Abstract
Here we describe a reagent system and robotic methods for the purification of PCR(a) fragments from other contaminating amplification reaction components. The methods use the MagneSil™ paramagnetic particle chemistry(b) to isolate double stranded DNA fragments from 150bp to 23kbp. Purified fragments are eluted in water ready for downstream applications such as cloning, fluorescent DNA sequencing and microarray printing. This method has been adapted to a number of liquid handling robotic platforms, including the Biomek® FX and Biomek® 2000 Laboratory Automation Workstations, in both 96 and 384-well formats.
Introduction
Many post-PCR applications require removal of excess nucleotides, salts, primers and small, non-targeted amplification products, like primer dimers from the PCR product. Contaminating primers and primer dimers will compete with the target PCR product for binding sites in microarray printing. Residual primers can also be extended in fluorescent DNA sequencing reactions, causing increased background.
Common methods for the purification of PCR products include precipitation with ethanol or isopropanol and gel filtration chromatography. These procedures are problematic for high throughput processing in that they require (multiple) centrifugation or vacuum steps. The MagneSil™ chemistry offers a high throughput alternative to these traditional methods. MagneSil™ is a silica-paramagnetic particle that acts as a mobile solid phase to capture double stranded DNA fragments using solution kinetics. The particle-adherent target complex is washed to remove residual contaminants and the purified DNA is eluted in water, ready for downstream cloning, fluorescent DNA sequencing and microarray applications.
The MagneSil™ system is scalable, allowing for scale-up for higher yield and scale-down for use in 384-well formats. PCR products greater than 150bp in size are selectively isolated from primers and primer dimers with high yields and purity. This technology has been adapted for use with a number of robotic workstations including the Biomek® FX and Biomek® 2000 Laboratory Automation Workstations. The plate processing times are less than 30 minutes with a single POD Biomek® FX. Typical recovery is greater than 90% for fragments above 300bp with negligible carryover of primers or nucleotides.
Two methods have been developed to purify either a single plate or multiple plates of PCR samples. The multi-plate method that optimizes usage of available deck positions allows for parallel processing of four plates before it is necessary to repopulate the deck. Cycle time for the maximized deck method is not significantly increased over the single-plate method. Higher order processing is achieved by performing iterations of the maximized deck method with an associated high capacity stacker carousel to deliver consumables.
Methods and Materials
PCR Amplification: Reactions (100ul) contained 20ng plasmid template, 2uM primers (fwd: 5'-GATGTGCTGCAAGGCGAT-TAA-3', rev: 5'-TAGCACCGCCTACATACCTCG-3'), and 50μl of 2X PCR Master Mix(c) (Promega). GeneAmp system 480 Thermal Cycler conditions were: initial two minute melt at 97°C, 30 cycles of 97°C, one minute, 64°C, two minute, 72°C, two minute, followed by 10 minute at 72°C, then storage at 4°C.
Gel Quantitation: Samples were run on 0.1% Agarose (TBE) gels and stained with ethidium bromide. Gels were scanned with a FluorImager® and individual bands were quantitated using the ImageQuant Software.
Fluorescent DNA Sequencing: Purified PCR products were stored at 4°C prior to shipment to a custom sequencing facility for analysis on an ABI Prism® 3700 DNA Sequencer. The sequencing primer was the −21M13 forward
(5'-TGTAAAACGACGGCCAGT-3'). Run parameters were: Injection Voltage: 1000V, Injection Time: 50 seconds, Post Injection Voltage: 5250V, Step Interval: 50V, Interval Time: 40 seconds. POP-6 polymer was used.
Microarray Analysis: Probe RT-PCR products and target cDNAs were prepared using the ImProm-II™ Reverse Transcription System according to the manufacturers instructions. For microarray printing, PCR products were purified using the Wizard® MagneSil™ PCR Clean-Up System, dried down in a SpeedVac® concentrator and resuspended in 6ul of 3x SSC. PCR products were arrayed on poly-L-lysine-coated slides using a Gene Machines® OmniGrid™ arrayer. Cy®3- and Cy®5-dUTP were incorporated into target in separate reactions. Labeled targets were purified and concentrated using Amicon® Microcon® microfilters. Concentrated target was resuspended in 25μl of hybridization buffer consisting of Perfect Hyb, salmon sperm DNA and yeast tRNA. Separate arrays were hybridized for 16 hours at 60°C with Cy®3- and Cy®5-labeled cDNA. After hybridization, arrays were washed following the Eisen and Brown method. Arrays were scanned on a Packard Biochip Technologies ScanArray® 5000. The scanned images were analyzed using QuantArray® Microarray Analysis Software.
Robotic methods described can be obtained from the Beckman Coulter eLab Notebook at www.beckmancoulter.com/resourcecenter, or by contacting Promega Technical Services at www.promega.com/techserv.
THE PCR FRAGMENT CLEAN-UP PROCESS
The Wizard® MagneSil™ PCR Clean-Up System uses a particle type designed for high recovery of double stranded DNA fragments from 150bp to 23kbp. This formulation has been optimized to selectively adsorb DNA fragments from the amplification reaction mix (Figure 1). After capture, an external magnetic field is applied to localize the particle and adherent sequencing extension products at the side of the well, and non-bound material is removed to waste. Residual contaminants are eliminated with a series of particle washes and purified target is eluted in water, ready for downstream applications.
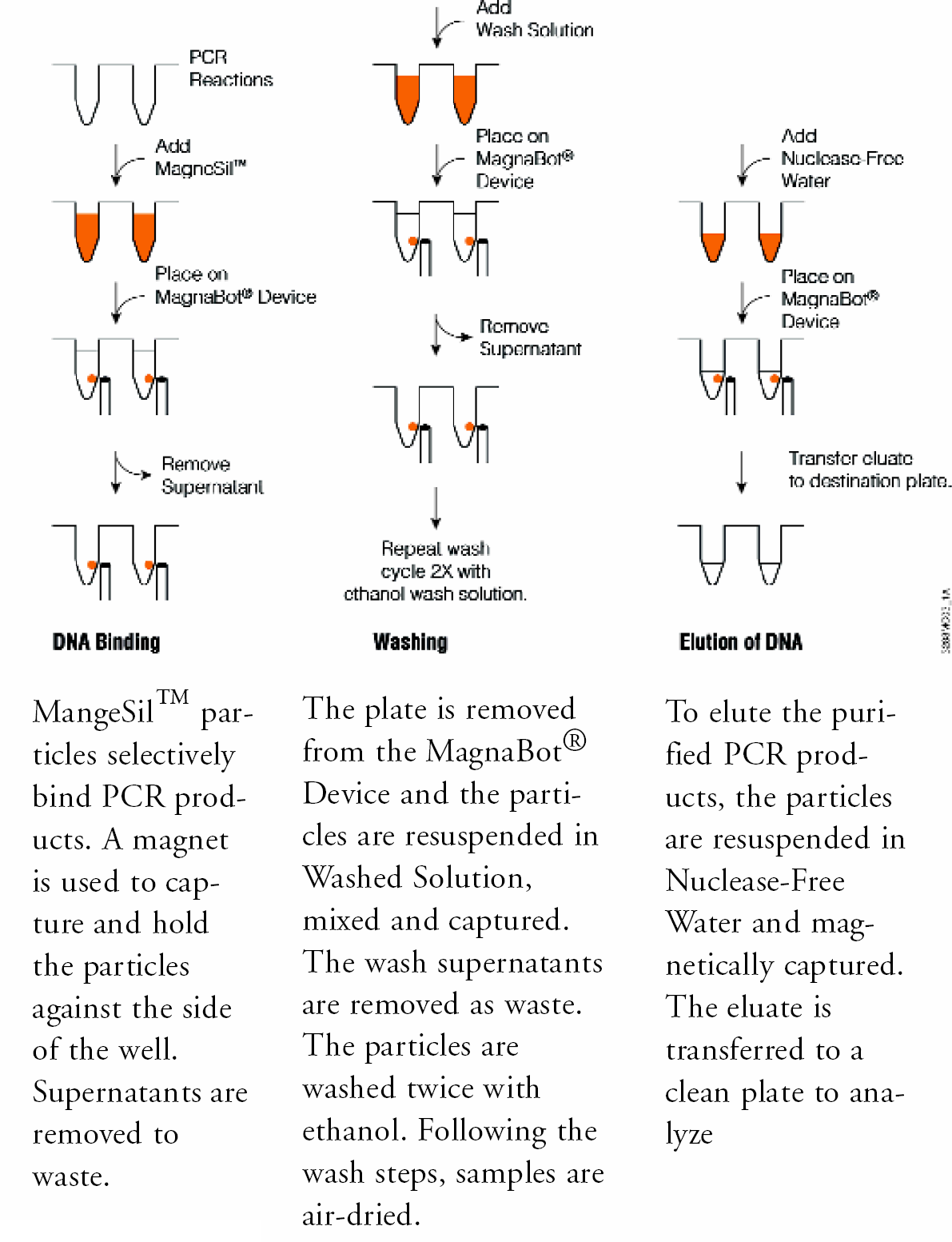
MAGNETIC ARRAY HARDWARE
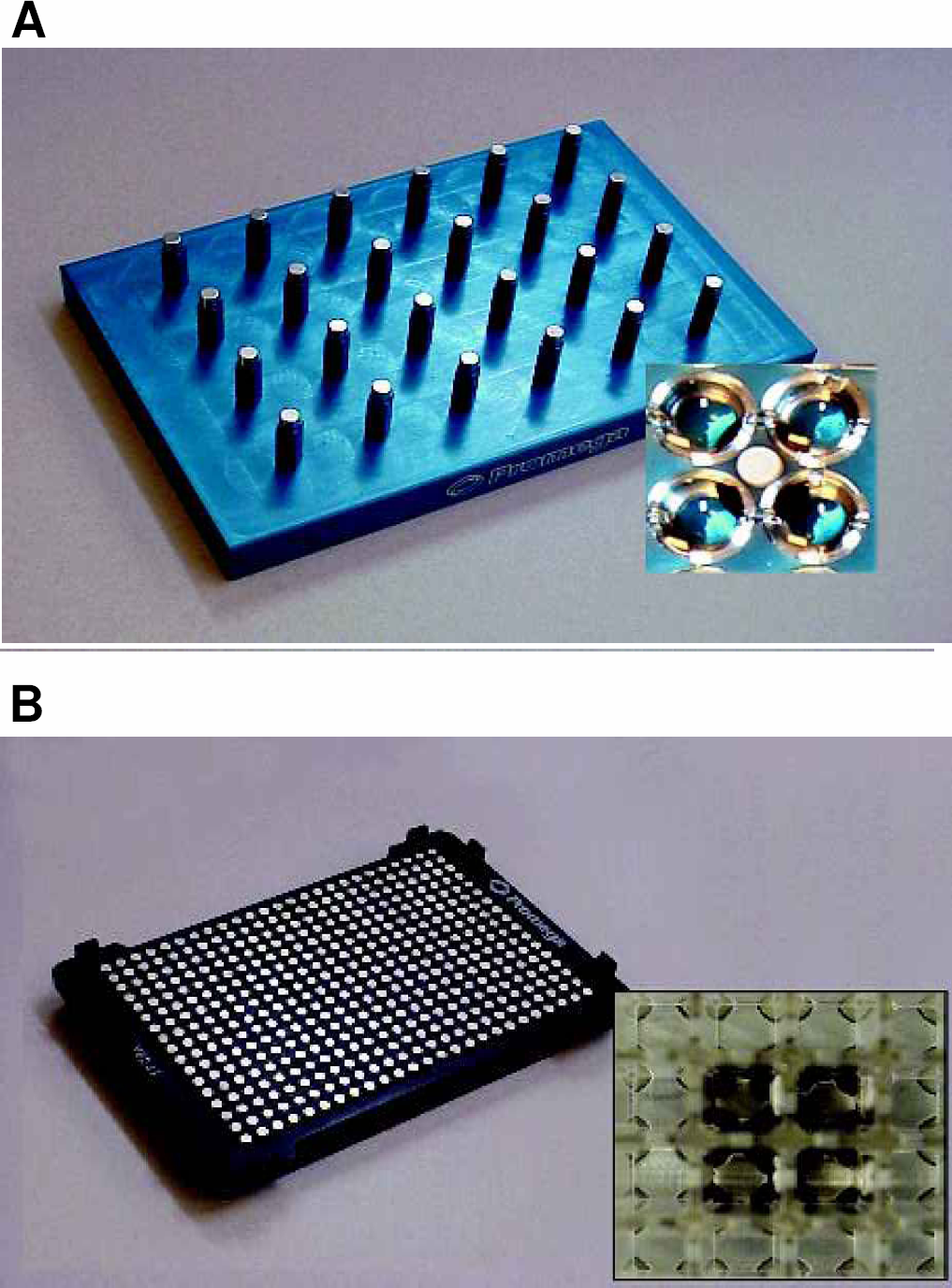
Custom magnetic arrays have been developed for high-throughput processing in 96- and 384-well plate formats (Figure 2). These magnetic stands incorporate S36-grade neodymium rare earth magnets. More than 90% of the MagneSil™ particles are cleared from solution in fewer than 15 seconds (5). These devices have an SBS (Society for Biomolecular Screening) standard footprint to allow for maximum usefulness among liquid handling workstations. The magnets are orientated such that the MagneSil™ particles are captured to the side of each well. This allows quantitative removal of liquids with minimal particle loss.

The plate is removed from the MagnaBot® Device and the particles are resuspended in Wash Solution, mixed and captured. The wash supernatants are removed as waste. The particles are washed twice with ethanol. Following the wash steps, samples are air-dried.
AUTOMATED METHODS
The MagneSil™ purification process has been adapted to a number of robotic platforms including the Beckman Biomek® FX (Figure 3). Maximizing usage of available deck positions with a single POD (processing arm) instrument configuration allows for parallel processing of four plates (Figure 4). Each four-plate run cycle takes approximately 45 minutes before re-populating the deck is necessary.

RDual POD Beckman Biomek® FX Laboratory Automation Workstation.
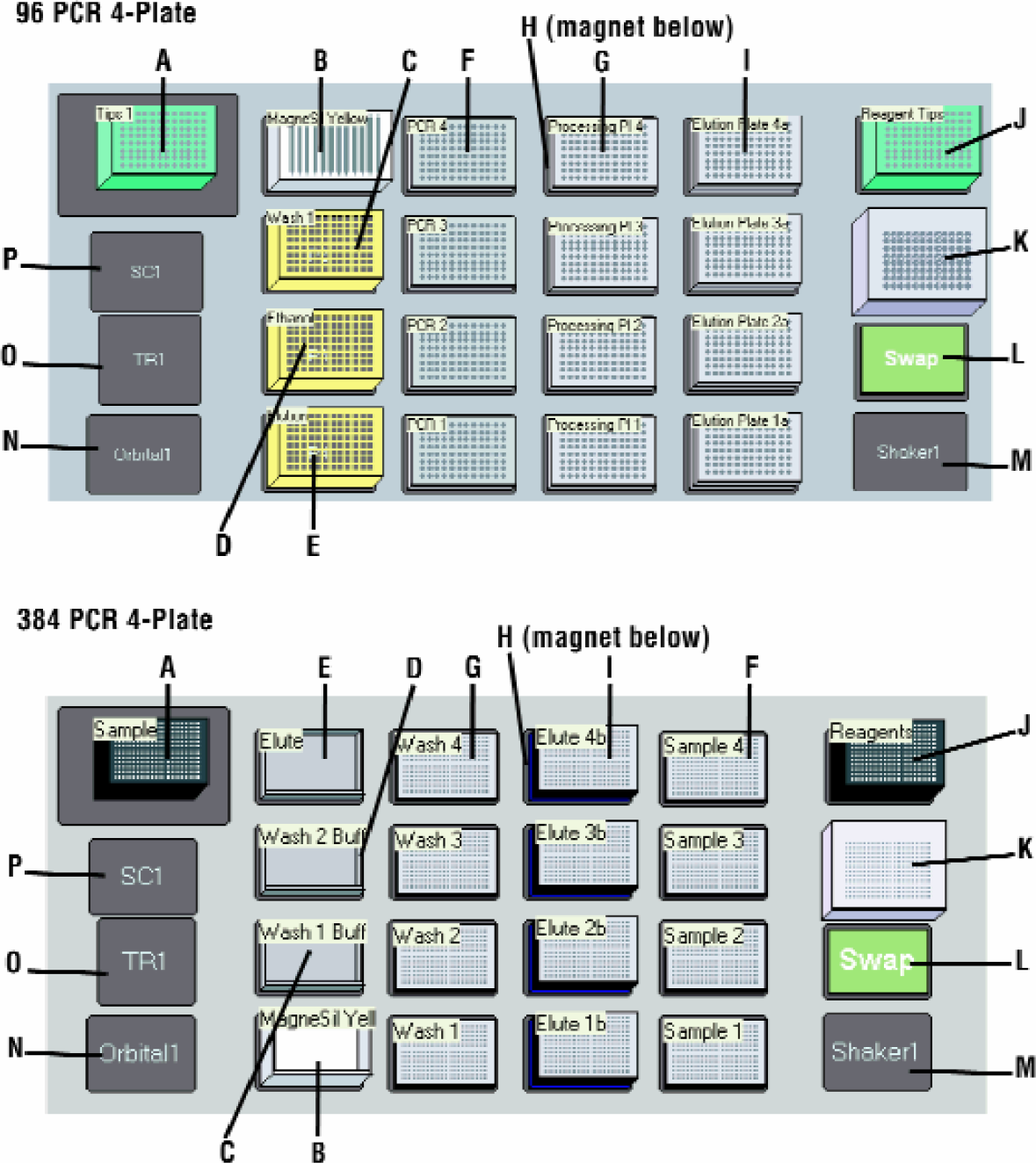
Biomek® FX Deck Configuration for 4-Plate Processing Methods. Panel A: 96-Well Panel B: 384-Well. (A) Sample processing tips, (B) MagneSil™ Yellow, (C) Wash 1, (D) EtOH Wash, (E) Water, (F) Sample Plates, (G) Processing Plate on (H) MagnaBot® 96, (I) 2x Elution Plates, (J) Reagent Tips, (K) Tip Wash ALP, (L) Swap Position, (M) Open 1×1 ALP, (N) Orbital Shaker ALP, (O) Trash ALP, (P) Stacker Carousel.
The following sequence of steps forms the basis for both 96 and 384-well robotic methods. Input sample volume is scaleable from 25–100ul for the 96-well method and 10–20μl for the 384-well format.
BIND
Resuspend and add 1 part MagneSil™ particle slurry to 1 part PCR reaction sample.
Incubate 4 minutes at room temperature. Mix by pipeting up and down four times at 0 and 2 minutes.
Resuspend and transfer slurry to a processing microtiter plate.
Place onto MagnaBot® magnetic separation device to capture the particles.
Remove and discard the supernatant.
WASH
Remove the plate from the MagnaBot® device.
Add 2 volumes Wash 1 solution.
Incubate 4 minutes at room temperature. Mix by pipeting up and down four times at 0 and 2 minutes.
Place onto MagnaBot® magnetic separation device to capture the particles.
Remove and discard the supernatant.
Remove the plate from the MagnaBot® device.
Add 1 volume ethanol wash solution.
Repeat steps 8–10 for a total of 2 ethanol washes.
Remove and discard the liquid.
Incubate 5–10 minutes to dry particles.
ELUTE
Remove the plate from the MagnaBot® device.
Add 1 volume of water. Mix by pipeting up and down four times at 0 minutes.
Incubate 2 minutes at room temperature.
Place onto MagnaBot® magnetic separation device to capture the particles.
Transfer the liquid containing purified PCR products to a collection plate.
Seal plate and store at 4°C (1–2 days) or −20°C
FRAGMENT PURITY
Removal of primer and primer dimer were assessed by agarose gel electrophoresis (Figure 5). Purified samples were run and analyzed as described in Methods. Primers and nonspecific amplification products smaller than 100bp are quantitatively removed (greater than 98% removal of primer / primer dimer). A band with an apparent size of approximately 100bp is visible in all un-purified samples (lanes 2–6) and not in the MagneSil™ purified samples (lanes 7–14).
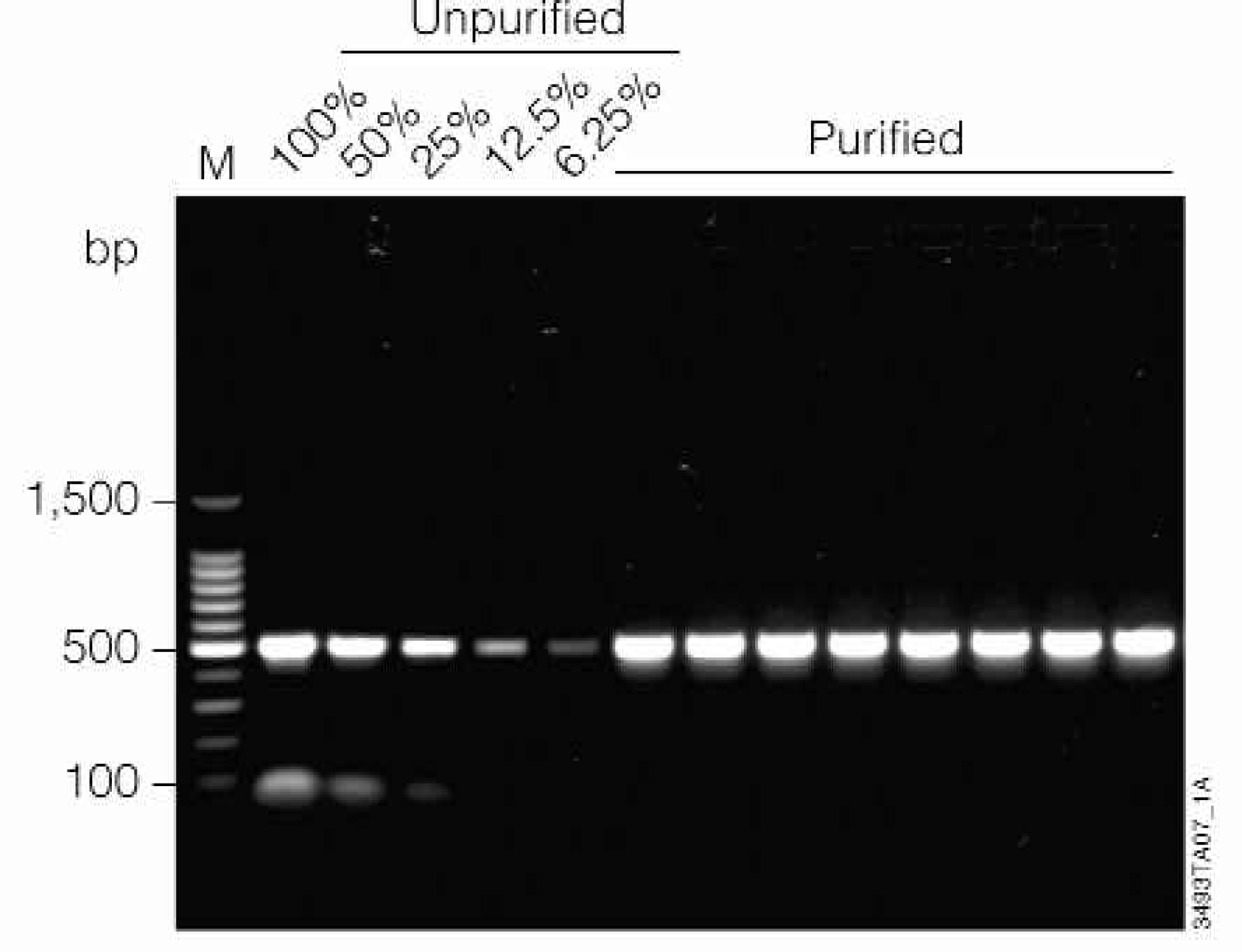
Removal of primer dimers from PCR products. Lanes 2–6: A titration of unpurified 506bp prothrombin PCR product is shown (100%, 50%, 25%, 12.5%, and 6.25% of the input volume used for MagneSil™ purification). Lanes 7–14: Eight replicates purified using the MagneSil™ PCR Clean-Up System. Lane 1: 100bp DNA Ladder (Promega).
FRAGMENT YIELD
PCR products in the range 300–1,500bp were generated as describe in methods and purified using the 1-plate 96 well MagneSil™ method for the Biomek® FX (Figure 6). Recovery was assessed by densitometric scan of MagneSil™ purified samples relative to unpurified amplification reactions after separation by agarose gel electrophoresis as described in Methods. PCR products in the range 300–1500bp were generated as described in Methods and purified using the 1-plate 96-well MagneSil™ method for the Biomek® FX (Figure 6). Recovery is greater than 90% for all size fragments greater than 300bp. Recovery is size dependent for smaller fragments (Figure 7). Results are unchanged when identical samples are purified using the 384-well method.
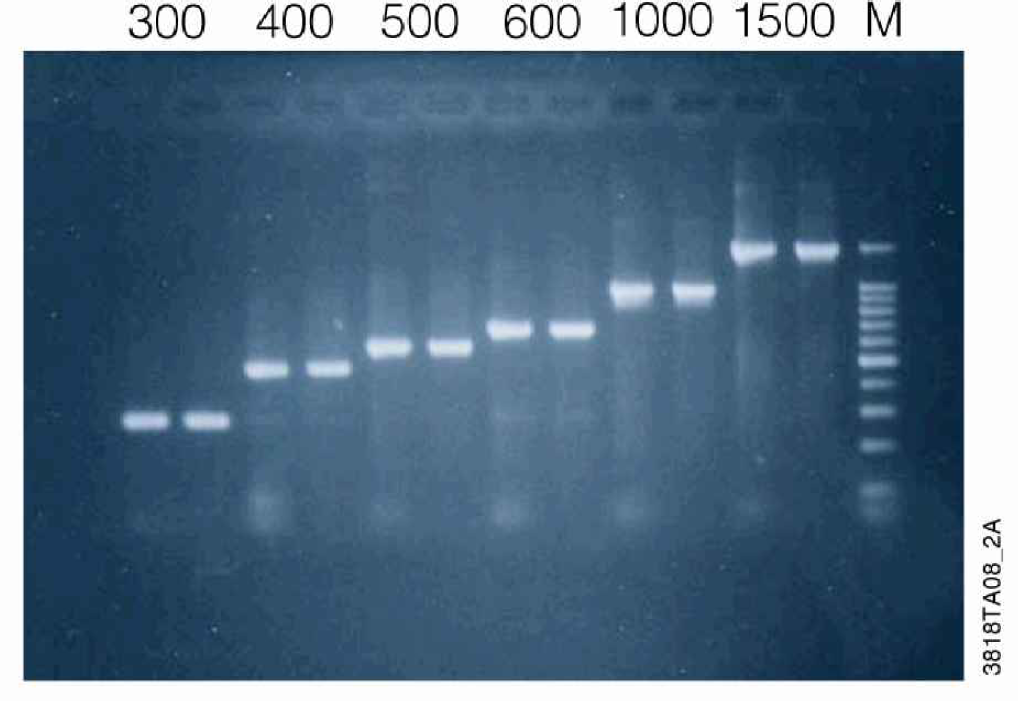
Fragment Purity. A series of modified pGEM®-luc vectors were constructed with deletions within the luciferase-coding region such that the same primer pair could be used to amplify the indicated size fragment. PCR products were generated from these templates as described in Methods and purified using the 1-plate 96-well MagneSil™ method for the Biomek® FX. Purified samples were separated by agarose gel electrophoresis and analyzed as described in Methods. Lanes 1,3,5,7,9,11: unpurified amplification reaction. Lanes 2,4,6,8,10,12: MagneSil™ purified samples. Lane 13: 100bp DNA Ladder (Promega).
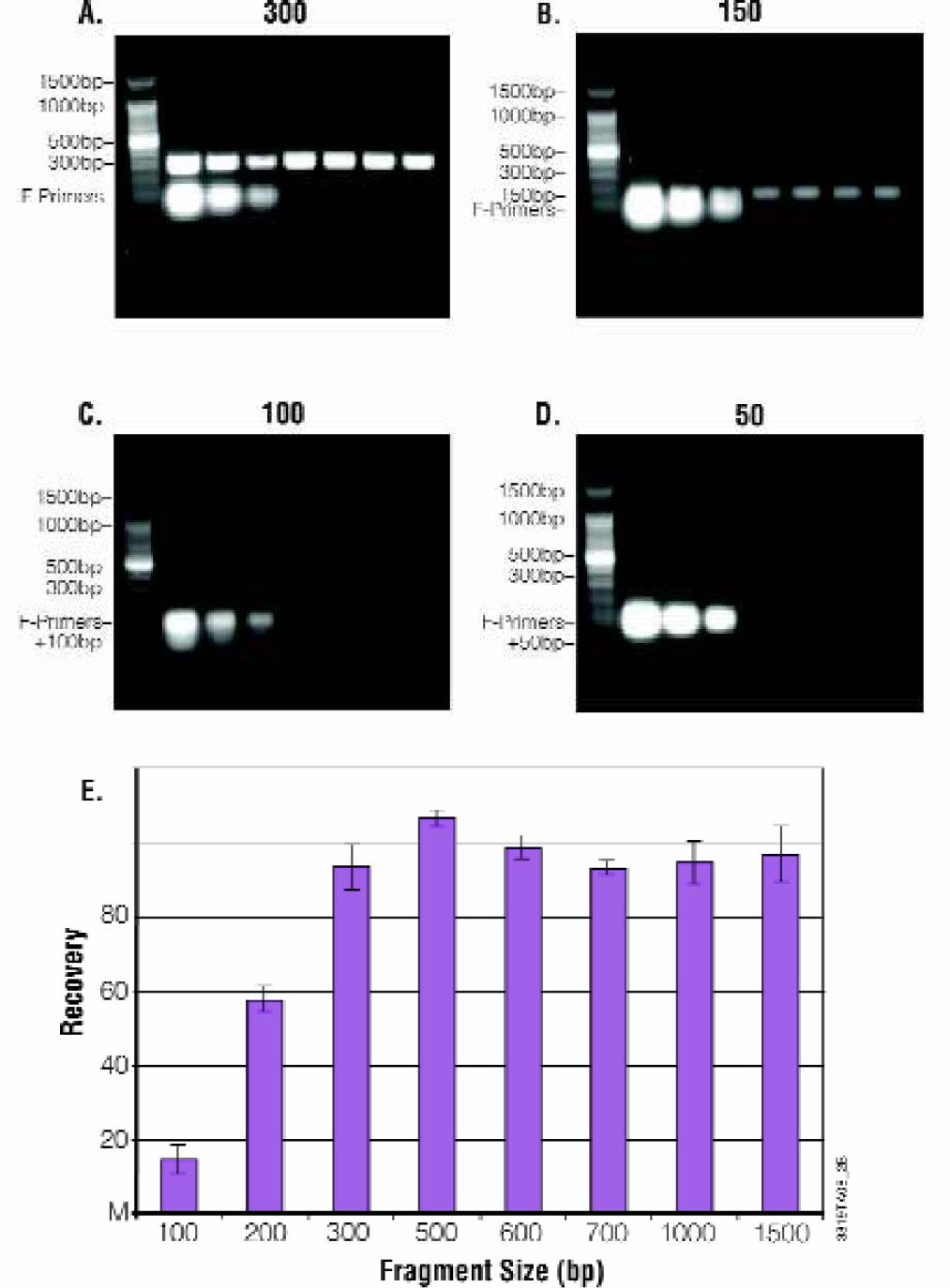
Fragment Purity. PCR products were prepared, purified and analyzed as described in figure 6. Panel A: 300bp PCR fragment, Panel B: 150bp PCR fragment, Panel C: 100bp PCR fragment, Panel D: 50bp PCR fragment. Lanes 2–4: A titration of unpurified PCR product is shown (100%, 50% and 25% of the input volume used for MagneSil™ purification). Four replicates purified using the MagneSil™ PCR Clean-Up System. Lane 1: 100bp DNA Ladder. Panel E: Recovers versus size for fragments 100–1000bp.
SEQUENCING APPLICATION
A 1000bp PCR product was generated from pGEM®-3zf (+) as described in Methods and purified using a 2-plate 96-well MagneSil™ method for the Biomek® FX. Purified samples were sequenced using the ABI BigDye® Terminator version 3 chemistry on an ABI Prism® 3700 DNA Analyzer (Figure 8). Average accuracy was greater than 98% with average Phred > 20 scores of over 600 for each plate. Results are unchanged when identical samples are purified using the 384-well method.
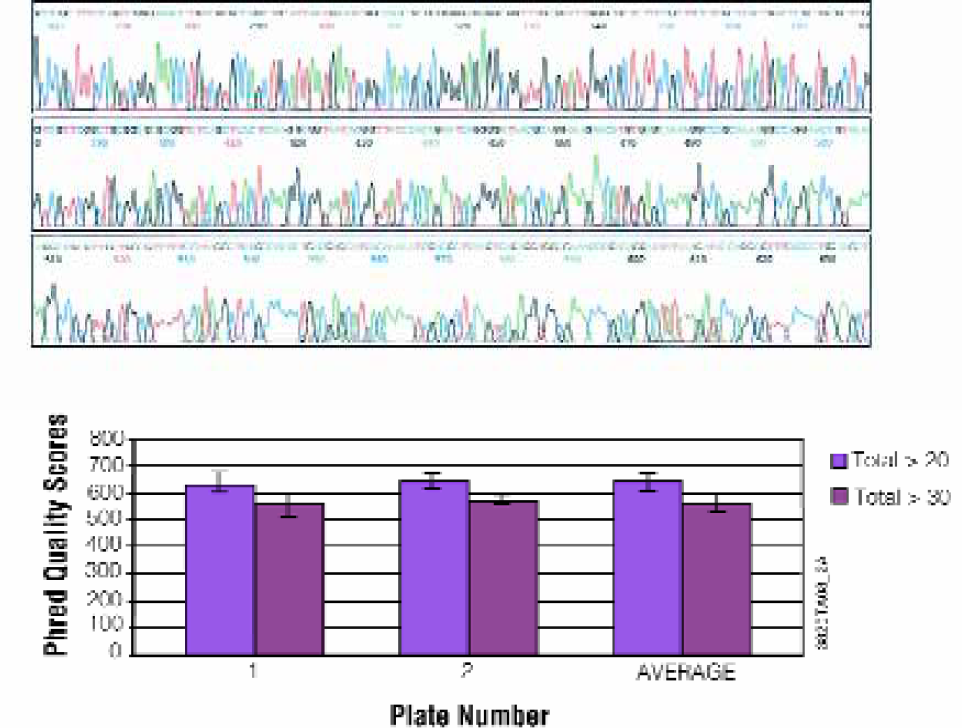
Fluorescent DNA Sequencing Application. A 1000bp fragment was prepared as described in Methods and purified using a 2-plate 96-well MagneSil™ method for the Biomek® FX. Purified samples were sequenced using the ABI BigDye® Terminator version 3 chemistry on an ABI Prism® 3700 DNA Analyzer. Panel A: Typical electropherogram. Panel B: Average Phred quality scores.
MICROARRAY APPLICATION
cDNA microarrays are typically prepared by spotting purified PCR products on a solid substrate, often a treated glass slide. The spotted PCR products serve as coordinately defined probes for interrogating mRNA expression levels. Target DNA's are typically fluorescent labeled during reverse transcription of RNA, prior to interrogation by hybridization.
Kanamycin RT-PCR products (undiluted, diluted 1:5, 1:25, 1:100) were purified using the Wizard® MagneSil™ PCR CleanUp System. The purified DNA was attached to poly-L-lysine coated glass slides by contact printing in replicate sets. Separate microarrays were hybridized with either Cy®3- or Cy®5-labeled kanamycin cDNA targets (Figure 9) produced using the ImProm-II™ Reverse Transcription System. The PCR products were purified independently in three plates using the stated dilutions. Four independent sample series were spotted from each of the three plates. Each sample set was arrayed in two identical sets to address intra-sample and spotting reproducibility (columns A and B). High inter- and intra-samples reproducibility were consistently seen (undiluted product c.v., 0.31%), indicating consistent purification yields and array spotting. Both Cy®3- and Cy®5-labeled cDNA gave similar results, with minimal background.
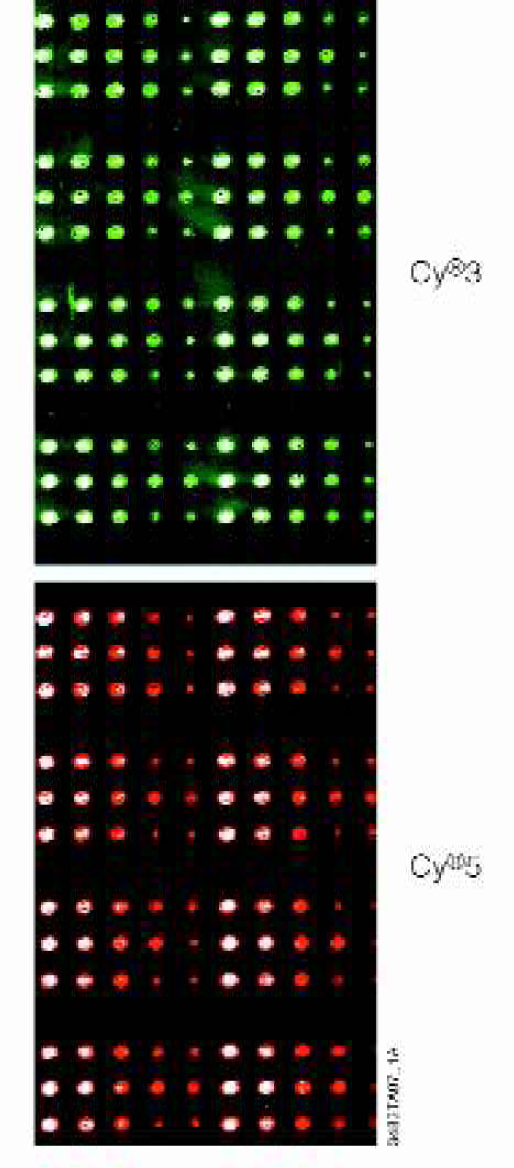
Microarray Application. A Sample quadrant of kanamycin-control reactions for both Cy®3- and Cy®5-target hybridizations. Each quadrant consists of four identical groups of three rows each. The three rows correspond to three 96-well plates containing replicates of the purified probes. Each row is a duplicate group of five spots corresponding to undiluted, 1:5, 1:25, and 1:100 diluted RT-PCR products, and a blank 3X SSC spot.
Conclusion
We have developed a fully automated procedure for the purification of PCR products using the Wizard® MagneSil™ PCR Clean-Up System. Methods have been developed in 96 and 384-well formats using a number of liquid handling robots. With the Biomek® FX allows as many as four plates can be processed in an unattended run of about 45 minutes. For higher order processing, the FX may be configured with a high capacity stacker carousel to re-supply consumables to the deck between processing cycles.
The MagneSil™ chemistry is highly reproducible and generates PCR products of high purity with minimal loss of target DNA. There is greater than 98% removal of primer and dsDNA fragments smaller than 100bp such as primer dimers. The system has utility for fragments from 0.15 to 23kbp. For fragments smaller than 500bp, yield is size dependent. Typical yields are greater that 80% for fragments above 500bp. Purified samples are suitable as templates for fluorescent DNA sequencing or as probes in microarray production. Typical sequencing results are greater than 98% accurate over 600–700 bases with Phred > 20 scores of greater than 600.
Footnotes
Acknowledgements
All trademarks are property of their respective owners.
