Abstract
Sequencing of the human genome has opened the way and provided the impetus for building a comprehensive picture of a mammalian cell. Significant efforts are underway in the fields of genomics and proteomics to identify all genes and proteins in a given organism. The goal is a complete map of the genes, gene products, and their interaction networks in a functioning cell. The next step in establishing a comprehensive picture of a cell will be to integrate the cell's metabolome with the rapidly developing genomic and proteomic maps. A cell's metabolome, however, is such an enormous and complex entity that characterizing it can only be approached in sections. Our group of laboratories, the LIPID MAPS consortium, has focused on the lipid section of the metabolome. We have implemented a Lipid Metabolites and Pathways Strategy, termed LIPID MAPS, that applies a global integrated approach to the study of lipidomics in cells and tissues. This paper describes key aspects of the design, implementation, and accessibility features of a Laboratory Information Management System (LIMS) which serves the LIPID MAPS consortium. This software serves as a model system for integrating experimental information obtained by laboratories participating in metabolomics studies. (JALA 2007;12:230–8)
Introduction
The goal of LIPID MAPS (http://www.lipidmaps.org) is to identify and quantify all lipids and their metabolic pathways in a single cell type, mouse macrophage, and to extend such efforts to related and associated cells and tissues. The effort is supported by recognition that major effort is required in this area, where an understanding of lipid composition and changes in a carefully defined model living system is of vital importance. This knowledge is intended to lead to construction of a reference guide to direct future work in the field. The RAW 267.1 mouse macrophage-derived line of cultured cells 1 was chosen, among other reasons, because macrophages carry out many functions that are related to lipid metabolism and cell signaling, and are pertinent to many noted disease states, such as atherosclerosis, diabetes, and Alzheimer's disease. More recently, studies with primary macrophages derived from mice by the consortium have begun.
The member laboratories of the LIPID MAPS comprise approximately 16 institutions geographically dispersed across the United States. The laboratories span the domains of chemistry, biochemistry, biophysics, pharmacology, and cell biology. Each contributes expertise in, and familiarity with, particular classes of lipids, encompassing eicosanoids, fatty acids, sphingolipids, phospholipids and glycerophospholipids, structural lipids, neutral lipids (di- and tri-acylglycerols), and sterols. Each of these classes is sufficiently unique and complex that specialized domain knowledge has come be a near prerequisite for study of the class. 2 This breadth of experience is rare, and justifies a multi-laboratory, multi-institutional approach.
To accomplish transfer, centralized storage, and sharing of data among its members, we have developed a Laboratory Information Management System (LIMS) to submit data to a central database and to obtain data from the same source. A previous publication described an earlier form of the LIMS, used by the Alliance for Cell Signaling. 3 That earlier work involved a wide-ranging variety of experiments and procedures. The current paper describes a modified system intended for a more focused use in lipidomics.
Metabolomics studies often involve inducing perturbations to the ongoing state of living systems and subsequently monitoring changes at specific time points. 2 The strategy taken by LIPID MAPS is to subject cells and tissues to carefully delineated agents of stress, such as the active component of the bacterial toxin lipopolysaccharide, Kdo2-lipid A. 4 Such agents typically initiate or participate in inflammatory processes within organisms. The lipids within the cells, as well as those that may be released to the extracellular environment, are then extracted, identified, and quantified by mass spectrometry (MS) and associated methods at defined time points after initiation of treatment.
In support of these aims, the design of the LIPID MAPS LIMS is based on several core concepts. The first is that the amount of information necessary to place data in context is often greater than the actual value of the measured data. This enveloping information, recognized as experimental metadata, typically comprises systematically varied parameters. Metadata thus represent a highly important consideration in laboratory experimentation, in which conditions are held constant and only a single parameter of the metadata is changed. A second principle lies in recognition that data are often structurally diverse in nature. For example, data may simply consist of concentrations consisting of numbers and differing units. Alternatively, they may consist of mass spectra in differing digital formats. These data formats, in particular, are difficult to capture in electronic form for analysis, and may be better left to be manually analyzed and curated in a database for subsequent mining. Thus, the focus of the LIMS is on capturing essential metadata.
The association of metadata with data for purposes of electronic storage for LIPID MAPS is manually performed and is the task of the Bioinformatics section, situated within the San Diego Supercomputer Center (SDSC) and the Department of Bioengineering at the University of California, San Diego. Experimental measurements not entered into the LIMS are forwarded to the section by email messages.
Additional design principles embedded in the LIMS are described in the following sections.
Implementation Overview
The LIPID MAPS LIMS is written in the Java programming language and is downloaded as a single file from a secure website, requiring a username and password to access. The download is implemented as a Java Web Start program and installs automatically on a user's desktop computer. Web Start is a feature of the Java Runtime Environment (JRE) distributed by Sun Microsystems (Sun Microsystems Inc., Santa Clara, CA, USA). Once installed, the program is called from within the Web Start graphical user interface (GUI). The current version requires prior installation of JRE v. 1.4.2_07 or later version. Web Start checks the originating web site for availability of updates automatically, and subsequently provides updates, with minimum interaction required of the user.
The LIMS was designed on a Microsoft Windows platform and was tested using Windows and Apple Macintosh operating systems.
Each user must enter his user identifier (user ID) when using the LIMS. Database connection information needed by the application (URL, database username, and database password) is contained in a configuration text file installed by the user in his/her home directory. An identical but separate downloadable LIMS that accesses a test LIMS database containing simulated data is furnished to encourage testing by users who might feel intimidated when confronted with a complex new program, without fear of untoward consequences. This second LIMS uses a second set of connection information in the configuration file.
The user interface of the LIMS consists of a number of discrete GUIs representing modules of functionality that are called from a single main window interface by clicking the appropriate button (Fig. 1). The modules allow users to enter information and browse the LIMS database. After entering information, the user clicks a button to send information to a central Oracle database (Oracle Corporation, Redwood Shores, CA, USA).
Main form of the LIMS. Pressing any of the buttons (shown in cyan) causes the corresponding module to be created and displayed.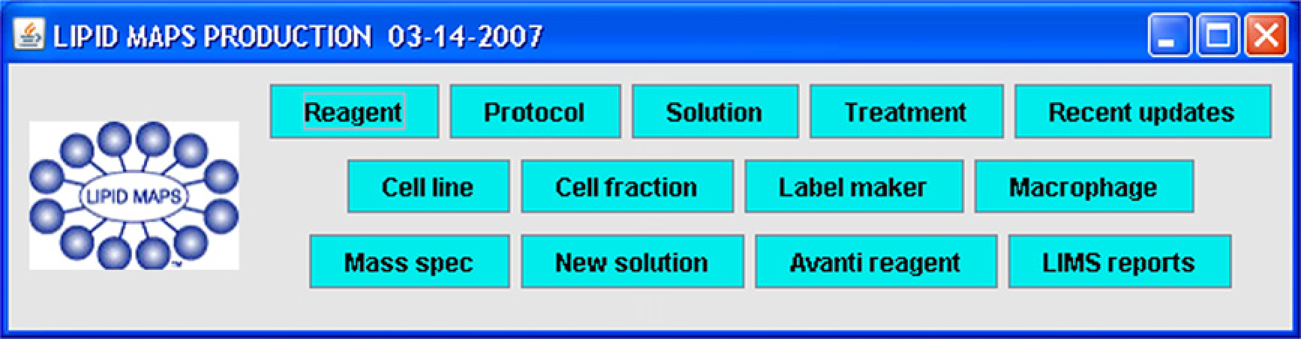
The LIMS also allows tracking of laboratory materials and protocols via printed labels that may be scanned into modules using barcode readers, thus minimizing typing errors.
Implementation Details
The LIPID MAPS LIMS is a two-tier client–server window form-based application which at its core enables entry and tracking of information pertinent to tracking of time-dependent changes in extracted metabolites. It is composed of 13 modules, selectable from a main form (Fig. 1). Packaging all modules together in this fashion enables integration of functionality; modules can thus avail themselves of views into the database provided by other modules, as discussed in the following paragraphs.
A conscious effort was undertaken to present a display that was distinct, uniform across modules, and as seamlessly connected between modules as possible. Details such as colors, button titles, tool tips, and placement of controls were considered. These are elements that are expected of modern, well-engineered interface design. Of particular note is the possibility of copying and pasting to and from desktop spreadsheet applications with all interfaces, increasing the comfort level of users who may be familiar with these generic applications. Each module contains detailed “Help” windows which explain the operations of a module, and its interactions with other modules. These features were included to foster the goal of achieving widespread user acceptance.
Updating of most data is allowed, with some restrictions as detailed below. In addition to easing usability, allowing updating is important from another standpoint. It is generally considered a “best practice” for LIMS to be used close to the point of the benchwork being performed. Introduction of a computer into a sterile environment may potentially cause problems for cell culture laboratories. The LIMS described here was purposefully engineered to allow obtaining identifiers and printing barcode labels in a single initial step before entry of cell passage information (such as cell counts) in a subsequent step.
Although preserving many features of the earlier Alliance for Cellular Signaling (AfCS) software, 3 the LIPID MAPS LIMS is organized around cellular treatments and MS experiments. The essential process of a simplified pattern of usage is shown in Figure 2. Each step in utilization ends typically with presentation of an identifier string, or ID, to the user. Entries can be made in other modules using these IDs. Entry of IDs can be accomplished by typing, by copying and pasting by means of keyboard commands, by scanning from barcode labels using a barcode reader, or lastly and most reliably, by selecting from lists.
Simplified sequential flow of LIMS usage. Rectangles represent modules of functionality as described in the text. Arrow labels indicate sets of one or more identifiers that are generated by the module at the base of the arrow and entered into the immediately after target module.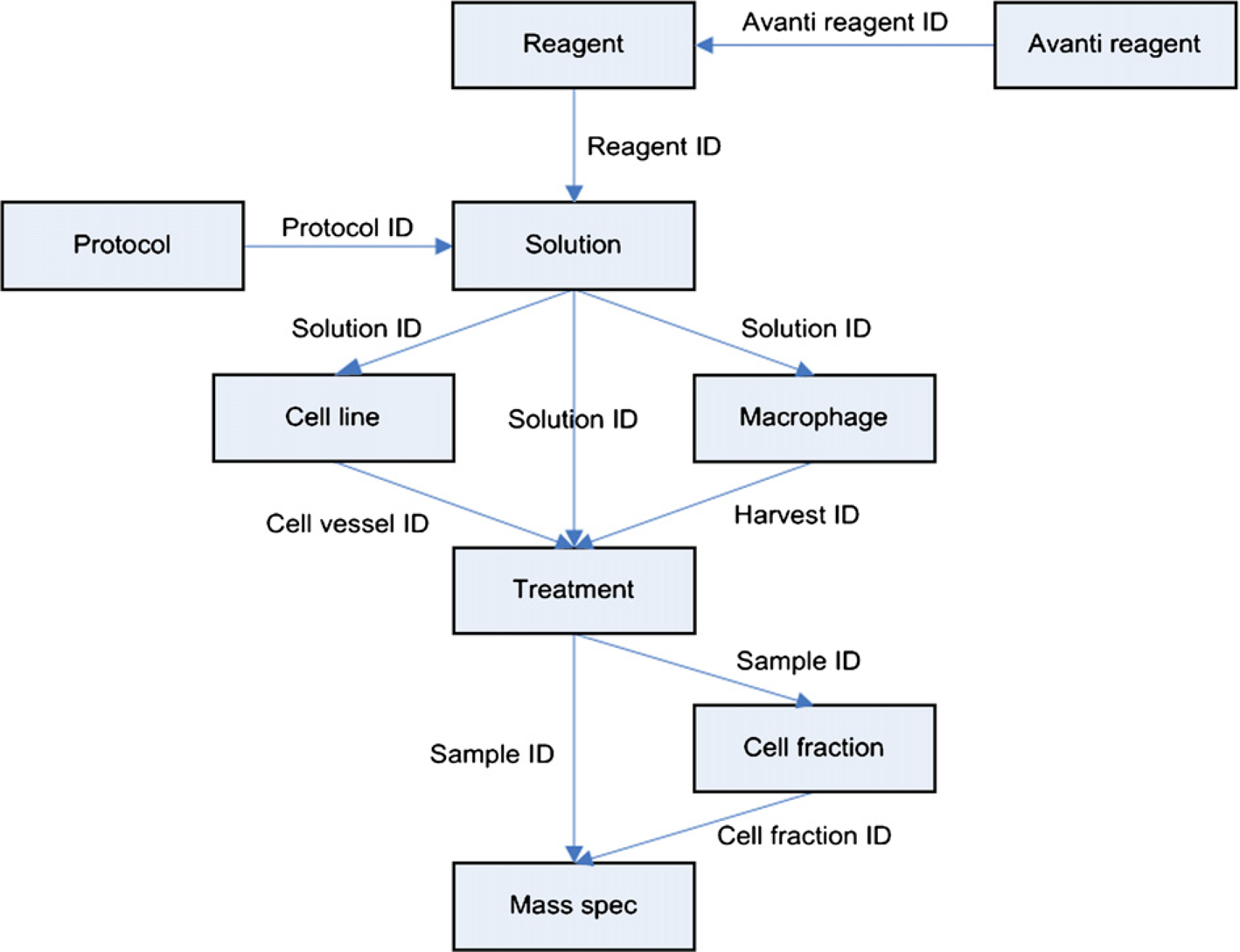
IDs are 12 or 13 characters long and are formatted differently by different modules. The first one to three characters are typically alphabetic, for example, “R” or “S” for entries made with the Reagent or Solution modules. A character designating the LIPID MAPS laboratory is usually included next. An eight-digit date is often added. Database sequence objects provide unique indexes that are also often used, particularly if the ID represents a primary key field for a database table. A final character (beginning with the letter “A”) provides up to 26 final ways of characterizing an ID, permitting, for example, up to 26 different experiments on a single day (Treatment module; see below).
A first step in using the software (Fig. 2) might begin with the Reagent module, where entry of information such as name, abbreviation, vendor, vendor's catalog number, date, and explanatory comments is performed. After saving information, a reagent ID is assigned and is displayed to the user. This ID can subsequently be used in the Solution module, where solution components are entered and a solution ID is assigned (Fig. 5 of Ref. 3). Solution IDs are used in turn in subsequent modules in the process flow.
Throughout the LIMS, solutions and reagents are checked for validity through structural database constraints and by client application code. For example, a check of reagent identity and stability is provided by the Avanti reagent module (Fig. 3). Avanti Polar Lipids (Alabaster, Alabama, USA) furnishes internal standards which are used to quantify the response of mass spectrometers to specific lipids or distributions of lipids. Users are not allowed to enter a particular reagent, or use an existing reagent ID, if the reagent, originally supplied by Avanti, is determined to be out of specification or to not possess an initial validation document. This issue is of great importance to the goal of lipid quantification.
Avanti reagent module. All table columns in the LIMS can be sorted alphabetically or by number or date, depending on the column data type, by clicking table column headers.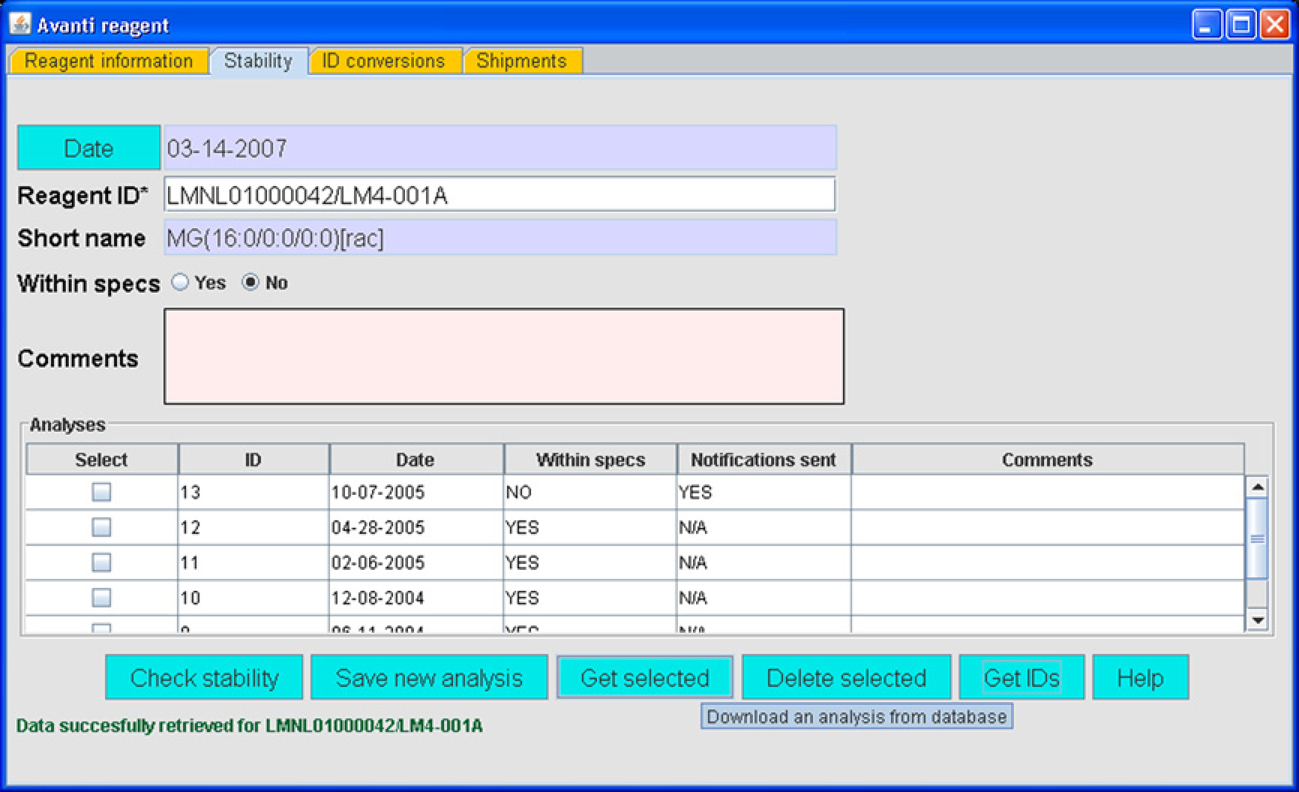
Avanti has sole responsibility for adding validation information, with the aid of a special username–password combination in the configuration file. All other laboratories have read-only access to Avanti-entered information.
Once a reagent has been declared out-of-specification in the LIMS database, an automated program (written in Java) running on an SDSC-based mainframe UNIX operating system periodically checks the relevant database tables. The program automatically emails all LIPID MAPS users who have been sent such reagents that the material is now outdated.
The LIMS enforces adherence to process controls in the form of exact control of experiments using strict solution and procedural protocols. A protocol ID is required by most modules. The protocol ID refers to a document in the LIMS database that describes a laboratory procedure or solution composition. For example, the dissolution of dry Kdo2-lipid A is described in a solution protocol, requiring use of phosphate-buffered saline and sonication under prescribed conditions. When entering solution information, the user must select a protocol by which solutions are prepared. The user may use one of the protocol documents that are already within the LIMS for this purpose. The LIMS database currently contains 52 current protocols. In addition, any of the participating LIPID MAPS laboratories may upload a new protocol and generate a new protocol ID. However, the Macrophage Biology laboratory (George Palade Laboratories for Cellular and Molecular Medicine, UCSD) has responsibility for the content of all protocols entailing cells and related materials. This oversight ensures uniformity of experiment design. Cell treatment protocols from the Macrophage Biology laboratory, once added to the LIMS, are modifiable only by laboratory members who possess a special username and password combination whose distribution is limited to that laboratory.
Another example of a constraint that must be addressed by the LIMS user is that a referenced protocol must ordinarily be current. Current and all obsolete (superseded) versions of protocols are available to users from the database. Generally, within any given span of time, the plans of the LIPID MAPS consortium require that all laboratories perform identical treatments. However, because analyses in some laboratories may require longer periods than analyses in other laboratories, the option to elect to continue to use an obsolete protocol is possible. If the user attempts to enter an obsolete ID, the user is informed that the protocol is obsolete and a current version is available. The user then has the choice of correcting the ID he/she has entered, or of explicitly opting to use the obsolete version of the protocol.
A list of solution and procedure protocols is available from any module by means of double-clicking on any surface not occupied by a control. The user may then select a protocol from the list that appears, after which the document will be downloaded and displayed by Adobe Reader (Adobe Systems Incorporated, San Jose, CA, USA) installed on the computer of the user. The list of documents available for downloading by the user includes a sample form describing animal experiments that is to be submitted to an institutional animal safety committee for its approval. The LIMS thus serves as a centralized document storage facility.
The Cell line module (Fig. 4) follows growth of cultured macrophage cell lines that are used as targets of treatments. Of particular interest when using the Cell line module is recognition that the characteristics of cultured cells change with time. This module tracks passage number and growth after thawing of frozen cells, thus ensuring that all laboratories perform treatments of cells within the span of a certain number of cell passages. The Macrophage Biology laboratory possesses and provides vials of RAW 267.1 cells frozen at a specific passage number to other LIPID MAPS laboratories for this purpose. The Cell line module provides a cell vessel ID that is used in the Treatment module (Fig. 6).
Cell line module. Text entry areas are coded white if an entry is required, pink if entry is optional, and blue or purple if the field is read-only. All buttons and combo boxes (drop-down lists) are cyan. A tool tip (i.e., “Usable only once”) providing a hint as to functional use appears if the cursor rests over a control or area for more than 2–3 s.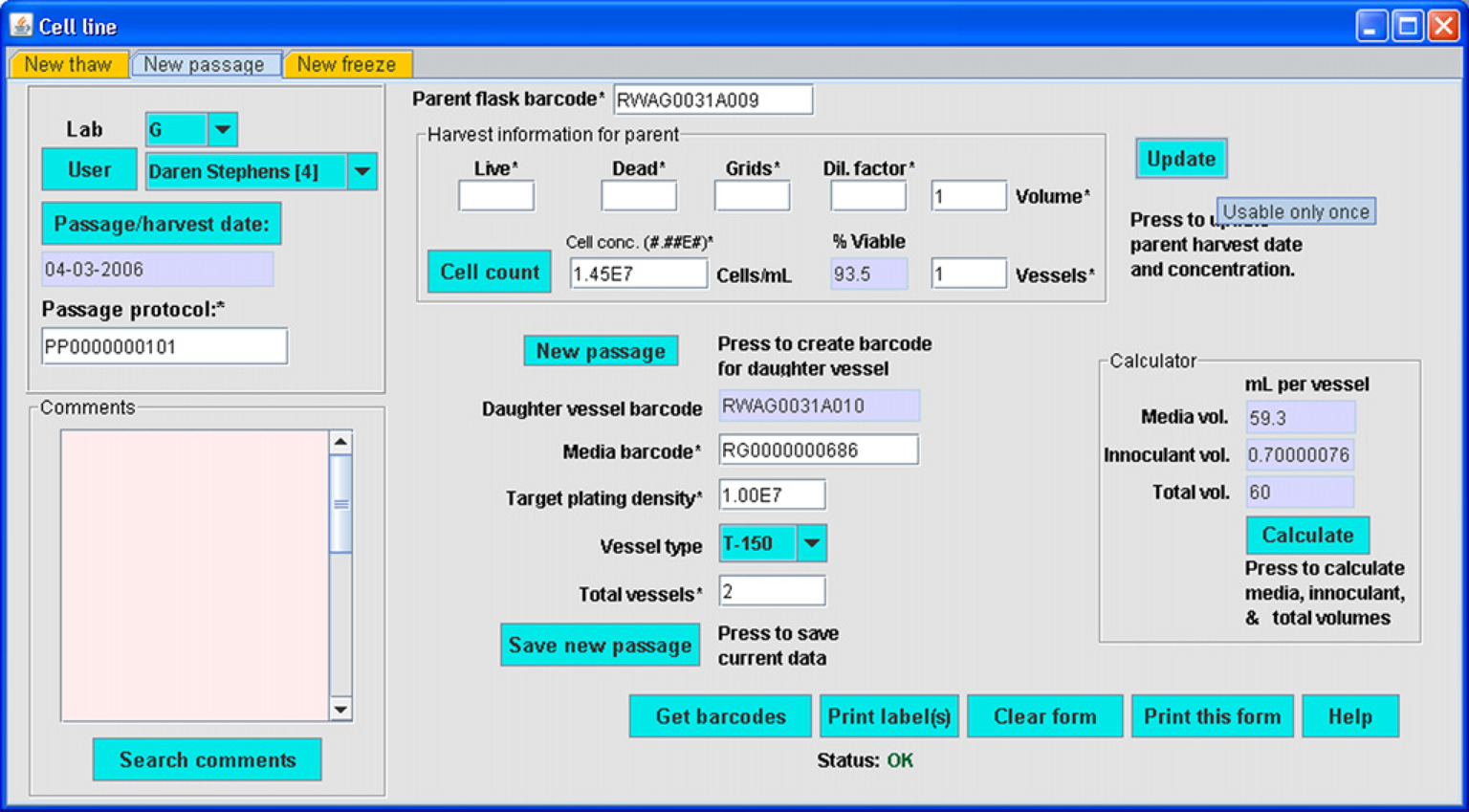
The Macrophage module (Fig. 5) is similar in concept to the Cell line module. However, passaging is not performed, as macrophages do not reproduce in vitro. They are obtained directly from a number of different organs in mice. This form allows entry of details relevant to mouse strain, sex, age, and cellular plating medium. This module (Fig. 5) will be additionally useful when mice containing genes with specific inactivated genes are used. As with the cell vessel ID, the harvest ID so obtained is entered into the Treatment module.
Macrophage module.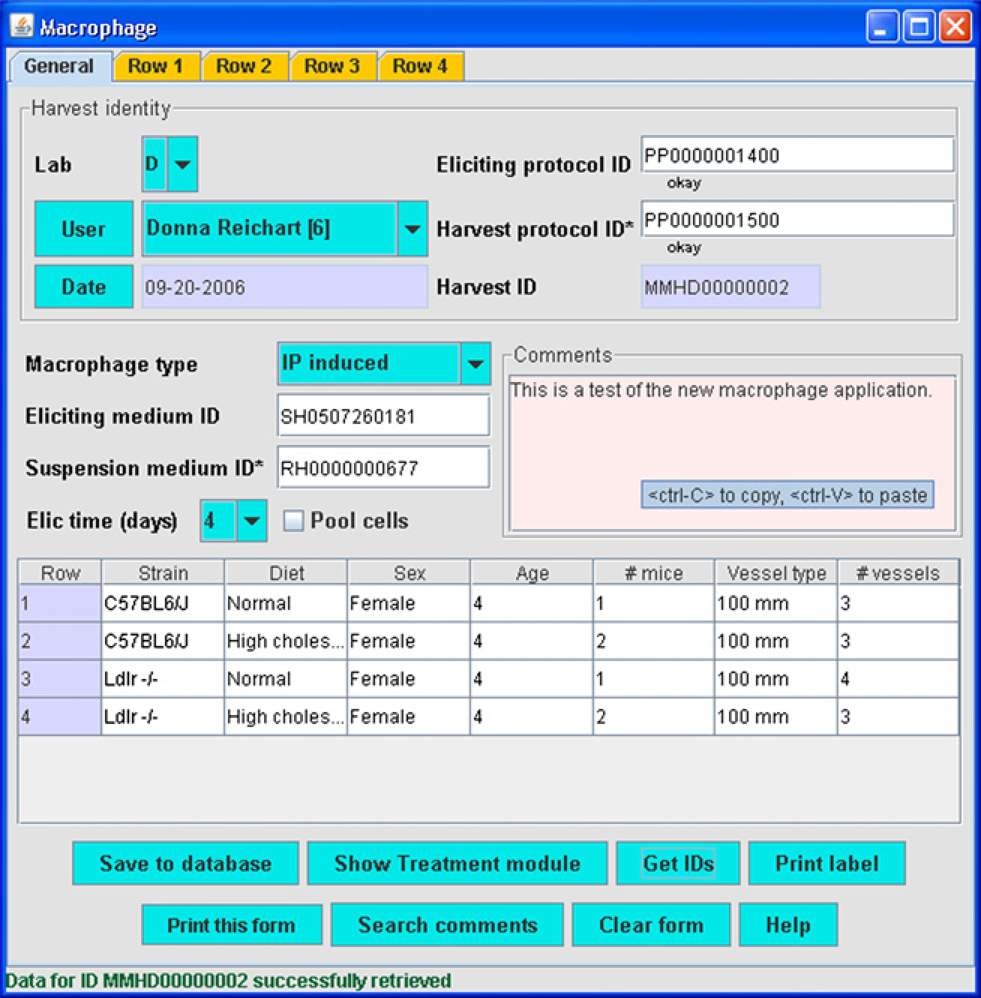
The Treatment module provides the essential lipidomics functionality of the LIMS (Fig. 6). Into this form, details of treatment conditions are entered. These include reagent or solution IDs, concentrations, and the start time, end time, and durations of both current treatment and pretreatment during an experiment with a particular cell preparation. These data are vital for studies of stimulus- and time-dependent alterations to lipid composition. The end result of use of the Treatment module is an overarching experiment ID associated with the experiment. Individual sample IDs are associated with cells receiving different treatments within an experiment.
Treatment module.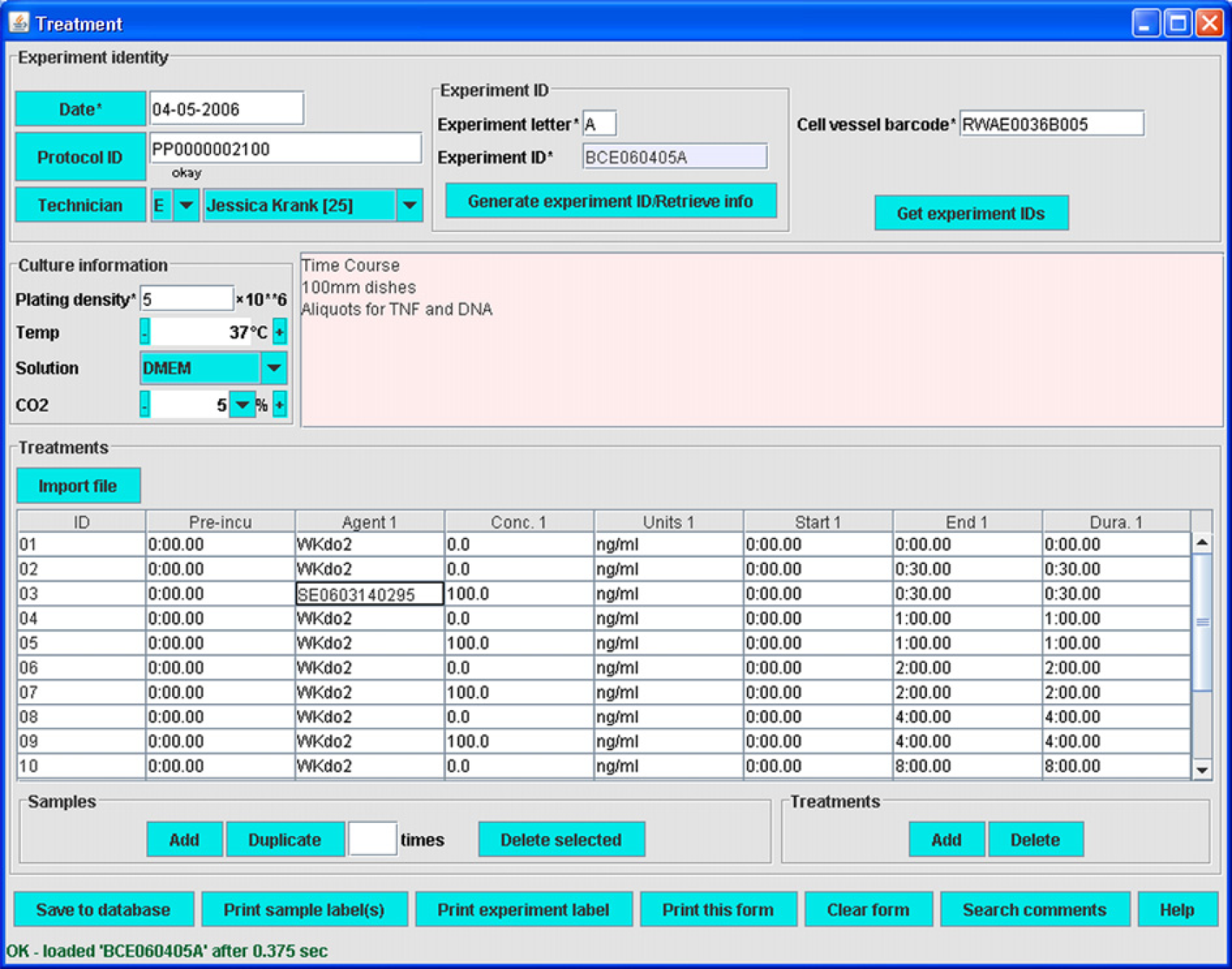
After treatment, cell samples may either be subjected to immediate disruption and lipid extraction and characterization, or be broken into fractions enriched in cellular organelles before analysis. The Cell fraction module (not shown) allows assignment of cell fraction IDs to these different preparations and thereby associates cell fractions with samples.
A “Mass spec” module allows selection of sample and cell fraction IDs to receive one or more mass spec IDs (MSIDs; Fig. 7). These MSIDs are used to relate experimental conditions such as MS acquisition and liquid chromatography conditions (
Mass spec module.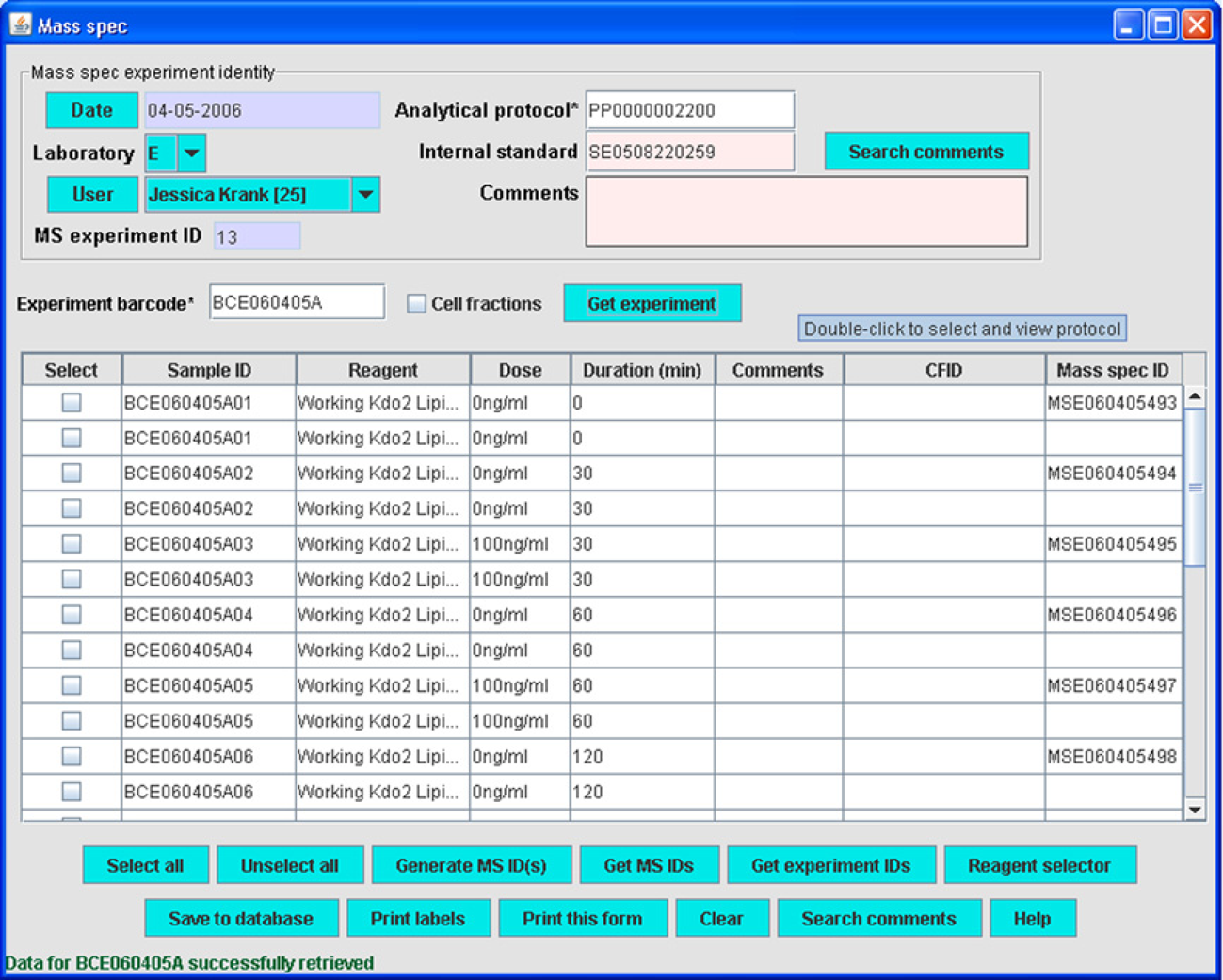
As mentioned previously, a significant contribution to the functionality in the LIMS arises from close integration of modules. Each module has search functions that search database tables for information entered by that module. Another implementation of searching and user interaction occurs in the case of the Reporter, or the LIMS Reports, module. The Reporter module allows the user to construct high-level reports summarizing overall database content using certain key parameters as search terms. For example, the user may obtain a summary table of cell vessel IDs that originate in a thaw of a particular vial of frozen cells used by a laboratory, along with the protocol ID that was used for thawing and passaging and the ID of any experiment in which a cell passage deriving from that vial was used (Fig. 8). The history of a cell line from freezer to experiment is thus obtained. Selecting a table cell containing a cell vessel ID, a protocol ID, or an experiment ID (Fig. 8) brings up the corresponding module, filled with information pertaining to the ID that was contained in the form when the data were submitted. This allows the user to construct a visual representation of database content, presented within the context of data entry. The same secondary module form is used to report on each subsequent cell selection. On other tabs of the Reporter entry form, the user may obtain reports on reagents, solutions, and protocols used in treatments, mass spectrometry experiments, and on macrophage preparations.
LIMS reporter (reporting tool) module.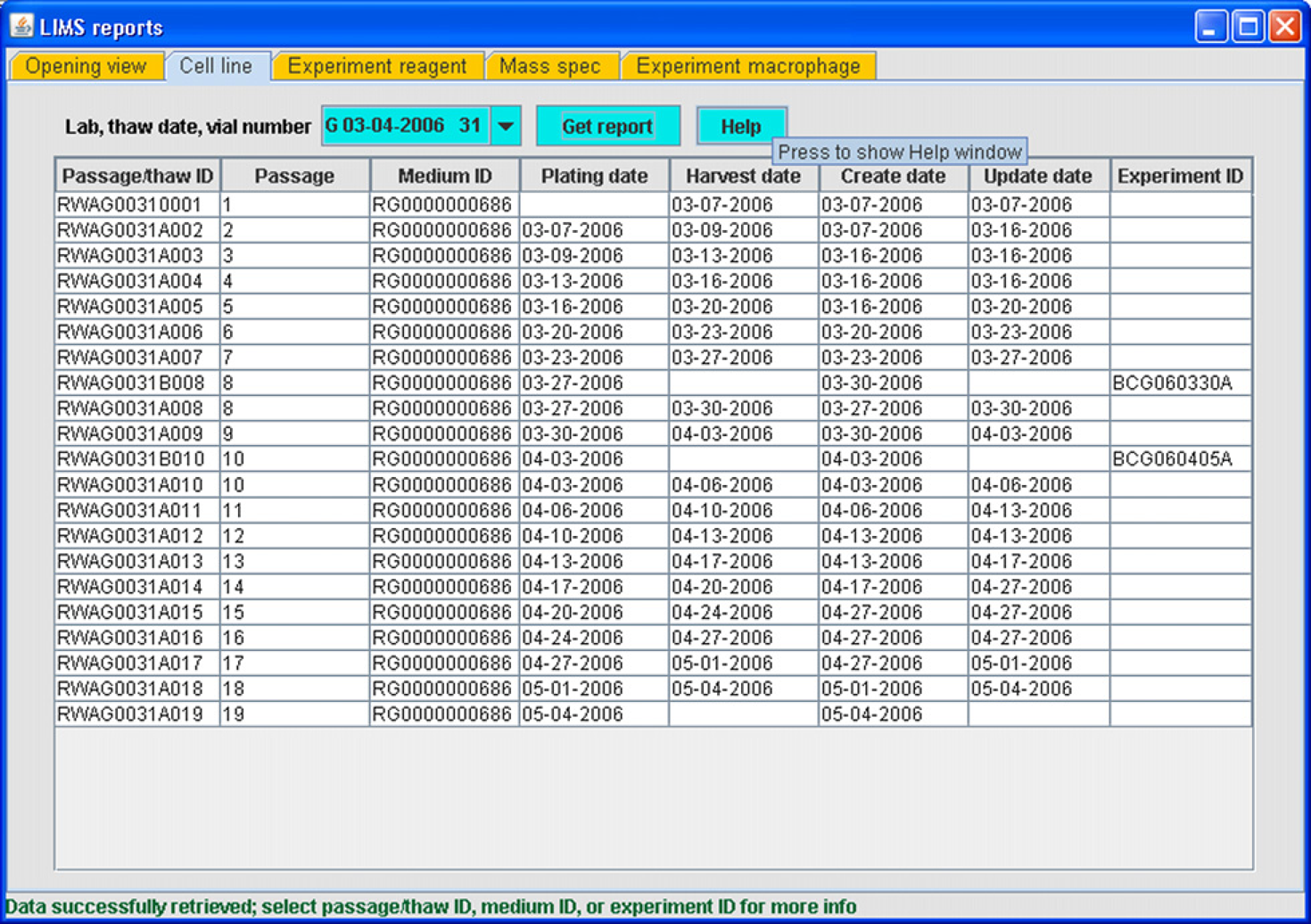
Updating is easily achieved in LIMS modules. Data entered into the LIMS can be accessed by means of the Reporter module, or by entering the corresponding ID into the correct module. Because the retrieved data are shown using the same forms as were used to input the data, users can easily update by typing into the fields that need to be modified and saving the data.
Summary
The LIPID MAPS LIMS is a specialized yet powerful data tracking system for lipidomics studies. It provides functions that encompass many of the goals of Systems Biology, spanning the intracellular, organellar, and extracellular milieus. Each LIMS module represents an idealized abstraction of an experimental step performed in a laboratory and records information pertinent for that step. The LIMS retains flexibility because of a careful selection of item tracking requirements, leaving entry of measured parameters into the database to be accomplished via other means. These individually tracked items and their usage are described generically or at length in referenced protocols, as desired.
The LIMS modules evolved over the span of several years. They rely extensively on human interaction for data input, rather than automated data transmission, which may comprise a large part of other LIMS systems. We believe that the modules have been developed to a state that they may be usable by a variety of laboratory personnel, not all of whom possess an extensive familiarity with computer-based data entry or an inherent understanding of databases. Several means to increase user acceptability in this regard were used and have been discussed in this report. We also found it important to provide prompt support in response to user questions and requests for additional functionality. Many of the enhancements described here resulted from this user feedback, which is a key element of the software development process.
Reporting standards for metabolomics experiments are currently under development.6, 7 Insofar as lipidomics is concerned, at a minimum, a lipidomics experiment must be annotated with descriptions of (1) the materials used, such as, reagents, solutions, and nature of animal nutrition; (2) the biological system; (3) conditions of treatment (experimental protocol), including duration; (4) quantities of lipids measured; (5) descriptions of procedures used for data analysis; and (6) a traceable path of association of these data, as can be provided by a relational database schema with tables linked by keys or identifiers (IDs). This may euphemistically be known as Minimal Information required for the Analysis of Lipidomics Experiments (MIALE).
The LIPID MAPS LIMS currently does not meet all of these requirements. The LIMS does, however, meet requirements 1, 2, 3, and 6, with additional features as noted previously in the text. Lipid measurements (requirement 4) are provided outside the LIMS as described below; data analysis (requirement 5) will be described elsewhere (to be submitted). The LIMS may thus serve as a reporting system suitable for internal use within an organization. The LIMS may meet the broader reporting standards of the general scientific community with this supplemented information.
Analysis and mining of the metadata and associated data obtained with the assistance of this LIMS is conducted offline at the Bioinformatics core. LIMS metadata and the experimental data described by the metadata are available on the Internet for browsing, and are directly linked to a public database of lipid structures that is curated by experts, 8 and to a database of proteins known to be involved in lipid metabolism in mice and in humans. 9 Both are accessed on our informatics server (http://www.lipidmaps.org; Fig. 9). The availability of solution and procedure protocols as well as tools allowing searching and drawing of lipid structures are also featured at this site.
Representative views of the LIPID MAPS web site.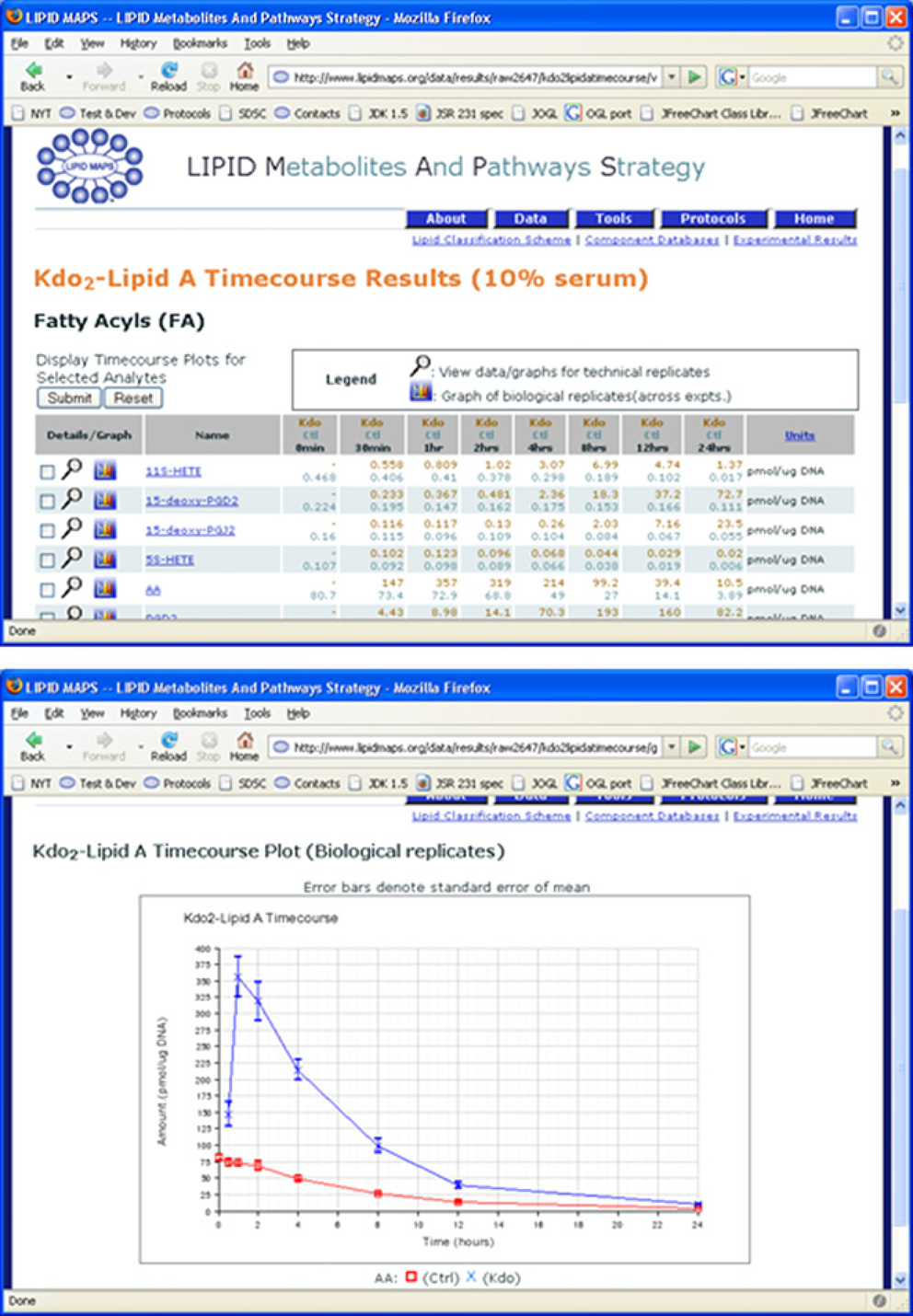
The goal of our web presence is to allow and encourage utilization of lipidomics tools by researchers on the large amounts of data being acquired. Currently (October 2006), time-dependent changes in the levels of 175 different lipid metabolites, defined at varying degrees of structural characterization, occurring in response to Kdo2-lipid A have been tracked in RAW 267.1 cells. With the aid of microarray data from Kdo2-lipid A-treated cells, 10 we are constructing networks of the interaction pathways in which these structures and associated proteins participate, to acquire a greater understanding of the processes occurring in the lipidome. A desktop network analysis tool with capability to directly download pathway data from LIPID MAPS databases is also being developed. These projects will be described at length elsewhere.
In summary, the LIMS described here can serve as a useful component of a management program for data generated by laboratories participating in studies of metabolomics.
Footnotes
Acknowledgments
The authors thank the individual members of the LIPID MAPS consortium for helpful suggestions during the development of the LIMS. This work was supported by National Institutes of Health (NIH) National Institute of General Medical Sciences (NIGMS) Glue Grant NIH/NIGMS Grant 1 U54 GM69558.
