Abstract
Selection of the optimum cell line from large populations can be time consuming and laborious. Compromises are often necessary to meet challenging timelines, but can limit the parameters or cell line construction strategies that can be evaluated. This article describes a new automated cell culture system for cell line selection and characterization. The technologies enable data to be obtained from large numbers of cells lines allowing users to evaluate, in parallel, a wider range of molecular approaches and cell culture processes. This facilitates more rapid and efficient cell line development. Also, presented are initial biological testing data for the new system and descriptions of how highly parallel processing can contribute to significantly reduced cycle times and better decision making through the rapid identification of optimum cell lines and culture conditions.
Keywords
Introduction
One major problem that affects both discovery and manufacturing in the pharmaceutical industry is the amount of time required to fully evaluate cell lines. Testing cell lines to determine optimum conditions for increasing biopharmaceutical protein production or culturing specific cell types for drug discovery research can be both costly and time consuming. For example, to determine optimum conditions for protein production, many parameters such as clone selection, media formulation, and feed regimes need to be investigated. Currently, most research for these evaluations with mammalian or yeast bioproduction cell lines is performed in shake flasks. 1 –3 Flask handling and sample processing are carried out manually and it is only the shaking incubators that are automated. This means that cell feeding, expansion, harvesting, and cell counting are all performed by laboratory staff. This is not only labor intensive but also requires good aseptic technique. Staff availability and their working hours can limit the numbers of experiments, or the numbers of cell lines that can be maintained in parallel and means that a skilled laboratory worker can process only 50 flasks per day. Additionally, with the traditional manual approach it is also difficult to ensure all cells are processed appropriately and consistently. This can lead to issues with the quality of the data obtained. When all these factors are considered, this can mean fewer clones are assessed or it can take longer to complete all the experiments associated with this stage of cell line development and therefore create a bottleneck in either downstream assays or manufacturing scale-up.
To overcome this issue, some experimental flow cytometric cell sorting methods are available 4,5 to predict which cells will have the desired properties; however, to identify the optimum growth conditions for these cells, manual culturing is still required. The only commercial method available for determining optimum culture conditions is the SimCell System (BioProcessors, Woburn, MA). The SimCell uses a miniaturized bioreactor in a multiplexed platform and is automated to allow users to test hundreds of different culture conditions simultaneously. It has been adopted by pharma and biotech companies for bioprocess optimization and by life sciences reagent suppliers for media development. This miniaturization and modeling method is a new approach to this problem and there are some data for its validity 6,7 with mammalian cell lines, but there is also evidence that scale-up is not linear due to bubble formation in the microbioreactors. 8 Another issue in using the SimCell system is the requirement for specialist equipment and labware.
In this article, we describe an alternative method for optimizing cell culture conditions using a new system called Sonata. Sonata automates all cell culture processes for mammalian and insect cells growing in shake flasks, enabling the processing of many flasks, cell lines, and experiments concurrently and can operate unattended, even overnight and at weekends. We have also included performance data, which show how the system compares to manual processing and illustrate how Sonata can improve productivity without compromising the quality of cells for specific downstream cell-based applications.
Experimental
System Description
Sonata (Fig. 1) automates all the processes that shake flasks containing mammalian or insect cells undergo. It enables tasks such as incubation, media feeding, cell counting, cell splitting, and supernatant harvesting to be carried out completely unattended. Users can program Sonata to retrieve specific shake flasks from the incubator at specified time points, automatically remove the cap, pipette out media, and replace it with fresh media. The system can also be programmed to harvest cells and automatically check cell density and viability before seeding new flasks or plates with specific cell numbers. The system tracks flask locations and identifies which flask to select, it correctly orientates and checks the unique barcode before processing the flask. Each flask can be processed with a set of experiment specific parameters including timings, media, and volumes required. The system software is capable of scheduling and performing multiple different experiments in parallel. Output can be either as a harvested cell suspension or specified cell numbers dispensed into 6- or 24-well plates.
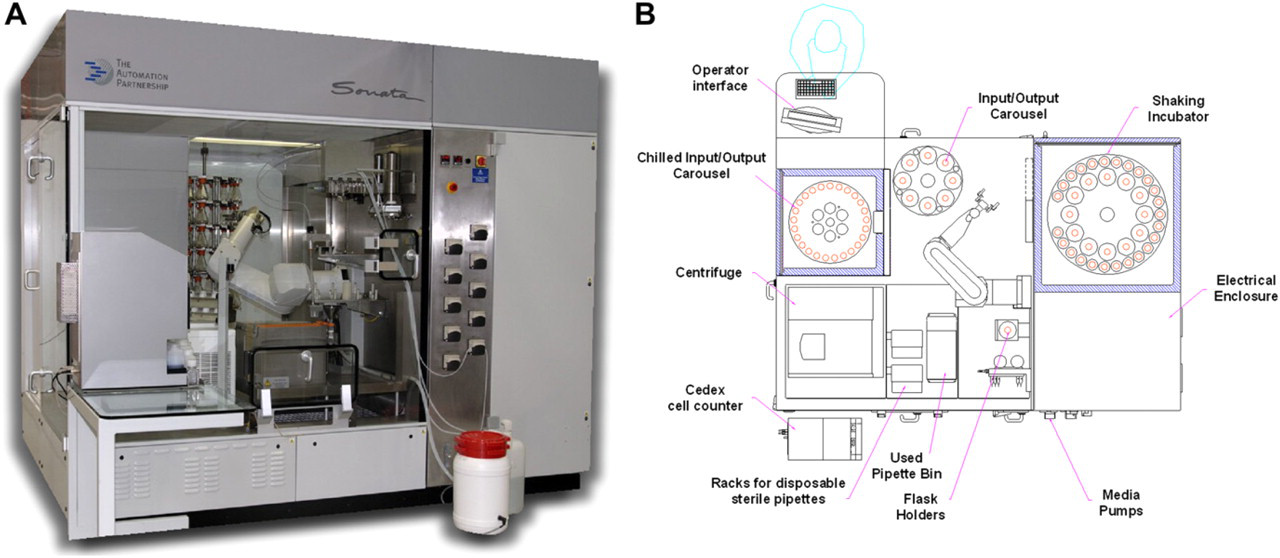

Sonata automated shake flask cell culture system. (A) The Sonata-automated cell culture system and (B) plan view showing layout of core modules.
To maintain aseptic processing, the system is contained within an interlocked high efficiency particulate absorbing (HEPA)-filtered enclosure maintained at negative pressure with a laminar down flow to remove aerosols. The liquid-handling module uses disposable sterile pipettes for labware-to-labware transfers and bulk media addition occurs by non-contact dispensing. These precautions eliminate cross contamination and provide operator protection. Should users wish to decontaminate the system, a vapor hydrogen peroxide procedure has been developed (BioQuell, UK).
System Design
Sonata is based around a six-axis RX60L Transfer Robot (Stäubli, Faverges, France) capable of moving labware between six processing modules (liquid handling, shaking incubator, input/output carousel, chilled input/output carousel, centrifuge, and cell counter). The hardware design builds on elements such as the flask processing and liquid handling that has already been successfully proven in The Automated Partnership's (TAP's) other cell-culture systems, SelecT and CompacT SelecT.
Core Modules
Sonata is fitted with a shaking incubator module comprising two concentric rotary axis units, which can hold combinations of 125, 250 mL, or 1 L shake flasks (Table 1). Flasks can shake continuously at 110 rpm (25 mm throw) and only stop during pick/place moves or when users add/remove labware. Additionally, this module can hold up to 40 stationary 6- or 24-well microtiter plates. There is also a single rotary axis input/output carousel to provide storage for new or spent flasks and a refrigerated (6 ± 2 °C) input/output carousel capable of storing up to 300; 50 mL centrifuge tubes and 24, 6- or 24-well plates.
Sonata capacity illustrations
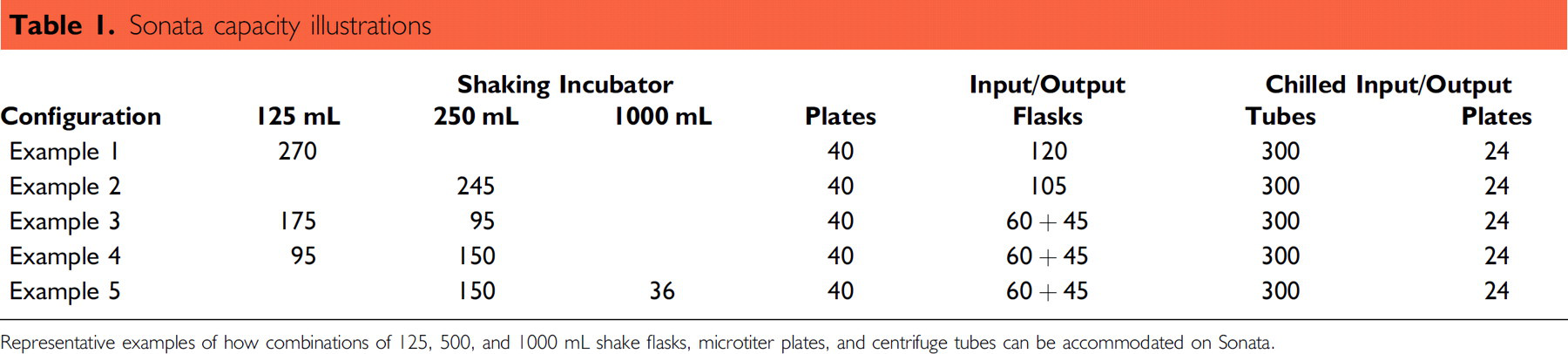
Representative examples of how combinations of 125, 500, and 1000 mL shake flasks, microtiter plates, and centrifuge tubes can be accommodated on Sonata.
A robot accessible Sigma 4K15 refrigerated centrifuge (Sigma, Germany) is integrated within the system and allows experiments to generate cell pellets and, in combination with the liquid-handling system, isolate cell supernatants. The centrifuge can process up to four 50 mL centrifuge tubes and can substitute “balance” tubes if an uneven number of tubes needs to be spun. Throughput of the centrifuge is maximized by interleaving the pre- and post-centrifuge processing steps while the centrifuge is running.
The liquid-handling module consists of three subsystems: bulk media addition using any combination of liquids dispensed from the 10 media pumps; labware-to-labware transfers with the pipetting head; and labware-to-labware transfers by pouring contents between flasks, centrifuge tubes, or a combination of both. Bulk media liquid handling is by non-contact dispensing and flask-to-flask pipette transfer is carried out using a clean disposable pipette for each flask or cell line.
Using the pipetting system, cell suspensions can be transferred to an integrated Cedex cell counter (Innovatis, Bielefeld, Germany), which can automatically perform a trypan blue exclusion assay. Cell counting uses the Cedex software and results are returned to Sonata where they can be used to dynamically drive further flask processing such as whether to passage the cells or return to the incubator to continue growing. Additional output ports will be incorporated into Sonata to allow for integration with analytical detectors to provide enhanced dynamic experimental processing and this is a future development project.
System Software
The robot and associated modules are controlled via bespoke software developed by TAP. The software has a user-friendly graphical interface allowing creation, planning, modification, and deletion of multiple experiments (Fig. 2). The user interface guides operators through the process of loading/unloading labware, as well as providing realtime feedback on experiments in progress. The software also contains a powerful scheduling engine that frequently reconsiders its forward workload according to any input received (e.g., cell viability data returned from the cell counter) and can delete, regenerate, and replan experiment as necessary. The scheduler uses an iterative approach to generation and planning that starts a few minutes into the future and extends further into the future with each round of processing. Using this approach, Sonata can drive its modules in parallel and execute tasks on multiple labware items while continually adapting its schedule. To assist operators in planning forward workload, a separate software tool can be used to download the current schedule and then model “what-if” scenarios to see what impact additional tasks may have on the system.
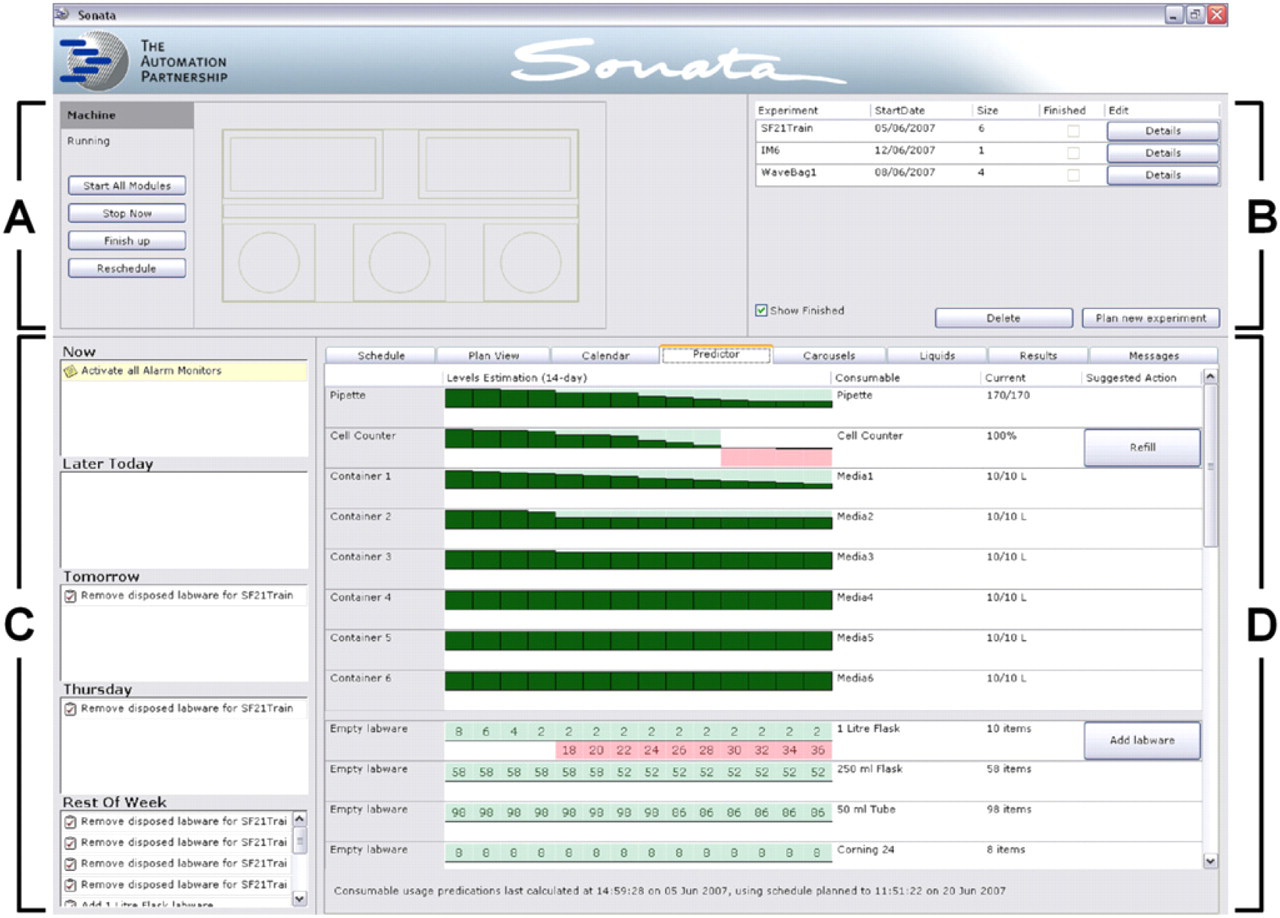
Sonata software graphical interface. The user interface is divided into four areas: (A) A simple representation of the Sonata system that shows the status of each subsystem in real time and buttons to start, stop, pause, and reschedule the system. (B) A record of all in-progress and completed experiments and buttons that quickly allow users to access and edit experimental parameters, create new experiments, or delete experiments no longer required. (C) A “to-do” list that displays the tasks required of the operators and when they need performing. (D) A tabbed interface that allows users to navigate between screens displaying short- and long-term scheduling, labware and consumable tracking (illustrated), labware locations within carousels, liquid and media management, results returned from integrated cell counters or metabolite analyzers, and messages reporting on system status.
System Application
To demonstrate that Sonata is capable of growing and maintaining cell cultures in a manner similar to a standard shaking incubator, replicate flasks of the insect cell lines Sf21 and T.ni were seeded (50 mLs; 1 × 106 cells/mL) into 250 mL disposable shake flasks containing SFM9000-II or ESF921 media, respectively, and sealed with gas-permeable filter caps (Corning, USA). Half of these flasks were then processed manually and the other half processed on Sonata.
Manually processed flasks were maintained at a constant temperature and shaking speed (27 °C, 110 rpm) in a Stuart SI500 orbital shaking incubator (Fisher Scientific, Loughborough, UK) and a viable cell count on each flask was carried out every 48 hours using a trypan blue exclusion assay on a Cedex cell counter. Cells were passaged into new flasks (50 mLs; 0.5 × 106 cell/mL) if the viable cell count was greater than 90%. Once viable cell counts reached a consistent level for three successive passages, cells were expanded into shake flasks (1 L) and the passage routine continued. Growth curves were generated by seeding multiple 250 rnL flasks (50 mLs; 0.5 × 106 cell/mL) and a viable cell count was performed every 24 hours for 7 days. The total time taken for the user to perform these actions was approximately 14 hours and required weekend working.
On Sonata, flasks (250 mL) containing Sf21 or T.ni cells were registered onto the system as was the new labware required for the duration of the experiment. This registration process involved creating association between labware and storage location by scanning the attached barcodes and assigning the labware to the desired experiment. Flasks were automatically counted and passaged using the same routine as the manually processed flasks until authorization from the operator allowed the flasks to be assigned to a maintenance experiment. This maintenance experiment automatically expanded the cells into flasks (1 L), which were then counted and passaged without further operator intervention. To generate growth curves, flasks (250 mL) were automatically created from the maintained flasks (1 L) and the cell counts were automatically performed on the integrated Cedex cell counter every 24 hours for 7 days. Total operator interaction time with the system for this process was approximately 1 hour.
To determine how Sonata performed during a typical cell-culture application, a Sf21 baculovirus inoculation, amplification, and harvest experiment were conducted. Briefly, 6- and 24-well microtiter plates were prepared with half the wells containing a baculovirus stock and the other half containing only sterile buffer. Flasks (250 mL) containing Sf21 cells were registered, maintained, and expanded into flasks (1 L) as described previously. An experiment was created whereby 48 flasks (250 mL) were automatically created from the maintained flasks (1 L) and inoculated with either baculovirus or underwent a “mock inoculation” with buffer alone. Each flask was maintained for 3 days before the contents were transferred to centrifuge tubes, spun (20 min, 2000 G), and the supernatant collected in a new tube. Both the cell pellet and supernatant were then returned to the chilled input/output carousel for off-line processing. Samples of the supernatant were sent for viral titer analysis (Oxford Expression Technology, Oxford, UK). In parallel to this, 6 flasks (250 mL) were created, 3 inoculated with baculovirus and 3 mock inoculated. Flasks were counted every 24 hours for 6 days to obtain growth curves for comparison. With the exception of initial/final labware handling and inputting experimental parameters, this entire experimental process was carried out without further operator intervention. Total user interaction time with the system was approximately 2 hours for an experiment that lasted 3 weeks.
Results and Discussion
System Application
The results obtained indicate that cells grown on Sonata show growth properties similar to those obtained when cell culture was performed manually (Fig. 3). Maximum viable cell count for the Sf21 cells was reached in 144 hours both with manual and automated culturing and cell viability was not significantly different. For the T.ni cell line, maximum viable cell count was achieved between 72 and 96 hours. Viability declined after this time with both manually and automatically grown cultures and is probably due to the cells exhausting their media or being sensitive to oxygen limitation, as has been reported in shake flask cultures of Escherichia coli. 9 During routine growth and maintenance, viability of cells grown in automated culture remained around 90% and cells retained a normal morphology.
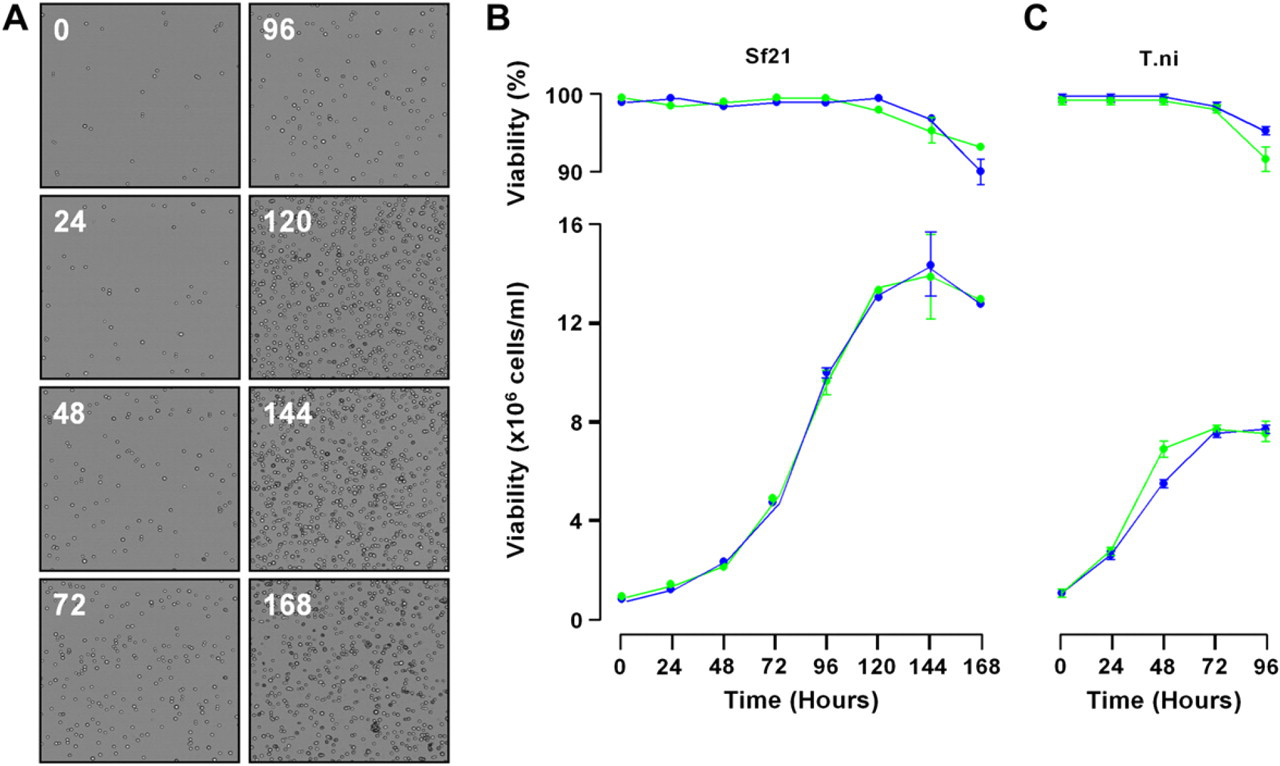
Comparison of manual and automated culture of insect cell lines. (A) Representative images captured by the integrated Cedex cell counter of Sf21 cells at various time points during growth. Growth curve comparison of (B) Sf21 and (C) T.ni insect cell lines showing cell viability (top) and viable cell count (bottom). Cells cultured on Sonata are shown in green and those cultured manually are shown in blue. Data shown are mean ± SD, n = 4.
In general, there is little difference between growth rates and cell viability between Sf21 and T.ni cells cultured on Sonata and those cells cultured manually and are within acceptable limits for cell culture and protein production. 10,11 As no other automated shake flask systems are discussed in the literature, the performance of Sonata cannot be compared to published data.
Sf21 cells cultured on Sonata and automatically inoculated with baculovirus showed a detectable change in cell viability, cell size, and growth rate (Fig. 4). Cell growth was severely affected with cell counts only reaching 4 × 106 cells/mL compared to 12 × 106 cells/mL with the mock-inoculated cells. At the same time as this change in growth rate, cell viability also decreased to 35% at 144 hours, whereas the mock-inoculated cells retained over 90% viability. Baculovirus inoculated cells also showed a marked increase in cell size compared to mock-inoculated cells. These changes in cell health correlated well with the levels of baculovirus expression seen after quantitative polymerase chain reaction (QPCR) analysis of cell supernatants created as part of the Sonata experiment. Flasks inoculated with baculovirus typically showed levels of virus, five orders of magnitude higher than mock-inoculated flasks. These results show that using Sonata, cells can be effectively cultured, expanded, and inoculated with baculovirus. Additionally, the amplified baculovirus is of high enough quality for use in subsequent inoculations.
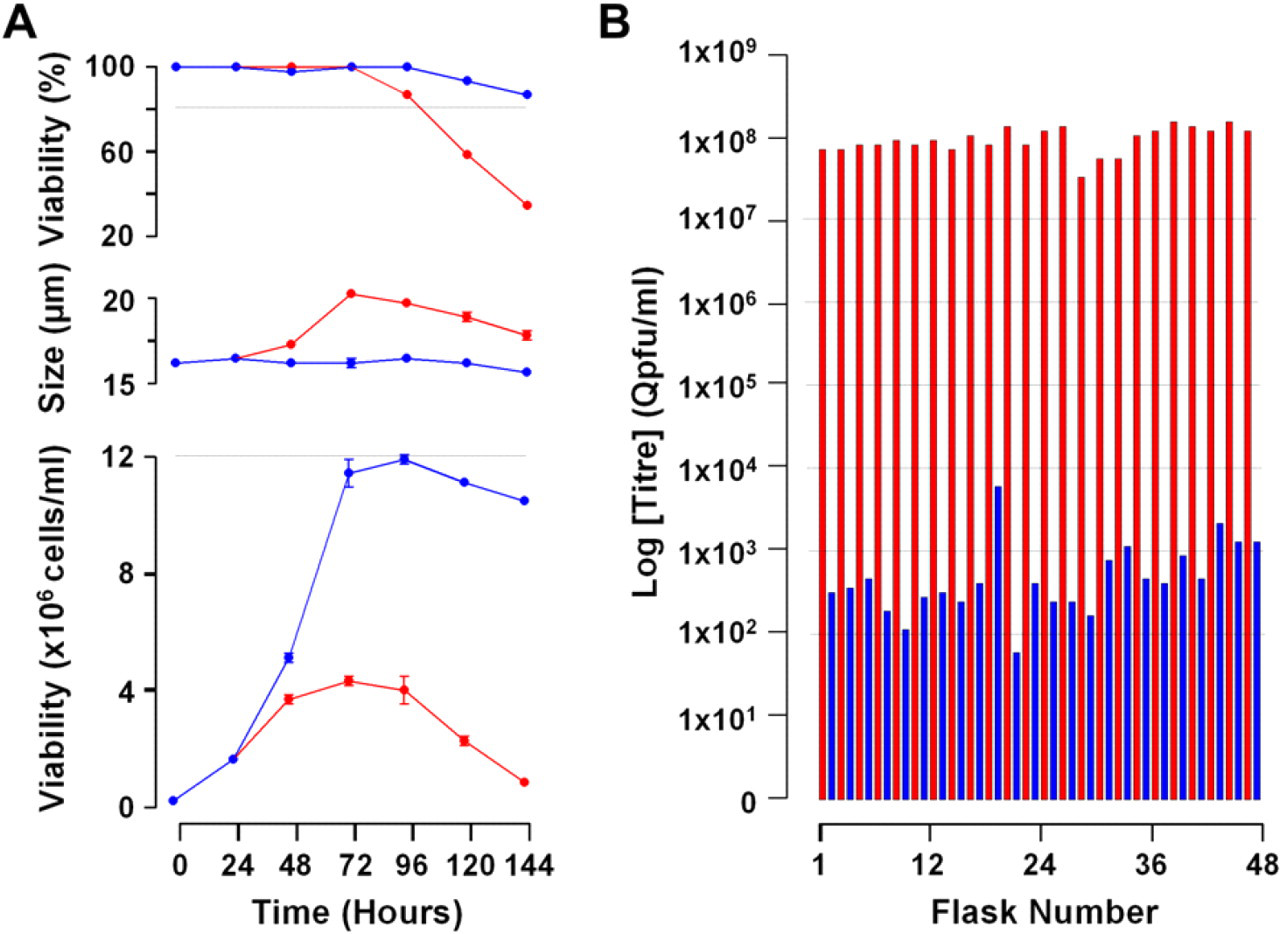
(A) Graphs showing cell viability (top), cell size (middle), and viable cell count (bottom) of Sf21 inoculated with baculovirus (shown in red) compared to Sf21 cells that underwent a mock inoculation (shown in blue) cultured on Sonata. Data shown are the mean ± SD, n = 3. (B) Baculovirus titer from the media of Sf21 cells inoculated with either baculovirus (red) or a mock inoculation (blue) showing several orders of magnitude difference between inoculated and mock-inoculated cells.
Conclusions
TAP's Sonata technology has been conceived to automate the manual shake flask process as closely as possible using the standard labware and reagents to maintain comparable consumable costs. It provides the potential for rapidly and simply optimizing cell growth and also supports applications such as viral inoculation.
A high throughput, parallel approach means that cell line development capacity is significantly increased and this stage ceases to be a bottleneck. As the Sonata system uses standard labware, there is also no need for major infrastructure changes or investment in expensive consumables. Additionally, because manual and automated results will be similar, scientists can make comparisons with their historical shake flask data.
The time taken for individual process steps is similar for manual and automated processing as the speed of operation is frequently constrained by the need to protect sample integrity and also ensure good aseptic technique. As Sonata produces similar results to manual shake flask methods it enables a single operator to manage a larger number of flasks. This makes the operator significantly more productive and makes a wider range of experimental designs possible (Table 2). Hence, the main efficiency gain with Sonata is its ability to operate unattended at any time during a 24-hour period and also over weekend and public holidays thereby reducing the Full Time Equivalent (FTE) requirement to complete a given series of experiments.
Comparison between manual and Sonata in a process development environment

On the basis of the performance data observed in culturing insect cells, it is possible to predict how the system could be used in a mammalian cell line development process. If the manual shake flask processes were transferred to the Sonata platform for a typical mammalian cell line development activity, the number of FTEs required to carry out the relevant process development activities would be reduced. It would also be expected that there would be a corresponding reduction in required lab space and other infrastructure.
In summary, Sonata can meet the growing need to supply cell-based reagents to support drug discovery, structural biology, and screening applications in drug discovery. Additionally, Sonata also meets the demand for cell culture optimization and characterization during development of biologics-based drugs and vaccines making it appropriate for the use as a tool for helping to scale-up bioprocesses where time to market and cost of goods are vital for producing profitable therapies.
Acknowledgments
The Automation Partnership would like to thank the DTI for supporting this collaborative program (funding reference TP/4/BIO/6/I/22287).
