Abstract
The demand for high-throughput RNA isolation has been dramatically increasing with wide applications of RNAi, expression profiling, and molecular diagnosis. A comparison of various RNA isolation methods that have been adapted to high-throughput platforms, focusing on consistently high yield and quality of isolated RNA, reduction of cross-contamination, and simplicity and robustness of the protocol is presented. The streamlining of RNA isolation with RNA quantification by qRT-PCR and amplification for microarray analysis is also discussed. In general, a microspheric bead-based approach results in more consistent RNA recovery than glass fiber filter-based RNA isolation method, and RNA can be eluted in a smaller volume. This is because beads can be fully re-suspended in solution to enable more thorough mixing, washing, and elution, whereas the glass fiber matrix is fixed in a filter plate. (JALA 2004;9:140-5)
Introduction
The genome wide profiling of gene expression, which was made possible by the development of DNA microarray technology and made more powerful by the sequencing of the human genome, has been widely used for tumor classification and biomarker discovery, 1 as well as in target validation and toxicogenomics. 2 RNA interference or RNA mediated mRNA degradation, facilitates the high-throughput functional analyses of gene targets identified during drug discovery. 3 These advances in molecular biology over the past decade have created tremendous opportunities for high-throughput sample preparation. The automation of the tedious sample preparation process helps avoid human error. However, the traditional manual sample preparation kits cannot simply be slapped onto a robotic platform and be expected to provide optimal performance. Before any kit can be used on a robot, the entire process has to be examined. In this article, Ambion's approach of streamlining each step of the sample preparation process is discussed.
Automation guidelines
In theory, with modern advances in robotic control, every single step in sample preparation that a human being is capable of performing can be achieved by a robot. In practice, automation has its limitations with constraints of space and budget. The most important question is whether automation is cost effective for your particular needs. Here are two examples: (1) One can easily load a filter plate onto a centrifuge; however, it is quite expensive to integrate a plate centrifuge into a robotic platform. (2) One can easily mix a solution by inverting a plate a few times; however, it may cost a fortune to purchase a robot arm that can mimic this action. Based on our experiences of high-throughput method development and customer service, we have delineated at least three fundamental steps for any sample process: defining the right approach, simplifying the process, and choosing the right robotic platform.
Defining the right approach: Sample preparation can vary widely for different applications. For biomarker discovery and in new drug clinical trials, genome-wide express profiling is achieved by DNA microarray analysis, where high-quality total RNA has to be isolated and amplified. For molecular diagnosis and target validation, gene expression (mRNA level) is often determined by real time reverse transcriptionpolymerase chain reaction (RT-PCR). In general, the requirement of RNA quality (purity and integrity) for RT-PCR is much lower than that for microarray analysis. This is because only a handful of targets (genes of interest) are evaluated and often both gene specific primers and probes are used that deliver very high specificity in target mRNA quantification. 4 As a result, RNA isolation may be unnecessary for quantitative RT-PCR (qRT-PCR) (cf. the section entitled “Skip RNA Isolation”). The simpler the approach, the more reliable the outcome.
Simplifying the process: Statistically, the more steps in a process, the more likely the final output will deviate from your expectation. Therefore, minimizing steps involved in the whole process is very important. As discussed previously, the most effective way is to analyze your application and define a practical approach that has minimal steps involved in the whole process. Then maximizing master mix is desirable to further reduce pipetting steps. It is also very important to simplify pipettor movement to reduce cross contamination. As a reagent company, we spend a lot time studying the stability of master mixture of reagents/enzymes and make appropriate recommendations, as well as how to lay out plates and reagents to minimize pipettor crossing plate/ reservoirs and to shorten travel time.
Choose the right robotic platform: A well-defined approach for your applications will help you find the right robotic platform and essential accessories. Unless you have a well-defined application and will probably never change the assay, a generic robot with an open platform is usually the best choice. It is very difficult to find a company who has expertise in both robotic platform and reagents/assays. Most people are trapped in fully work-away automation solutions at their initial approach. In general, full automation means more sophisticated integration, which results in a dramatic rise in cost and would probably reduce the input per hour. In a lot of cases, you will find that semi-automation is a better solution for the most robust high-throughput sample preparation, which is discussed in more depth in the following sections.
Skip RNA Isolation
The fluorescence-based real-time reverse transcription PCR (RT-PCR) is widely used for the quantification of steady-state mRNA levels and is a critical tool for basic research, molecular medicine and biotechnology. Assays are easy to perform, capable of high throughput, and can combine high sensitivity with reliable specificity. As discussed previously, the high specificity and sensitivity of qRT-PCR enables gene expression (mRNA concentration) to be determined with minimal sample preparation. Ambion's Cells-to-cDNA™ technology (U.S. patent pending) completely bypasses RNA isolation thereby increasing the throughput potential, improving the precision of the final results and decreasing overall costs.
In Cells-to-cDNA technology, cultured cells from tissue culture are lysed and the ribonucleases are inactivated in a single step with a novel cell lysis buffer. The resulting cell lysate is competent for reverse transcription and is added directly into a one-step RT-PCR reaction. The sample preparation is simplified to removing culture media, adding lysis solution, and heating, therefore very consistent results can be obtained for anyone without training on specially handled RNA samples. The coefficient of variation (CV) for all 96 samples from one 96-well plate is less than 3% and typically 1–2%. 5 It is very suitable for quantifying mRNA expressed at moderate to high level (Figure 1). The starting material can be cultured cells, LCM samples, and white blood cells.
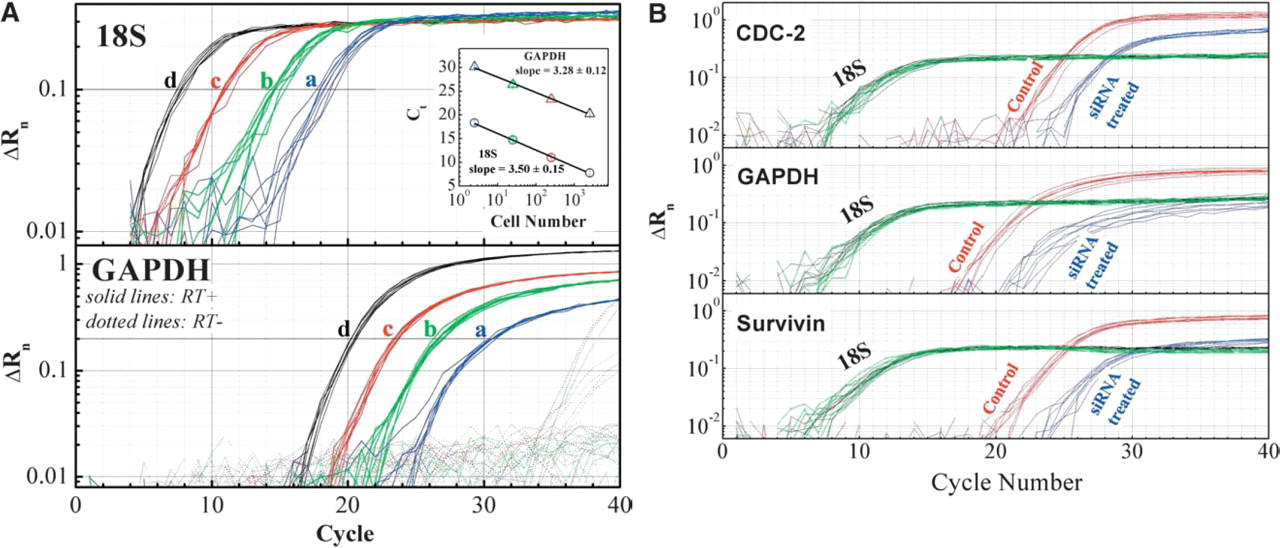
mRNA quantification without RNA isolation by Ambion's Cells-to-cDNA™ technology. Panel A: 100 (a); 1,000 (b); 10,000 (c); and 100,000 (d) HeLa cells were processed with Cells-to-cDNA technology. 2.5% lysate was used for qRT-PCR. Both GAPDH and 18S levels were linear with cell numbers. RT- indicates negligible genomic DNA contamination. Panel B: siRNA validation by qRT-PCR using Cells-to-cDNA technology. After siRNA transfection, target gene expression was lowered by ∼10 fold. Target expression was normalized to 18S rRNA expression.
You may seamlessly integrate sample preparation and one step RT-PCR setup on a generic robot with an integration of a heating tile/oven. The automated processes take about 1 hour, from removing the cell culture media to lysing cells and then finally to the setup of the one-step RT-PCR (protocol and automation guidelines are available at Ambion's automation website 5 ). In reality, most of our customers choose semi-automation to achieve higher throughput by a semi-automation approach: remove media and add lysis buffer (6–10 plates at a time) on a 96-channel robot → manually transfer plates to a convection oven for the lysis step → manually move back to robot for the qRT-PCR setup. As such, 6–10 plates of samples can be prepared (including qRT-PCR setup) in 2 hours.
RNA Isolation with Filter Plate
For most applications, RNA needs to be amplified before detection, such as microarray analysis, which requires high-quality RNA. Although only mRNA is to be analyzed, common approaches isolate total RNA (mRNA, rRNA, and tRNA), which usually gives better quality of RNA because properties of the entire RNA molecule can be utilized for isolation instead of only using the poly(A) tail of eukaryotic mRNA molecules. Traditionally, total RNA is isolated by phenol extraction, which is difficult for automation. 6 Solid phase binding methods were thus developed to replace phenol extraction for high-throughput RNA isolation. As exemplified by Ambion's RNAqueous® technology, the process usually contains the following steps: lyse sample → bind RNA/DNA to matrix → optional DNase treatment → wash away contaminates → elute RNA. 5 There are many choices of RNA/DNA binding matrixes, such as glass fiber filter, ion-exchange resin, and beads with various binding surfaces. Ion-exchange resin is less favorable because it usually requires a high-salt eluent for final RNA elution, which creates potential problems for downstream RNA amplification/detection. Glass fiber filters and beads (especially para-magnetic beads, which will be discussed in the next section) are among the best choices of binding matrixes.
Filtration can be performed with either a vacuum manifold or a centrifuge with a 96-well plate rotor. A vacuum manifold is cheap and easy to integrate, but usually needs extra care to control the vacuum level. Too high of a vacuum setting is the major cause of cross contamination, whereas a lower vacuum setting may not generate enough pull strength to create a seal upon applying vacuum to successfully pull samples through the filter. If the vacuum level can be controlled within the robotic controlling software, we recommend middle level of vacuum (10–15″ Hg) to minimize failure in binding and washing steps, and low level of vacuum (5–7″ Hg) to avoid liquid splashing during the elution step. If the robot has no control over the vacuum setting, a low level of vacuum should be used for the whole process. When a vacuum manifold is used for RNA isolation, gasket(s) should be checked routinely to make sure that they create a good seal between the filter plate and the vacuum manifold. When very precious samples are to be processed, people usually babysit the whole process to be ready to correct any malfunctions of the robot.
In contrast to vacuum filtration, centrifugation usually generates more consistent data and may require less intensive care. However, a centrifuge with a plate rotor is expensive to integrate, thus most people choose to do off-line centrifugation when they decide to use a centrifuge instead of a vacuum manifold. Another advantage of centrifugation is that 4 plates can be processed in parallel for high throughput or a partial plate for low throughput. A typical run takes less than 1.5 hours. RNA obtained is of high quality and the recovery rate is consistent. However, RNA yields generally vary more at lower cell number input (Figure 2).
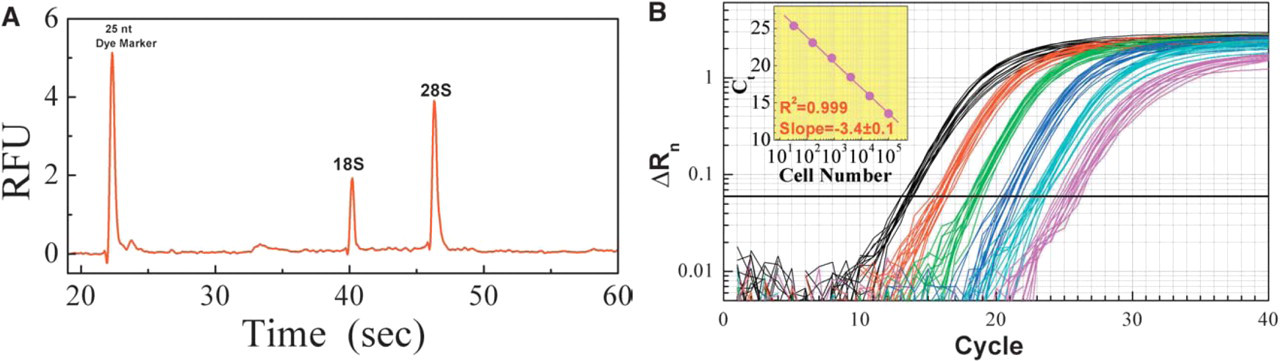
Glass fiber filter plate-based total RNA isolation with Ambion RNAqueous®-96 Automated kit. Panel A: High-quality total RNA was evident by the sharp peaks of 28S rRNA and 18S rRNA and their 2:1 ratio (analyzed on Agilent 2100 bioanalyzer with RNA LabChip). Panel B: Linear recovery of high-quality total RNA quantified by Real-time qRT-PCR. Total RNA was extracted from 32, 160, 800, 4,000, 20,000, and 100,000 K562 suspension cells. 5 μL of 100 μL eluted total RNA was used for a 25 μL one-step qRT-PCR reaction. Human GAPDH primer/probe set was used on an ABI7900 sequence detection system.
RNA Isolation with Magnetic Beads
Solid phase RNA isolation with magnetic beads provides the same convenience and automated capability as filter-based RNA isolation. The procedure is very similar to the filter plate-based method. As exemplified by Ambion's RNAqueous®-MAG technology, the protocol consists of (1) sample lysing and DNA/RNA binding, (2) DNA digestion, (3) washing, and (4) RNA elution. 5 A magnetic field is used for pelleting beads to allow supernatant removal instead of applying vacuum/centrifugation to pass through liquid as in filter-based RNA isolation.
Micro-spherical beads have large available binding surfaces and can be fully dispersed in solution, allowing more thorough RNA binding, washing, and elution, and therefore delivering more consistent results (Figure 3). The process is very easy for total automation, and tends to be more robust than the filter-based method because vacuum failures are more likely to occur than magnetic field failures. Because a 96-well magnetic stand has a small footprint as a standard SBS plate and an orbital shaker can hold 4 plates a time, higher throughput (4 plates in 1.5 hours) can be achieved. One Ambion customer has very successfully applied this technology for Exotic Newcastle Disease diagnosis with chick swab samples. 7 With over 100,000 samples analyzed so far, not a single case of cross contamination or false positive/negative result has been observed. Another advantage of bead-based RNA isolation techniques is that RNA can be eluted in very small volume (<20 μL), that can be easily streamlined with RNA amplification.
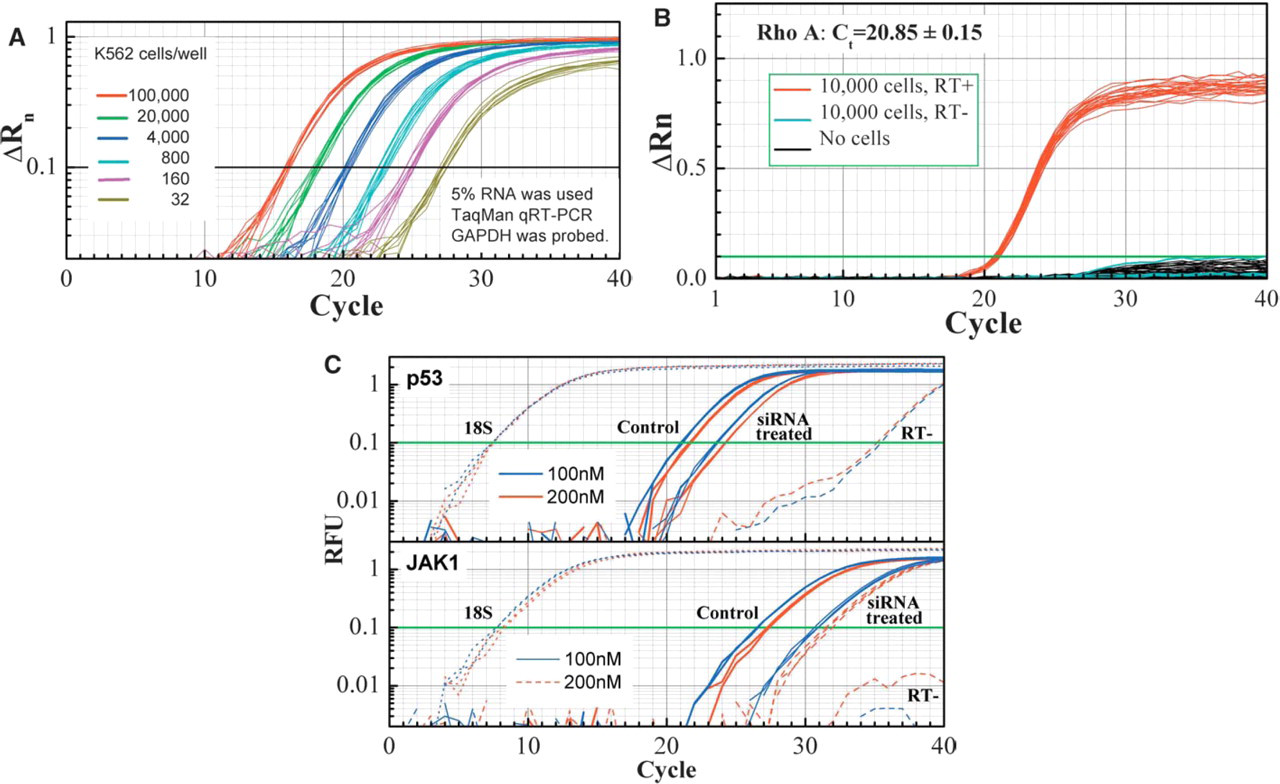
Magnetic bead-based total RNA isolation with Ambion RNAqueousR-MAG 96 Automated kit. Panel A: High consistency and linear recovery of total RNA quantified by real-time qRT-PCR. Total RNA was extracted from 32, 160, 800, 4,000, 20,000, and 100,000 MCF-7 cells. 5 μL of 100 μL eluted total RNA was used for a 25 μL one-step qRT-PCR reaction. Rho A primer/probe set was used on an ABI7900 sequence detection system. Panel B: No cross contamination was observed in the sample preparation process. 10,000 K562 cells were spiked in 24 random wells of a 96-well plate and remaining wells are filled with water. Rho A mRNA was consistently detected in final elutions from wells with cell input, while consistently absent in the elutions of wells without cells. Panel C: RNAqueous®-MAG's application in siRNA evaluation. Human umbilical vein endothelial cells were transfected with siRNAs, mRNA was isolated 24 hours post transfection and quantified by qRT-PCR with SYBR Green.
Sample Preparation for Microarrary Gene Profiling
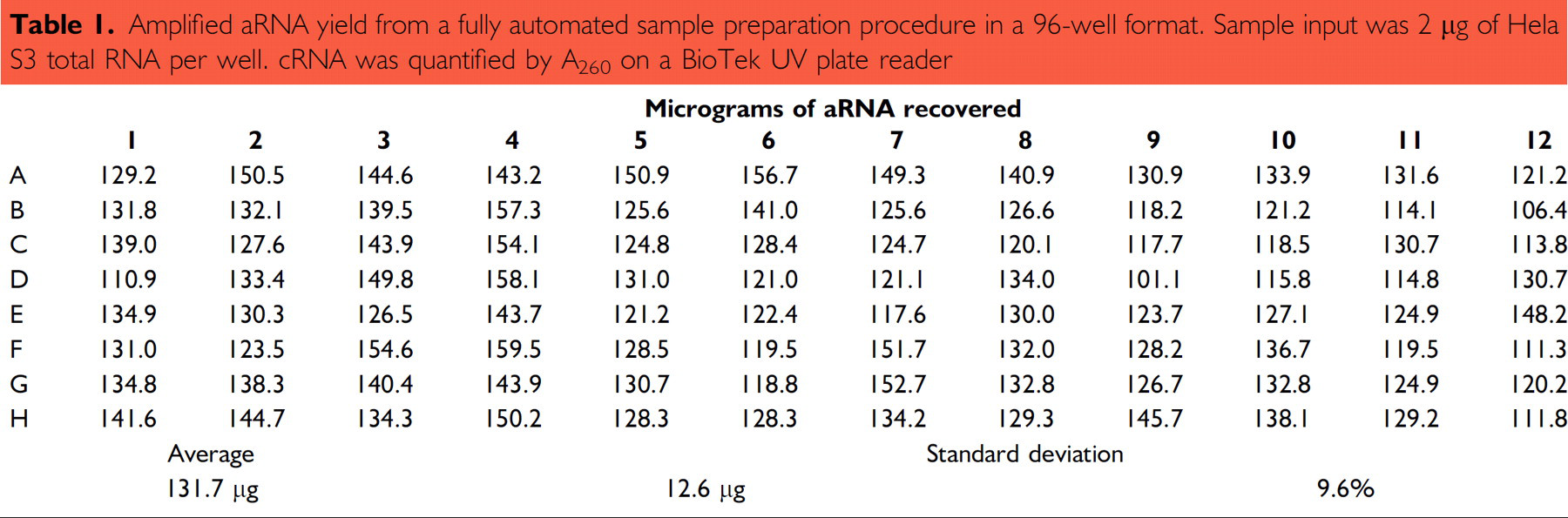
Sample preparation for microarray analysis is the most tedious process: (1) high-quality RNA is isolated and quantified (and concentrated if too diluted); (2) RNA is reverse transcribed to make a double-stranded cDNA template and incorporate a T7 promoter; (3) each cDNA is transcribed to thousands of cRNA; and (4) cRNA is labeled, cleaned, and quantified for array hybridization. The whole sample preparation takes about 2 days and many steps are involved in the sample preparation, therefore a lot of training is required to perform the whole protocol to achieve expected results. Full automation is highly desirable to get better consistency. In this case, because the downstream array analysis is very expensive, the cost of automation integration is easily justified. The big challenge here is to develop a protocol to seamlessly streamline the whole process. At Ambion, we fully automated the lengthy protocol by choosing a magnetic bead-based RNA isolation method to get high quality of concentrated RNA, and use magnetic beads for cDNA and aRNA purifications. Compared to the filter-based clean up procedures, the bead-based clean up process does not alter the expression profile (Figure 4), and has no bias in recovery RNA of different sizes (Figure 5). The aRNA generated is consistently of high yield and quality (Figure 6 and Table 1). The array data produced with the generated aRNA had very low background and a high-percentage presence call (data not shown).
Amplified aRNA yield from a fully automated sample preparation procedure in a 96-well format. Sample input was 2 μg of Hela S3 total RNA per well. cRNA was quantified by A260 on a BioTek UV plate reader
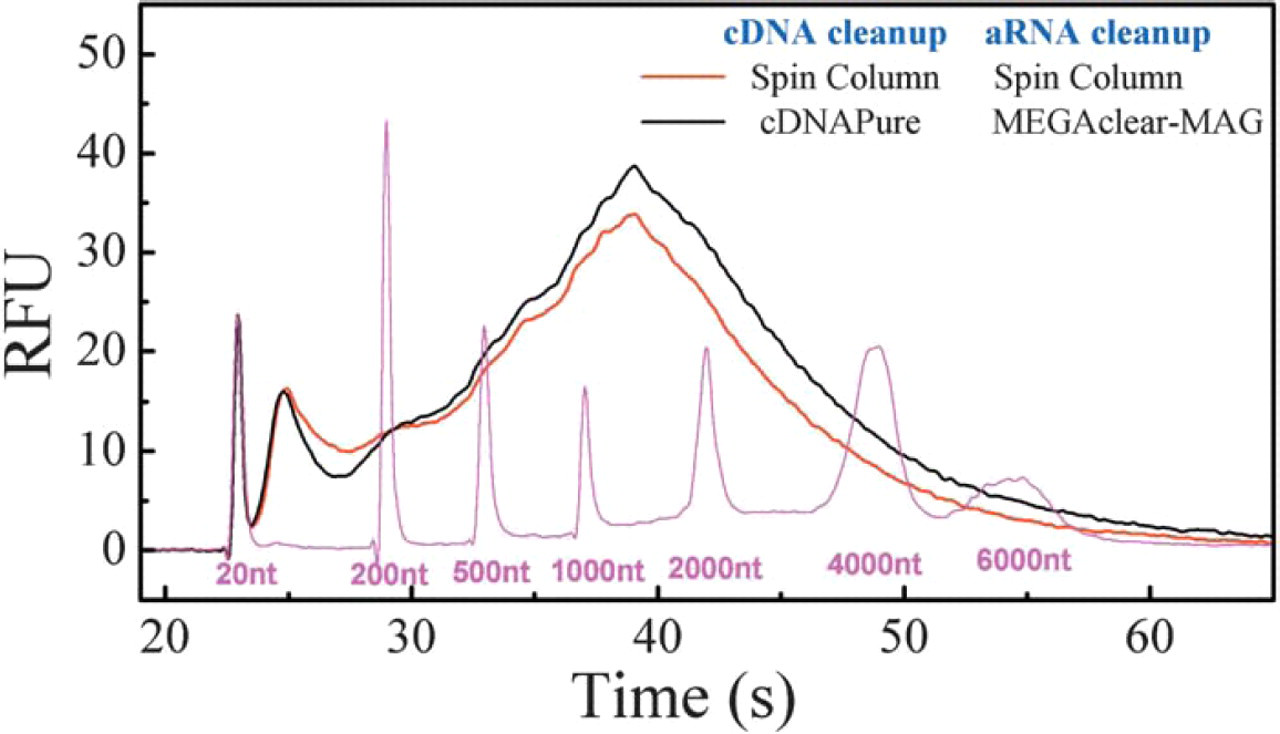
Consistently high yields and quality of RNA obtained using Ambion's MessageAmp™ kit with different clean up methods. With both clean up methods (either with glass fiber spin column or with magnetic beads) high-quality aRNA was evident by the high yield and consistent size distribution with a maximum size around 1500 nucleotides.
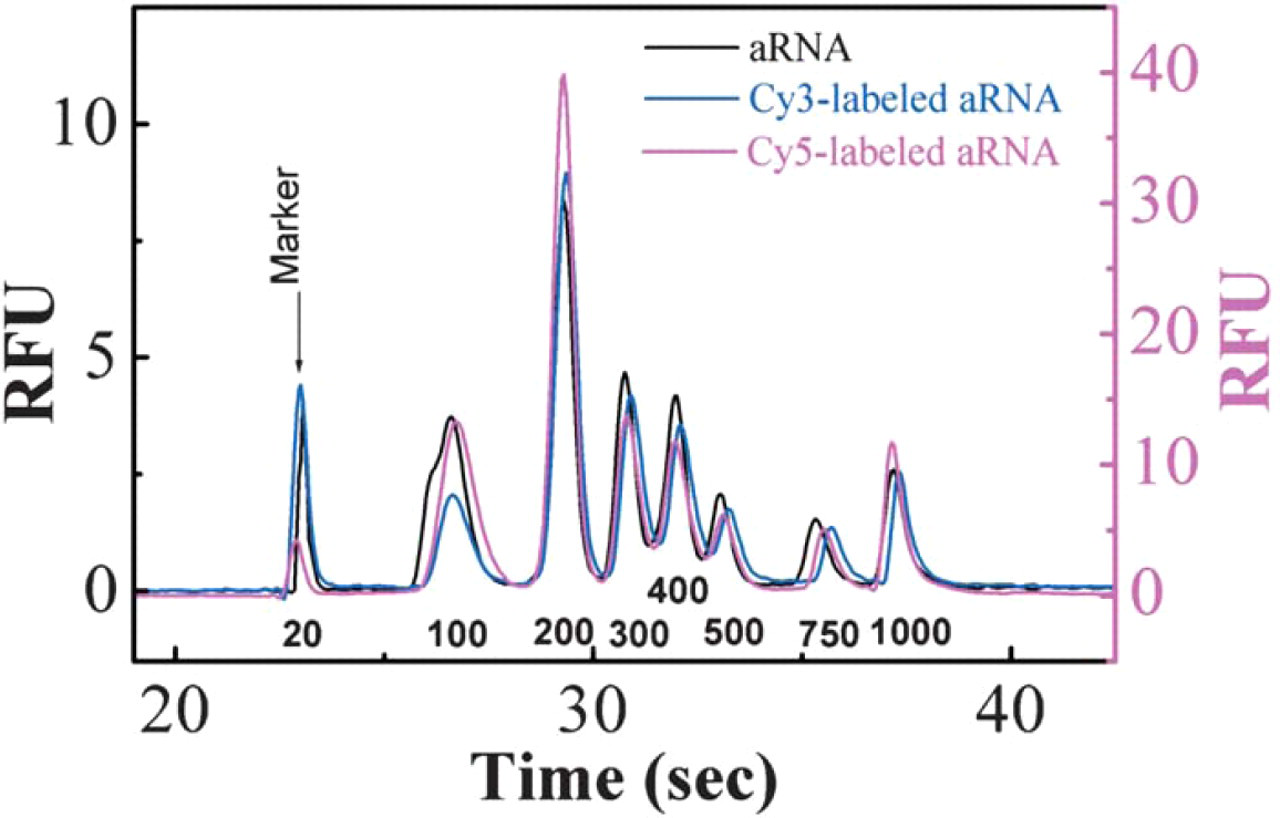
Dye coupling bead-based and cleanup of coupled amino allyl RNAs. The amino allyl MessageAmp™-96 kit protocol was used to generate amino allyl RNA mixture using Ambion Century™-Plus Marker Templates and to clean up the dye-coupled RNA mixture. The RNA mixture, before and after dye coupling, was analyzed on an RNA 6000 chip with Agilent bioanalyzer. Cy5-labeled RNAs (magenta curve, right Y-axis) have much stronger signal because the fluorescence spectrum of Cy5 overlaps with the detection wavelength. This figure demonstrates that dye coupling is efficient for both long and short RNA with various GC contents, and that the bead-based clean up method has no bias in recovering RNA of various sizes.
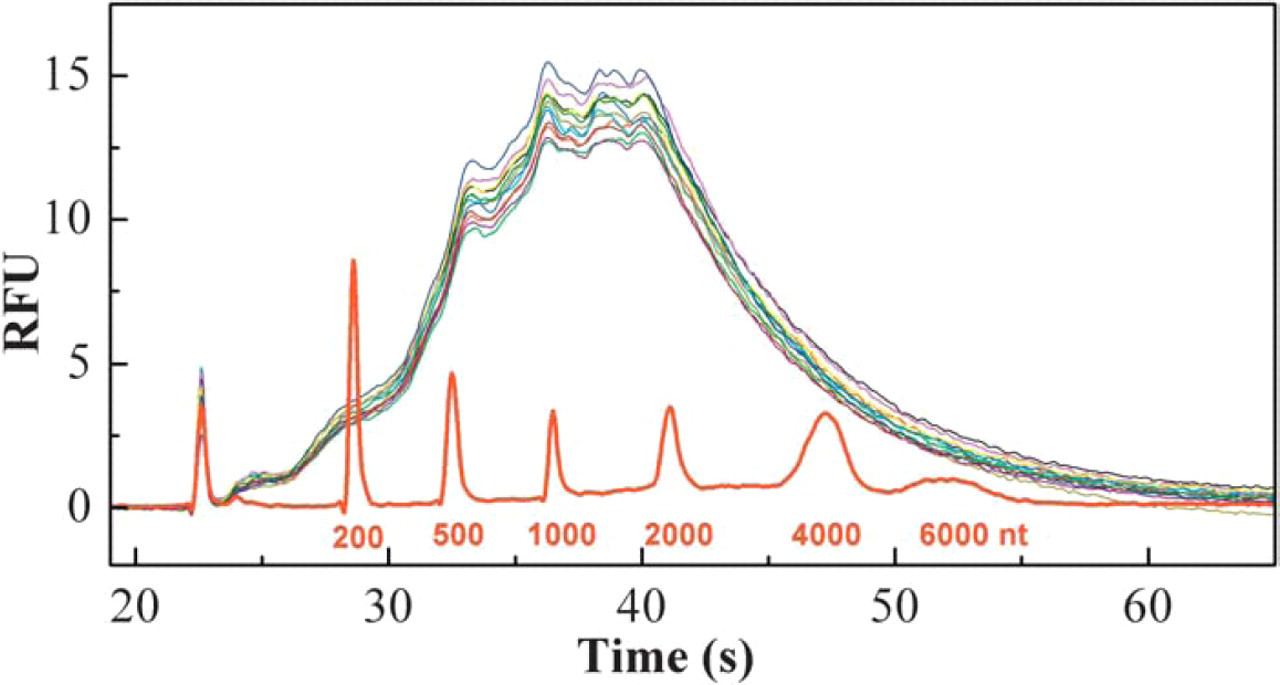
Size profile of amplified aRNA generated by a fully automated process in a 96-well format (cf. Table 1 for aRNA yield and consistency). Twelve randomly selected samples of the 96 were run on an RNA 6000 chip with Agilent bioanalyzer. High-quality aRNA was evident by the high yield and consistent size distribution with a maximum around 1500nt.
Acknowledgment
The authors would like to thank Dr. Mangkey Bounpheng for her comments and scientific editing of the manuscript.
