Abstract
Development of a flexible, open-architecture modular robotic workstation consisting of a Calipers' Twister II robotic arm, several Labsystems Multidrop dispensers, a Kendro Cytomat incubator, and a multimode BMG Polarstar Optima reader has been reported. Our new, fully functional scheduling software developed under LabWindows allows the integration of an unlimited number of devices from diverse suppliers using native communication protocols.
Introduction
The convergence of traditional workstations and fully integrated robotic systems resulted in integrated screening solutions as a new generation of workstations. 1 These systems are typically compact to save laboratory space, and more importantly, to enable nonspecialized staff with moderate training to run screens in their own lab. Following the era of random screening, these workstations can now be used for primary screening of diverse and/or focused libraries, as well as secondary screening purposes. Workstations can handle several hundreds of plates daily using different kinds of limited axis robotic arms, liquid handlers, and readers operated by simple scheduling software.
Almost all suppliers developed such workstations. Tecan introduced Genesis Freedom (Tecan Group Ltd., Männedorf, Switzerland), Beckman Coulter developed a solution on Biomek FX platform (Beckman Coulter Inc., Fullerton, CA), Perkin Elmer has SmartStation (PerkinElmer Inc., Wellesley, MA), Staccato Cell Station and Elisa Station are available from Caliper (Caliper Life Sciences, Hopkinton, MA), while Thermo Labsystems (Thermo Electron Corporation, Waltham, MA) offers workstations using Catalyst Express which is the robotic arm of Thermo CRS. These solutions are common at least in one point: since major suppliers feature virtually all necessary components, most of the items are provided by a single manufacturer. The emphasis on such a compact, ready to use solution might ensure that vendor-supported applications would demonstrate the significant gain in precision and reproducibility. On the other hand, the flexibility and adaptation of new technologies might be limited by suppliers' access to assay technology.
Although these workstations are modular in nature, the reconfiguration of the system requires device drivers and interfaces. Most of the companies offer a large number of device drivers for a wide variety of instruments, including robots, pipetting stations, washers, detectors, and incubators. Since heterogeneous device environment is more typical in biological laboratories, device drivers are essential for the installation of a workstation in a predefined instrument environment. Although some drivers might be available from commercial sources, all of them are specific to the scheduling software environment used. Custom development of unavailable drivers and implementation of newly added instruments could make the development time-consuming and particularly expensive. An open-architecture system, however, would add significant flexibility with maintaining the beneficial features of a bench-top solution. Instrument drivers in such a system could be readily created since they are based on the native communication protocol of the given device. Drivers in this approach are not “black boxes” anymore, that is, their multiple usage, reconfiguration, and update could be done without additional cost. Our flexible and modular solution, AssayStation, realizes this concept using the National Instruments Lab-Windows environment (National Instruments Corporation, Austin, TX). 2 The scheduling software developed under LabWindows allows rapid integration of new devices from diverse suppliers.
Materials and Methods
Our AssayStation consists of the following devices: the central Twister II robotic arm with three standard stackers, three Thermo Labsystems Multidrop Dispensers, a BMG Polarstar Optima multimode plate reader (BMG Labtechnologies GmbH, Offenburg, Germany), a Heraeus Cytomat 2C plate thermostat (Kendro Laboratory Products, Asheville, NC), a bar-code reader, and an ambient temperature storage place for six microplates (Fig. 1). A central PC controls the workstation with an eight-port RS232 interface card and Microsoft Windows 2000. Our new scheduling software was developed under the National Instruments LabWindows C programming environment (www.ni.com).
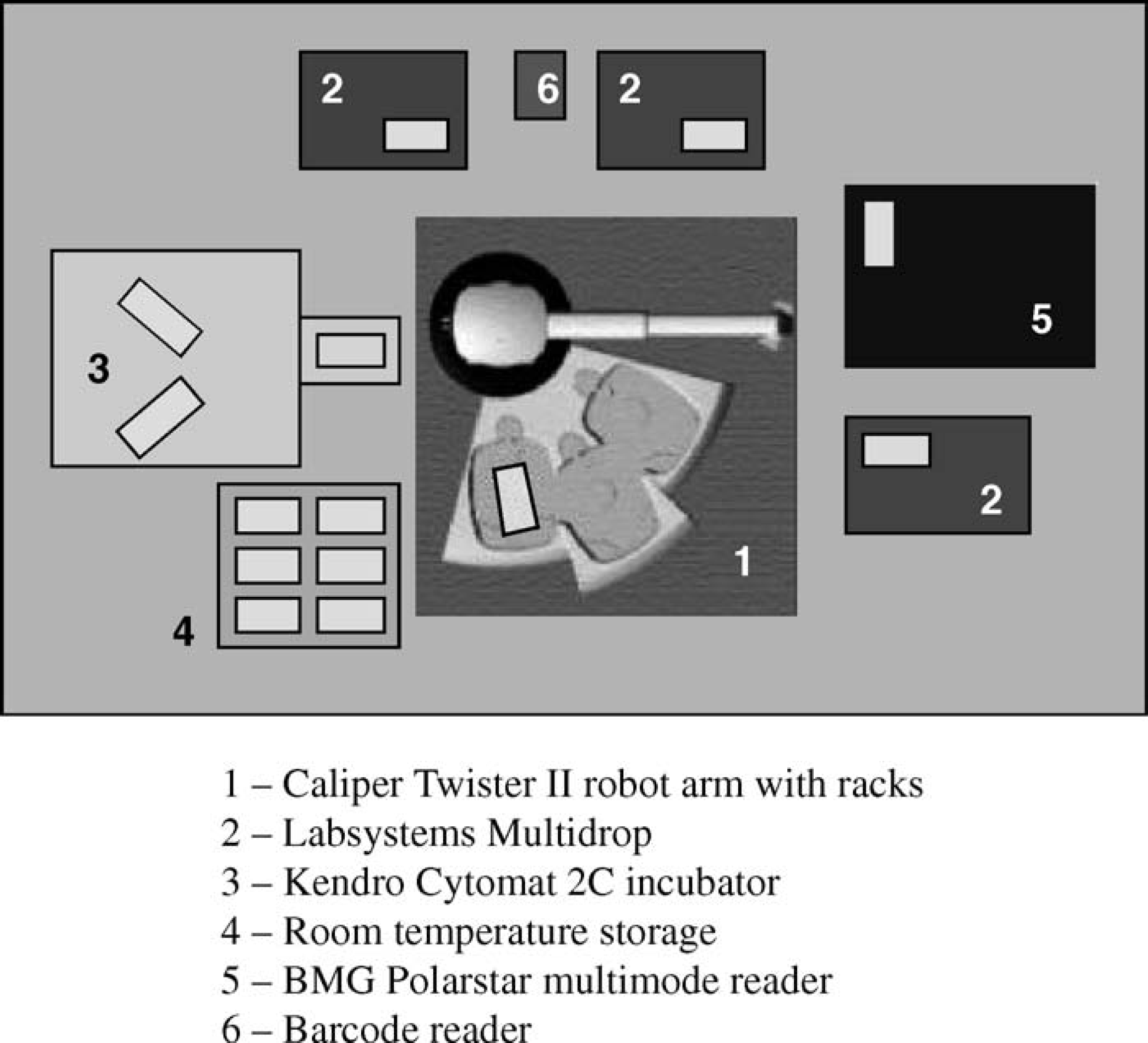
Schematic illustration of AssayStation.
Results and Discussion
To control the various components of our workstation, we divided our central kernel into three main units: (I) robot module, (II) programming and program execution unit, and (III) the general device interface (Fig. 2).
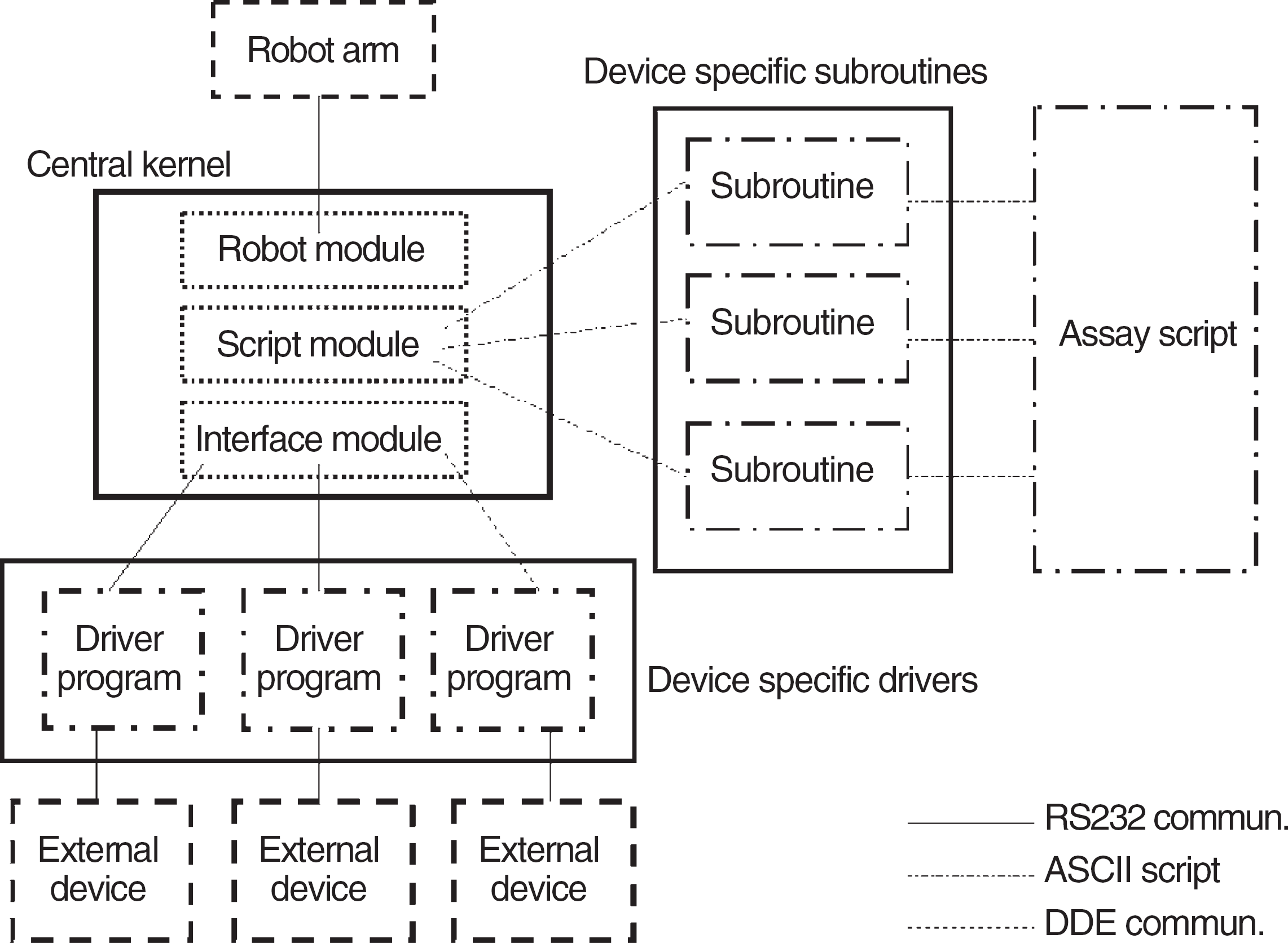
Software components and connections realized in LabWindows environment.
The robot module handles the entire communication between the robot and the computer. It also helps the user to train the device positions to the robot. It can be considered as a special device driver. We implemented the Twister II robotic arm's native serial communication protocol in our program.
The programming and program execution unit of the central kernel implements the programmability of the workstation and makes it possible to record a series of related movements (subprogram), store, and replay them. The movements thus stored can be combined into an assay protocol program and can then be replayed as many times as the number of plates to screen. Fix delays can also be included between any of two individual movements or subprograms. When the arm picks up a plate, the assay protocol program starts immediately for this particular plate. Consequently, the program execution unit runs the assay protocol program simultaneously exactly as many times as the number of plates already entered into the system. The step-by-step execution of assay protocol programs assigned to each plate is triggered by acknowledgments from the external units (e.g., one of the dispensers send the message when finished dispensing the reagent). The scheduler algorithm switches between assay protocol programs running parallel and lets the robot pick up the next plate and be ready to move to the next place as defined in the protocol. Priority can be assigned to each step encoded in the assay protocol program. If the program execution unit has more than one plate ready at the same time, the scheduler selects by the priority assigned to the actual step of these plates. The highest priority wins, therefore the robot first serves the highest priority step. Using the above-described priority logic, we can assure that time-critical steps—start and stop of incubation in our case—have prioritized over other steps, and therefore, the total time of the critical step remains constant.
The device interface of the central kernel is the connection point that communicates with all devices taking part in the assay. This communication can be split into two basic sections. The first section is the starting of an external device (e.g., the start of the feeder or placing a plate into the incubator). The start command may contain an auxiliary code that describes that the operation has to be started (e.g., taking out a plate from the fifth position of the incubator). The second section of the dialog is the reply of the external device, which can be a simple acknowledgment, a detailed answer containing information, or an eventual error message (e.g., successful feeding, plate placed at the 9th position of the incubator, or “there was no plate at the 5th position”). The communication between the central kernel and the device drivers is based on the standard message-queuing protocol of Microsoft Windows, and therefore, device drivers can be made using any programming languages that are able to use this protocol.
These three units (I, II, and III) together form the operational program that will be able to run an assay described in the assay protocol program.
Device drivers were created to be able to run independently for testing purposes and manual operations. If, however, the given program is started by the central kernel, it should only carry out the command from the kernel. It should stop automatically after having performed the prescribed operation and having sent a message to the central kernel.
We have chosen the LabWindows product of National Instruments for the whole software development process. Selection of LabWindows was supported by the advantages summarized below:
strong graphical support for generating the user interface; automatic frame program generation for the developed operational interface; libraries with simplified structure for various communication standards (ActiveX, DDE, serial protocols etc.); backing for the communication between the various programs (central kernel, device managers); having a style oriented for virtual instruments; can be used with standard C language; fast and comfortable software development; existing reference for system integration capabilities.
The most important feature of LabWindows, in our case, was clearly the serious support for creating standard instrument drivers that makes the operation of totally different devices possible in a similar style. The standard instrument driver is a set of high-level universal routines that hide the lowlevel commands and is generic and modular enough to serve for different applications used by the same instrument. Such a device driver contains an interactive surface that allows (1) easy search in the hierarchical structure of instrument handling functions, (2) the descriptions of selected functions to be available by a mouse click, (3) functions to be parameterized via a graphical interface (Fig. 3), and (4) the driver to be tested prior to integration.
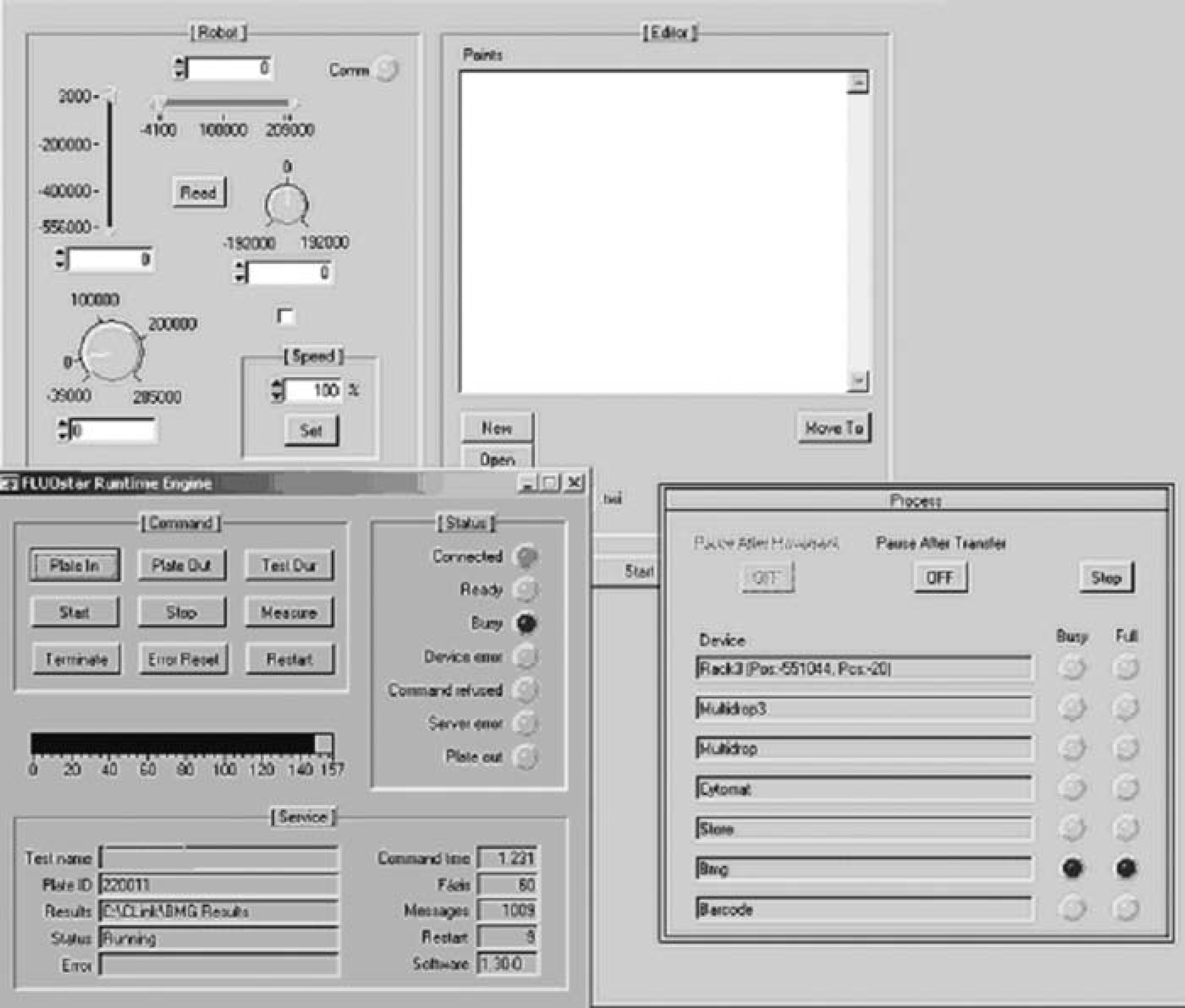
A snapshot of the scheduling software developed for AssayStation.
This LabWindows-based scheduler combined with suitable device drivers was applied to control our screening solution, AssayStation. AssayStation is equipped with Labsystems Multidrop Micro/Multidrop 384 and a BMG reader, all compatible with 384-well plate format. In this configuration we use Twister II, the telescoping arm and rotating wrist joint make it possible to access the x-y index positions of virtually all microplate instruments. Due to the wide reach area, we could integrate Kendro Cytomat 2C automatic incubator that enables performing fully automated assays. Compared to the previous Twister base solution of Tanabe, 3 AssayStation (as a commercial workstation) has the ultimate advantage of scheduling that enables it to process almost 340 plates per day in our fully automated environment. The BMG reader integrated to AssayStation supports five different measurement principles: fluorescence polarization, fluorescence intensity, time-resolved fluorescence, high-performance luminescence (flash and glow), and absorbance that makes the system available for a wide range of detections.
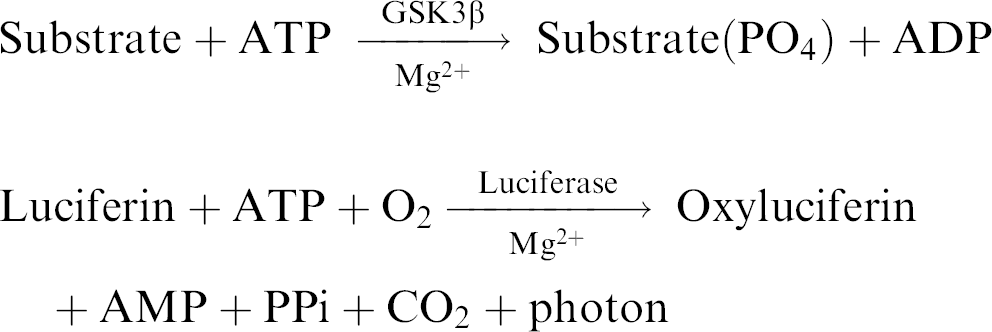
Capabilities of AssayStation have been demonstrated in a kinase assay targeting glycogen synthase kinase 3-beta (GSK3β). To determine the GSK-3β activity, we adopted the Kinase-Glo Luminescent Kinase Assay, a homogeneous method of measuring kinase activity provided by Promega (Promega Corporation, Madison, WI). 4 The Kinase-Glo assay is a homogeneous nonradioactive method of determining the activity of purified kinases by quantifying the amount of ATP remaining in solution following a kinase reaction. Remaining ATP is quantified by the monooxygenation of luciferin catalyzed by UltraGlow luciferase in the presence of Mg2+ and molecular oxygen that produces one photon of light per ATP molecule.
In our case, the assay was performed in 96-well plates by adding a volume of Kinase-Glo reagent equal to the volume of solution in the well of a completed kinase reaction and luminescence was measured. The luminescence signal was correlated with the amount of ATP present and inversely correlated with the amount of kinase activity. Luminescence was recorded using a BMG Polarstar Optima reader.
We first confirmed the linear relationship between the luminescence measured and the amount of ATP in the kinase reaction buffer in the range of 0,003-3 μM (r2 = 0,999; data not shown). The Kinase-Glo reagent relies on the properties of a thermostable luciferase that generates a stabile “glow-type” luminescent signal and has a half-life greater than 4 h. In our experiments, the measured signal decreased by ∼ 15% in 90 min at room temperature.
We used human recombinant GSK-3β, active, (expressed in Sf21 insect cells) obtained from Upstate (Upstate Group LLC., Charlottesville, VA) and a pre-phosphorylated polypeptide substrate provided by American Peptide Incorporation (American Peptide Company, Sunnyvale, CA). The peptide was650 His-Ser-Ser-Pro-His-Gln-Ser(PO3H2)-GluAsp-Glu-661 Glu-Glu that refers to C-terminal fragment of GS-2 peptide extended by C-terminal Glu642 instead of Asp from the original sequence of human muscle glycogen synthase.
In order to optimise the kinase reaction, we performed a set of experiments to determine the exact amount of ATP, substrate and kinase, necessary for the reaction (data not shown). The best signal/noise ratio was obtained under the following condition: 25 μM substrate, 1 μM ATP, 20 ng enzyme/assay in a final volume of 40 μl at 37 °C. The reaction was stopped after 30 min by addition of 40 μl of Kinase-Glo reagent. The luminescence was recorded after 10 min, which was found to be necessary to stabilize the luminescence signal. Enzyme activity was expressed as a ratio of the consummated and total ATP present in the reaction.
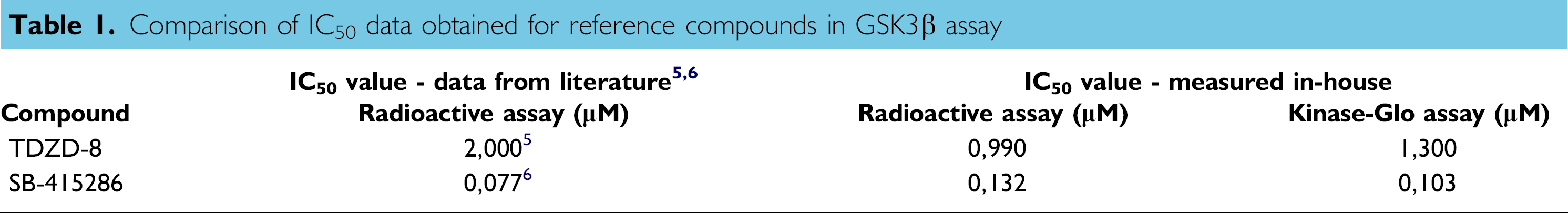
Two reference compounds were tested for assay validation: TDZD-8 and SB-415286. Both compounds inhibited GSK3β at IC50s, which correlates well with data published in the literature (Table 1). 5,6
Comparison of IC50 data obtained for reference compounds in GSK3β assay

HTS experiments started from preassembled, bar-coded microplates containing screen compounds at the concentration of 5 μM/L dissolved in assay buffer. Positive controls were without any compounds added (100% enzyme activity, no inhibition), and negative controls contained 5 μM SB415286 (100% inhibition, no enzyme activity). In the first dispensing step (Step 1) we added 20 μL mixture of the substrate and ATP. The next step is to add the enzyme to the reaction mixture. Step 3 is an incubation period at 37 °C for 30 min in the Cytomat 2C incubator, followed by Step 4 where we add 40 μL Kinase-Glo reagent to the reaction mixture and shake the plate for 30 s. Step 5 is an ambient temperature incubation for 10 min just before reading the plates in Step 6 by BMG Polarstar Optima multimode reader. The measured plate then moves to Rack 1 in Step 7. The assay steps are described in Table 2 and Figure 4. It was crucial in our enzyme assay to execute Step 3 with minimal deviation in order to keep the consistency of the results (time critical step). To achieve this goal, we applied the priority logic feature of our software, assigning high priority flag to Step 3 that allowed the duration of Step 3 to be kept constant. As it can be seen in Figure 4, there are two devices with relatively long program duration and multiplate storage capacity (Cytomat 2C incubator can store 40 microplates and the ambient temperature spot can store 6 microplates) and one device without multiplate storage capacity but with long operation time (BMG Polarstar Optima reader). To permanently assure the precise duration of incubation, a delay appropriate to the duration of plate reading should be inserted into the assay protocol. Consequently, we applied a 170 s delay at the end of Step 1.
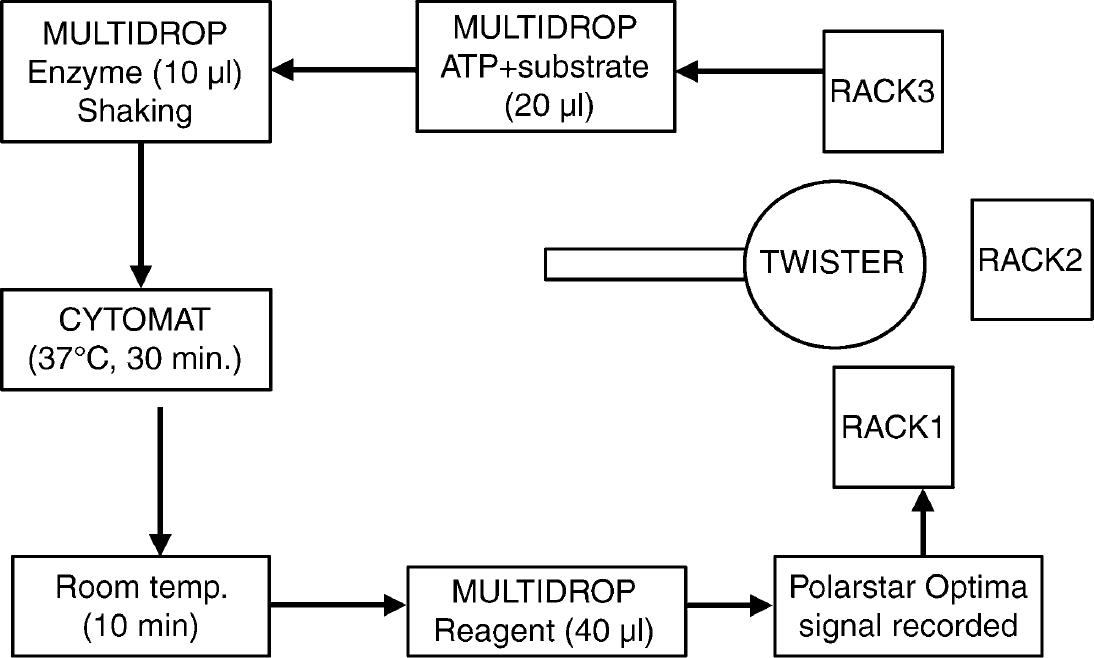
Flowchart of our assay protocol realized on AssayStation.
Steps of the assay performed by AssayStation

To demonstrate the capabilities of our software and AssayStation, we analyzed an approximately 10 h run with 141 measured plates, 13536 wells in total. Averages of unit operation times including plate transfer, priority delay, and standard deviations are summarized in Table 3.
Unit operation time (s) measured on AssayStation

Reproducibility of the assay was quantified by the calculation of Z' factors 7 for all 141 plates. For this purpose, we used six positive and six negative controls measured at each plate and calculated corresponding averages (AVG) with standard deviations (SD) that were used in eq 1.
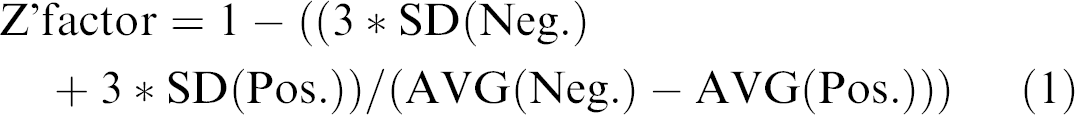
Distribution of Z' factors is depicted in Figure 5. Average Z' factor for the investigated plates was found to be Z' = 0.77 with a SD of 0.08. Reasonable Z' factors obtained in this set suggest the assay, and also the system, are capable for HTS operations against GSK3β.
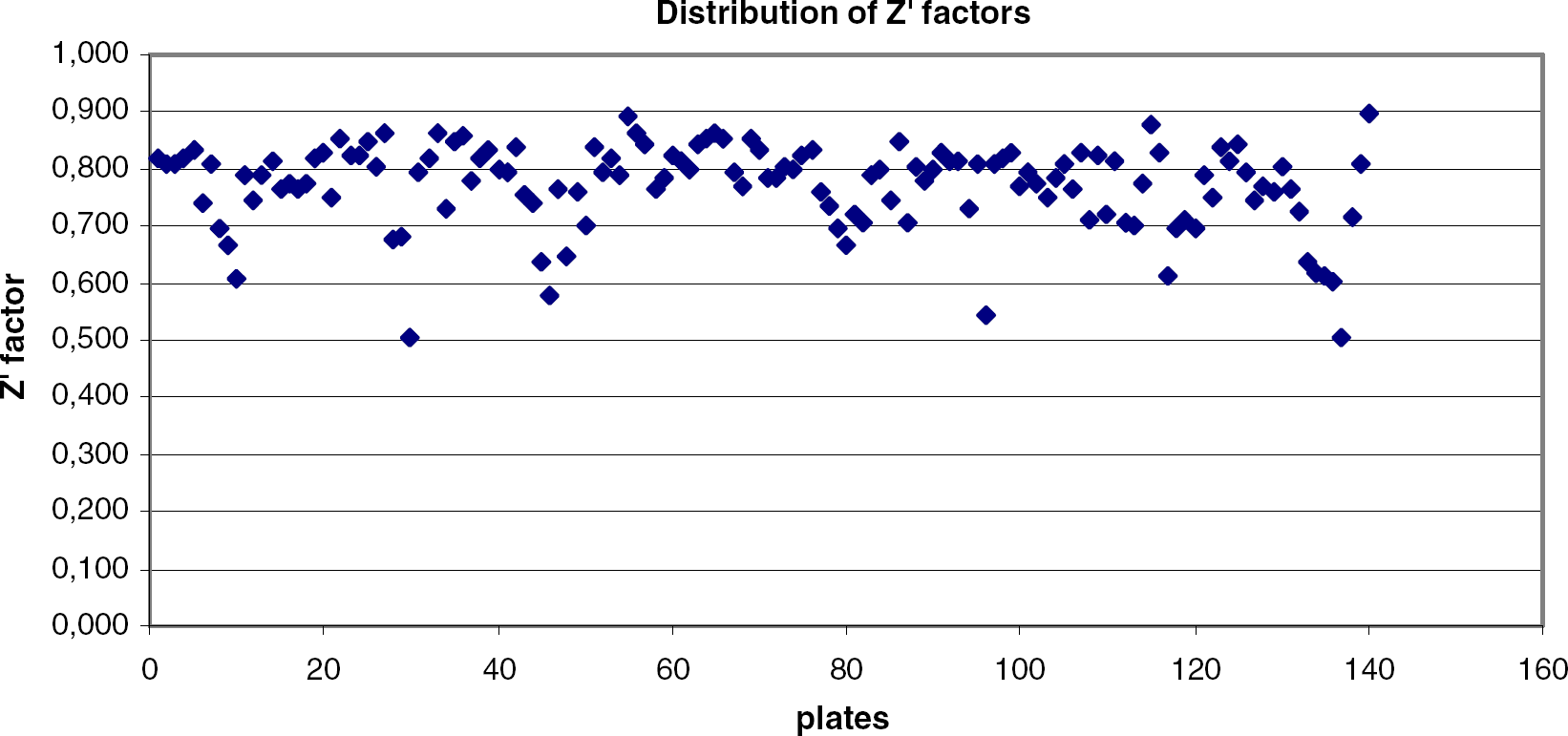
Distribution of Z' factors along the test run in GSK3β assay.
In summary, we developed a LabWindows-based scheduling program and AssayStation with key features that included scheduling capabilities, priority handling, 384-well plate compatibility, and integration of incubation steps. Our software was written in C under the LabWindows development suite and communicates with all but one of the devices through RS-232C using the native communication protocol of the components. DDE interface was exclusively used for communication to the BMG reader. All of the Active X components of Twister II (iLink) were omitted because of limited performance in our experiments. Due to the script module of the central kernel, our software meets all requirements of a scheduler, including simultaneous multidevice handling and priority logic. The LabWindows environment allowed us to develop a vendor-independent, fully flexible, open-architecture solution for homogeneous assays.
Acknowledgments
The authors are grateful to Andrea Baki for developing the assay, Krisztina Recska and Eva Horvath for technical assistance, and Timea Polgáir, Dóra Menyhárd, and James Peterson (Amgen) for helpful comments.