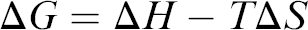Abstract
We report the extension of the microcantilever platform to study the thermal phase transition of biomolecules as they are heated. Microcantilever-based sensors directly translate changes in Gibbs free energy due to macromolecular interactions into mechanical responses. We observe surface stress changes in response to thermal dehybridization of double-stranded DNA oligonucleotides that are attached onto one side of a microcantilever. Once the cantilever is heated, the DNA undergoes a transition as the complementary strand melts, which results in changes in the cantilever deflection. This deflection is due to changes in the electrostatic, ionic, and hydration interaction forces between the remaining immobilized DNA strands. This new technique has allowed us to probe DNA melting dynamics and leads to a better understanding of the stability of DNA complexes on surfaces. (JALA 2006;11:222–6)
Introduction
The detection of conformational changes and hybridization of biomolecules such as DNA has great importance in biotechnology. Detection of biomolecules has traditionally been done using immunoassays, such as enzyme-linked immunosorbent assay or radio-immunoassays.1, 2 These methods use some form of label, such as a fluorescent tag, enzymegenerated chromophore, or radiolabel. Oftentimes, labeling imposes significant restrictions because it modifies the chemistry or structure of the molecule. Additionally, the labeling process increases reagent cost and involves multiple reaction steps, which increases assay time. For these reasons, label-free detection methods are highly desirable. In recent years, advances in the technology of microelectromechanical systems have been used to create label-free detection methods.
3
These devices rely on a transduction mechanism caused by molecular recognition to detect target analytes with high sensitivity and specificity. There is now a large effort being devoted to creating systems that require small volumes, rapid measurement, and the capability of detecting several compounds in a parallel format.

Micromechanical cantilever sensors are being used increasingly for label-free sensing of biomolecules. 4–6 Their extremely high surface-to-volume ratio allows detection of surface stresses that are too small to observe on a macroscale. Many groups have shown that processes like adsorption and desorption of molecules, and surface reorganization induce a stress that causes the cantilever to bend. 7 The cantilever thereby converts a biochemical signal into a mechanical one. One can follow surface processes by measuring the deflection of the cantilever tip. We have used this phenomenon to study phase transitions in macromolecules while scanning the sample temperature. In this article, we explore the role of the microcantilevers as more than a traditional detection device. We use the microcantilevers to provide additional information such as conformational changes like those identified by calorimetery. 8
To demonstrate the use of the microcantilever to detect thermally induced phase transitions, we have studied the melting of DNA molecules. There is a large motivation in understanding the properties of DNA bound to a surface because of the popularity of the gene chip.
9
Two strands of DNA are held together with the hydrogen bond. These bonds are weak in nature (approximately 5 kcal/mole).
10
DNA melting is the process of separating the two strands wound in a double helix into two single strands. Helix stability is the result of the interplay of two thermodynamic quantities: enthalpy, ΔH, and entropy, ΔS. Enthalpy represents the amount of heat released or absorbed during the reaction, while entropy relates to the orderliness of the system. Free energy change of the system depends on these two parameters in the following way:
The entropic change tends to destabilize the duplex, whereas the enthalpic contribution tries to stabilize the duplex. The stability of the DNA duplex is sensitive to the solution temperature. It is difficult to characterize the temperature dependence of surface-bound DNA because in this configuration, DNA has physical properties different from those of the bulk material due to interfacial interactions and 2-D confinement.
Materials and Methods
Fabrication of the microcantilever array has been described previously. 11 Briefly, a chemical vapor deposition of low-stress silicon nitride and thermal evaporation of chromium and gold layers is performed on a silicon wafer. Each microcantilever is a released silicon nitride beam that is 500 nm thick, 40 μm wide, and 200 μm long. At the end of the beam, a rigid paddle structure is fabricated to provide a flat surface for the optical detection of the beam's reflection. On one side of the beam, there is a 5-nm chromium layer, which is used to support the adhesion of a 25-nm thick gold layer. The gold layer provides a surface for attaching thiolated probe molecules.
A pyrex wafer containing wet-etched microfluidic chambers is bonded to the cantilever chip using a UV curing adhesive (Norland 121). The glass wafer has a 0.1-μm thick film of amorphous Si (A-Si) deposited in a low-pressure chemical vapor deposition (LPCVD) process. The A-Si is then patterned and used as the mask in the following HF etching process. The glass wafer is cut into nine chips, and individual glass chip is etched for 20 min in concentrate HF bath. The trenches that form during the HF etching process provide sufficient clearance for the initial and reaction-induced deflections of the cantilevers. The fabrication process is shown in Figure 1. After HF etching, A-Si mask is removed using silicon etchant to keep the glass cover transparent for future optical readout. The glass cover is bonded to the silicon chip using a “stamp and stick” method.
12
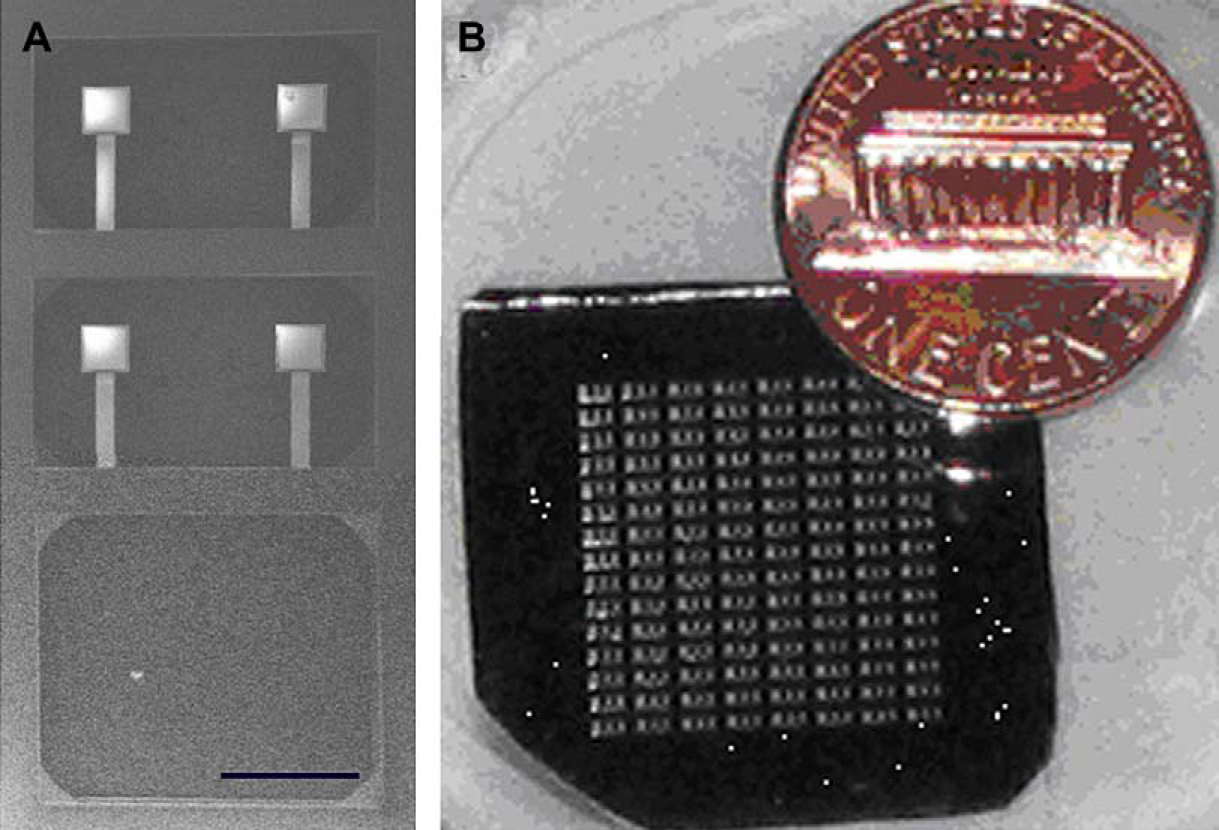
UV curing adhesive is first spin-coated onto a glass slide. The etched pyrex cover is carefully brought into contact with the adhesive-coated slide and then removed in a “stamping” process. During this process, a layer of adhesive is transferred to the pyrex cover. Next, the silicon chip and the pyrex chip are aligned and bonded, and then checked under the microscope to assess the degree of bonding. The 2 cm2 cantilever chip shown in Figure 2 is made up of a 15 × 8 array of individual fluid chambers, each of which contains four cantilevers.
Cantilever array fabrication process. Further details can be found in reference 11. (A) SEM image of SiN/Au microcantilevers. Scale bar: 200 μm. (B) Picture of cantilever array chip.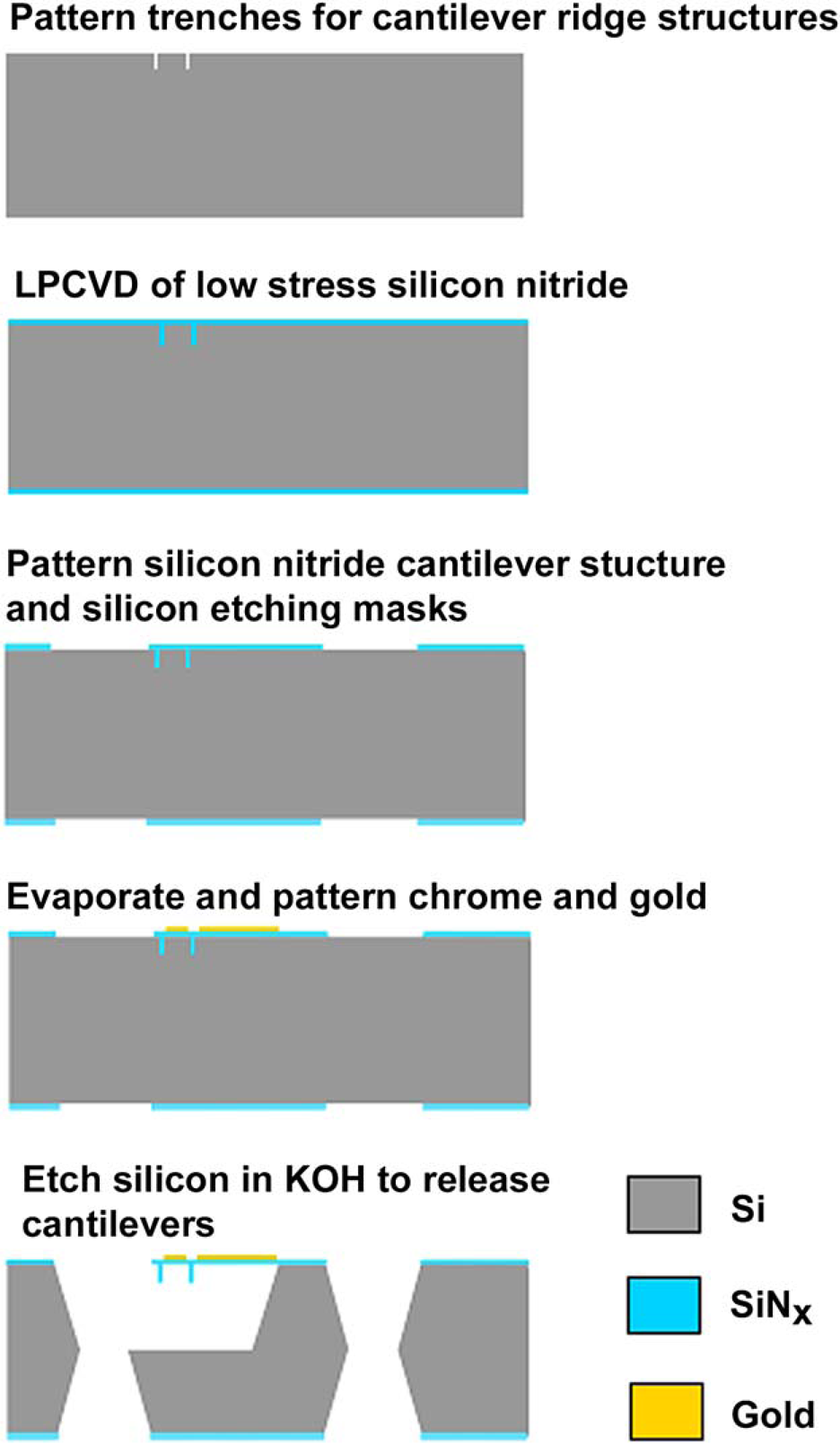

The cantilever array is imaged using an expanded helium–neon laser (632.8 nm, 15 mW) to illuminate the entire chip as it sits in a temperature-controlled holder as shown. The bending of a cantilever results in a change in angle of the light deflected from its paddle. This light then reflects from the beam splitter and passes through the lens at an angle different than that from the undeflected cantilever as shown in Figure 3A. The reflections from each of the cantilevers are captured onto a CCD camera chip (Apogee Inc, 3072 × 2048 pixels, 16 bit) and appear as spots on the CCD image, shown in Figure 3B. The frame capture rate is 1 frame every 10 s. A Matlab particle tracking program tracks each cantilever paddle spot in CCD pixels.
(A) Schematic of optical readout system used to detect cantilever detection. As the cantilever deflects, there is a shift in the reflection captured by the CCD. (B) CCD image of cantilever array. Each spot represents a single cantilever.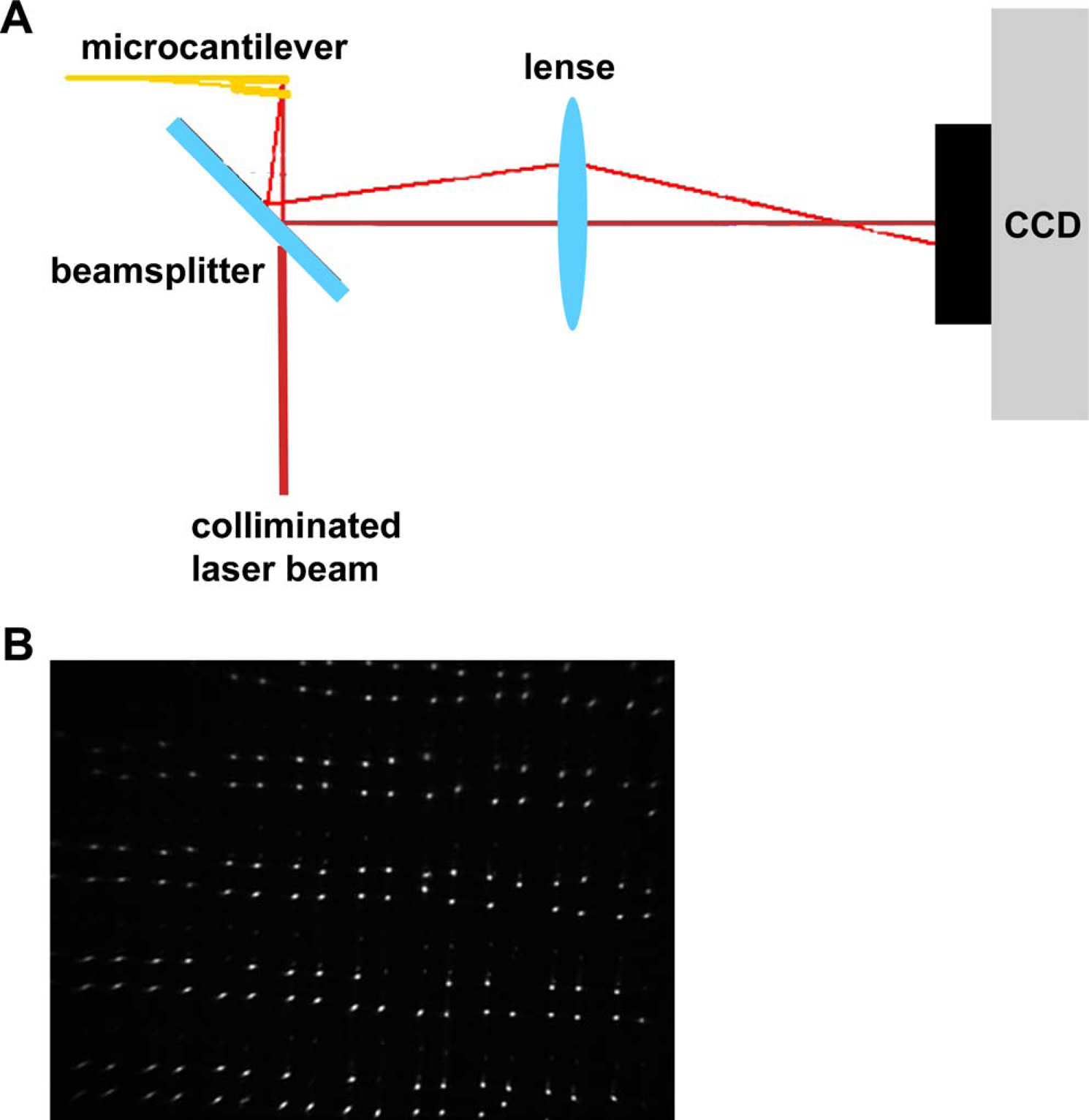
The microcantilevers will bend as the temperature is raised due to mismatch between the thermal expansion of the gold and silicon nitride layer. This temperature-dependent deflection is used to calibrate the optical readout system. The temperature of the chip is controlled and monitored using a thermocouple, which is placed on the glass side of the chip. The thermomechanical response of the cantilevers in the array is measured in units of CCD pixels/°C as shown in Figure 4. A white light interferometer (Veeco, NY) is used to measure the thermomechanical response of the cantilevers in nm/°C. As described previously,
11
this interferometry provides the calibration (pixel/nm) between motion of the spot on the CCD image and the cantilever deflection in nanometers. The thermomechanical response of the cantilevers is then used to calculate the stress caused by biomolecular reactions. Stoney's formula to relate the surface stress in the gold layer to the cantilever deflection is given in Eq. (1):
13
Thermomechanical response of cantilevers due to temperature changes measured in CCD pixels.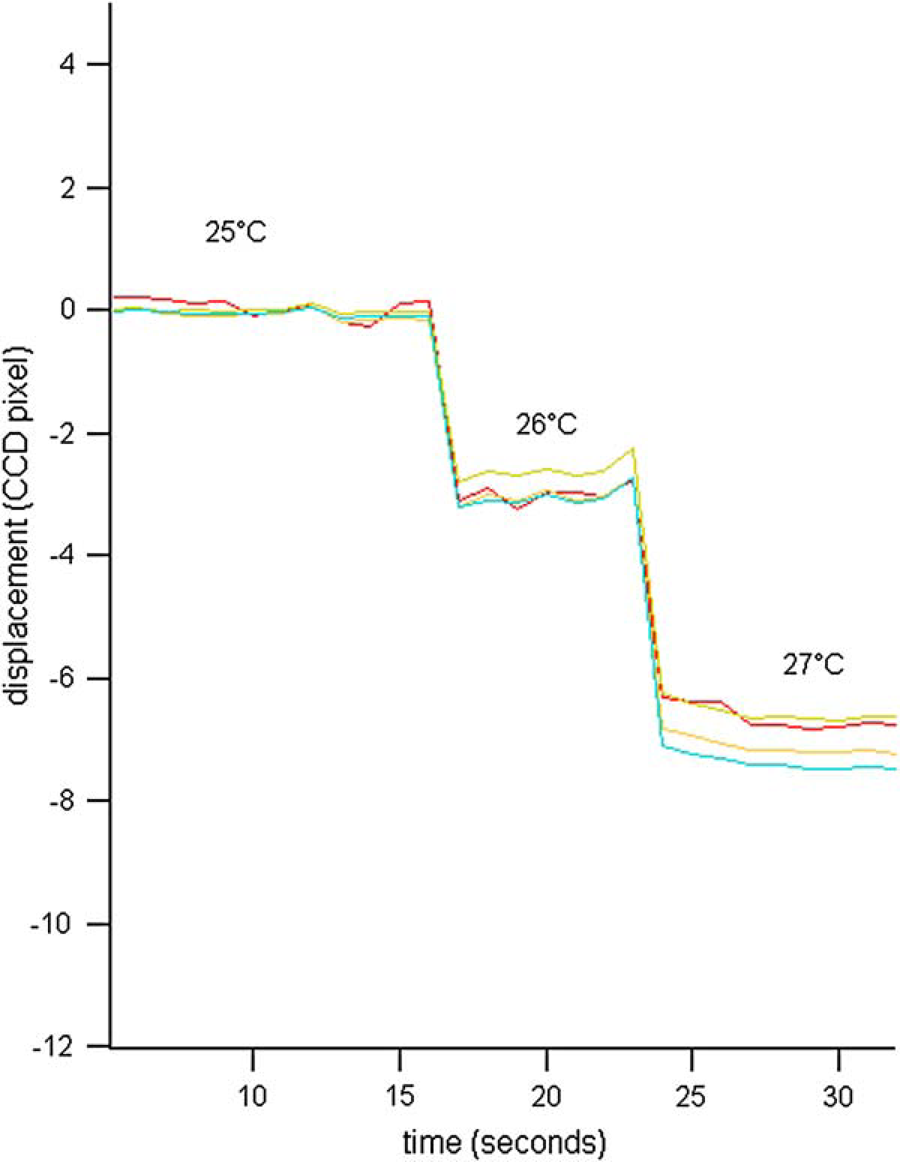
where Δγ is the change in surface stress, ν1 is Poisson's ratio of the thick layer, E1 is Young's modulus of the thick layer, t1 is the thickness of the gold layer, and L is the length of the cantilever. The measured thermomechanical sensitivity of the cantilevers is 208 nm/°C with a standard deviation of 14 nm/°C. 11 A 1°C change in temperature corresponds to a stress change of 25 mJ/m2.
To demonstrate the use of the microcantilever to detect thermally induced phase transitions in biomolecules, a model system based on synthetic oligonucleotides was used. A 20-base (5′-thiol-TAGATGAAGGTGAGAG-3′) oligonucleotide sequence was used as the immobilized target molecule. The thiol group attached to the 5′-end through a (CH2)6 linker allows for covalent attachment to the gold surface. To prevent nonspecific binding to the backside of the cantilever, a polyethylene glycol-silane coating is placed on the silicon nitride.
After immobilization, each well was rinsed with buffer to remove any unattached oligonucleotides. The wells were then injected with a complementary 20-base oligonucleotide sequence. As controls, a DNA-free buffer and a solution with a noncomplementary sequence were injected into different wells. The cantilever response was monitored for approximately 1 h to ensure successful hybridization had occurred. DNA melting curves were obtained by ramping up the temperature from 20 to 50 °C at a rate of approximately 1 °C/min.
Results and Discussion
As previously shown, cantilever deflection results from an increase in surface stress due to immobilization and hybridization of DNA strands to complementary DNA strands on the surface.11, 14 Cantilever deflection as a function of time for the hybridization of the complementary ssDNA to surface immobilized probes is shown in Figure 5. The matching sequence hybridizes and produces a change in surface stress of close to 20 mJ/m2, which results in bending of the cantilever.
Change in surface stress (mJ/m2) due to hybridization of ss20 DNA. When buffer alone is added, the cantilever does not deflect. When noncomplementary strands are added, there is a small change in surface stress, but when complementary strands are added, there is a large surface stress change.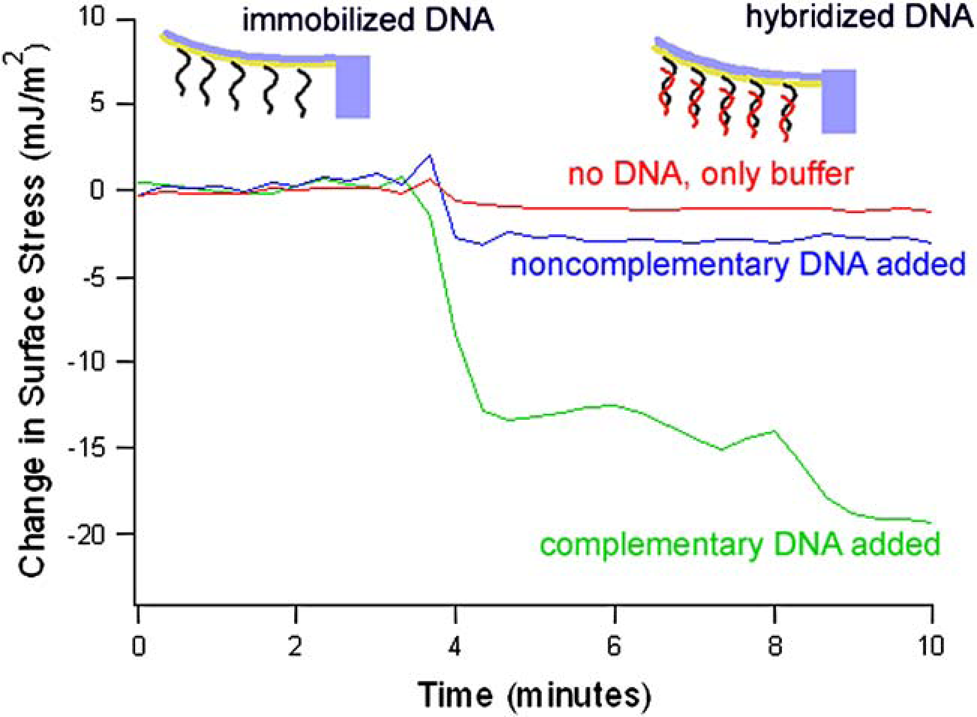
After hybridization, the cantilever temperature is ramped up to induce melting. Our experiments were performed in a 25 or 50 mM phosphate-buffered saline (PBS) buffer. Figure 6 displays a typical melting response. At 50 mM PBS, the otherwise linear deflection response shows an abrupt discontinuity at a temperature of T ≈ 47 °C. If we lower the salt concentration to 25 mM, we see a shift in the discontinuity to a lower temperature, T ≈ 44 °C. These measurements have a variation of 0.7 °C in the 20 cantilevers that were measured. DNA melting is often thought to be cooperative so that the AT-rich regions melt first, destabilizing the neighboring helical structure. As the temperature of the cantilever is raised, there is a conformational change of the DNA molecules, resulting in a change in the surface stress of the layer. It is interesting to note that the melting temperature observed on the microcantilevers is lower than the theoretical bulk melting temperatures, which was calculated using OligoAnalyzer 3.0 in SciTools (http://www.idtdna.com) at which we expect the double helix to separate. This result indicated that DNA is less stable on a surface compared to solution because of the difference between DNA dynamics in a bulk environment compared to dynamics of DNA near an interface.
Thermal scans of hybridized DNA. For a blank cantilever, the change in surface stress with respect to temperature is caused by the bimetallic effect. For the cantilevers with hybridized DNA attached, there are abrupt transitions as the DNA melts.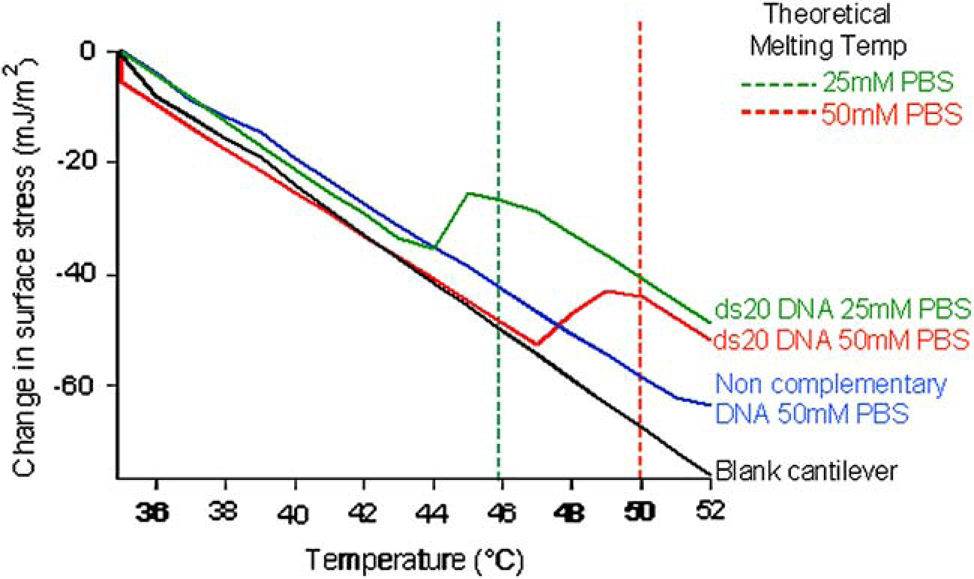
The melting properties of DNA are strongly dependent on two main variables: the salt concentration in the solution and base composition. It is well known that GC-rich DNA melts at a higher temperature than AT-rich DNA, because GC base pairs have three hydrogen bonds, while AT base pairs have only two. 15 Additionally, it is thought that DNA is stabilized at higher salt concentrations because the salt brings about a screening effect that minimizes electrostatic repulsion between the bases. It is less well known, and not well understood, that the stability decreases with even higher ionic strength. 16
We speculate that for DNA bound on a surface, the melting temperature is also dependent on the DNA density on the gold surface, which affects the interactions between molecules. In the bulk solution, interactions between neighboring strands are oftentimes insignificant. However, on a surface there are interactions between neighboring oligonucleotide strands that must be taken into account. Additionally, the attachment of the DNA to the surface may also destabilize the helix at the point of surface contact.
Conclusions
The stability of DNA under a variety of solution conditions can be studied in an automated format using a microcantilever array. Conformational changes due to differences in the lengths and intermolecular interactions of single- and double-stranded DNA are detected as variations in cantilever deflection. This new technique has allowed us to probe DNA melting dynamics associated with the stability of DNA complexes on surfaces.
Various technologies have been developed for biomolecular analysis in a high-throughput manner. Most of these detection methods are based on the detection of certain labels, fluorophores, enzymes, or radioisotopes, which are directly or indirectly attached to the analytes. A major disadvantage of these detection methods is the lengthy and costly labeling process. An easy label-free detection method compatible with microarray format is thus more attractive now. A cantilever array that enables multiple experiments to be performed simultaneously will expedite the experimental process, and thereby rapidly provide a large database to study the underlying physics.
Footnotes
Acknowledgment
This work is funded through the Center for Information Technology Research in the Interest of Society (CITRIS) by a grant from HP Labs.