Abstract
We describe automated nucleic acid (NA) isolation from diverse sample types using MagMAX kits (Ambion, Inc.) on KingFisher Magnetic Particle Processors (Thermo Scientific). The MagMAX-96 Blood RNA Isolation Kit is designed for total RNA isolation from whole blood from several species, without white blood cell fractionation, in about 45 min (including genomic DNA removal). The MagMAX-96 Total RNA Isolation Kit is designed for total RNA isolation from up to 2 × 10 6 cultured cells and up to 10-mg tissue. The isolated total RNA is highly intact and pure, ready to use in downstream applications such as microarray analysis or real-time reverse transcription (RT)-PCR for gene expression profiling or pathogen detection. The MagMAX-96 Viral RNA Isolation Kit is designed for viral RNA and DNA isolation from cell free or nearly cell-free samples such as swabs, serum, and plasma; it takes about 15 min. Total NA of high quality and purity is recovered at >75% efficiency, providing high sensitivity for pathogen detection by real-time RT-PCR. Unlike automated liquid handling systems that move reagents into and out of a single well of a multiwell plate to perform the different steps of an RNA isolation procedure, the KingFisher Magnetic Particle Processors use permanent magnetic rods to collect magnetic beads from solution and release them into another well containing reagent for the subsequent step of the procedure. The effectiveness of bead collection and transfer lead to superior washing and elution efficiency, as well as rapid processing. It is a very effective strategy for automation of magnetic-bead-based NA isolation kits. (JALA 2007; 12:195–201)
Introduction
RNA isolation is a tedious process, and is often the bottleneck for gene expression profiling using DNA microarrays. 1 The demand for high-throughput RNA isolation is steadily growing because techniques that require RNA become more widely used, such as quantitative reverse transcription (RT)-PCR for pathogen detection 2 and functional analyses of gene targets identified by RNA interference. 3 Traditional methods for total RNA isolation use phenol: chloroform extraction, which is difficult to automate or even to manually perform on many samples in parallel. 4 Solid-phase nucleic acid (NA) isolation techniques have been developed to enable high-throughput processing. Among the most widely used are methods that rely on glass-fiber filters and magnetic microspherical beads. Magnetic-bead-based technology yields NA of higher purity with more consistent recovery than glass-fiber filter technology, because the agitation of beads in the NA isolation reagents enables more efficient binding, washing, and elution. 5 Furthermore, it is not subject to filter clogging that can complicate filter-based methods, and it does not require vacuum filtration or centrifugation, making walk-away automation easier to implement. Finally, magnetic-bead-based methods allow for recovery of NA in a small elution volume with a relatively high NA concentration, which is desirable for downstream applications.
We have developed the MagMAX product line of magnetic-bead-based kits for isolation of total RNA from cultured cells, animal and plant tissues, and whole-blood samples, as well as for isolation of viral NA from tracheal and cloacal swabs, plasma, serum, urine, and milk. These methods are designed for high-throughput sample processing that can be easily automated, and they have been used with several open-platform liquid handlers. In this article, we describe the performance of MagMAX kits automated using the KingFisher Magnetic Particle Processors.
Materials and Methods
All MagMAX kits were from Ambion, Inc. RNA isolation protocols are described in detail in the individual kit instruction manuals, which are available at the Ambion web site. 6 Briefly, cells and/or viral particles were lysed in a chaotropic solution that inactivates nucleases, and NAs were selectively bound to magnetic microspherical beads. The beads were washed to remove any residual impurities that may inhibit downstream applications such as RT and PCR. Finally, the purified NA was eluted in a small volume of low-salt buffer. In some experiments, genomic DNA was digested with TURBO DNase (included with the MagMAX-96 Total RNA Isolation Kit and the MagMAX-96 Blood RNA Isolation Kit). TURBO DNase is active, even at high salt concentrations that are inhibitory to most ribonucleases, minimizing any RNA degradation during the genomic DNA digestion.
RNA integrity was assessed by capillary electrophoresis with an RNA LabChip Kit on an Agilent 2100 bioanalyzer. NA was quantified by direct UV absorbance measurement with a UV–Vis spectrophotometer (NanoDrop Technologies). Quantitative RTPCR (qRT-PCR) was also used to evaluate NA recovery.
RNAlater Solution (Ambion, Inc.) was used as indicated for stabilization of RNA in cell or tissue samples before RNA isolation. ArrayScript Reverse Transcriptase (Ambion, Inc.) was used for RT, and SuperTaq DNA Polymerase (Ambion, Inc.) was used for quantitative PCR on a 7900HT Fast Real-Time PCR System (Applied Biosystems, Inc.).
We constructed an artificial RNA, called XenoRNA-01 to serve as an internal positive control for RNA quantification. XenoRNA-01 is 1000 bases and has a poly(A) tail. The sequence was designed to lack homology with any sequences in the GenBank database. To use it as a control for viral RNA isolation, we packaged the XenoRNA-01 into Escherichia coli bacteriophage MS2 coat proteins to form Armored RNA, a viral particle mimic that is resistant to nuclease degradation. 7 XenoRNA-01 Armored RNA was added to samples before starting the MagMAX procedure to monitor the efficiency of viral particle lysis and viral RNA recovery.
The MagMAX procedures were automated using KingFisher Magnetic Particle Processors (Fig. 1; Thermo Scientific), which feature permanent magnetic rods that fit into disposable tip combs, with independent control of the movement of each. These processors capture magnetic beads from solutions in multiwell plates by lowering the magnetic rods, sheathed inside a tip comb, into the solution and waiting for the magnetic beads to collect on the bottom of the tip comb. The beads can then be moved to different wells containing the next reagent in a workflow. Beads are released into solution by simply moving the magnetic rods up and away from the tip comb. The tip comb can also be moved up and down to mix reagent with the magnetic beads, and to facilitate sample lysis. In contrast, when magnetic-bead-based methods are implemented on a liquid handler, reagents must be aspirated out of the wells slowly to avoid losing beads and to remove the reagent to completion. Even with these precautions, however, there is always some residual reagent left behind. Using the KingFisher processors, on the other hand, the magnetic beads are fairly dry when pulled out of the solution, resulting in more efficient washing and elution, and accelerated processing speeds. For moderate throughput, the KingFisher Magnetic Particle Processor was used to process up to 24 samples in two 96-well plates; each sample was processed in one column of a plate, with each step of the procedure occurring in a different well of that column.
KingFisher (A) and KingFisher 96 (B) Magnetic particle processors. The KingFisher Magnetic Particle Processors use permanent magnetic rods and disposable tip combs, with independent movement control. When the magnetic rod, sheathed inside the tip comb is lowered into solution, magnetic beads collect at the bottom of the tip comb (C). The rods/tip comb can then be positioned in a different well or plate and the beads can be released by moving the magnetic rods out of the tip comb. Furthermore, the tip comb can be moved up and down to mix reagent with the magnetic beads. The KingFisher processes up to 24 samples in two 96-well plates, the KingFisher 96 processes up to 96 samples at a time.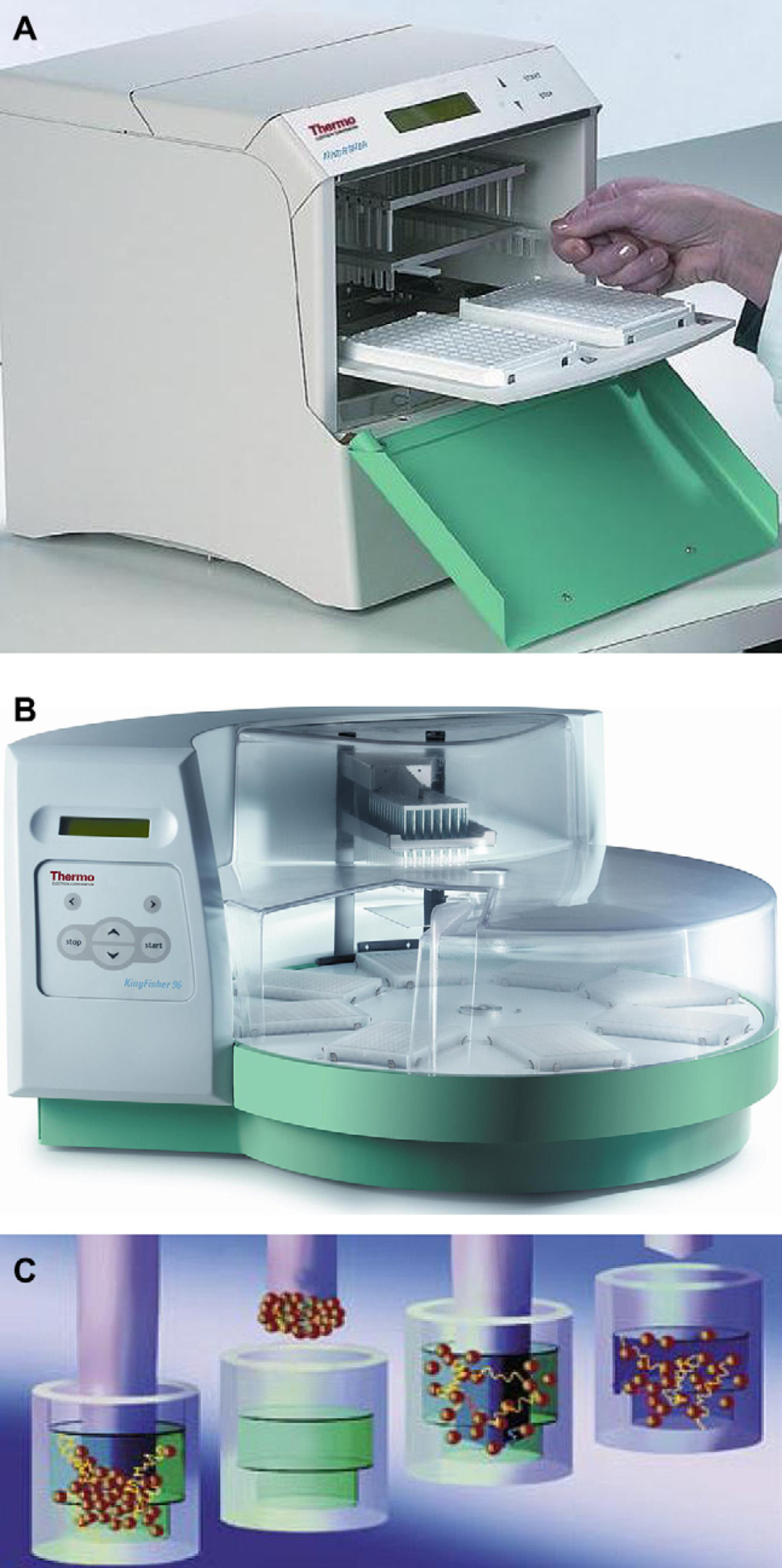
For higher throughput processing, the KingFisher 96 Magnetic Particle Processor was used to process one 96-well plate at a time, with each step of the procedure occurring in a different plate. MagMAX kits are scalable and can be adapted to process sample volumes both smaller and larger than the default volumes in the instructions. The KingFisher 96 processor can process sample volumes up to 300 μL using 2-mL deep-well plates. The programming is very straightforward; it consists of simply selecting the first desired function, for example mixing, binding, or elution; then, defining the parameters for that function, for example, position, speed, and time; and repeating the process to program each step of the protocol. Automation protocols for the MagMAX kits are available at Ambion's automation resource web page. 8 Furthermore, once the automation protocol is configured, it can be used on the different machines of the same model without calibration. In contrast, liquid handling systems typically require fine-tuning for each individual machine and every protocol.
Results
Total RNA Isolation from Whole-Blood Samples
Most whole-blood RNA isolation methods require a pretreatment of whole blood to enrich for white blood cells, either by preferentially lysing red blood cells or by fractionating white blood cells (buffy coat) by centrifugation. This makes them difficult to automate. In contrast, the MagMAX-96 Blood RNA Isolation Kit is designed for isolation of total RNA, which includes RNA from blood-borne pathogens, directly from 50 μL of whole blood in a 96-well plate.
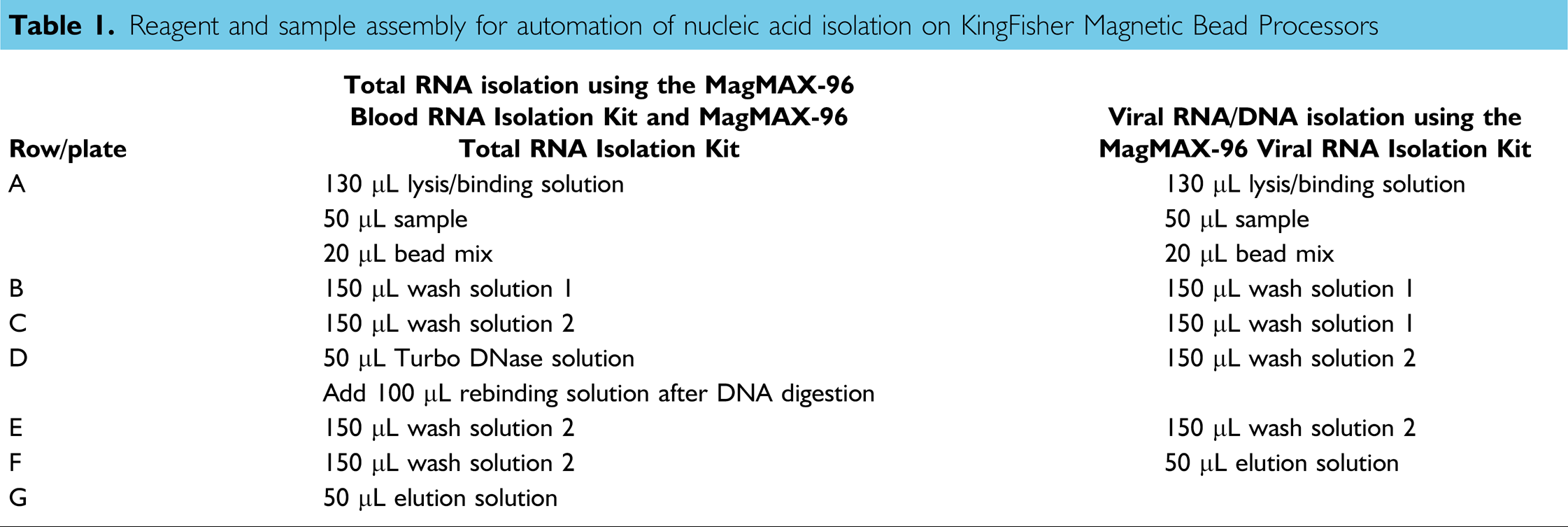
The MagMAX kit reagents and blood samples were dispensed into the wells of 96-well plates as described in Table 1, and RNA was isolated using KingFisher and KingFisher 96 Magnetic Particle Processors. The procedure was complete in about 45 min when the procedure included a genomic DNA digestion, or in less than 15 min when DNase digestion was unnecessary. RNA isolated from fresh human whole blood typically had an A260/A280 > 2.1, and 28S:18S ribosomal RNA (rRNA) ratios >1.0, indicating that the RNA was very pure and had a high degree of integrity (Fig. 2). The average RNA Integrity Number (a metric developed by Agilent, which incorporates information from both the rRNA peaks and the area outside the peaks to provide a complete picture of RNA degradation states) was >7.3, indicative of intact, high-quality RNA. RNA yield varied according to species (Table 2). The typical RNA yield from 50-μL normal human whole blood was 530 ng or ~ 10 μg/mL.
Reagent and sample assembly for automation of nucleic acid isolation on KingFisher Magnetic Bead Processors

Total RNA yield from whole-blood samples
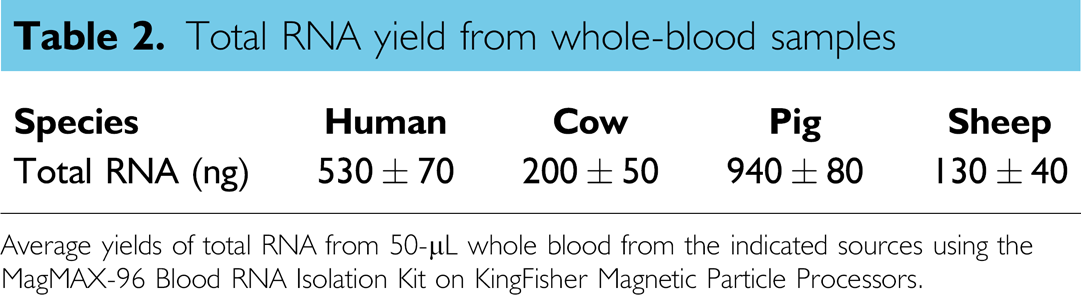

Average yields of total RNA from 50-μL whole blood from the indicated sources using the MagMAX-96 Blood RNA Isolation Kit on KingFisher Magnetic Particle Processors.
Integrity of RNA isolated from human whole blood. Total RNA was isolated from fresh human whole blood using the MagMAX-96 Blood RNA Isolation Kit. RNA integrity was assessed with an RNA LabChip on an Agilent 2100 bioanalyzer. The 28S:18S ribosomal RNA ratio was > 1.0, and RNA Integrity Number was >7.3 for all samples analyzed.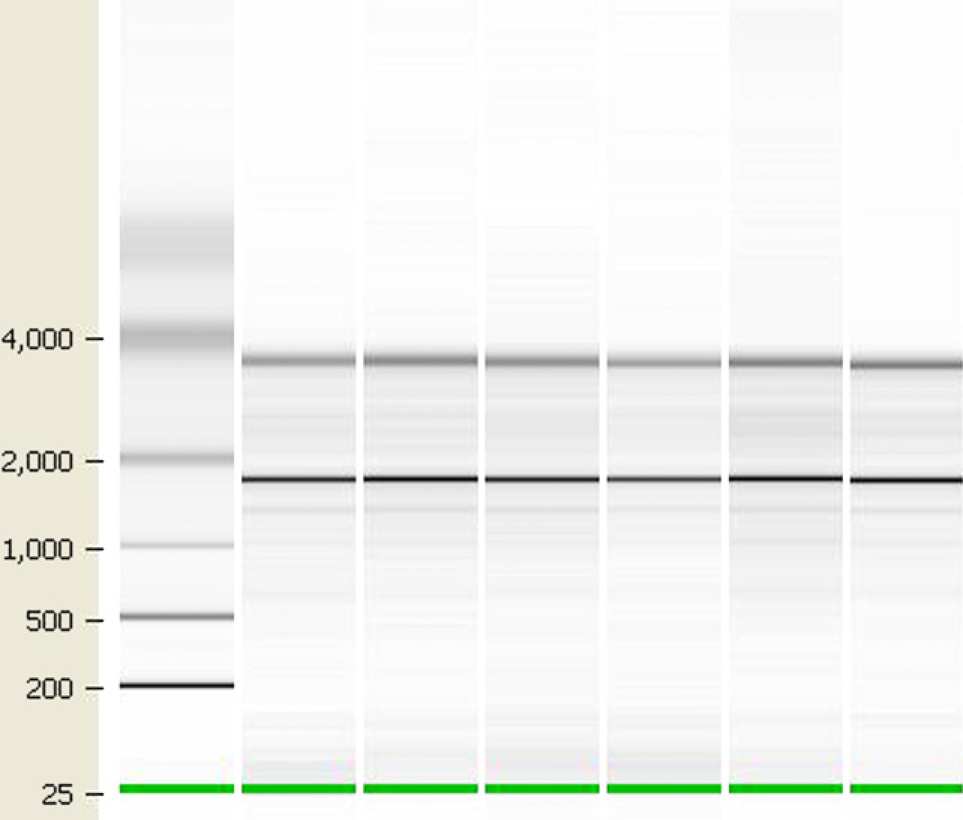
The consistency of RNA recovery and the potential for sample cross contamination are the major concerns in pathogen detection. To examine these issues, we distributed twenty-four 50-μL aliquots of a human blood sample across a 96-well plate, and put water in the remaining wells. We then isolated RNA using the MagMAX-96 Blood RNA Isolation Kit on the KingFisher 96 processor. The purified RNA was used in qRT-PCR targeting the RNA polymerase II mRNA. Amplification of this target was highly consistent: the average threshold cycle (Ct, the PCR cycle when amplification signal becomes detectable above background) was 20.4 ± 0.26 (Fig. 3). No amplification signal was detected in samples from wells that contained water, indicating that no cross contamination had occurred.
Assessment of RNA recovery and cross contamination. A total of 50 μL human whole-blood samples (green) or water (white) were distributed into separate wells of a 96-well plate as indicated in the figure, and processed using the MagMAX-96 Blood RNA Isolation Kit on the Kingfisher 96 processor. The purified RNA was used for quantitative reverse transcription-PCR targeting human RNA polymerase II mRNA, and the average threshold cycle (Ct) from the 24 positive samples is shown.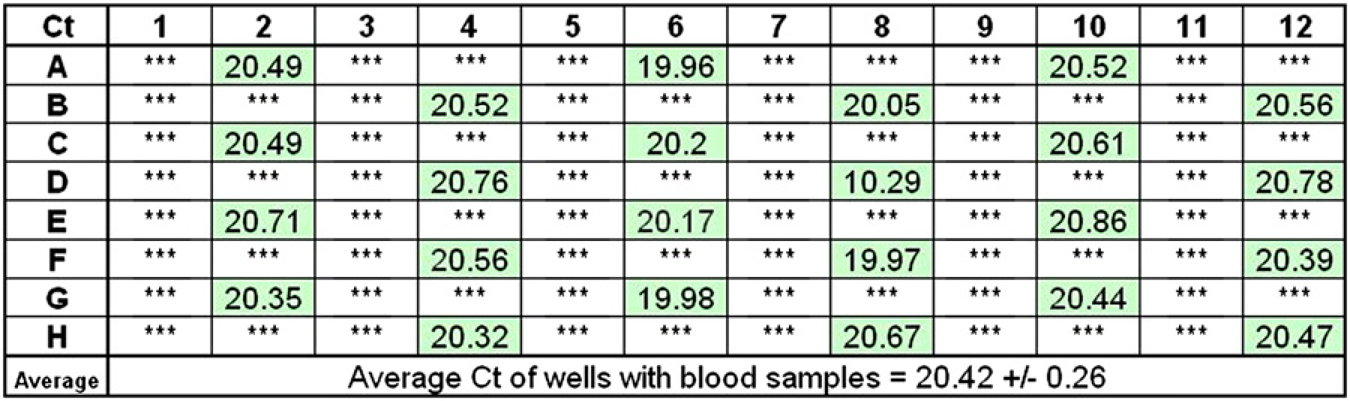
To demonstrate the effectiveness of RNA recovery from blood-borne pathogens, we used the MagMAX-96 Blood RNA Isolation Kit on a KingFisher processor to isolate RNA from blood samples from a healthy cow and a cow persistently infected with Bovine Viral Diarrhea Virus (BVDV). Aliquots of bovine blood from the BVDV-positive cow were distributed to 24 wells across a 96-well plate, and blood samples from the BVDV-negative cow were distributed to the remaining wells of the plate (Fig. 4). XenoRNA-01 Armored RNA was spiked into each sample before lysis to monitor sample processing consistency across the plate. RNA from all of the samples was used in qRT-PCR to detect both BVDV and the XenoRNA-01 internal control sequence. BVDV was detected in all of the wells containing BVDV-positive samples but not in any of the wells with BVDV-negative blood samples, indicating that there was no cross contamination (Fig. 4). Both BVDV and XenoRNA-01 target amplification were highly consistent, with an average Ct of 26.27 ± 0.27 and 31.4 ± 0.24, respectively, across the 96 samples, indicating that sample processing was robust and reliable.
Viral RNA detection with the MagMAX-96 Blood RNA Isolation Kit. A total of 50 μL aliquots of cow blood samples from Bovine Viral Diarrhea Virus (BVDV)-positive (red) and BVDV-negative (gray) animals were distributed into separate wells of a 96-well plate, as indicated in the figure. XenoRNA-01 Armored RNA was spiked into each sample well and the samples were processed using the MagMAX-96 Blood RNA Isolation Kit on a KingFisher processor. The purified RNA was used for quantitative reverse transcription-PCR targeting BVDV RNA and XenoRNA-01 Control RNA to monitor full plate sample processing consistency.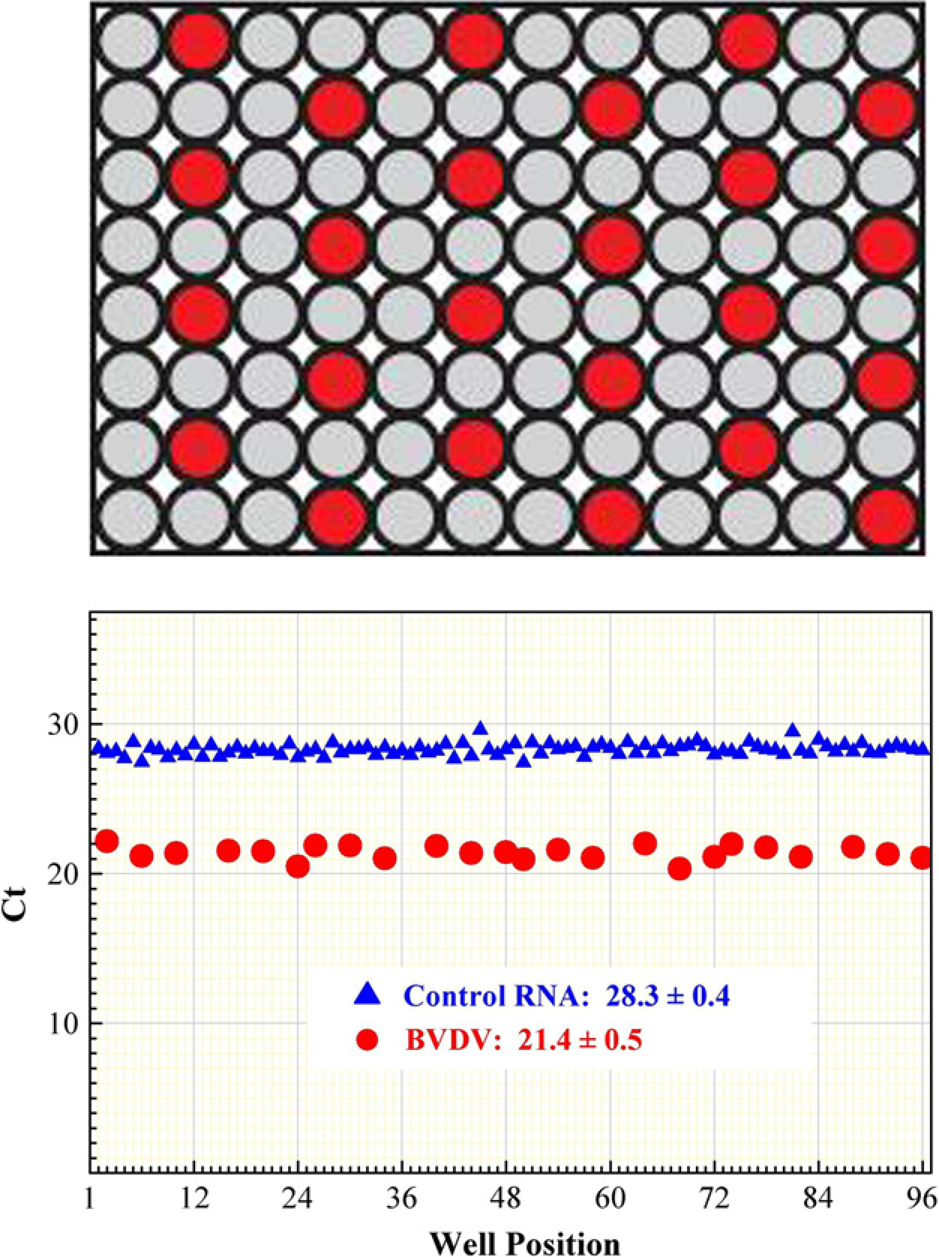
The sensitivity of the MagMAX-96 Blood RNA Isolation Kit for pathogen detection was evaluated by isolating NA from 50 μL human whole-blood samples that contained a serial dilution of XenoRNA-01 Armored RNA control. Samples were processed using the MagMAX-96 Blood RNA Isolation Kit protocol on the KingFisher processor and analyzed by qRT-PCR targeting XenoRNA-01 Control RNA. As few as 20 input copies of XenoRNA-01 Armored RNA were detected, and recovery was linear across a broad input range from 100 to 100,000 copies, indicating that recovery efficiency was consistent (Fig. 5). These capabilities make this technology suitable for sample preparation for detection of most blood-borne pathogens.
Efficiency of recovery of XenoRNA-01 Armored RNA spiked into whole blood. XenoRNA-01 Armored RNA was spiked into human whole blood, and then diluted serially in whole blood that did not contain Armored RNA. After RNA isolation using MagMAX-96 Blood RNA Isolation Kit on the KingFisher, recovery of XenoRNA-01 Control RNA was analyzed by quantitative reverse transcription-PCR using 5 μL of the eluted sample (10% of eluted volume) in a 15 μL reaction.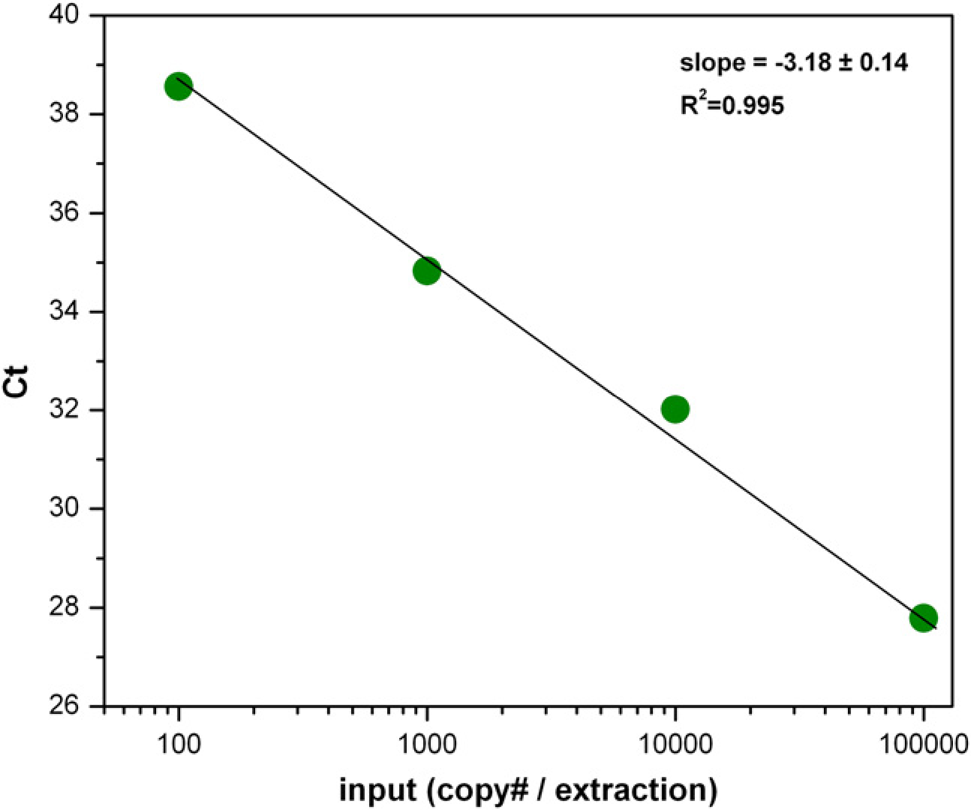
Total RNA Isolation from Cultured Cells and Tissues
The MagMAX-96 Total RNA Isolation Kit is designed for isolating high-quality RNA from cultured cells and small tissue samples. The reagent layout on KingFisher processors for this kit is the same as for the MagMAX-96 Blood RNA Isolation Kit (Table 1). Intact RNA was isolated from HeLa cells, mouse kidney, corn leaf, and corn kernels (Fig. 6, panel A). Typical RNA yields from a variety of sample sources are compiled in Table 3.
Total RNA yield using MagMAX-96 Total RNA Isolation Kit
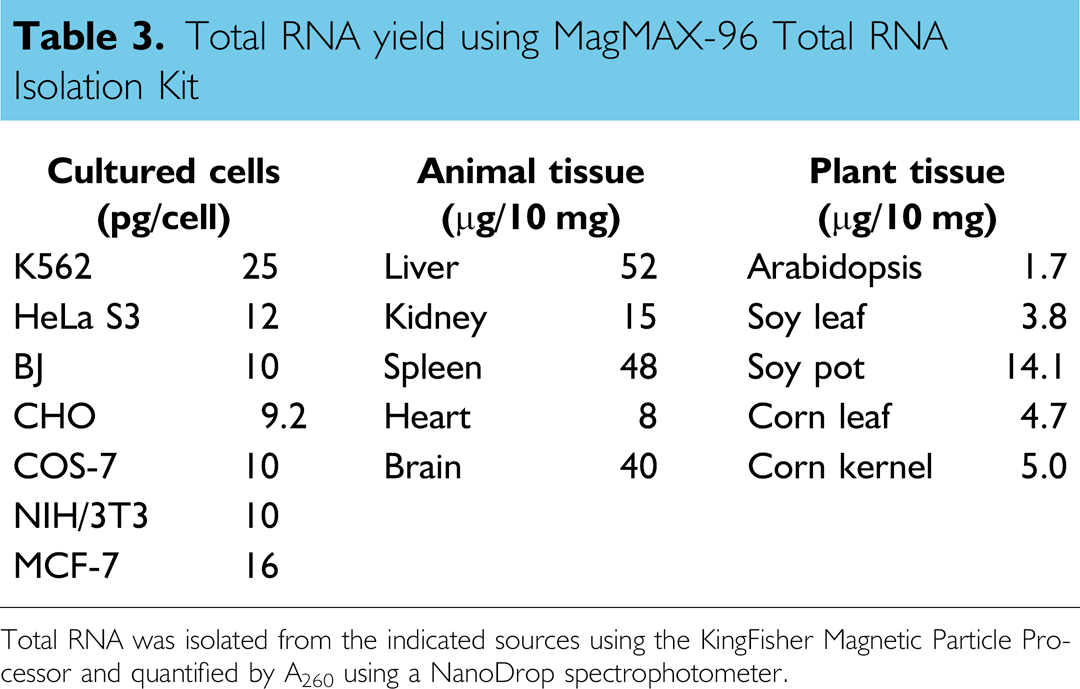
Total RNA was isolated from the indicated sources using the KingFisher Magnetic Particle Processor and quantified by A260 using a NanoDrop spectrophotometer.
Efficiency of recovery and integrity of RNA isolated using MagMAX-96 Total RNA Isolation Kit. (A) Agilent 2100 bioanalyzer profiles of RNA isolated from indicated sources. (B) RNA was isolated from 25 to 2 × 10
6
K562 human cells in eight replicate wells using the MagMAX-96 Total RNA Isolation Kit and a KingFisher processor. Equivalent volumes of the recovered RNA (4% of the eluant) were used for quantitative reverse transcription-PCR (10 μL reaction) targeting human RNA Polymerase II mRNA. The coefficient of variance among Ct values for the eight replicate samples of each cell number input was <3%.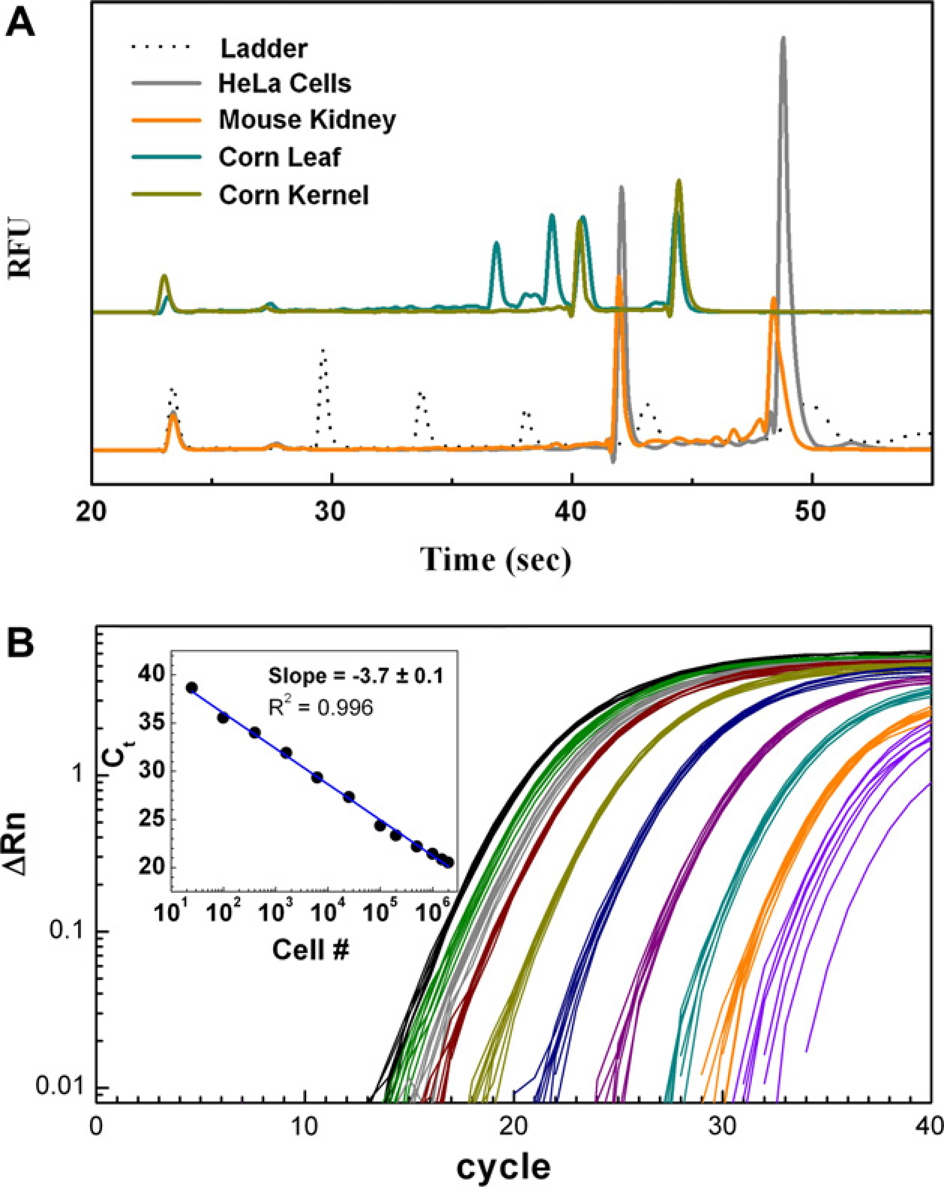
To examine the consistency of RNA recovery and the dynamic range of sample input for this method, human K562 cells were harvested and resuspended in RNAlater Solution to stabilize the RNA. The cells were then serially diluted, and RNA was isolated using the MagMAX-96 Total RNA Isolation Kit on a KingFisher processor. The RNA was used in qRT-PCR targeting human RNA polymerase II mRNA. The recovery of RNA among the replicates was very consistent, with a coefficient of variance of <3% among Ct values for eight replicate samples at each input level. The qRT-PCR analysis also demonstrated that RNA recovery was linear across a dynamic range of sample input from 25 to 2 million cells; Fig. 6, panel B).
Viral RNA/DNA Isolation from Swabs, Plasma, Serum, and Milk
The MagMAX-96 Viral RNA Isolation Kit is designed for viral NA isolation from cell-free or nearly cell-free samples such as swabs, plasma, serum, and milk. It can also be used in some cases to isolate bacterial NA, depending on whether the bacterial cell wall/membrane can be efficiently disrupted by the lysis reagents included in the kit.
To demonstrate consistent recovery of viral RNA and DNA from several different sample matrices, serial dilutions of XenoRNA-01 Armored RNA and Hind III-digested bacteriophage Lambda DNA were added to water, plasma, serum, and milk samples. The MagMAX-96 Viral RNA Isolation Kit reagents and samples were distributed into the wells of 96-well plates as shown in Table 1, and NA was isolated using the KingFisher processor. NA recovery was evaluated using qRT-PCR targeting XenoRNA-01 Control RNA or qPCR targeting Lambda DNA. Both RNA and DNA were recovered at >75% efficiency. Using 16% of the recovered NA in a qRT-PCR, we could easily detect as few as 20 input copies of XenoRNA-01 Armored RNA in all of the sample types tested (Fig. 7).
Recovery efficiency and detection limit of RNA and DNA isolated from biological fluids using the MagMAX-96 viral RNA Isolation Kit. Serial dilutions of HindIII-digested Lambda DNA and XenoRNA-01 Armored RNA were spiked into water, plasma, serum, and milk samples. After isolating nucleic acids with the MagMAX-96 Viral RNA Isolation Kit, the recovery of DNA (A) and RNA (B) was analyzed by qPCR and quantitative reverse transcription-PCR, respectively, using 8 μL of the eluted sample (16% of eluted volume) in a 15 μL reaction.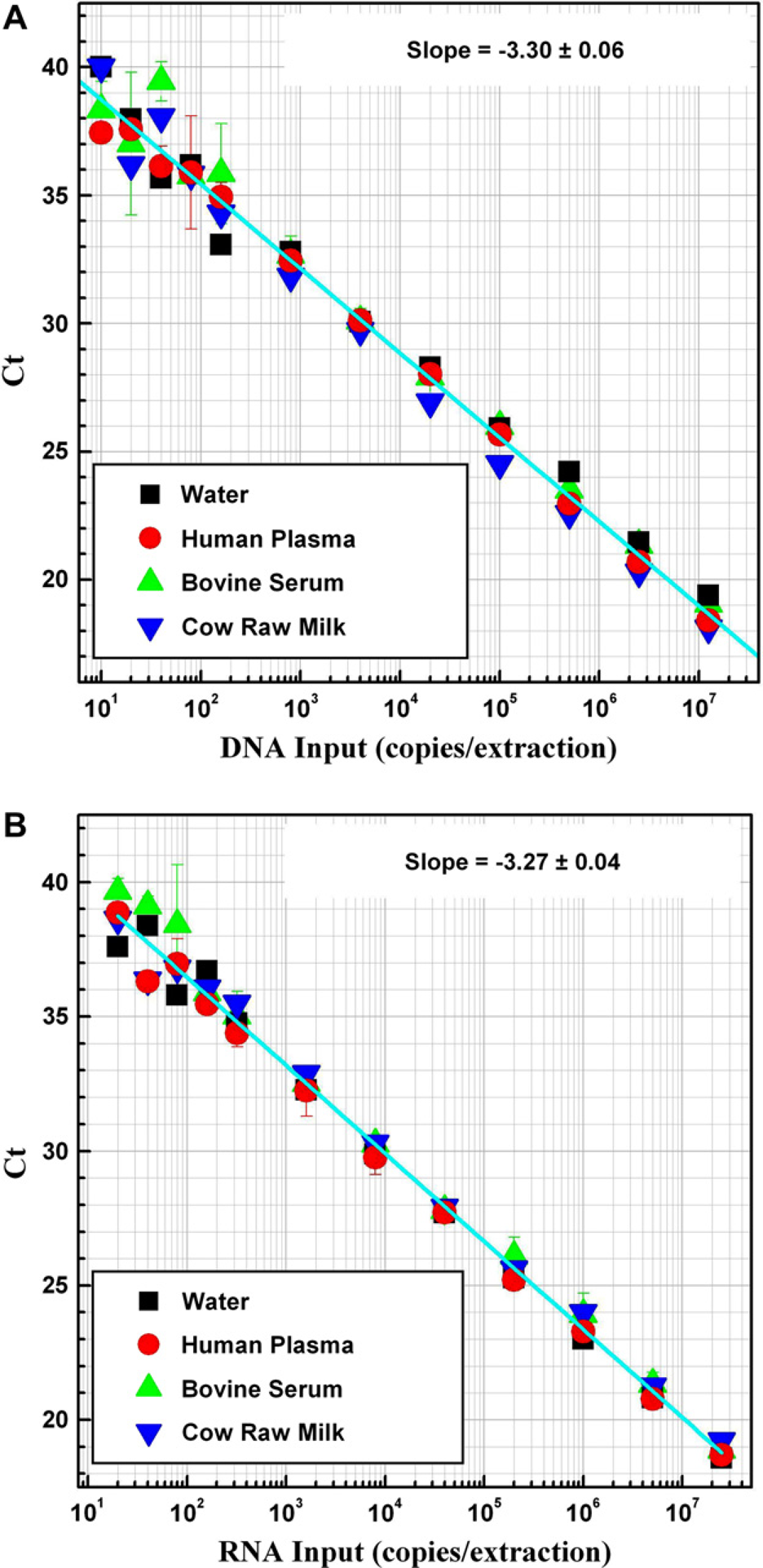
The MagMAX-96 Viral RNA Isolation Kit is widely used for viral pathogen detection, but because most animal diagnostic laboratories do not use automation, we compared results using the MagMAX-96 Viral RNA Isolation Kit manually, with multichannel pipettors, or robotically on the KingFisher processor. RNA was isolated from human plasma samples containing a serial dilution of XenoRNA-01 Armored RNA. The manual RNA isolation process took ~35 min, whereas the KingFisher protocol took 15 min. As monitored by qRT-PCR, recovery was equally efficient for both methods at all control RNA input levels (Fig. 8).
Comparison of manual versus automatic sample processing. XenoRNA-01 Armored RNA was serially diluted into human plasma, and RNA was isolated using the MagMAX -96 Viral RNA Isolation Kit, processing samples either manually or on the KingFisher processor. Equivalent portions (16%) of the eluted volume were analyzed by quantitative reverse transcription-PCR targeting XenoRNA-01 Control RNA.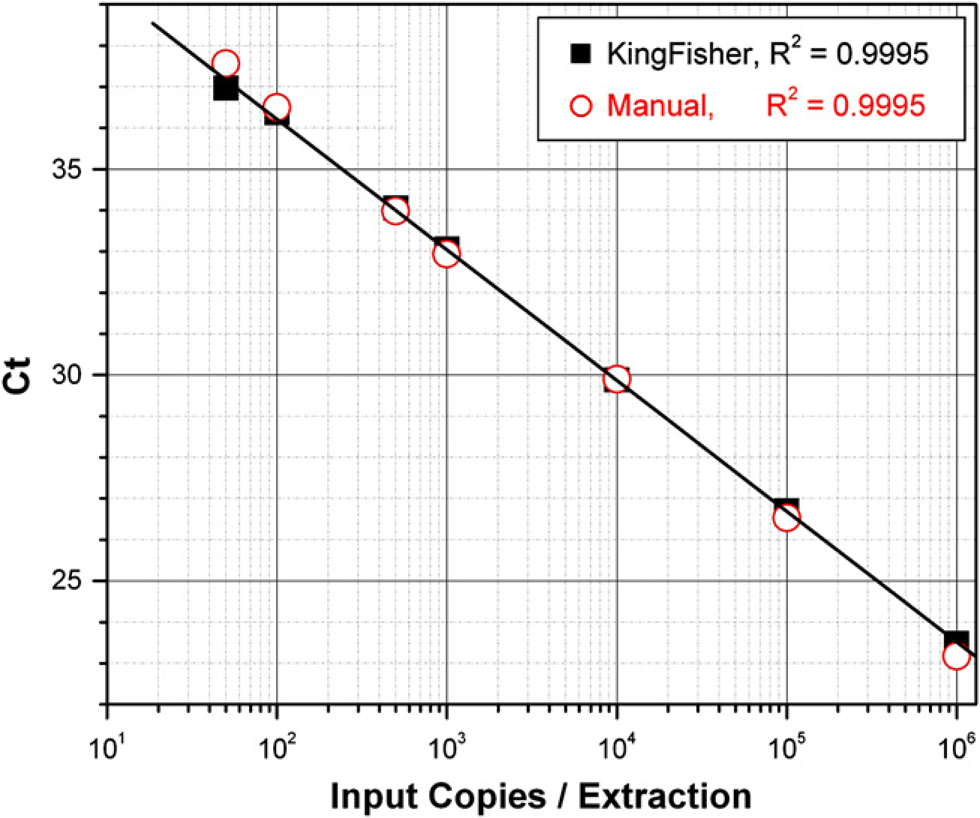
Summary and Conclusion
MagMAX magnetic-bead-based NA isolation methods are very robust and can be used with a wide variety of sample matrices. The isolated NA is pure and intact, making it suitable for a variety of downstream applications such as pathogen detection, gene expression profiling, and microarray analysis. The MagMAX methods are specifically designed for high-throughput processing.
KingFisher Magnetic Particle Processors automate magnetic-bead-based processes with the coordinated move of the permanent magnetic rods and disposable tip combs. Captured magnetic beads are lifted out of solution, minimizing reagent carryover from one step of a protocol to the next. In contrast, automated liquid handling systems aspirate liquid away from captured magnetic beads, a process that invariably leaves some residual liquid in the well. The unique KingFisher processor approach makes washing and elution more efficient, accelerates processing, and also makes walk-away automation feasible. At a fraction of the cost of automated liquid handling systems, KingFisher Magnetic Particle Processors provide a quick, efficient, and reliable platform for NA isolation using Ambion MagMAX NA isolation products.
Footnotes
Acknowledgments
We thank Lisa Albright and Lori Martin for editing the manuscript.
