or will be expected to rely increasingly on automation to propagate, transfect, and coculture cells that require increasingly complex conditions to maintain cellular assay efficacy. Automating cellular assay efficacy we find, requires parity between three components: (1) Sustaining the most archetypal cell model, (2) Establishment of cell robustness to withstand unpredictable assay demands, and (3) Sustaining phenotypically disparate cells in a finite, automated environment.
The Archetypal Cell Model
Mammalian Cells as Physiological Models
Of the three components comprising automated cellular assay efficacy, the cell model is perhaps the most difficult to characterize, design, and sustain accordingly. It is therefore worthy of the most discussion.
Mammalian cells are mercurial, volatile entities, and their inherent peculiarities are still being unveiled. A useful context that envisages how immature the science of in vitro cell culture really is, can be made by adapting a model put forth by the futurist and sociologist, Alvin Toffler.
What Toffler embraced was contextualizing the entire evolution of human technology and achievement into his cogent “800th lifetime” 9 analogy. To quantitate human achievement, Toffler divided the past 50,000 years of the human experience into equal lifetimes of 62 years each. Using this time line, of these 800 lifetimes one sees that the first 650 were spent by humans living in caves. In only the last 70 of these 800 lifetimes has it been possible for humans to communicate from one lifetime to another using the written word. Only in the last four, have humans been able to measure time with any precision, and only in the last two lifetimes has anyone used an electric motor.
Transferring the evolution of cellular biology into Tofflers' model, of the first 798 lifetimes in vitro cell culture did not even exist. A cells genetic constitution consigned it to a healthy finite existence or perhaps to cancer and immortal proliferation. Regardless of fate, the cell was completely dependent upon its “mammalian host.” For every cell from zygote, to stem cell, to the full complement of differentiated somatic cells that constitute an organism, interphase through telophase, metabolic convention was relegated to the boundaries of the mammalian host, a “host island.” It was not until the 798th lifetime that Wilhelm Roux was able to excise living cells from a host—a live chick—and maintain chick embryo cells in saline for 3 days. Furthermore, it was not until the second half of the 800th lifetime, that the first continuous human cancer line was grown in vitro. 10 Nearly, every cellular automation contrivance used has been developed in the last hours of the 800th lifetime.
It has been a remarkable ascension of technology that has lead to the in vitro culture of living cells. In vivo, a mammalian cell programmed by its 3 billion nucleotide code is not only destined to a particular path based on its genotypic composition, but it is also subject to the whims of cytokines, hormonal excitations, antibodies, and myriad extraneous influences. Author and scientist, Richard Dawkins paraphrased this point by stating “… the true utility function of life, that which is being maximized in the natural world, is DNA survival. But DNA is not floating free; it is locked up in living bodies and it has to make the most of the levers of power at its disposal.” 11 Hence, removing live cells from the host, placing them into an external environment, and molding them into tools for performing cell-based assays contrary to what is dictated by their genetic propensity presented certain challenges.
A cell or cell colony is not static and it is moving in three dimensions. It is changing in mass, changing in density, and changing in phenotype. The best-laid experimental design for a cell-based assay may come to naught, for cells will invariably attempt to endeavor as they please (Fig. 2).
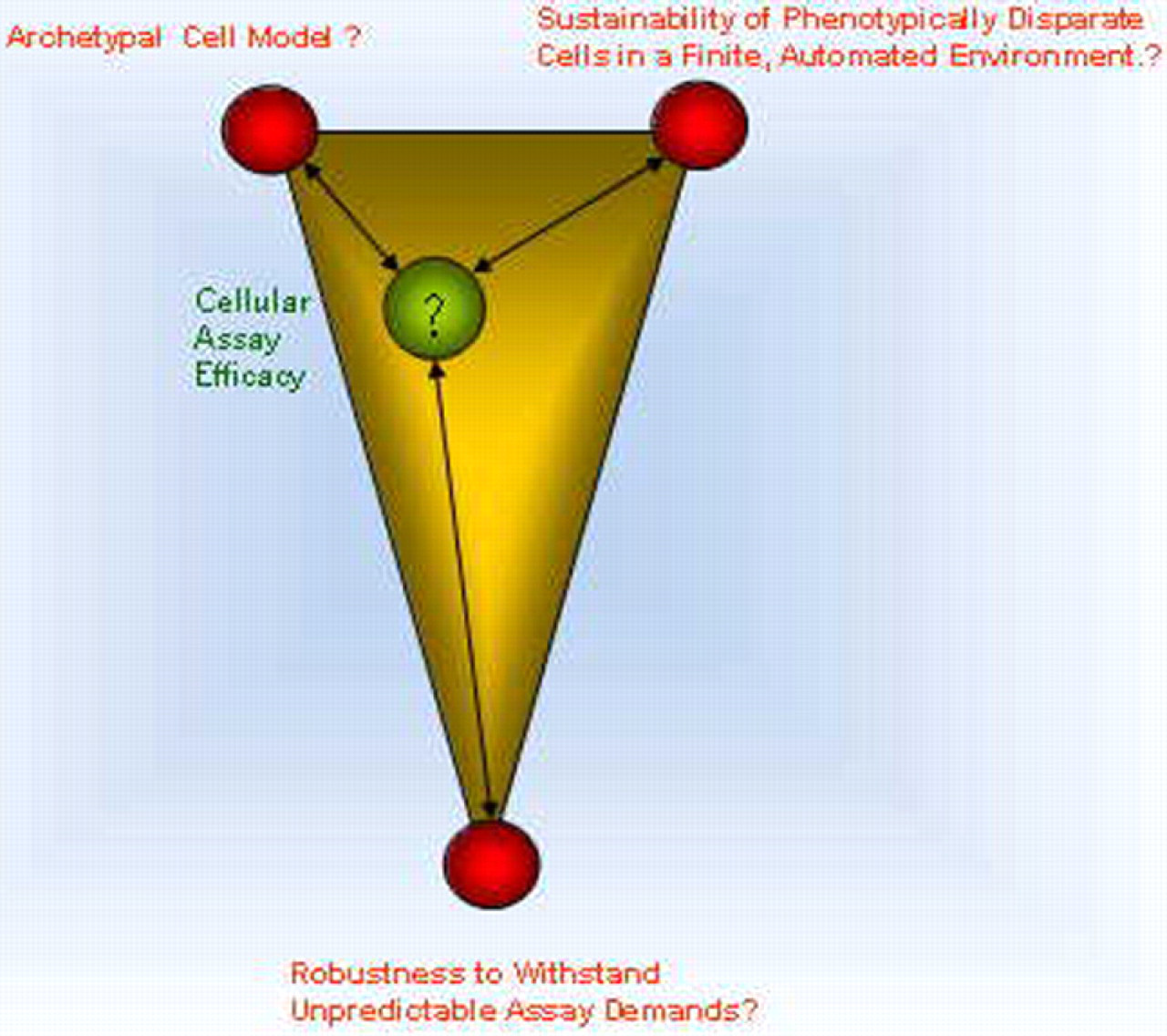
Cellular assay efficacy may be fairly balanced between archetypal cell models and the automated cell environments. Adding robustness however, to decouple cell automation from unforeseen downstream assay automation introduces challenges to cellular assay efficacy.
Modeling Cells Manually as a Predecessor to Automated Cell Culture
It should be kept in mind that the cell line of choice for a physiological model is merely a subset—an incomplete sampling of the tissue or organism being studied. Often, the cell line of our choice only bears partial resemblance to the human model it strives to emulate. For example in our laboratory, we use a Chinese Hamster Ovary cell line expressing the human ether-a-go-go related gene (hERG) channel for studying cardiac toxicity of experimental compounds. 12 Contributing factors to such decisions may include robustness of the cell line, ease of transfection, superior signal-to-background characteristics of the assay, or a clearer understanding of the signaling pathways of that line.
It is possible to partition cell culture into at least two broad categories. One category being the use of cells as tools to routinely express protein for further processing. Here, the cell model is not so important—so long as the secreted product displays the correct biological activity (i.e., antibody production). Performed in mammalian, insect cells, or bacteria, these cells are harnessed in bioreactors and fermentation vessels at steady state. They are outside the scope of this discussion. The other category uses cells as models to study cellular pathways, responses to stimuli, and to emulate a particular mammalian physiology. This requires application of smaller, though more diverse cell-culture methods and conditions. Adapting these models to an automated system is an additional challenge.
Before one can effectually begin the process of automating cell biology models, we find it an irrefutable asset to the individual considering automated cell culture, to have a firm grasp of the biology of in vitro cell propagation. Understanding cell biology and anticipating potential perversions to the cell model should presage large-scale automation commitments. Absolute authority on the biology of every cell model being considered for automation is not imperative. However, the biology is the greatest variable and it generally drives programming design on the system.
Many automation processes have arisen by applying an automated paradigm to a skilled, manual technique. For example, introducing the mechanical sewing machine to the labor force in 1790 perhaps removed tedium from the sewing process that might have been associated with producing bulk yardage of industrial linens. This same device in the hands of a skilled tailor, however, could introduce flexibility and increased output without compromising his craft and expertise as a tailor. Allegorically, we have extracted enormous utility from our SelecT (The Automation Partnership, Hertfordshire, England) in “bulk yardage” cell production. However, tailoring live cell biology in vitro strikes a balance between science and intuition. Automating cell culture further challenges the intuitive component and presents the unique challenges, which will follow.
Intuition versus Automation and the Cell Model
An experienced cell biologist navigates cell culture and cellular biology in an approach comprising part science and part intuition. The subtle changes in an in vitro cell culture are often hard to quantitate and if not addressed promptly, culminate in rapid deterioration or phenotypic change to the cell line of interest. Deterioration is often manifested as shed plasmid, decreased assay window, low viability, poor growth, decreased protein secretion, etc. 13 When it comes to automating the science of cell culture, it is translating this intuitive component into machine code to sustain a living thing that is intriguing and challenging. Using the sewing machine analogy, the science and craft of cell biology will generally be learned by scientists before exposure to large-scale automation. It is the subtle applications of techniques eventually rendered as obvious to manual handling of cells by a skilled person that need to be translated into equivalent robot articulations. Articulations capable of performing the same action on an automated system. For example, the simple hand application of a quick “knock” to dislodge the cells from a flask requires several lines of program code to achieve the same result using automation.
Another difference between manual and automated handling is the ability to process multiple cell flasks in parallel. Parallel processing becomes second nature to the scientist executing the tasks of counting, splitting, and feeding cells in a hood. The investigator may line up as large a number of flasks that they feel comfortable with addressing, then aspirate or dispense liquid among them. On completion, the individual might move a stack to the incubator in one motion. Our instrument however performs tasks serially. The arm is only capable of manipulating a single flask at one time. Attempting to emulate the efficiency of parallel processing has required development of some clever code and the application of new consumables to process flasks. For example, the NUNC triple flask, and Cornings' 10-tiered Hyperflask permit the system to emulate parallel flask handling actions.
Another example of the effort needed to transition from manual flasking to automated cell handling is the simple act of reseeding the same flasks after a split. Reseeding may be performed to avoid flask disposal if a scientist relying on the cells for some downstream application was not ready to use them. This mundane maneuver, so instinctively performed by hand required some rather counter-intuitive programming of the SelecT to work efficiently. With a little insight, automation adds freedom and flexibility, but the learning curve may be steep.
When attempting to harness full flexibility of the system, translating the instinct and science of cell biology into program code of the automated arena is the crux of fully exploiting automation potential. A way to appreciate the enigma of intuition versus automation with in vitro cell propagation, is to imagine catching a ball tossed to you from some distance. You are not consciously analyzing a complex trigonometric identity stipulating that the motion of an object on the Earths surface is determined by the acceleration and deceleration of gravity, its launch speed, and the launch angle. However an individual with reasonable prowess, can position themselves down range of the lofted ball, raise their hand and catch it, never lamenting that at some point they would need to analyze that Range = V 0 2 sin 2θ/g (where V 0 is the initial velocity, and g is of course the gravitation constant on earth, which is 9.8 m/s2). It is an intuitive assessment of the problem at hand. It is this type of intuition that should be encoded into the automation, line by line of code, to ensure that the cell model translates into the automated environment. Once this has been successfully achieved the precision of repetition, freedom of scheduling, and multitasking potential that can be tailored to the system becomes realized. This has not yet been completely attained, hence we must always balance our intuition with the automation capabilities.
Efficiency and the Matter of Scale
Our automated system does not work faster than people. It does, however, perform 24/7 and permits the scientist to pursue other endeavors while the machine performs. A logistical problem illustrating the relationship between robot processing speed and the compromise between serial versus parallel processing was encountered when translating the care regimen of some projects into the SelecT. We noted early with certain scenarios that a disproportional amount of valuable robot time was required per number of assayable cells returned. Large robot commitments are particularly evident during cell model optimization. For example, during assay development of a cell-based screening campaign, small test quantities of cells and flasks need to be processed under conditions of varying cell concentration, dissociation conditions, and plating volumes to optimize assay conditions. It was our observation that a threshold exists whereby transferring a cell model onto the robot becomes economically unwieldy and inefficient using the automated system. What may be described as Manual Time Commitment (MTC), is the overhead user intervention required to operate the machine for a unit of work. Alternatively, MTC describes the time needed to perform the same task manually. The change of efficiency is attributed to the difference between MTC needed to operate the SelecT versus the MTC needed to perform the same work manually. Our platform, like any automated cell-culture system, executes a set number of programmed mechanical articulations to perform a job. Additionally, however, a number of user-mandated steps are required (i.e., loading consumables, programming and scheduling the job, etc.) to complete an automated procedure (i.e., trypsinizing and counting cells to make one plate from one flask of cells). We have found that a simple arithmetic progression can be used to estimate the MTC, which is required to achieve certain work output. We find that for smaller cell projects placed on the system, the MTC is disproportionately high for the amount of output recovered compared to nonautomated MTC.
For example, we estimate the MTC for any number of flasks to be processed for generation of any number of plates can be expressed in terms of an arithmetic series:
where n = 1, 2, 3, 4 … = the number of flasks processed, d is the constant difference between each term, and p is the number of plates produced.
We found that MTC in minutes allocated by an individual preparing the instrument to process n flasks to generate p plates is thus approximately described by the equation:
Therefore, the user interaction needed in preparing the instrument to process 1 flask to generate 1 plate can be described by:
Therefore, the MTC for preparing the SelecT to process 10 flasks for say, a 100-plate job is expressed as:
therefore:
So using the SelecT, we can process 10 times the number of flasks to yield 100 times the plate output, though conceding only 36.2 min/20 min = 1.81 times the MTC of preparing a single plate from a single flask on the system.
However, a formula modeling purely manual preparation of flasks and plates generally adheres to rules of proportion.
So to process 1 flask by hand to prepare a single plate:
which at MTC =10 min is twice as efficient, requiring only half the MTC of using the robot for the same task.
However, processing 10 flasks manually to yield 100 plates manually is expressed as the proportion:
Solving for X:
Therefore, 10 times the manual flask processing to yield 100 times the number of plates requires 100 min/10 min =10 times the MTC of preparing a single plate from a single flask. For this larger job to be performed manually, MTC at 100 min is 2.76 times greater than the MTC of 36.2 min using the robot.
Even though the flasks can perhaps be processed serially by hand faster than the robot, the manual-plating component requires extensive time resources.
This can be summarized in Figure 3.
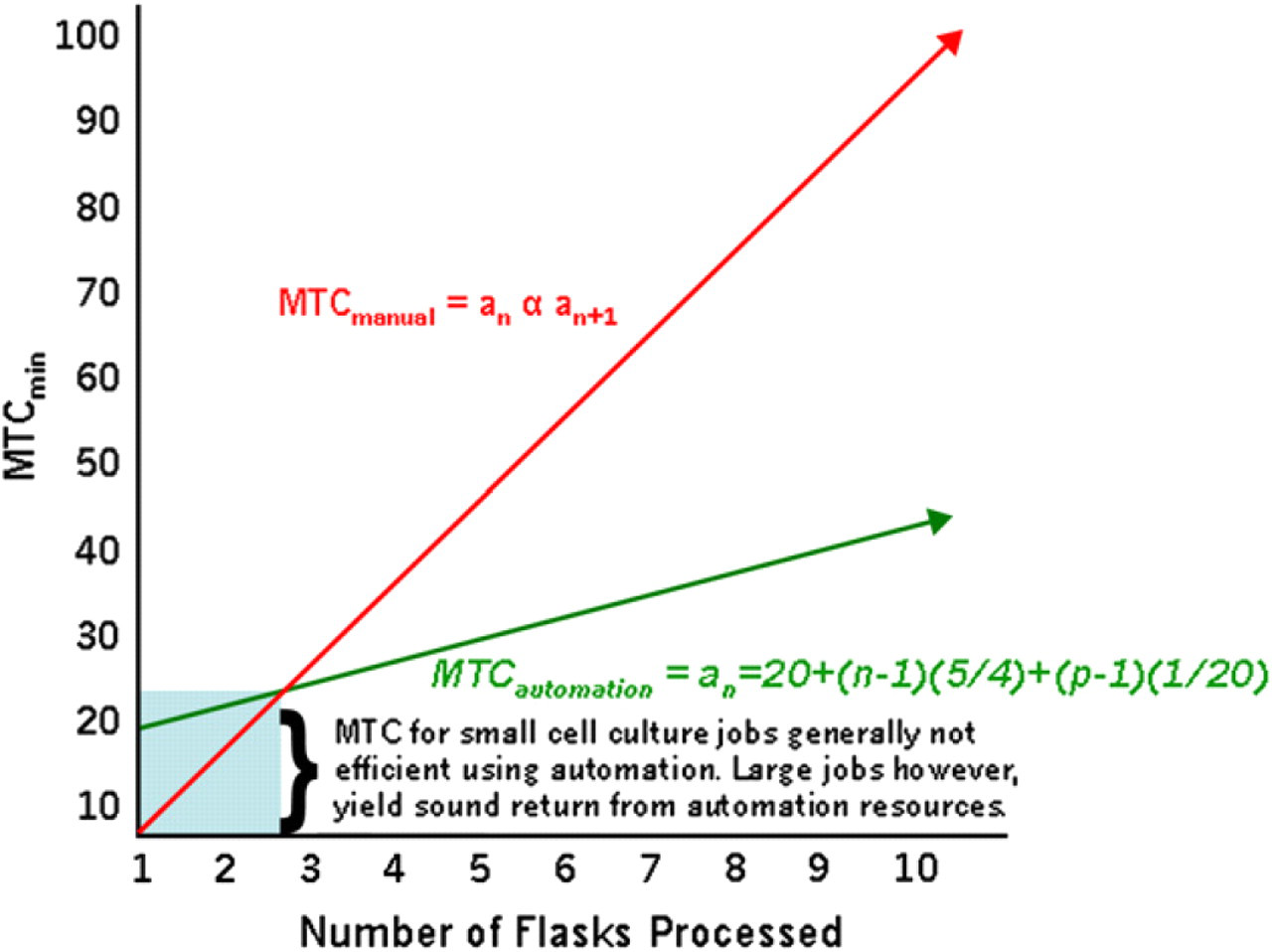
Manual Time Commitment- the human intervention required to secure a unit of output (i.e. cell plates for an assay), is greatly reduced as automated cell culture resources are fully exploited.
Though the constants for this formula vary somewhat between cell models based on the specifics of the cell line being used, optimal use of the automated system is attained when large numbers of cells and plates are needed from a comparatively few number of individual lines. When preparing small amounts of plates from multitudinous cell lines or cells cultured under various conditions, MTCautomation tends to be higher than MTCmanual, as indicated in the blue shaded area of Figure 3.
Robustness of the Cell Model
Interfacing Cells from Automation with Downstream Robotics
In an automated cell-culture environment, another part of the paradigm is reconciling an equitable balance between the archetypal cell model and robustness of the assay. Among other things, assay robustness refers to z scores, 14 day to day reproducibility of a strong signal window, and a long working window (the amount of time cells can remain in the plates without deteriorating). To the cell biologist working with cells in a scaled-up environment, “robustness” refers to the resiliency of the cell to withstand conditions of over confluence, media depletion, and repeated passaging. These cell-culture conditions can affect overall assay robustness. Robustness must be taken into account if a scientist on a cell-based assay platform wishes to “de-couple” the cell-culture automation from the assay automation. 15 As afore mentioned, cell lines, and particularly cells derived of tumor lineage, are living in a three-dimensional environment comprising exponential increases. Cell lines exhibit increases in mass, density, and media consumption over time, while confined to a finite environment. Our SelecT robot with the triple flask upgrade can hold 184 flasks for a combined growth area of 96,600 cm2 for adherent cells to proliferate. However, a single flask of HCT116th cells could be expanded to occupy this entire space in less than a week. If the cells are not used for whatever reason, machine resources must be used to cull and maintain cells, producing little valuable return on invested machine and consumable resources. For one particular cell model in our drug discovery portfolio, the cells needed to be maintained at strict subconfluent levels. The cells were sensitive to contact inhibition and they promptly and irreparably lost signal if they were over grown.
We could not accede to the whims of cell proliferation. These cells needed to be promptly harvested for use when 50% confluent or else split and passed for future use, taxing resources of both robot time and incubator space.
Striking a Balance between Exponential Cell Growth and a Finite System
Cells consume resources and generally grow exponentially while the supporting environment is of finite capacity. In laboratories studying stem cells, these progenitor cells must be constantly maintained under exacting conditions. Cell density must be monitored. Over density of cell cultures may challenge laboratories because if stem cells are allowed to clump together, thereby forming embryoid bodies, they begin to differentiate spontaneously. 16,17 They can form muscle, neuronal, or hepatic cell lineages. The National Institutes of Health indicates that spontaneous differentiation of embryoid bodies is reflective of general cell health. However, the capricious differentiation of an investigators' stem cells into muscle cells while using said stem cells to study a target in neurons for example, would not present a convincing neuronal cell model for say, safety profiling, compound screening, etc. This hypothetical example illustrates one example of spurious effects resulting from overcrowding and why the operator must be vigilant in assuring the system maintains cells in homeostasis.
The problem of containing exponential growth is exacerbated when using continuously growing cell lines (i.e., of tumor lineage). To avoid overcrowding-related perturbations to the cell model such as fluctuations in gene expression of a particular target, 18 a cell biologist in general must assay the cells during the narrow inflection point of log growth, or split cells back to remain subconfluent. Worse, over confluent cells may lead to undesirable gross cellular changes, depleted media resources, and death.
In vitro, mammalian cells succumb to similar population pressures, as do the human hosts from which they are derived. The automation platform, if viewed as an island, has a finite carrying capacity much as does an island nation of human dwellers. The 17th century population demographer Thomas Malthus outlined in his Essay on the Principles of Population, that human populations increase at an exponential rate, whereas the ability to increase food supply increases at geometric rate. 19 Eventually, a point is reached where the population outpaces food production (what was later referred to as reaching the “carrying capacity” of the land) for a given population. On exceeding carrying capacity, the unfortunate scenarios of famine, war, and disease were Malthus' predicted outcome. Thus, a population will check itself or colonize neighboring lands and secure necessary means to return said population below the carrying capacity. Cells dwelling in a confined environment are incessantly expanding toward the carrying capacity of a flask.
As earlier indicated, primary cells or stem cells must be maintained below the carrying capacity of the system to avoid over confluence—possibly leading to phenotypic changes and cell death. 20,21 Immortalized cells must also be used either for an assay, split below the flask carrying capacity at confluence, or alternatively expanded into more flasks. The later scenario can rapidly saturate system capacity.
We have found that approximately 50% of the SelecT programs we design and approximately 35% of robot time consumed is dedicated to the sole effort of harnessing exponential growth in a finite system. It is during log growth where cells are usually healthiest, 22 and best typify the archetypical cell model. To accommodate exponential cell growth, our initial practice was to repeatedly expand cells into new flasks then discard the flasks when cells were harvested. Such program design may be referred to as Boom-Bust design. “Boom” referring to the rapid expansion of new flasks, whereas the “Bust” cycle denotes culling of flasks due to confluence or for other use. The general paradigm was that robot flask number tended to follow the growth and dismissal of cells within our system. Boom-bust design tends to cause wide fluctuations in flask number. During the boom cycle, it may not be possible to expand new flasks without eliminating existing flasks first. To avoid crossing the carrying capacity of the machine while expanding new flasks, the operator is forced to harvest jobs or dispose of flasks that might be retained for other uses. This potential scheduling faux pas resulted in inconvenient or unacceptable capacity overrides. Flexibility was lost by forcing “bust” jobs to run first to make room for future flasks, whether the culled flasks were at optimal density or not.
Confining Log Growth to the Flask versus the Entire Automation Environment
Although the basic SelecT software was not set up for this, one technique we have used on our system has been to redesign our programs to reuse flasks. In theory, with flask reuse programming you can run the machine constantly at a point hovering just below the carrying capacity of the environment. Once the desired flask number is achieved, cells are removed for study or disposal, and the cells returned to the original flasks at an appropriate split ratio. There is very little fluctuation in total number of flasks. The number of flasks remains flat and the cells acquiesce in standard sigmoid growth within each flask until collected and reseeded. Reseeding confines the sigmoidal characteristics of cell growth to the environment within the flask itself. By detaching swings in flask number from the growth characteristics of the cells, a static flask number in the machine can be established. By abolishing the boom-bust paradigm, this permits easier scheduling, generates less waste, and reduces MTC (Fig. 4).
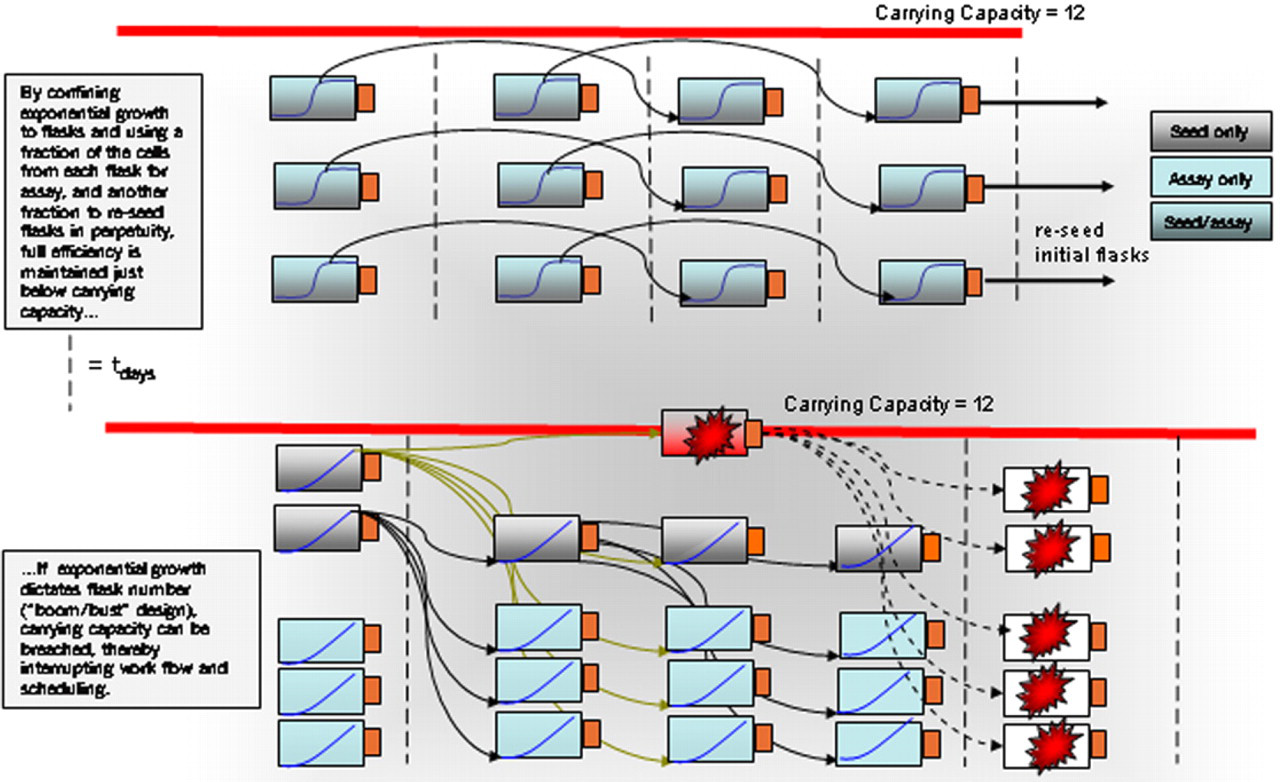
Linear versus Boom-Bust program design.
Decoupling Cell Kinetics from Downstream Assay Automation
As illustrated, the previous strategy can help economize on consumables and robot resources by confining log growth to the flasks.
However, the problem associated with exponential growth segueing into over confluence is not limited to the cell-culture flasks. Once the cells are transferred to the assay plate of choice, the plates are at the mercy of the downstream assay equipment. The phenomena of logarithmically expanding population will invariably lead to cell senescence in each well, for the exigencies of plated cells require prompt attention. Finite media and growth surface resources in the well will tax the cells, gradually closing the assay window. Any number of variables may arise with downstream assay automation or with the personnel scheduled to perform the assay. If a downstream assay platform crashes and the failed assay instrument cannot be repaired before cell senescence in the plate, the days cell-culture production is lost.
Several methods of decoupling cell-culture automation from cell-assay automation are available to the investigator. All of them regrettably, may compromise the cell model or thwart existing automation capabilities. Decoupling approaches expand the narrow cell plating to cell-assay window by slowing log growth or stopping growth altogether thereby allowing time for the offending instrumentation to be repaired. The strategy is to delay the growth trend toward carrying capacity in the plate as long as possible. We have investigated two decoupling methods. Both methods present benefits in respect to increasing assay robustness and detractions that compromise the cell model.
The trend in cell biology has been toward building better cell models. These include the use of 3D cell culture, 23 primary cells, and specialized biological coatings such as collagen on the plates. However, decoupling the SelecT cell automation from the downstream assay robotics to add robustness to the process also adds to the cell model.
One example of decoupling is placing cells in stasis. This can be performed by expanding cells with the robot, and bulk freezing them into a cryo-bank for future thawing and plating. 24 If the downstream assay equipment tests favorably, and all personnel are prepared, then the cells are thawed and plated. We have used this technique with cells supporting certain ion-channel programs. Bulk cryopreservation we found, though a suitable decoupler, yields two disadvantages to the automated HTS paradigm. The least of which is that the cryopreservation strategy on our system negates part of the automation fluidity inherent with automated cell culture and plating. This handicap is realized by the requirement of the operator to manually aliquot and slowly freeze cells, then later retrieve, rapidly thaw, and load into the machine on a daily basis. Though these inconveniences might be reconcilable with additional automation modifications. The other concern is preserving the ever-tenuous cell model. In some cell lines, cryopreserved cells will not recover from preservation to physiologic normalcy. No freezing process preserves 100% of the cells initially aliquoted. Many cells are typically lost during the cryopreservation process. Although recovered cell number can be indexed for assay optimization, the initial loss implies that the cells that do survive are perturbed. Surviving cells may exhibit an activation or deactivation of ubiquitous stress pathways (i.e. apoptosis related pathways), thereby making those cells more resistant to the rigors of preservation. 25
Cryopreservation methods have not changed much in 20 years. Cryopreservation protocols were not designed envisioning the effects that the freezing process might have on targets involved with for example, stress pathways. No single process works best for all cell models. The widely used cryopreservation agent DMSO has numerous toxic effects on cells. DMSO is known to activate heat shock protein (HSP 70) 26 and is also known to be toxic to hematopoetic stem cells. 27 Furthermore, cotreatment of several human myeloid cell lines (KG1, HL60, U937, and THP1) with DMSO was found to potentiate the cytotoxic and antiproliferative effects of Tumor Necrosis Factor (TNF). 28 Different targets and different cellular or histological models may be affected by cryopreservation with spurious results.
If cell recovery by cryopreservation is too laborious, alternative decoupling methodologies giving license to direct flask-to-plate convenience are available.
The second prophylactic approach decouples cell automation from assay automation by slowing or halting cell proliferation, thereby impeding over confluence in the plate. This strategy may delay phenotypic changes or senescence associated with over confluence by 1–2 days. Presumably, this approach would permit time allocation to repair a failed assay instrument. For example, delaying cell senescence can be achieved by irradiation of cells with a cesium source. 29 This procedure damages DNA thereby disrupting the cell cycle.
Other easily automatable methods we have demonstrated using the SelecT include thermal or chemical approaches. For example lowering the temperature in the cell incubator from a physiological 37 °C to 30 °C will inhibit Chinese hamster ovary (CHO) cell growth without observable stress to the cells. Chemical approaches to inhibiting cell division include the addition of genotoxic compounds (i.e., mitomycin C) 30 or tubulin depolymerizing compounds (i.e., vincristine sulfate salt) 31 to the cell media just before cell plating. These approaches will respectively disrupt the cell cycle by damaging DNA or by preventing spindle formation critical to cell division. These approaches preserve automation flexibility and can be programmed into the SelecT platform.
These methods work well in our own experiments, with control compounds preserving assay windows up to 72 h. However, mitomycin C can induce apoptosis via the Fas/FasL dependent pathway in cervical carcinoma cells. 32 The tubulin-targeting compound vincristine sulfate salt can activate Raf-1, phosphorylate blc-2 family proteins, and induce apoptosis. 33 Inadvertent exacerbation of these genes may present potential perturbations to the model.
Antagonism between the archetypal cell model and robustness gained from senescence-delay strategies should be weighed carefully in context with targets being investigated.
Maintaining Phenotypically Disparate Lines in the Automated Environment
Finally, to further augment the challenges facing the cell automation hub, we have found that the system needs to be malleable enough to accommodate multiple cell lines as robust cell models. In our laboratories, rare is there a point in the pipeline where only a single cell line is being maintained at any one time. In addition to the large screening campaigns being supported, sundry cell projects are supported for early toxicity profiling, electrophysiology, and radioligand binding assays. These cell lines are characteristic of many organ lineages representing multitudinous species. There are four tissue types in the human body comprising diverse cell lineages: epithelial tissue (i.e., hepatocytes, kidney cells, keratinocytes), connective tissue (i.e., adipocytes), muscle tissue, and nerve tissue (i.e., neurons and glial cells). A 12 sample list of organs comprising these tissues might include the circulatory, digestive, endocrine, excretory, integumentary, lymphatic, muscular, nervous, reproductive, respiratory, skeletal, and urinary system. In theory, cell lines representing tissue models of these 12 various organ systems could be easily accommodated on our SelecT providing that the requisite maintenance regimens are translatable into program code. Again, we have found it helpful to have familiarity with each cell lines' capriciousness and robustness before transferring them onto the system. At the level of sustaining multiple lineages, the “automation island” mimics a mammalian host providing the basic components for sustaining multiple tissue models programmed into the system. A recurring theme, though, is that the cells and cell-handling techniques currently need to be conformed to the system, and programs controlling the automation environment written for each cell line. Accommodating different growth dynamics and culture conditions in a static system requires considerable knowledge of the tissue models. Applying this knowledge to intertwining jobs appears to be key to successful deployment of an automated cell-culture platform. An off-the-shelf automation platform capable of handling all of the myriad permutations for cultured cell modeling simply does not exist yet. It is for this reason that it will remain difficult to design a system to completely automate and drive the biology.
The evolution of instruments capable of hosting multiple human tissue components is a remarkable engineering achievement. However, much basic in vitro cell biology is learned in small industrial or academic environments. The knowledge of experimental cell biology in these settings is often accrued in the absence of automated platforms. Here, small diverse quantities of cells are the norm and the underlying biology is learned before the automation. History presents a reliable pattern though. Smaller, more powerful and economical systems will invariably arise. The potential of such systems will be empowered to greater numbers of biologists. In time, tissue-modeling paradigms may be initially developed with these platforms in mind, abating the current challenges associated with forcing the existing biology into an automated system.
Returning to Tofflers' 800th lifetime model, it would not be until the last few minutes of the last 800th lifetime of the human experience that automated sustenance of organ tissues has been accomplished. As the recent seconds of the 800th lifetime accrue, we look to the developing possibilities of stem cells—our own progenitors, being further nurtured in vitro by automated systems. Although mammalian cells may endeavor to do as they please in vitro, in time many more optimized cell systems, which maximize robustness and the natural, physiological accuracy of various models will be established. This will further enable large-scale application of automated cell culture and automated assay capabilities.