Abstract
Simple sequence repeats (SSR) or simple sequence length polymorphisms (SSLP) are PCR-based molecular markers that have been widely used in genomic mapping and marker-assisted selection. SSR markers have made it possible to establish a high-density genetic map and evaluate genes of interest via tight association between markers and phenotypes. Large-scale markerassisted breeding studies in plants have created the need for a high-throughput system for genomic DNA preparation and analysis from thousands of samples in a segregating population. However, the standard methods for purifying DNA from plant tissues can be laborious, time-intensive, and not readily amenable to automation. Thus, an automated system for the rapid extraction and subsequent amplification and analysis of plant genomic DNA has been developed to facilitate high-throughput genome mapping and marker-assisted breeding studies. This system utilizes Sigma's Extract-N-Amp Plant PCR kit, a novel system for the rapid extraction and subsequent amplification of genomic DNA from plant tissues, and the maize SSR primer set. This extraction system eliminates time-consuming steps such as organic extractions and mechanical disruption, releasing sufficient genomic DNA from plant tissues for direct use in SSR marker analysis.
Introduction
Simple sequence repeats (SSR) are small segments of DNA, such as CAACAACAA, that are found randomly in almost every plant and animal genome. The number of repeats for each SSR marker often varies among different species and varieties, allowing for their use as molecular markers. SSR markers are normally detected by PCR amplifications with primers that flank sequences located at both ends of the specific sequence repeat. SSR primer sequences are conserved among different varieties; however, the length of SSR marker varies, resulting in polymorphisms among different varieties.
SSR markers have been widely used in genetic mapping, gene localization, and marker-assisted selection because of their simplicity and high rate of polymorphisms. 1 Applications utilizing SSR markers require a large number of samples for effective analysis. For example, to construct a fine genetic map or to identify a tight linkage between two genes of interest, hundreds or thousands of plant samples are often needed to reveal a rare recombination event. In marker-assisted selection studies, a large number of plants need to be screened in order to find the desired genotype. Because of these challenges, a high-throughput and automated process for DNA sample preparation and PCR amplification is required to process the large number of samples necessary for these types of applications.
In plants, genomic DNA extraction is the bottleneck associated with sample processing for marker-assisted selection and genetic mapping studies. Most existing methods for preparing DNA from plant tissues are time consuming, tedious, and labor intensive. Virtually all require mechanical disruption such as grinding in liquid nitrogen 2 or reciprocal shaking 3 to break the plant cell wall. Furthermore, many methods use multiple extraction steps with organic solvents such as phenol and chloroform, detergents, 2 salts, 4 and poly-vinylpyrrollidone (PVP) 4 to remove polysaccharides and poly-phenolic components that can inhibit enzymatic reactions. Clearly, none of these methods are amenable to automation.
In this study, an automated system for high-throughput marker-assisted breeding studies using the Sciclone ALH 3000 Liquid Handling Workstation is described. This system utilizes Sigma's Extract-N-Amp Plant PCR kit, which is a simple and rapid genomic DNA extraction method. This extraction system releases genomic DNA from leaf punches in a few minutes, and does not require freezing or mechanical disruption of leaf tissue. Without further cleanup, the extracted DNA is ready for direct use in PCR applications using the master mix supplied in the Extract-N-Amp Plant PCR kit. To demonstrate this system's utility in marker-assisted breeding studies, genotypes of 3 generations of maize using an SSR primer library were profiled.
Materials and Methods
Automated Method Description
Genomic plant DNA was extracted with the Extract-N-Amp Plant PCR kit (Sigma-Aldrich, St. Louis, MO) utilizing a validated procedure developed for the Sciclone ALH 3000 Workstation (Caliper Life Sciences, Hopkinton, MA). Prior to the start of the method, a 96-well PCR plate containing 0.5 to 0.7 cm disks of leaf tissue was placed onto the deck of the Sciclone. Genomic DNA was extracted by incubating leaf tissue in extraction solution at 85°C for 10 min, followed by neutralization. Extracts were then ready for PCR amplification without further cleanup. A 4 μL aliquot of the extract was combined with the 2x Extract-N-Amp PCR Ready Mix and 0.4 μM of the universal chloroplast primers (Sigma-Genosys, Woodlands, TX) for a total reaction volume of 20 μL. Amplification was performed using the GeneAmp PCR 9700 System (Applied Biosystems, Foster City, CA). The GenElute Plant Genomic DNA Miniprep Kit (Sigma-Aldrich, St. Louis, MO) was used to isolate genomic DNA from maize leaf for use as positive controls. An aliquot of 6 μL from each PCR reaction were analyzed on a 2% agarose gel.
Screening Maize Populations for SSR Marker Polymorphisms
All the maize populations in this study were obtained from the Department of Agronomy at the University of Missouri-Columbia (Columbia, MO). Two parental maize lines (P1: B73, P2: MO17) were crossed and DNA was extracted from the P1, P2, and the resulting F1 plants using the automated method for the Sciclone ALH 3000 Workstation. PCR reactions were set up using the SSR primer set (Sigma-Aldrich, St. Louis, MO). Polymorphisms occurring between the parental lines were observed via gel electrophoresis. F1 plants were then crossed and DNA was extracted from the 21 randomly selected plants of the resulting F2 generation. These plants were then screened with the selected SSR markers as identified from parental generations. Polymorphisms in the F2 generation were observed via gel electrophoresis.
Results and Discussion
Verification of the Automated Method for Extract-N-Amp Plant PCR Kit
To confirm the use of a high-throughput method to study marker-assisted breeding, a system was developed and evaluated for the performance of both the Extract-N-Amp Plant PCR kit and the Sciclone ALH 3000 Workstation. An automated method was developed on the Sciclone for the Extract-N-Amp Plant PCR kit and was used to extract genomic DNA from 88 maize leaf samples. Post extraction, PCR reactions were subsequently set up to amplify a fragment of the universal chloroplast gene in all 88 DNA extracts, as well as genomic DNA controls. As shown in Figure Figure 1, amplification of the targeted genomic DNA region was consistent and specific across the 88 maize DNA extracts. The template DNA in the extracts was of sufficient quality to allow for efficient PCR amplification. To assess for potential cross-contamination resulting from automated processing on a liquid handler, blanks were placed into alternating wells of a 96-well plate containing the maize leaf punches. The entire plate was processed and amplification was performed, followed by analysis on an agarose gel. The results shown in Figure Figure 2 demonstrate that no PCR product was detected in the blank wells. Additionally, the Extract-N-Amp Plant PCR kit can be used with a variety of plant types. In Figure Figure 3, the automated method was successfully used to extract and amplify target DNA sequences from a variety of leaf samples such as soybean, tobacco, and tomato.
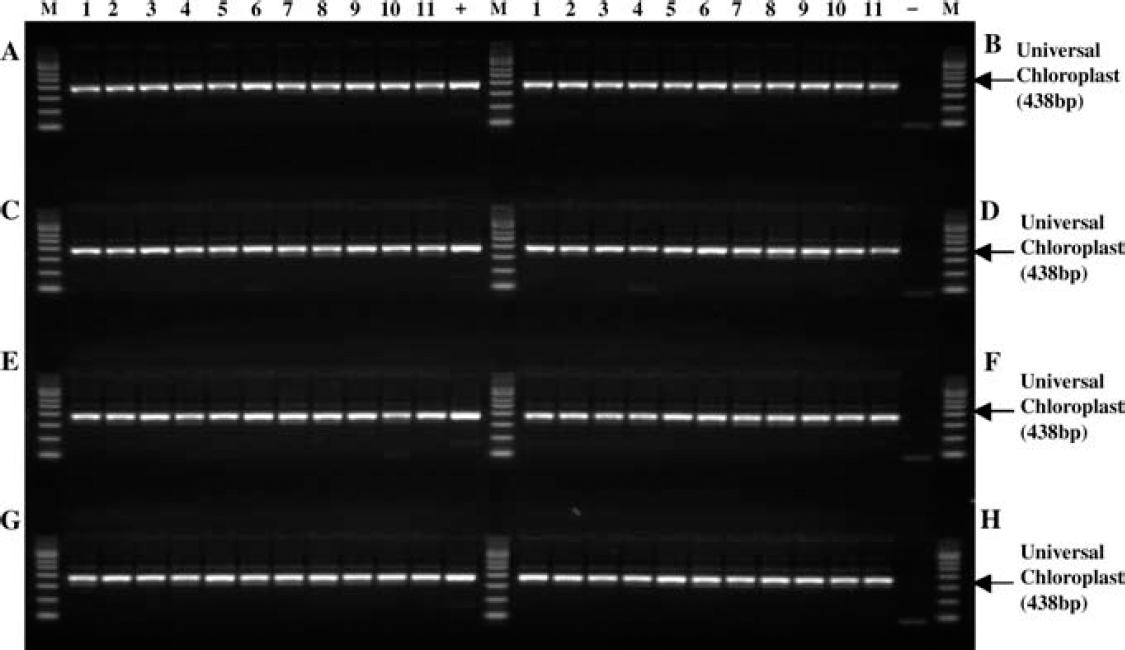

Genomic DNA was extracted from 88 maize leaf samples and subsequently amplified by the PCR reaction using the automated Extract-N-Amp Plant PCR method. Amplification of the 438 bp fragment of universal chloroplast genomic DNA is indicated by the arrow. M: PCR marker (Sigma-Aldrich Product No. P9577); (+): maize genomic DNA control; (-): no DNA template control.
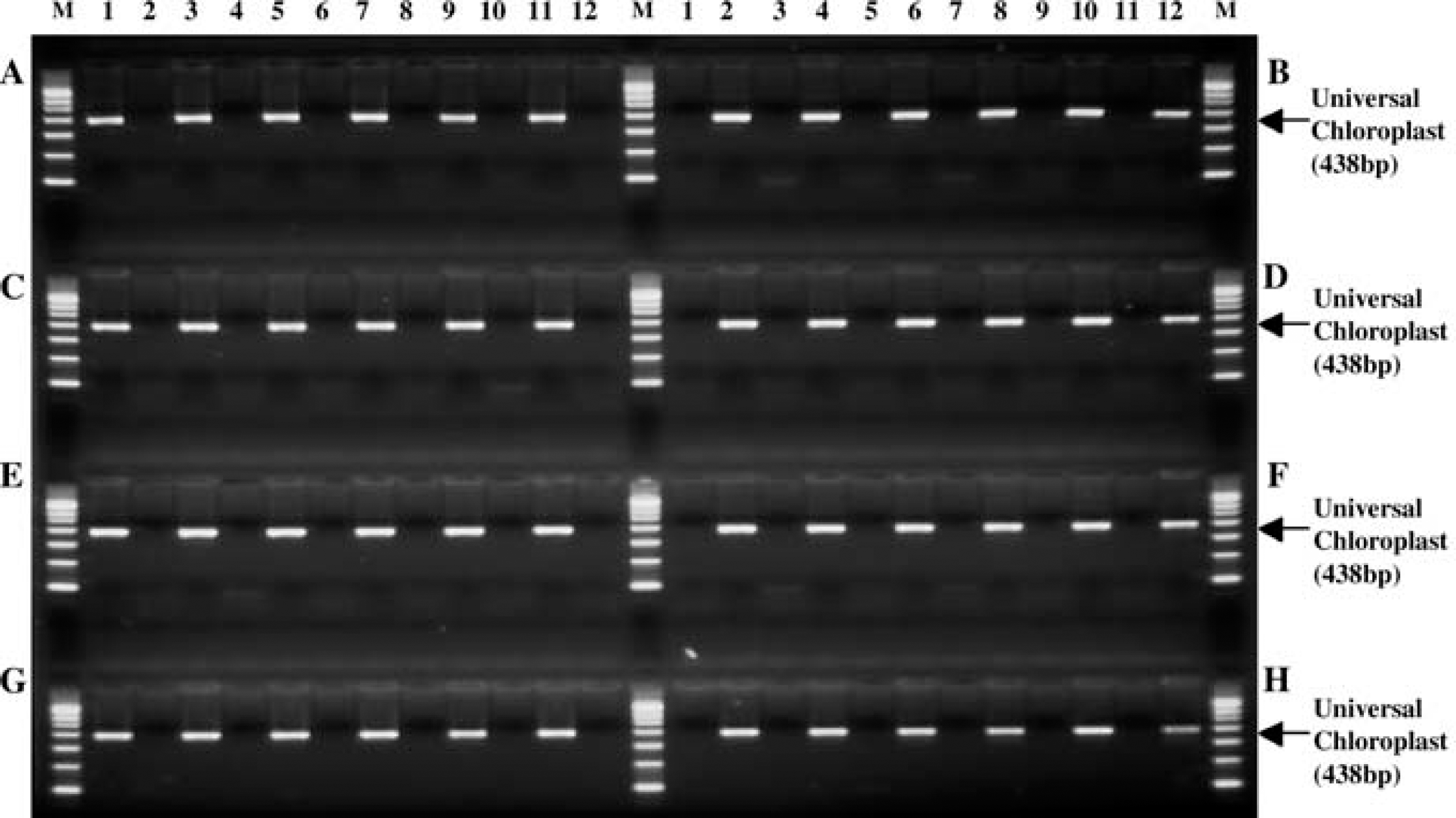

Maize leaf punches and blanks were placed in alternating wells of a 96-well plate. Genomic DNA extraction, amplification, and analysis were performed. No PCR products were detected in the wells without plant tissue samples. M: PCR marker.
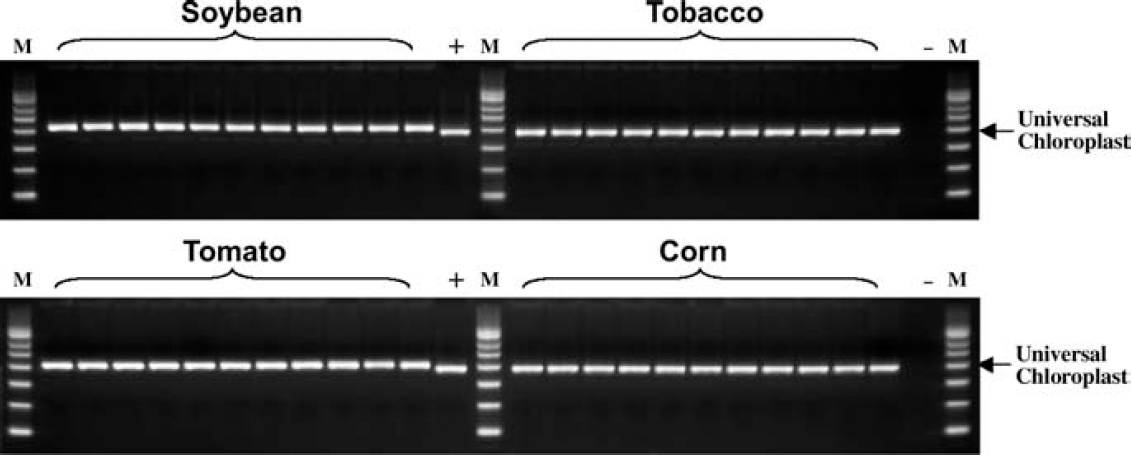
Genomic DNA was extracted from soybean, tobacco, tomato, and maize leaves, and subsequently amplified by the PCR reaction using the automated Extract-N-Amp Plant PCR method. Amplification of 400–500 bp fragment of the universal chloroplast genomic DNA is indicated by the arrow. M: PCR marker; (+): maize genomic DNA control; (-): no DNA template control.
Screening for Polymorphic Primers in a Maize Population
The validated method developed for the Extract-N-Amp Plant kit on the Sciclone ALH 3000 was then used to screen a maize population for SSR marker polymorphisms. The maize SSR primer set consisted of 384 pairs of SSR primers selected from the existing maize genome database. 5 The 384 pairs of primers were selected based on their genome location and covered the entire maize genome with an average map distance of 20 cM between two SSR markers. Each primer of the 384 primer pairs was synthesized in a separate well of eight 96-well plates. Equal concentrations of a forward and reverse primer set were combined into the same well of a 96-well plate and then lyophilized. To demonstrate the use of this library, one plate of 96 SSR primer sets was randomly selected to screen for SSR marker polymorphisms in three maize populations (P1, P2, and F1). The gel picture in Figure Figure 4 shows the amplified products for thirty-two of the 96 SSR markers evaluated. The data in Table 1 provides specific information for each of the 32 SSR markers evaluated in Figure Figure 4. The presence of a polymorphism was described as a size difference in PCR products generated by the P1 and P2 populations. This was further confirmed by the presence of hybrid P1 and P2 PCR products in the F1 generation. Of the 96 SSR markers analyzed, 12 were identified as polymorphs in the two parental lines. Three representative primer sets were then selected to study the segregation pattern in the F2 population.
Maize SSR primer set
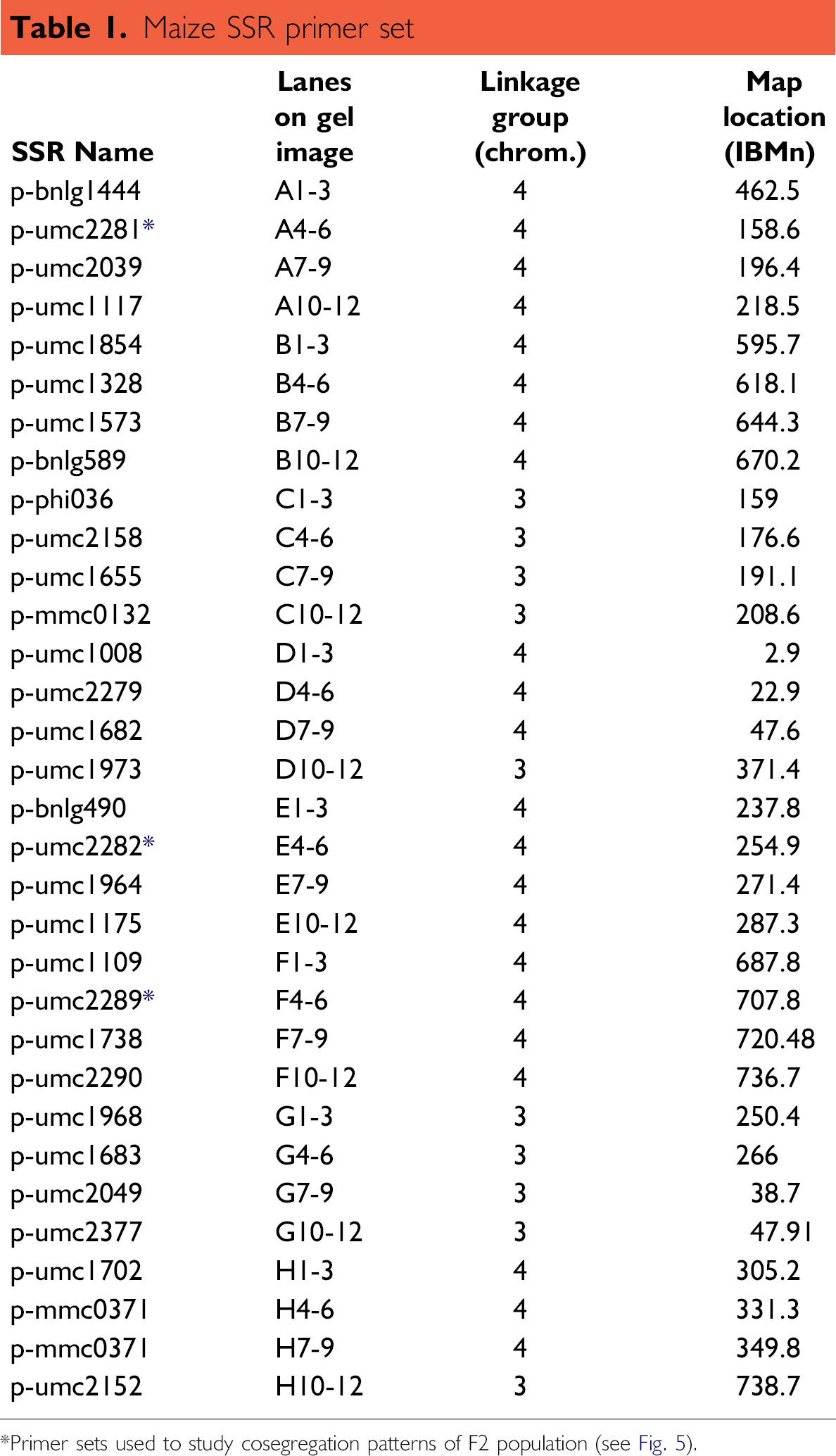
Primer sets used to study cosegregation patterns of F2 population (see Fig. 5).
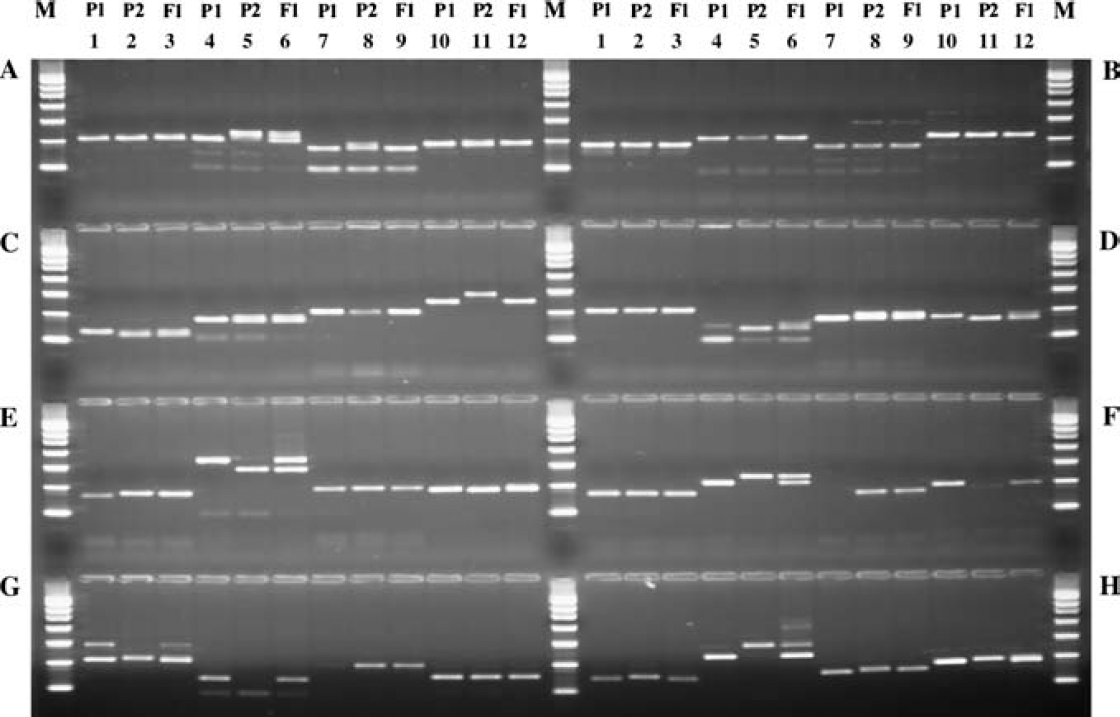
Genomic DNA was extracted from the leaves of two parental lines (P1 and P2) and the resulting hybrid line (F1) using the automated Extract-N-Amp Plant PCR method. The SSR fragments were amplified and analyzed. M: PCR marker.
The Use of Selected Polymorphic SSR Primers to Identify Segregation Patterns in an F2 Population
The segregation patterns of 21 strains within an F2 generation were observed using agarose gel analysis of the amplified SSR markers with three selected polymorphic SSR primers (Fig. 5). The ratio of the number of strains in the F2 generation corresponding to genotypes of the P1, P2, or F1 population can be calculated accordingly for each SSR primer. For an example, the ratio of P1:F1:P2 for the SSR primer shown in panel A is 3:10:8. However, a large number of F2 populations must be screened to draw statistically significant conclusions.
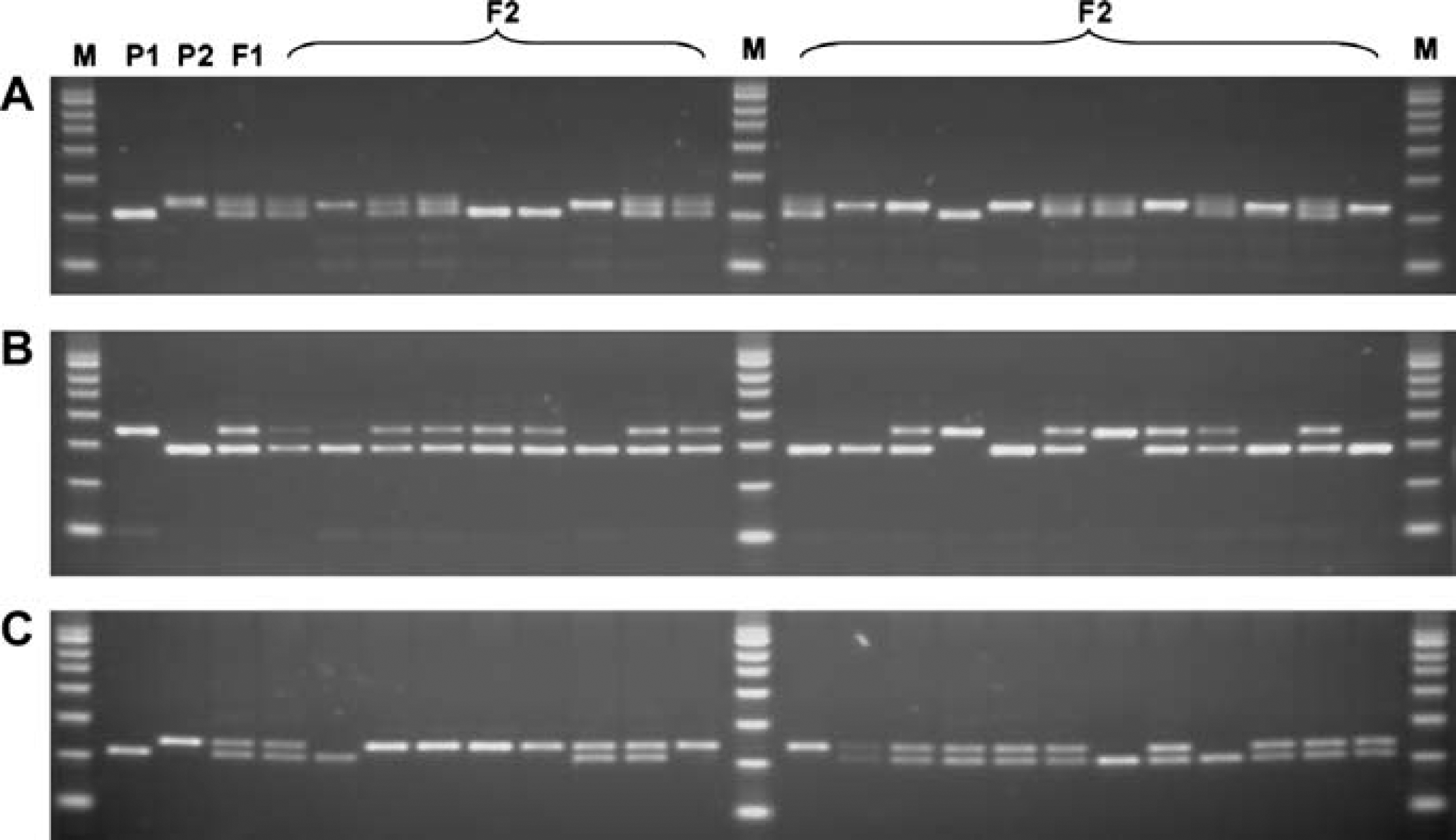
Genomic DNA was extracted from the leaves of two parental lines (P1 and P2), one hybrid line (F1), and 21 samples from the F2 line, using the automated Extract-N-Amp Plant PCR method. The SSR fragments were amplified and analyzed. The amplified SSR fragments from three different SSR primer sets identified from Figure 4 (Lanes A4–6, E4–6, and F4–6) are shown in panel A, B, and C respectively M: PCR marker.
Conclusions
A high-throughput system for genomic DNA extraction and subsequent PCR amplification from leaf tissue for use in marker-assisted breeding and genome mapping studies has been successfully developed. The use of the Extract-N-Amp Plant PCR kit eliminates the bottlenecks associated with plant tissue preparation and DNA extraction. In this report, an example of the effectiveness of this system for screening a maize population for SSR marker polymorphisms is provided. The entire process was automated from leaf punch through PCR reaction setup. In only 30 minutes the Sciclone ALH 3000 Workstation completed the genomic DNA extraction and PCR reaction setup for 96 leaf tissue samples. Template DNA in the extracts was shown to be of sufficient quality to allow for efficient PCR amplification using the SSR primer set.
The automated method presented was developed for the Sciclone ALH 3000 Workstation, but could easily be transferred to another liquid handling system. All program files associated for the automated method are available for download at www.sigma-aldrich.com/automation.
Acknowledgments
The authors would like to thank Dr. Georgia Davis, University of MissouriColumbia, for providing the maize populations in this study.
