Abstract

OVERVIEW
Automated isolation of total RNA from cultured cells was accomplished by adapting Ambion's solid-phase total RNA isolation kit, RNAqueous-96, for use with the MultiPROBE II liquid handling system from Packard Instruments.
Cultured cells (HeLa S3 adherent cells and K562 cells in suspension) were lysed, DNase treated, washed, and then eluted from the 96-well filter plate.
Total RNA recovered was analyzed for yield, intactness, purity, and ability to serve as a substrate for real-time RT-PCR.
INTRODUCTION
Methods for high-throughput isolation of total RNA are required to maximize the information that can be obtained from assays such as microarray expression profiling and quantitative RT-PCR. The present study was undertaken to evaluate the use of Ambion's solid-phase total RNA isolation kit, RNAqueous-96, with the Packard MultiPROBE-II HT liquid handling system. The total RNA recovered from human cell lines was analyzed by several methods to check for quality and yield of the samples. RNA integrity was assessed on a denaturing agarose gel, and realtime RT-PCR was used to determine consistency of expression levels between high and low abundance genes within a sample set. Results indicate this system provides a robust method for the automated preparation of high-quality total RNA for use in the quantitative analysis of target mRNAs.
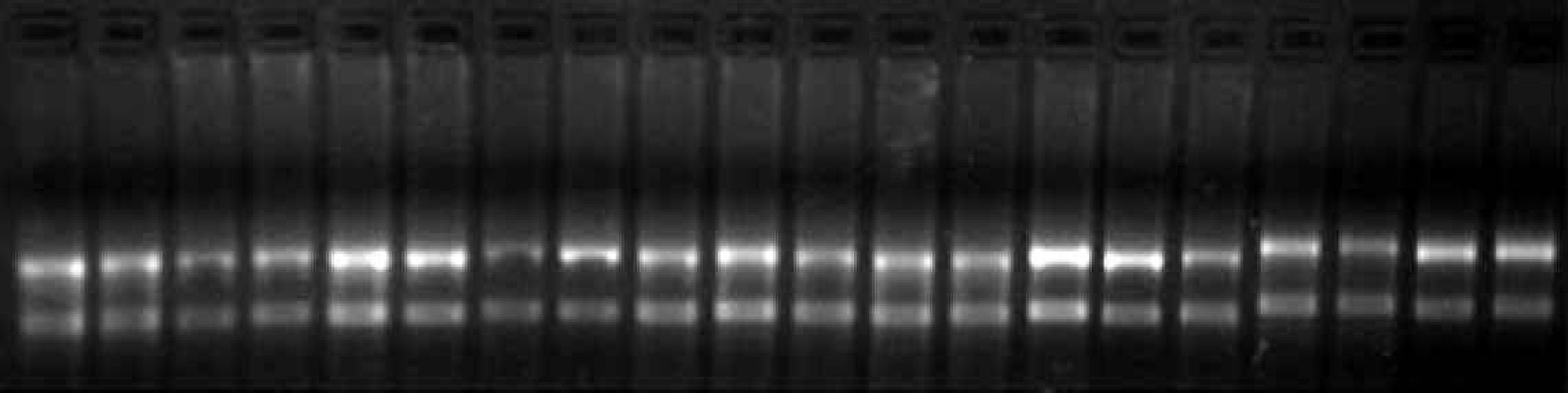
Analysis of total RNA isolated using the RNAqueousTM-96 kit with the MultiPROBE II® robotic liquid handling system. Total RNA was purified from 80,000 HeLaS3 cells using the protocol for automated RNA isolation described. One-fourth of each sample was mixed with gel-loading solution containing ethidium bromide, denatured by heating for five minutes at 65°C, and separated by electrophoresis at 80 volts on a 1% agarose/formaldehyde gel.
EQUIPMENT AND MATERIALS
MultiPROBE II HT with Gripper Integration Platform
Eight probe VariSpan™ positive displacement pipetting using VersaTip™ pipet tips
Gripper relies on Gantry-based system with five axes of motion
Syringe pumps used for aspiration of liquids
Peristaltic pump used for fast and rigorous washing of pipet tips
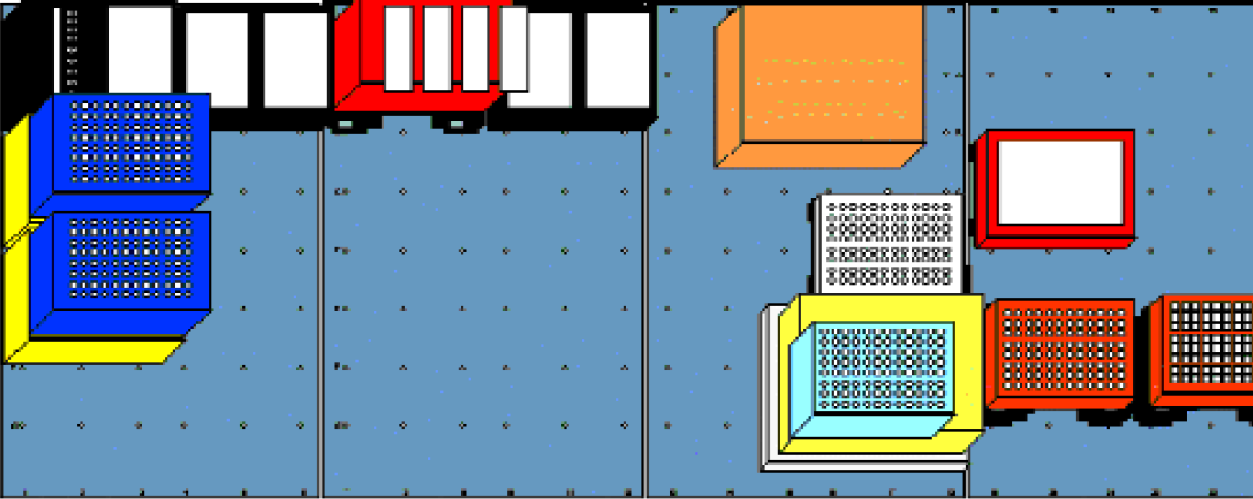
Three-dimensional deck layout of the Multiprobe II HT adapted for RNAqueous-96TM Protocol
RNAQUEOUS-96™
Phenol-free RNA isolation technology based on RNA binding to silica matrix
Kit includes reagents and supplies to perform 192 isolations
Reagents for on-the-filter DNase treatment included
RNA obtained is intact and free of contaminating genomic DNA
METHODS
PREPARATION OF CELLS
HeLa S3 and K562 cells were cultured in DME/10% FBS in 5% CO2. Before use, the K562 cells (human leukemia line, grown in suspension) were pelleted and re-suspended in 1X PBS, counted, and distributed manually into a 96-well deep well plate at ∼3.75×105 cells/well, then 150 μl of Lysis Solution from the RNAqueous-96 kit was added to each well robotically. The HeLa S3 cells (human cervical carcinoma line, adherent) were washed by removing the culture media and flooding the plate with 1X PBS. The PBS was removed and ∼8×104 cells per well were lysed by adding Lysis Solution from the RNAqueous-96 kit directly to each well in the culture plate robotically.
AUTOMATED RNA ISOLATION PROTOCOL
150 μL of Lysis Solution is added to cells in each well
An equal volume of 64% ethanol is added to the cell lysate in each well, and the samples are mixed and transferred to the filter plate
Vacuum suction (20 mm Hg for three minutes) is applied tothe filter plate. The liquid passes through the filter while the RNA remains bound to the silica matrix
400 μL of Wash Solution A is dispensed to each well, and filtered by applying vacuum at 20 mm Hg for two minutes
40 μL of DNase I solution containing 100 units of RNasefree DNase I in 1X buffer is added to each well, and the plate is incubated for 20 minutes at room temperature
400 μL of Wash Solution B is dispensed to each well and filtered by applying vacuum as above
400 μL of Wash Solution C/D is dispensed to each well and filtered by applying vacuum as above
400 μL of 100% ethanol is dispensed to each well and filtered by applying vacuum as above
The filter plate is then moved to a drying heat block at 42°C for three minutes to remove any residual wash solution left on the bottom of the filter plate. The heat block is covered with fresh perforated absorbent material which allows the filter plate to dry without the drip directors contacting the heat block to avoid RNase contamination
A Spacer and a Collection Plate are moved to the inside of the vacuum manifold and then the filter plate is returned to the top of the vacuum manifold
80 μL of Elution Buffer is added to each sample and allowed to stay on the filter for two minutes. The RNA is then collected by vacuum filtration at 10 mm Hg for 30 seconds followed by 20 mm Hg for two minutes
(optional) User manually disassembles the vacuum manifold and checks for any fluid remaining on the drip directors of the filter plate. If fluid is observed, the user may centrifuge the filter plate/collection plate assembly to recover residual sample.
REAL-TIME RT-PCR
The following PCR profile was used with the ABI 7700:
42° C for 15 minutes (reverse transcription; MMLV 100 units/reaction, Ambion)
94° C for 3 minutes (denaturation)
40 cycles: 94° C for 20 seconds
60° C for 40 seconds (Supertaq™ Polymerase, five units/reaction, Ambion)
Selected RNA samples from the filter plate were run on the Agilent Bioanalyzer 2100 for quality and quantity (data not shown), then half of the samples (48 samples, rows D-G) were used for realtime RT-PCR to detect both GAPDH (glyceraldehyde phosphate dehydrogenase) and hTERT (reverse transcriptase subunit of human telomerase) in each sample, in separate amplification reactions. 5 μL of each sample was used in each reverse transcription reaction.
RESULTS AND ANALYSIS
The Ct value corresponds to the x-axis and indicates the threshold cycle where PCR product amplification surpasses a selected “critical value”. The Ct value corresponds to the amount of RNA originally present in the sample. The standard deviations for the sample sets indicate that the quantity of RNA was approximately the same for each sample. Relative fluorescence of the samples is represented by the y-axis.
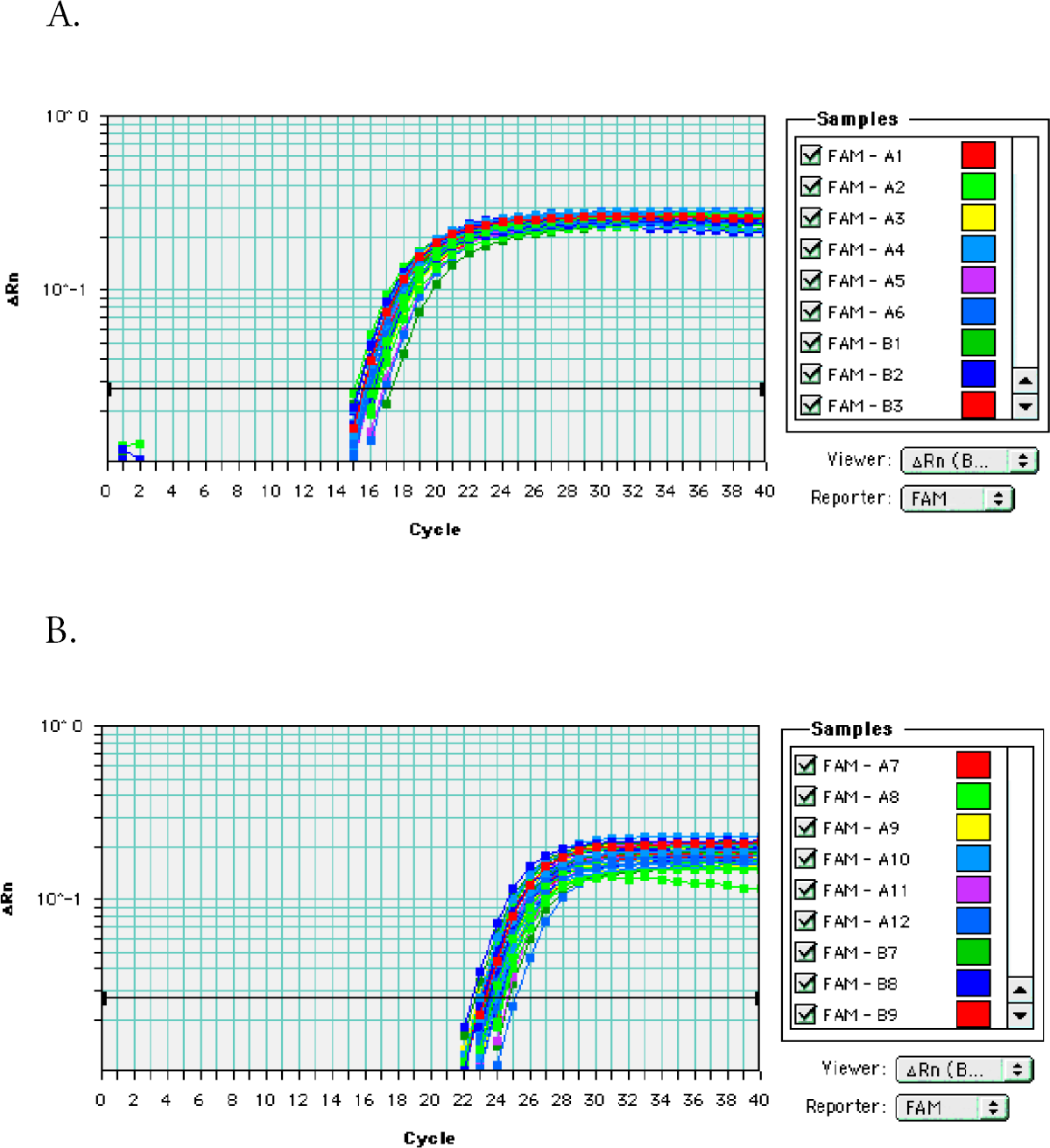
Detection of GAPDH (A) and hTERT (B) mRNA targets using real-time RT-PCR. Amplification products were detected using dual-labeled fluorogenic probes (FAM/TAMRA). The 48 RNA samples were isolated from K562 cells by using the RNAqueous-96 automated protocol described. Details of the PCR step are as described. Different colored lines indicate different samples.
K562 cells:
average GAPDH Ct 15.86, standard deviation 0.45
average hTERT Ct 23.63, standard deviation 0.54
average of Ct ratios hTERT/GAPDH 1.49, standard deviation 0.03, CV of ratios 1.79%
(real-time data for HeLa cells not shown) HeLa cells:
average GAPDH Ct 16.18, standard deviation 0.80
average hTERT Ct 26.00, standard deviation 1.01
average of Ct ratios hTERT/GAPDH 1.61, standard deviation 0.03, CV of ratios 2.1%
CONCLUSIONS
The PerkinElmer MultiPROBE-II HT liquid handling system was evaluated for high-throughput isolation of total RNA in a 96-well microtiter plate format using the RNAqueous-96 kit from Ambion, Inc. Run time required to complete the procedure was approximately one hour and twenty minutes, including the on-the-filter DNase treatment used to eliminate contaminating genomic DNA. Yield of total RNA from HeLa S3 cells was ∼5 pg per cell, and for K562 cells ∼8 pg per cell, based on Agilent 2100 Bioanalyzer analysis (data not shown). The total RNA recovered is intact as judged by denaturing agarose gel electrophoresis and microfluidic capillary electrophoresis. Real-time RT-PCR using dual labeled fluorescent probes shows highly reproducible absolute Ct values and ratios of Ct values for endogenous mRNA targets of high and low abundance. The PerkinElmer MultiPROBE-II HT liquid handling system provides an efficient method for automating the isolation of total RNA in a 96-well microtiter plate format, using the RNAqueous-96 kit from Ambion, Inc.
